Abstract
A bacteriochlorophyll-containing bacterium, designated as strain N10T, was isolated from a terrestrial hot spring in Nagano Prefecture, Japan. Gram-stain-negative, oxidase- and catalase-positive and ovoid to rod-shaped cells showed the features of aerobic anoxygenic phototrophic bacteria, i.e., strain N10T synthesised bacteriochlorophylls under aerobic conditions and could not grow anaerobically even under illumination. Genome analysis found genes for bacteriochlorophyll and carotenoid biosynthesis, light-harvesting complexes and type-2 photosynthetic reaction centre in the chromosome. Phylogenetic analyses based on the 16S rRNA gene sequence and 92 core proteins revealed that strain N10T was located in a distinct lineage near the type species of the genera Tabrizicola and Xinfangfangia and some species in the genus Rhodobacter (e.g., Rhodobacter blasticus). Strain N10T shared < 97.1% 16S rRNA gene sequence identity with those species in the family Rhodobacteraceae. The digital DNA–DNA hybridisation, average nucleotide identity and average amino acid identity values with the relatives, Tabrizicola aquatica RCRI19T (an aerobic anoxygenic phototrophic bacterium), Xinfangfangia soli ZQBWT and R. blasticus ATCC 33485T were 19.9–20.7%, 78.2–79.1% and 69.1–70.1%, respectively. Based on the phenotypic features, major fatty acid and polar lipid compositions, genome sequence and phylogenetic position, a novel genus and species are proposed for strain N10T, to be named Neotabrizicola shimadae (= JCM 34381T = DSM 112087T). Strain N10T which is phylogenetically located among aerobic anoxygenic phototrophic bacteria (Tabrizicola), bacteriochlorophyll-deficient bacteria (Xinfangfangia) and anaerobic anoxygenic phototrophic bacteria (Rhodobacter) has great potential to promote studies on the evolution of photosynthesis in Rhodobacteraceae.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacteriochlorophyll (BChl)-producing aerobic bacteria, known as aerobic anoxygenic phototrophic bacteria (AAPB), have photosynthetic ability but cannot grow phototrophically (Shimada 1995; Yurkov and Hughes 2017). AAPB are widely found in natural environments and have attracted attention due to the ecological importance of their aerobic heterotrophic metabolism (Kolber et al. 2001). Culture-dependent and culture-independent studies have identified AAPB in the phyla Proteobacteria, Acidobacteria and Gemmatimonadetes (Thiel et al. 2018). Some AAPB are phylogenetically closely related to typical anaerobic anoxygenic phototrophic bacteria which produce BChls under anaerobic conditions with light and grow photoheterotrophically and photoautotrophically (Yurkov and Csotonyi 2009). Comparative studies between AAPB and non-AAPB are of considerable interest in understanding the evolutionary diversification of phototrophic organisms and their photosynthetic ability (Yurkov and Hughes 2017).
The class Alphaproteobacteria is a representative group that includes AAPB. Of more than 300 genera within Alphaproteobacteria, 41 genera contain AAPB, and this number is increasing with the continuous discovery of AAPB (Thiel et al. 2018). Bacteria of the genus Tabrizicola in the family Rhodobacteraceae in the class Alphaproteobacteria were firstly described as non-AAPB in 2013 (Tarhriz et al. 2013). Recently, BChl-production ability and photosynthetic gene cluster were reported for bacteria in this genus (Tarhriz et al. 2019; Han et al. 2020). Photosynthetic gene cluster is composed of approximately 40 genes for photosynthetic reaction centre, light-harvesting complexes, BChl and carotenoid biosynthesis and regulatory factors (Zsebo & Hearst 1984). The description of the genus Tabrizicola has been emended as “some species of this genus produce bacteriochlorophyll a under aerobic, heterotrophic conditions”, and the type species, T. aquatica, is defined as an aerobic anoxygenic phototrophic bacterium (Tarhriz et al. 2019). Bacteria of the genus Tabrizicola is phylogenetically related to some bacteria of the genera Xinfangfangia and Rhodobacter (Hu et al. 2018; Suresh et al. 2019; Hördt et al. 2020). Bacteria of the genus Xinfangfangia does not have photosynthesis-related genes (Hu et al. 2018), and BChl-production ability was not reported. Rhodobacter is one of the most well-known groups of anoxygenic phototrophic bacteria (Imhoff 2015); the type species, Rhodobacter capsulatus was originally reported in 1907 (Molisch 1907). The genus Rhodobacter consists of phylogenetically diverse species, and their reclassification is often proposed (Suresh et al. 2019; Hördt et al. 2020). Phylogeny of the photosynthetic genes is actively studied to explain the patchy distribution of phototrophy in the family Rhodobacteraceae (Zheng et al. 2011; Imhoff et al. 2018; Imhoff et al. 2019; Brinkmann et al. 2018; Liu et al. 2019). To draw an elaborate picture of the evolutionary diversification of phototrophs and their photosynthetic ability in Rhodobacteraceae, it is desirable to acquire bacteria located at a phylogenetic lineage connecting anaerobic anoxygenic phototrophs, non-phototrophs and aerobic anoxygenic phototrophs.
In this study, we isolated a BChl-containing bacterium, strain N10T, from a hot spring in Japan. Cells produced BChl under aerobic conditions and could not grow under anaerobic conditions even with light. Phylogenetic analysis based on 16S rRNA gene sequences suggested that the isolate belonged to the family Rhodobacteraceae and was distantly related to the genera Tabrizicola, Xinfangfangia and Rhodobacter. The aim of this study was to determine the taxonomic position of strain N10T by polyphasic taxonomic analyses, and a novel genus and species are proposed for this aerobic anoxygenic phototrophic bacterium in the family Rhodobacteraceae.
Materials and methods
Sample collection and isolation
Microbial mats developed in hot spring water at Nakabusa Hot Springs, Nagano, Japan (36° 23′ 20″ N, 137° 44′ 52″ E) (Fig. S1) were collected. The temperature and pH of the hot spring water at the sampling site were 30 °C and pH 8.0, respectively. The greenish microbial mats are dominated by oxygenic phototrophs (Everroad et al. 2012). A piece of the mats was collected using a sterilized tweezers and brought to the laboratory in ice. Approximately 0.5 g of the sample suspended in 10 ml of sterile distilled water were aseptically homogenized on ice using POLYTRON PT10/35 (KINEMATICA, Switzerland), directly spread on 1/10 diluted PE agar plate (Hanada et al. 1995; Hirose et al. 2016) and aerobically cultivated at 30 °C in the dark. 1/10 diluted PE agar solidified with Bacto-Agar (1.5%, w/v) (Thermo Fisher Scientific, USA) contains (per litre, pH 7.5) 0.05 g each of sodium glutamate, sodium succinate, sodium acetate, yeast extract (FUJIFILM Wako Pure Chemical, Japan), Casamino acids, sodium thiosulfate and ammonium sulfate, 0.2 ml of vitamin mixture (Hanada et al. 1995), 5 ml of 1 mol/l phosphate buffer and 5 ml of a basal salt solution (Hanada et al. 1995). PE medium was originally developed by Hanada et al. (1995) as a phototrophic bacteria enrichment medium and the diluted versions have been widely used for isolation of aerobic and anaerobic anoxygenic phototrophic bacteria (Hirose et al. 2016). Colonies containing bacteriochlorophylls (BChls) were fluorescently detected with an imaging system: colonies on Petri dishes were illuminated with LED light at wavelengths of 375 nm and 590 nm, and the infrared fluorescence from BChl-containing colonies was observed using a CCD camera with a long-pass filter (> 850 nm) (Edmund Optics, USA) (Zeng et al. 2014). Colonies with BChl fluorescence were picked for isolation. Isolation was performed using the standard dilution plating technique at 30 °C under aerobic conditions in the dark with 1/10 diluted PE agar medium (Hanada et al. 1995; Hirose et al. 2016). Among four BChl-producing isolates in total six isolates, strain N10T which stably and strongly showed BChl-fluorescence was selected and used for further experiments. The purified strain was routinely cultured on 1/5 diluted PE agar (Hirose et al. 2016) or Reasoner’s 2A (R2A; FUJIFILM Wako Pure Chemical, Japan) (Reasoner and Geldreich 1985) agar solidified with 1.5% Bacto-Agar under aerobic dark conditions. R2A medium had been used for cultivation of the reference strains such as Rhodobacter blasticus (Hu et al. 2018). R. blasticus NBRC 16437T (= ATCC 33485T) was obtained from NBRC (Biological Resource Center, NITE, Japan) and cultivated under the same conditions for strain N10T.
Morphology and physiology
Morphological characteristics were examined using a phase-contrast microscope (Eclipse E600; Nikon, Japan). The cellular morphology was also observed by transmission electron microscopy (TEM) based on a rapid freezing and freeze-fixation method, performed at Tokai Electron Microscopy (Japan). For sample preparation for TEM, cells were frozen in liquid propane at − 175 °C and the frozen samples were substituted with 2% (v/v) glutaraldehyde, 1% (w/v) tannic acid in ethanol and 2% water at − 80 °C. After dehydration with ethanol at room temperature, the samples were infiltrated with propylene oxide and embedded in resin (Quetol-812, Nisshin EM, Japan). Ultra-thin section (70 nm) was prepared using an ultramicrotome (Ultracut UCT, Leica, Austria), stained with 2% (v/v) uranyl acetate and lead stain solution and observed using a TEM (JEM-1400Plus, JEOL, Japan). Gram staining was performed using the FAVOR-G kit (Nissui Pharmaceutical, Japan). Growth was tested on R2A agar at 4, 10, 15, 20, 25, 30, 35, 40, 45 and 50 °C. Salt tolerance was assessed in 1/5 diluted PE broth adjusted to 0%, 1%, 2% and 3% NaCl (w/v). Growth at various pH values (pH 5.0–9.0 at 1.0 pH intervals) was determined in 1/5 diluted PE broth. Anaerobic growth was tested using the AnaeroPack system (Mitsubishi Gas Chemical, Japan) on 1/5 diluted PE agar under illumination [photo-organoheterotrophic conditions, 2000 lx (tungsten light)] and on R2A agar in the dark (fermentative conditions). Anaerobic growth tests were also performed in screw-capped glass tubes fully filled with 1/5 diluted PE and R2A broth.
The in vitro absorption spectrum was obtained as follows: cells were harvested after a week of cultivation at 30 °C in 1/5 diluted PE broth under aerobic dark conditions and pigments were extracted from cells using acetone:methanol (7:2, v/v) (Tarhriz et al. 2019). The absorption spectrum in the range of 350–850 nm was determined using the UV-2600 spectrophotometer (Shimadzu, Japan).
Oxidase activity was detected using an oxidase reagent kit (BioMérieux, France). Catalase activity was determined by gas production using 3% H2O2 (w/v). Enzyme activities were evaluated using the API ZYM system (BioMérieux). The carbon source utilisation pattern was investigated using a GEN III MicroPlate (Biolog, USA).
Molecular phylogenetic analysis
Genomic DNA was extracted using the Wizard Genomic DNA Purification Kit (Promega). PCR amplification and sequencing of the 16S rRNA gene were performed as described previously (Hirose et al. 2016). The 16S rRNA gene sequence was compared with the sequence data from GenBank using the BLAST program. Phylogenetic trees based on 16S rRNA gene sequences were constructed with MEGA version 7.0 (Kumar et al. 2016) using the neighbour-joining (Saitou and Nei 1987) and maximum-likelihood (Felsenstein 1981) methods. The Kimura two-parameter model (Kimura 1980) was used to calculate evolutionary distances. The topology of the phylogenetic trees was evaluated using the bootstrap resampling method (Felsenstein 1985) with 1000 replicates.
Genome sequencing and analyses
The complete genome of strain N10T was sequenced by Bioengineering Lab. (Sagamihara, Japan) with GridION X5 (Oxford Nanopore Technologies, UK) and DNBSEQ-G400 (MGI, China). The sequences were assembled using Unicycler version 0.4.7 (Wick et al. 2017). Genes were annotated using Prokka version 1.13 (Seemann 2014) and DFAST version 1.2.4. (Tanizawa et al. 2018). From the genomic data of strain N10T and related species, 92 core gene sequences were extracted using the Up-to-date Bacterial Core Gene (UBCG) tool (Na et al. 2018). Concatenated amino acid sequences were prepared using the UBCG pipeline. A phylogenetic tree was constructed with MEGA version 7.0 (as described above) using the maximum-likelihood method. The average nucleotide identity (ANI) and digital DNA-DNA hybridisation (dDDH) values of strain N10T with its phylogenetic neighbours were calculated using the OrthoANI calculator (Lee et al. 2016) and the Genome-to-Genome Distance Calculator (GGDC 2.1; http://ggdc.dsmz.de/distcalc2.php) (Meier-Kolthoff et al. 2014), respectively. Average amino acid identity (AAI) values were calculated using AAI-Matrix (http://enve-omics.ce.gatech.edu/g-matrix/).
Chemotaxonomic characterization
The compositions of respiratory quinone, fatty acids and polar lipids were analysed by Techno Suruga Laboratory (Shizuoka, Japan). Cells for analyses were incubated at 30 °C on R2A agar under aerobic conditions in the dark for a week. Quinones were identified by HPLC as described previously (Hamada et al. 2010). Cellular fatty acids were identified using the Sherlock Microbial Identification System (version 6.0) with the TSBA6 database (MIDI, USA) (Sasser 2001). Polar lipid analysis was performed using TLC methods (Minnikin et al. 1979).
Results
Morphology
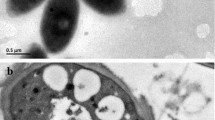
Colonies of strain N10T grown on R2A agar under aerobic dark conditions were beige, whereas those grown on 1/5 diluted PE agar were slightly purple. Cells were ovoid to rod-shaped, 0.7–1.0 μm in diameter and 1.3–1.8 μm in length (Fig. S2), non-motile and Gram-stain-negative (Fig. S3). TEM image indicated that cells of strain N10T had no lamellar internal membrane (Fig. 1), which are typically observed in Rhodobacter sp. (Imhoff 2015).
Phylogenetic analysis based on 16S rRNA gene sequences
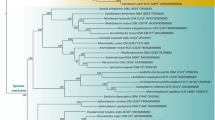
The nearly complete 16S rRNA gene sequence of strain N10T was obtained (1389 bp). BLAST analysis of the 16S rRNA gene sequence revealed that the highest sequence identities were 96.6% with Tabrizicola piscis K13M18T, 97.1% with Rhodobacter blasticus ATCC 33485T and 97.1% with Xinfangfangia soli ZQBWT. The maximum-likelihood tree based on 16S rRNA gene sequences demonstrated that strain N10T was distantly located at a distinct branch near three lineages containing the type species of Tabrizicola (i.e., Tabrizicola aquatica RCRI19T), Xinfangfangia (i.e., X. soli ZQBWT) and some Rhodobacter species such as R. blasticus ATCC 33485T (Fig. 2). This phylogenetic position of strain N10T was supported by phylogenetic analysis using the neighbour-joining method (Fig. S4).
Maximum-likelihood phylogenetic tree based on 16S rRNA gene sequences showing the phylogenetic position of strain N10T and closely related species. Roseobacter litoralis OCh114T (GenBank accession no. X78312) was added as an outgroup. Bootstrap values calculated from 1000 replications and the percentages (over 50%) from three algorithms are shown at the branch nodes (maximum-likelihood/neighbour joining/unweighted pair group method with arithmetic mean). Bar represents nucleotide substitutions per site. Bacteria known to produce BChl were marked with an asterisk (*) beside the accession numbers and among them, aerobic anoxygenic phototrophic bacteria were additionally marked with a hash (#)
Genome-based analysis
The complete genome of strain N10T consisted of a chromosome (4,154,010 bp) and four plasmids (119,227 bp, 70,477 bp, 32,096 bp and 16,643 bp). The DNA G + C content was 66.6 mol%. The genome encodes 4,242 protein-coding genes, 52 tRNAs and 6 rRNAs. The phylogenetic tree based on 92 single-copy core genes (UBCG) in the chromosome showed that strain N10T formed a separate branch from the lineage of T. aquatica, X. soli and R. blasticus (Fig. S5). The ANI and dDDH values between strain N10T and T. aquatica RCRI19T, R. blasticus ATCC 33485T and X. soli ZQBWT were 78.2%, 78.3% and 79.1%, respectively (for ANI), and 19.9%, 20.1% and 20.7%, respectively (for dDDH), which were all significantly lower than the cut-off values for discriminating bacterial species (95–96% ANI, 70% dDDH) (Rodriguez-R and Konstantinidis 2014; Goris et al. 2007). T. aquatica RCRI19T, R. blasticus ATCC 33485T and X. soli ZQBWT shared 72.4–83.3% of AAI; however, the AAI values between strain N10T and T. aquatica RCRI19T, R. blasticus ATCC 33485T and X. soli ZQBWT were 69.3%, 70.1% and 69.1%, respectively. Therefore, strain N10T could be differentiated at the genus level from Tabrizicola, Xinfangfangia and Rhodobacter.
The genome of strain N10T lacked the cbbS and cbbL genes that encode ribulose-1,5-bisphosphate carboxylase/oxygenase (RuBisCO). Strain N10T had the photosynthetic gene cluster, i.e., genes for BChl (bch) and carotenoid (car) biosynthesis, light-harvesting complexes and type-2 photosynthetic reaction centre (pufL, pufM and puhA) in the chromosome. PufLM sequence is a great tool for the phylogenetic analysis of anoxygenic phototrophic bacteria (Imhoff et al. 2018). As observed in the phylogenetic trees of 16S rRNA gene (Figs. 2 and S4) and core proteins (Fig. S5), PufLM amino acid sequence of strain N10T was closely related to those from a group of R. blasticus (e.g., Rhodobacter thermarum and Rhodobacter flagellates) and Tabrizicola (e.g., T. aquatica and T. piscis) but distantly related to those from the type species of Rhodobacter, i.e., R. capsulatus (data not shown). R. capsulatus and its phylogenetic relatives lacked the acsF gene that encodes the oxygen-dependent type of magnesium-protoporphyrin IX monomethyl ester cyclase for BChl biosynthesis and had the bchE gene for the oxygen-independent type (Boldareva-Nuianzina et al. 2013), but strain N10T had both the genes, acsF and bchE, in the photosynthetic gene cluster, as with R. blasticus, R. thermarum, R. flagellates, T. aquatica and T. piscis.
Phenotypic and physiological characterization
Strain N10T grew aerobically at 10–45 °C, at pH 7.0–8.0 and with the addition of 0–1% NaCl (w/v). Fermentative growth was not observed on R2A agar in the dark. Strain N10T did not grow in 1/5 diluted PE and R2A media under anaerobic light conditions. The in vitro absorption spectrum of strain N10T (Fig. S6) showed absorption peaks at approximately 360, 600 and 770 nm, indicating the presence of BChl a, and an absorption at approximately 480 nm, indicating the presence of carotenoids, which were similar to the spectra of phototrophic relatives (Eckersley and Dow 1980; Tarhriz et al. 2019).
Strain N10T was found to be oxidase- and catalase-positive. According to the API ZYM assay results, cells were positive for alkaline phosphatase, esterase (C4), esterase lipase (C8), leucine arylamidase, acid phosphatase, naphthol-AS-BI-phosphohydrolase, α-galactosidase, β-galactosidase, α-glucosidase, β-glucosidase and N-acetyl-β-glucosaminidase, and negative for lipase (C14), valine arylamidase, cystine arylamidase, trypsin, α-chymotrypsin, β-glucuronidase, α-mannosidase and α-fucosidase. In the GEN III MicroPlate, strain N10T was found to be positive for glucuronamide, Tween 40, acetoacetic acid, propionic acid and acetic acid, and negative for dextrin, d-maltose, d-trehalose, d-cellobiose, gentiobiose, sucrose, d-turanose, stachyose, d-raffinose, α-d-lactose, d-melibiose, β-methyl-d-glucoside, d-salicin, N-acetyl-d-glucosamine, N-acetyl-β-d-mannosamine, N-acetyl-d-galactosamine, N-acetyl neuraminic acid, α-d-glucose, d-mannose, d-fructose, d-galactose, 3-methyl glucose, d-fucose, l-fucose, l-rhamnose, inosine, d-sorbitol, d-mannitol, d-arabitol, myo-inositol, glycerol, d-glucose-6-PO4, d-fructose-6-PO4, d-aspartic acid, d-serine, gelatin, glycyl-l-proline, l-alanine, l-arginine, l-aspartic acid, l-glutamic acid, l-histidine, l-pyroglutamic acid, l-serine, pectin, d-galacturonic acid, l-galactonic acid lactone, d-gluconic acid, d-glucuronic acid, mucic acid, quinic acid, d-saccharic acid, p-hydroxy-phenylacetic acid, methyl pyruvate, d-lactic acid methyl ester, l-lactic acid, citric acid, α-keto-glutaric acid, d-malic acid, l-malic acid, bromo-succinic acid, γ-amino-butryric acid, α-hydroxybutyric acid, β-hydroxy- d,l-butyric acid, α-keto-butyric acid and formic acid.
Chemotaxonomic characteristics
The sole respiratory quinone was found to be ubiquinone-10, which is a typical characteristic of bacteria of the genera Tabrizicola, Xinfangfangia, and Rhodobacter (Imhoff 2015; Tarhriz et al. 2013; Hu et al. 2018). Strain N10T had C19:0 cyclo ω8c, C18:1 ω7c 11-methyl and summed feature 8 (C18:1 ω6c and/or C18:1 ω7c) as the major fatty acids (> 20%) and C18:0 and C18:0 3OH as the minor fatty acids (> 1%) (Table S1). The polar lipids of strain N10T mainly comprised phosphatidylglycerol (PG), phosphatidylethanolamine (PE), phosphatidylcholine (PC), a glycolipid (GL), an unidentified phospholipid (PL), an unidentified amino lipid (AL) and eight unidentified polar lipids (UL1 − UL8) (Fig. S7).
Discussion
The phylogenetic trees based on the 16S rRNA gene (Figs. 2 and S4) and core proteins (Fig. S5) showed that strain N10T was located at a distinct branch, which was related to Tabrizicola, Xinfangfangia and some Rhodobacter species in the family Rhodobacteraceae. The characteristics differentiating strain N10T from these genera are summarised in Tables 1 and 2. The ANI and dDDH values between strain N10T and T. aquatica RCRI19T, R. blasticus ATCC 33485T and X. soli ZQBWT were lower than the thresholds used for prokaryotic species delineation (Meier-Kolthoff et al. 2013). Tabrizicola, Xinfangfangia and a group of Rhodobacter containing R. blasticus shared AAI values of 72–83% (Hӧrdt et al. 2020), and the AAI values between strain N10T and these type strains were 69.1–70.1%, indicating that strain N10T could be differentiated at the genus level from Tabrizicola, Xinfangfangia and Rhodobacter. Moreover, strain N10T had C19:0 cyclo ω8c and C18:1 ω7c 11-methyl as major fatty acids and a glycolipid as a major polar lipid, which were not detected as major components in the closely related species (Table 2). Therefore, strain N10T is considered to represent a novel species in a novel genus in the family Rhodobacteraceae, for which the name Neotabrizicola shimadae gen. nov., sp. nov. is proposed.
Strain N10T had photosynthesis-related genes, synthesised BChl a under aerobic conditions and did not grow under anaerobic conditions even with light, indicating that strain N10T was categorised as an aerobic anoxygenic phototrophic bacterium (Shimada 1995; Yurkov and Hughes 2017). Strain N10T lacked complex intercellular membrane structure (Fig. 1), similar with T. aquatica (Tarhriz et al. 2019). Within the microbial mats dominated by oxygenic phototrophs in water stream, strain N10T heterotrophically grows mainly by oxygen respiration. Strain N10T could not utilize many organic compounds including d-glucose and sucrose, similar to Xinfangfangia sp. (Kämpfer et al. 2019) but different from Tabrizicola sp. (Sheu et al. 2020) and R. blasticus (Imhoff 2015). The growth of strain N10T may be partially supported by energy supply through photophosphorylation (Beatty 2002) in the mats. Positive reaction of N-acetyl-β-glucosaminidase of strain N10T could differentiate between strain N10T and Xinfangfangia sp. (Hu et al. 2018; Kämpfer et al. 2019). N-acetyl-β-glucosaminidase activity may help to obtain organic compounds from other bacterial cells in the mats (Jørgensen et al. 2003).
As estimated by phylogenetic analysis (Brinkmann et al. 2018), the photosynthetic ability was vertically evolved (e.g., Tabrizicola) (Tarhriz et al. 2019) and frequently lost (e.g., Xinfangfangia) (Hu et al. 2018; Kämpfer et al. 2019) in Rhodobacteraceae. The phylogenetic trees shown in Fig. 2, Fig. S4 and Fig. S5 suggest that strain N10T is located at the transition between aerobic anoxygenic phototrophic bacteria, Tabrizicola and typical anoxygenic phototrophic bacteria, Rhodobacter sp. such as R. blasticus. Further phylogenetic and biochemical studies on strain N10T and the relatives could provide beneficial knowledges about the evolutionary diversification of phototrophic organisms and their photosynthetic ability.
Description of Neotabrizicola gen. nov.
Neotabrizicola (Ne.o.ta.b.ri.zi’co.la. Gr. masc. adj. neos new; N.L. fem. n. Tabrizicola a bacterial genus name: N.L. fem. n. Neotabrizicola a new Tabrizicola).
Cells are Gram-stain-negative and ovoid to rod-shaped. The respiratory quinone is ubiquinone-10. The major fatty acids are C19:0 cyclo ω8c, C18:1 ω7c 11-methyl and summed feature 8 (C18:1 ω6c and/or C18:1 ω7c). The polar lipids are phosphatidylglycerol, phosphatidylethanolamine, phosphatidylcholine and glycolipid. The genus Neotabrizicola belongs to the family Rhodobacteraceae within the class Alphaproteobacteria. The type species is Neotabrizicola shimadae.
Description of Neotabrizicola shimadae sp. nov.
Neotabrizicola shimadae (shi.ma’dae. N.L. gen. n. shimadae, of Shimada, named after Dr. Keizo Shimada, Professor Emeritus, Tokyo Metropolitan University, Tokyo, Japan, in recognition of his many contributions to the biology of aerobic anoxygenic phototrophic bacteria).
Displays the following properties in addition to those given in the genus description. Cells are 0.7–1.0 μm in diameter and 1.3–1.8 μm in length, non-motile and obligate aerobe. Colonies are beige and purple and oxidase- and catalase-positive. Growth occurs at 10–45 °C, pH 7.0–8.0 and with 0–1% (w/v) NaCl. Extracted pigments from cells show absorption peaks at approximately 480 nm and 770 nm, corresponding to carotenoids and BChl a, respectively. Growth does not occur under anaerobic conditions even with light. In addition to the major fatty acids mentioned in the genus description, C18:0 and C18:0 3OH are present in relatively low proportions. The G + C content of the genomic DNA of the type strain is 66.6 mol%.
The type strain, N10T (= JCM 34381T = DSM 112087T) was isolated from a hot spring sample from Nagano Prefecture, Japan. The GenBank/EMBL/DDBJ accession number for the 16S rRNA gene sequence of strain N10T is LC350018. The genome sequence data of this strain are publicly available under the GenBank/EMBL/DDBJ accession numbers, CP069370 (chromosome) and CP069371–CP069374 (four plasmids).
References
Beatty JT (2002) On the natural selection and evolution of the aerobic phototrophic bacteria. Photosynth Res 73:109–114
Boldareva-Nuianzina EN, Bláhová Z, Sobotka R, Koblízek M (2013) Distribution and origin of oxygen-dependent and oxygen-independent forms of Mg-protoporphyrin monomethylester cyclase among phototrophic Proteobacteria. Appl Environ Microbiol 79:2596–2604
Brinkmann H, Göker M, Koblížek M et al (2018) Horizontal operon transfer, plasmids, and the evolution of photosynthesis in Rhodobacteraceae. ISME J 12:1994–2010
Eckersley K, Dow C (1980) Rhodopseudomonas blastica sp. nov.: a member of the Rhodospirillaceae. Microbiology 119:465–473
Everroad CR, Otaki H, Matsuura K, Haruta S (2012) Diversification of bacterial community composition along a temperature gradient at a thermal spring. Microbes Environ 27:374–381
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P, Tiedje JM (2007) DNA–DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol 57:81–91
Hamada M, Iino T, Iwami T, Harayama S, Tamura T, Suzuki K-I (2010) Mobilicoccus pelagius gen. nov., sp. nov. and Piscicoccus intestinalis gen. nov., sp. nov., two new members of the family Dermatophilaceae, and reclassification of Dermatophilus chelonae (Masters et al. 1995) as Austwickia chelonae gen. nov., comb. nov. J Gen Appl Microbiol 56:427–436
Han JE, Kang W, Lee JY et al (2020) Tabrizicola piscis sp. nov., isolated from the intestinal tract of a Korean indigenous freshwater fish, Acheilognathus koreensis. Int J Syst Evol Microbiol 70:2305–2311
Hanada S, Hiraishi A, Shimada K, Matsuura K (1995) Chloroflexus aggregans sp. nov., a filamentous phototrophic bacterium which forms dense cell aggregates by active gliding movement. Int J Syst Bacteriol 45:676–681
Hirose S, Matsuura K, Haruta S (2016) Phylogenetically diverse aerobic anoxygenic phototrophic bacteria isolated from epilithic biofilms in Tama River, Japan. Microbes Environ 31:299–306
Hördt A, García LM, Meier-Kolthoff JP, Schleuning M, Weinhold L-M, Tindall BJ, Gronow S, Kyrpides NC, Woyke T, Göker M (2020) Analysis of 1,000+ type-strain genomes substantially improves taxonomic classification of Alphaproteobacteria. Front Microbiol 11:468
Hu Q, Zhang L, Hang P, Zhou XY, Jia WB, Li SP, Jiang JD (2018) Xinfangfangia soli gen. nov., sp. nov., isolated from a diuron-polluted soil. Int J Syst Evol Microbiol 68:2622–2626
Imhoff JF (2015) Genus Rhodobacter. In: Whitman WB, Rainey F, Kämpfer P, Trujillo M, Chun J, DeVos P, Hedlund B, Dedysh S (eds) Bergey’s manual of systematics of archaea and bacteria. Wiley, New York
Imhoff JF, Rahn T, Künzel S, Neulinger SC (2018) Photosynthesis is widely distributed among Proteobacteria as demonstrated by the phylogeny of PufLM reaction center proteins. Front Microbiol 23:2679
Imhoff JF, Rahn T, Künzel S, Neulinger SC (2019) Phylogeny of anoxygenic photosynthesis based on sequences of photosynthetic reaction center proteins and a key enzyme in bacteriochlorophyll biosynthesis, the chlorophyllide reductase. Microorganisms 7:576
Jørgensen NOG, Stepanaukas R, Pedersen AGU, Hansen M, Nybroe O (2003) Occurrence and degradation of peptidoglycan in aquatic environments. FEMS Microbiol Ecol 46:269–280
Kämpfer P, Busse HJ, McInroy JA, Criscuolo A, Clermont D, Glaeser SP (2019) Xinfangfangia humi sp. nov., isolated from soil amended with humic acid. Int J Syst Evol Microbiol 69:2070–2075
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kolber ZS, Plumley FG, Lang AS, Beatty JT, Blankenship RE, VanDover CL, Vetriani C, Koblizek M, Rathgeber C, Falkowski PG (2001) Contribution of aerobic photoheterotrophic bacteria to the carbon cycle in the ocean. Science 292:2492–2495
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Lee I, Kim YO, Park SC, Chun J (2016) OrthoANI: an improved algorithm and software for calculating average nucleotide identity. Int J Syst Evol Microbiol 66:1100–1103
Liu Y, Zheng Q, Lin W, Jiao N (2019) Characteristics and evolutionary analysis of photosynthetic gene clusters on extrachromosomal replicons: from streamlined plasmids to chromids. mSystems 4:e00358-19
Meier-Kolthoff JP, Auch AF, Klenk HP, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform 14:60
Meier-Kolthoff JP, Klenk HP, Göker M (2014) Taxonomic use of DNA G+C content and DNA-DNA hybridization in the genomic age. Int J Syst Evol Microbiol 64:352–356
Minnikin DE, Collins MD, Goodfellow M (1979) Fatty acid and polar lipid composition in the classification of Cellulomonas, Oerskovia and related taxa. J Appl Bacteriol 47:87–95
Molisch H (1907) Die Purpurbakterien: nach neueren untersuchungen. Gustav Fischer Verlag, Jena, pp 1–95
Na SI, Kim YO, Yoon SH, Ha SM, Baek I, Chun J (2018) UBCG: up-to-date bacterial core gene set and pipeline for phylogenomic tree reconstruction. J Microbiol 56:280–285
Reasoner DJ, Geldreich EE (1985) A new medium for the enumeration and subculture of bacteria from potable water. Appl Environ Microbiol 49:1–7
Rodriguez-R LM, Konstantinidis KT (2014) Bypassing cultivation to identify bacterial species. Microbe 9:111–118
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sasser M (2001) Identification of bacteria by gas chromatography of cellular fatty acids, MIDI technical note 101. MIDI Inc, Newark
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30:2068–2069
Sheu C, Li ZH, Sheu SY, Yang CC, Chen WM (2020) Tabrizicola oligotrophica sp. nov. and Rhodobacter tardus sp. nov., two new species of bacteria belonging to the family Rhodobacteraceae. Int J Syst Evol Microbiol 70:6266–6283
Shimada K (1995) Aerobic anoxygenic phototrophs. In: Blankenship RE, Madigan MT, Bauer CE (eds) Anoxygenic photosynthetic bacteria. Advances in photosynthesis and respiration, vol 2. Springer, Dordrecht, pp 105–122
Suresh G, Lodha TD, Indu B, Sasikala C, Ramana CV (2019) Taxogenomics resolves conflict in the genus Rhodobacter: a two and half decades pending thought to reclassify the genus Rhodobacter. Front Microbiol 10:2480
Tanizawa Y, Fujisawa T, Nakamura Y (2018) DFAST: a flexible prokaryotic genome annotation pipeline for faster genome publication. Bioinformatics 34:1037–1039
Tarhriz V, Thiel V, Nematzadeh G, Hejazi MA, Imhoff JF, Hejazi MS (2013) Tabrizicola aquatica gen. nov. sp. nov., a novel alphaproteobacterium isolated from Qurugöl Lake nearby Tabriz city, Iran. Antonie Van Leeuwenhoek 104:1205–1215
Tarhriz V, Hirose S, Fukushima SI, Hejazi MA, Imhoff JF, Thiel V, Hejazi MS (2019) Emended description of the genus Tabrizicola and the species Tabrizicola aquatica as aerobic anoxygenic phototrophic bacteria. Antonie Van Leeuwenhoek 112:1169–1175
Thiel V, Tank M, Bryant DA (2018) Diversity of chlorophototrophic bacteria revealed in the omics era. Annu Rev Plant Biol 69:21–49
Wick RR, Judd LM, Gorrie CL, Holt KE (2017) Unicycler: Resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput Biol 13:1–22
Yurkov V, Csotonyi JT (2009) New light on aerobic anoxygenic phototrophs. In: Hunter CN, Daldal F, Thurnauer MC, Beatty JT (eds) The purple phototrophic bacteria. Advances in photosynthesis and respiration, vol 28. Springer, Dordrecht, pp 31–55
Yurkov V, Hughes E (2017) Aerobic anoxygenic phototrophs: four decades of mystery. In: Hallenbeck PC (ed) Modern topics in the phototrophic prokaryotes. Springer, Cham, pp 193–214
Zeng Y, Feng F, Medová H, Dean J, Koblížek M (2014) Functional type 2 photosynthetic reaction centers found in the rare bacterial phylum Gemmatimonadetes. Proc Natl Acad Sci USA 111:7795–7800
Zheng Q, Zhang R, Koblížek M, Boldareva EN, Yurkov V et al (2011) Diverse arrangement of photosynthetic gene clusters in aerobic anoxygenic phototrophic bacteria. PLoS ONE 6:e25050
Zsebo KM, Hearst JE (1984) Genetic-physical mapping of a photosynthetic gene cluster from R. capsulata. Cell 37:937–947
Acknowledgements
We thank Mr. Takahito Momose (the owner of Nakabusa hot springs) for allowing us to collect samples from the hot springs. We also thank Prof. Aharon Oren (The Hebrew University of Jerusalem) for helpful corrections of the proposed name and its etymology.
Funding
This research was supported by a research grant from the Institute for Fermentation, Osaka.
Author information
Authors and Affiliations
Contributions
SM, SHn and SHr planned the research. SM and SHi carried out experiments. SM, TI and SHr analysed the data and drafted the manuscript. MO and SHn supervised the research and SHn provided the research funding. All authors proofread the manuscript.
Corresponding author
Ethics declarations
Conflict of interests
The authors declare that there are no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Muramatsu, S., Hirose, S., Iino, T. et al. Neotabrizicola shimadae gen. nov., sp. nov., an aerobic anoxygenic phototrophic bacterium harbouring photosynthetic genes in the family Rhodobacteraceae, isolated from a terrestrial hot spring. Antonie van Leeuwenhoek 115, 731–740 (2022). https://doi.org/10.1007/s10482-022-01728-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-022-01728-6