Abstract
Sedimentary environments in the Arctic are known to harbor diverse microbial communities playing a crucial role in the remineralization of organic matter and associated biogeochemical cycles. In this study, we used a combination of culture-dependent and culture-independent approaches to understanding the bacterial community composition associated with the sediments of a terrestrial versus fjord system in the Svalbard Arctic. Community-level metabolic profiling and growth response of retrieved bacterial isolates towards different carbon substrates at varying temperatures were also studied to assess the metabolic response of communities and isolates in the system. Bacterial species belonging to Cryobacterium and Psychrobacter dominated the terrestrial and fjord sediment retrievable fraction. Amplicon sequencing analysis revealed higher bacterial diversity in the terrestrial sediments (Shannon index; 8.135 and 7.935) as compared to the fjord sediments (4.5–5.37). Phylum Proteobacteria and Bacteroidetes dominated both terrestrial and fjord sediments. Phylum Verrucomicrobia and Cyanobacteria were abundant in terrestrial sediments while Epsilonbacteraeota and Fusobacteriia dominated the fjord sediments. Significant differences were observed in the carbon substrate utilization profiles between the terrestrial and fjord sediments at both 4 °C and 20 °C incubations (p < 0.005). Utilization of N-acetyl-D-glucosamine, D-mannitol and Tween-80 by the sediment communities and bacterial isolates from both systems, irrespective of their temperature incubations implies the affinity of bacteria for such substrates as energy sources and for their survival in cold environments. Our results suggest the ability of sediment bacterial communities to adjust their substrate utilization profiles according to condition changes in the ecosystems and are found to be less influenced by their phylogenetic relatedness.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The Arctic is considered as a 'global hotspot' that is highly sensitive to climate change (IPCC 2013). Reduction in Arctic sea ice, glacier ice and snow, thawing of permafrost, and increase in productivity of the vegetation are some of the most direct impact of globally rising temperatures on Arctic ecosystems (Descamps et al. 2017). The Svalbard archipelago is one of the regions which is experiencing the fastest temperature increase within the circumpolar Arctic, along with the highest rate of sea ice loss and glacier retreat over the past three decades (Nuth et al. 2013).
The tidal and land terminating glaciers in Svalbard deliver sediment-laden freshwater into the downstream fjord systems, which in turn show strong physico-chemical as well as biological gradients along the head-to-mouth (glacier-to-ocean-influenced) axis. Our previous study on a glacio-marine transect in Svalbard Arctic reported significant differences in the bacterial community structure associated with a land terminating glacier (ice and snow), its pro-glacier melt waters and downstream fjord waters indicating the unique nature of these systems and the influence of environmental variables in determining the patterns of bacterial diversity (Thomas et al. 2020). The sediments associated with such glacier influenced and fjord environments also represents complex microbial habitats supporting distinct bacterial and archaeal communities with complex metabolism (Stibal et al. 2012; Anesio et al. 2017; Zeng et al. 2017; Teske et al. 2011). These microbial fraction comprise the bulk of the biomass and are crucial in remineralization of organic matter and associated biogeochemical cycles (Stibal et al. 2012; Lin et al. 2017). So far, there are numerous reports on microbial diversity, physiology and functions associated with supraglacial cryoconite sediments (Edwards et al. 2011; Lutz et al. 2017; Poniecka et al. 2020), subglacial systems (Skidmore et al. 2005; Stibal et al. 2012; Sułowicz et al. 2020) and glacier forefield soils (Kim et al. 2017; Nash et al. 2018; Malard et al. 2019), but studies on glacier snout and forefield associated sediments are limited to a single culture-dependent study from the sediments of a meltwater stream from Midtre Lovénbreen glacier (Reddy et al. 2009). They have reported the cold-active enzyme production and pigmentation properties of pro-glacier sediment bacteria and indicated the presence of Proteobacteria, Cytophaga-Flavobacterium-Bacteroidetes and high G + C Gram-positive bacteria as common inhabitants of these habitats. Since the glacier snout and forefield sediments are relatively unexplored in terms of microbial studies, it would be interesting to understand their bacterial community structure and how their metabolic signature reflects the glacier environment.
However, there are many studies highlighting the microbial community structure (Zeng et al. 2017; Conte et al. 2018; Fang et al. 2019) and functions (Teske et al. 2011; Buongiorno et al. 2019) associated with Svalbard fjord sediments. To highlight a few, Fang et al. (2019) have observed significant differences in the bacterial community structure between sediments of inner, middle and outer fjord basins and reported the influence of location depth, temperature and salinity on sediment community. The increased influx of Atlantic waters into the Kongsfjorden system influencing sediment community composition from the outer to the inner fjord has been reported by Zeng et al. (2017). Teske et al. (2011) reported higher rates of hydrolytic activities for Kongsfjorden sediment bacterial communities to hydrolyze polysaccharides and algal extracts as compared to the seawater communities. They also reported the presence of some microbial key populations (Bacteroidetes) having specific enzymatic capabilities, for example to readily hydrolyze substrates which were found to be recalcitrant in surface and bottom waters and suggested the need to test the linkages between enzymatic potential and phylogenetic identity using cultured isolates.
These studies are indicative that the bacterial communities associated with the Arctic glacio-marine sediments tend to exhibit distinct physiological properties and metabolic functions driven by the substrate availability and the various environmental conditions of each system. Therefore, assessing the metabolic potential of microbial communities along with their community structure is crucial in understanding their ecological role in such complex environments. The cultivable bacterial fraction, although accounting for as little as 1% of the total bacterial community, provides opportunities in exploring the metabolic diversity and traits within microbes inhabiting these environments.
Furthermore, in the light of enhanced warming scenario in the Arctic, with observed and projected annual average Arctic warming approximately twice the global mean (Overland et al. 2019), it is also imperative to understand the influence of temperature on the metabolic response of the total community as well as on different bacterial species of the terrestrial and fjord system. Kritzberg et al. (2010) studied the bacterial response in terms of production and respiration from the Fram strait waters by experimental manipulations of temperature and resources in combinations. They reported enhanced bacterial production and respiration following a temperature increase of + 6 °C from the in-situ conditions and the response to temperature was higher in resource amended treatments, indicative of a substrate-temperature interaction regulating the bacterial metabolism.
Keeping these aspects in mind, the following questions were posed 1) whether the total sediment community metabolic response towards ecologically important substrates in the terrestrial and fjord systems are different from the metabolic response exhibited by individual bacteria isolated from such systems (2) whether phylogenetically-related species tend to exhibit similar metabolic responses or whether sediment-type and origin influence their metabolic responses and (3) whether temperature variations have an influence on the metabolic profiles of the total community as well as on different bacterial species of the terrestrial and fjord system. In order to address these questions, in the present study we aim to explore the taxonomic diversity and metabolic potential of the total community as well as the retrievable fraction of a terrestrial versus fjord sediment system in the Svalbard. Cultivation-dependent and cultivation-independent high throughput amplicon sequencing approach was followed accompanied with measurement of physico-chemical parameters associated with each system. Community-level physiological profiling (CLPP) of the sediments using EcoPlates™ and growth response of retrievable bacteria towards different carbon substrates, with varying temperatures were studied. Such studies would help in estimating and eventually predicting the bacterial responses to varying substrate availability in the highly changeable Arctic ecosystems.
Materials and methods
Study area and sampling strategy
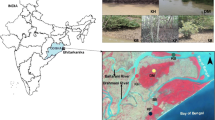
The study area covered Vestrebroggerbreen (VB) glacier which is a land terminating polythermal glacier in Ny-Ålesund (4.7 km2 long) and the downstream fjord system–Kongsfjorden (Fig. 1). The meltwater channels starting from the glacier snout drains through the Broggerbreen valley and finally discharges into the Kongsfjorden. The Kongsfjorden is a glacio-marine influenced fjord system located between 78° 04′ N–79° 05′ N and 11° 03′ E–13° 03′ E (Fig. 1). Kongsfjorden is influenced by Atlantic and Arctic water masses as well as by the glacial influx of sediment-laden freshwaters which in turn affect the ecosystem dynamics by creating strong environmental gradients and changes in community structure (Hop et al. 2002).
Map of the glacierized terrestrial regions and Kongsfjorden with the sediment sampling points (Vestrebroggerbreen (VB) glacier snout, foreland (BR) and fjord (KNBR, KNS1 and KNS9)) marked (https://github.com/MikkoVihtakari/PlotSvalbard)
A total of 3 sediment samples were collected from the Vestrebroggerbreen glacier snout region (VB1–VB3) and pooled together to form a single representative sample (VB). Similarly, 3 sediment samples were collected from the glacier foreland region (BR1–BR3), and a homogenous single sample was obtained as BR (Fig. 1). Surface sediment samples were collected across Kongsfjorden from the point where the meltwater channels empty into the fjord (KNBR), from the outer fjord mouth (KNS1), and the inner fjord (KNS9) region (Fig. 1) using Van-veen grab deployed from MS Teisten (https://nyalesundresearch.no/infrastructures/ms-teisten/). VB and BR sediments are representatives of the Arctic terrestrial system while KNBR, KNS1, and KNS9 sediments represent the fjord system. The overlying water temperature and salinity were measured using a Waterproof Portable meter (Cyberscan series 600) (Eutech instruments, Thermo Fisher Scientific, USA) for all the sampling locations. All the samples were collected in sterile sampling bags (Nasco Whirlpaks, Himedia, India) and were immediately transferred to − 20 °C until further processing. An aliquot of the samples was maintained at 4 °C for cultivation-dependent analysis. The samples were then transported in a frozen state to the home laboratory at National Centre for Polar and Ocean Research, India for analysis.
Chemical analysis of sediment samples
The pH of the sediment pore water was measured by centrifuging the sediment sample and measuring the pH of the supernatant water using Cyberscan pH meter 510 (Eutech Instruments, Thermo Fisher Scientific, USA). Water content (5 g of each sample) was measured as gravimetric weight loss after drying the sediment at 105 °C until constant weight. Organic and inorganic carbon were analyzed as per Winkelmann and Knies (2005) using PRIMACSMCS TOC analyser (SKALAR, Netherlands). Sediment samples were analyzed for trace elements using an Inductively-coupled Plasma Mass Spectrometer (Thermo iCAP Q ICP-MS, Thermo Fisher Scientific, USA). Sample preparation involved microwave digestion as detailed by Lu et al. (2013) and the USEPA method 3051, followed by a final dilution of 25 times with MilliQ. Total Mercury was analyzed using Direct Mercury Analyser (Duel cell-DMA 80, Milestone, Italy).
Total microbial abundance
Total sediment microbial count was carried out according to Epstein et al. (1997) using 4, 6-diamidino-2-phenylindole (DAPI) with minor modification. In brief, 0.5 g of sediment in 50 mL of filter-sterilized saline (0.9%) was vortexed for 10 min followed by ultra-probe sonication (at 40% speed over 80 s with interruption of 2–4 times) using an Ultrasonic homogenizer (Model 3000, Biologics, Inc, USA) for dislodging cells from sediment particles. A 2 mL aliquot of the sample was then incubated with DAPI (20 μL of 1 μg mL−1 working solution per mL) for 5 min and filtered through black 0.22 µm membrane filters (Nucleopore Track-Etch Membrane, Whatman). The total count was enumerated under the Leica DM6 B microscope (Leica Microsystems, Wetzlar, Germany).
Isolation and identification of retrievable heterotrophic bacteria
Retrievable bacterial fractions from the terrestrial and fjord sediments were enumerated by the spread plate method (Sanders 2012). The sample suspension was prepared by sonicating 5 g of sediment in 45 mL of filter-sterilized saline (0.9%). After serial dilutions (10–2 and 10–4 dilutions), the sediment suspension (100 µL) was plated onto solid R2A media (1/2 strength), Antarctic Bacterial Medium (ABM), Tryptone Soy Agar (TSA) (1/10th strength), Actinomycete isolation agar (AIA) and Zobell Marine Agar (ZMA) medium (1/4th strength). The plates were then incubated at 4 °C and 20 °C for 2–3 weeks and bacterial isolation and purification was carried out based on unique morphological characteristics.
The bacterial genomic DNA was extracted from the pure bacterial cultures using ChargeSwitch gDNA mini bacteria kit (Invitrogen, USA). The extracted DNA was subjected to Rep-PCR amplification with the BOX A1R primer (Yang and Yen 2012) and based on their unique banding patterns, isolates were chosen for 16SrRNA gene amplification (Sinha et al. 2017) followed by sequencing using 3500 Genetic Analyzer (Applied Biosystems, Thermofisher Scientific, MA, USA). The obtained sequences were trimmed and assembled using Codon code Aligner software Version 8.0.2 and the consensus contigs were chimera checked using Pintail 1.1 software. The 16S rRNA gene nucleotide sequences were analyzed on EzBioCloud server against validly published type strains of bacterial species (Yoon et al. 2017). The identified 16S rRNA gene sequences (104 sequences) were submitted in GenBank under the accession numbers MN080149–MN080222, MT309496–MT309525.
Bacterial community analysis using amplicon sequencing and downstream bioinformatic processing
Environmental DNA from the sediment samples (0.5 g of sediment) was extracted using DNeasy Power Soil kit (Qiagen, USA). The DNA concentration and purity were measured using the Qubit fluorometer (Thermo Fisher Scientific, USA) and by gel electrophoresis on 1% agarose. The DNA samples were sequenced on the HiSeq platform (Illumina, USA) using Pro341F/Pro805R primers for V3–V4 hypervariable regions of the 16S rRNA gene (Takahashi et al. 2014). The raw nucleotide sequences were deposited in the NCBI Sequence Read Archive (SRA) database under the SRA accession number: PRJNA475645. The downstream processing of raw DNA sequences was carried out according to the protocols detailed by Bolyen et al. (2019) using QIIME2 software. The demultiplexed raw sequence datasets were trimmed for the removal of 16S rRNA gene primers and multiplexing adapters by using QIIME2 Cutadapt plugin. Further, the sequences were subjected to denoising, dereplication, and chimera filtering using the DADA2 denoise-paired method of QIIME2. The clustering of reads into features was performed against SILVA_132 marker gene reference database (Quast et al. 2013) by training the database using the q2-feature-classifier plugin. Unique representative features/OTUs from each sample were further subjected to taxonomic classification using the classify-consensus-vsearch method in q2-feature-classifier plugin with 0.99% identity threshold. This was followed by taxonomy-based filtering to remove the sequences representing chloroplasts and mitochondria using the q2-taxa plugin and rarefaction of the OTU counts (rarefaction depth–9458, 10 iterations) before the downstream diversity analysis. The alpha diversity estimation was carried out using the q2-diversity plugin.
Community-level physiological profiling using EcoPlate
Biolog EcoPlate™ (Biolog Inc., USA) was used to evaluate community-level metabolic responses. The EcoPlate™ contains three replicate wells of 31 carbon substrates (Garland and Mills 1991) and control well (A1) with no added carbon substrate. The 31 substrates used in the EcoPlate™ represented carbohydrates, polymers, carboxylic acids, and amino acids, amines and amides. Carbon substrate utilization by microbes is indicated by the respiration-dependent reduction of the tetrazolium dye leading to a purple coloration which was quantified spectrophotometrically using a microplate reader (Synergy 2-multi mode microplate reader, BioTek Instruments, USA). Preparation of sediment samples for EcoPlate™ analysis was performed as detailed by Sinha et al. (2019) and the EcoPlates were incubated at both 4 °C and 20 °C to understand the influence of temperature on the metabolic potential of the sediment community. The intensity of color development was measured at a wavelength of 595 nm for 240 h (24 h time interval). The most intensive metabolism of carbon substrates was observed at 192 h for 4 °C and 216 h for 20 °C incubation. The results were expressed as Average Well Color Development (AWCD) and the Shannon–Wiener (H) and Richness (Rs) indices were computed. The AWCD was evaluated according to Garland and Mills (1991) following formula AWCD = ∑(ODi)/31; where ODi = C − R. C is the optical density measurement within each well and R is the absorbance value of the control well. The primary data were further normalized following the equation (C − R)/AWCD separately for each substrates using respective AWCD values. For the percentage utilization of each substrate category, the normalized values for each category were divided by the ∑(C − R)/AWCD values. The Shannon–Wiener (H) index was calculated using the formula H = − ∑pi(lnpi); where pi is the ratio of the activity on each substrate (ODi) to the sum of activities on all substrates ∑ODi (Gomez et al. 2006). Richness index (R) is the number of substrates metabolized by the microbes in each sample.
Understanding the growth yields of selected retrieved isolates of terrestrial and marine origin towards different carbon substrates
While the total sediment community metabolic profile was obtained from Ecoplate assay, it is imperative to understand how individual retrieved bacteria from terrestrial and marine samples could respond to different ecologically important substrates. For this purpose, from a total of 27 different species retrieved from the terrestrial sediments and 15 different species retrieved from the fjord sediments, we chose a total of 20 different species−10 belonging to the terrestrial sediments and 10 from the fjord sediments. Isolates retrieved from both 4 and 20 °C incubation were chosen based on their relative abundance (List of isolates given in Table 5). The fully grown bacterial cultures in the broth media were centrifuged and the resulting pellet was washed multiple times with 0.9% saline. The bacterial inoculum thus obtained was fixed to an optical density (OD) of 0.2 at 600 nm and the corresponding number of bacterial cells was measured using DAPI staining (protocol same as total microbial abundance). The substrate utilization experiment was carried out in microplates wherein 10 µL culture aliquot (with fixed OD of 0.2) was added to each well which was preloaded with 140 µL of a substrate minimum medium mix. Substrates tested included carbohydrates (D-cellobiose, sucrose, D-mannitol, erythritol, D-lactose, D-glucose, N-acetyl-D-glucosamine, and D-xylose), polymers (Tween 80, cellulose), amino-acids (L-phenyl alanine, L-serine, DL-Asparagine, L-glutamic acid), and carboxylic acids (D-malonic acid, D-galacturonic acid, p-aminobenzoic acid, DL-malic acid, and sodium pyruvate). Substrates were chosen based on the Biolog EcoPlate™ substrate list with few additional substrates included and were filter-sterilized before use. Each substrate was added to a Minimal media (MM, composition as described by Freese et al. (2010) and Pfennig (1974)) to produce a final concentration of 50 mg/L (Freese et al. 2010). The experiment was carried out in triplicates with controls comprising of MM with bacteria and without substrates, MM with substrates and without bacteria, and MM without bacteria and substrates. The experiment was tested at both 4 and 20 °C temperature incubations to check the effect of temperature on bacterial growth by the utilization of different carbon substrates. Absorbance (OD 600 nm) was measured for 15 days at regular intervals of 0, 3, 6, 9, 12, and 15 days.
For each of the selected bacterial isolates, the optimum temperature and temperature range for growth was determined on diluted R2A broth media (50% w/v of the original strength of 3.12 g/L) for terrestrial isolates and diluted ZMB (25% w/v of the original strength of 40.25 g/L) for fjord isolates at different temperature incubations (4, 10, 15, 20, 25, 30, and 35 °C) by measuring optical density at 600 nm.
Statistical analysis
The influence of geochemical properties in explaining the bacterial community structure associated with the terrestrial and fjord sites was done by Canonical Correspondence Analysis (using Bray Curtis dissimilarity) using CANOCO 4.56 statistical software. Pearson linear correlation was calculated using OriginPro 9.0 software to measure the correlation between geochemical properties and total bacterial community structure, bacterial community and Ecoplate substrate utilization as well as between geochemical properties and Ecoplate substrate utilization. Principal component analysis (PCA) was performed using the Canoco 4.56 package to ordinate the strains depending on the carbon sources they were able to metabolize. One-way Analysis of Variance (ANOVA) was performed for examining significant differences between the metabolic profiles of terrestrial and fjord sediment communities as well as between the retrievable bacterial isolates of the two systems, at 4 °C and 20 °C incubations.
Results and discussion
Geochemical properties of sediment samples
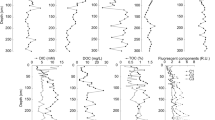
The overlying water temperature for the terrestrial sediments varied between 0.9 and 1.3 °C and salinity was 0.08 PSU, while that of the fjord sediments varied between 3.8 and 5.9 °C and 34.3–34.9 PSU (Table 1). The highest value of sediment pore water pH was observed in VB and BR sediments (~ 8.4) while the pH of the fjord sediment pore water ranged from 7.9 to 8.1 (Table 1). The total organic carbon (TOC) content among the terrestrial and fjord sediments varied between 1.2 and 2.9% with the highest value recorded in the outer fjord sediment (KNS1) and the lowest value noted in the foreland sediment (BR). A reduction in the concentration of TOC was observed from the outer to the inner fjord sediments while an increase in Inorganic carbon (IC) concentration was noted towards the inner fjord sediments (Table 1). The highest TOC values observed in the outer fjord sediment with a reduction noted from outer to the inner fjord sediments corresponds to the previous reports (Lu et al. 2013; Kumar et al. 2016). High turbidity towards the fjord head (inner fjord) and steep gradient in sedimentation rate along with the supply of terrigenous carbonates and organic matter deprived glacier materials towards the fjord head may be critical in producing such a clear spatial gradient in TOC and IC values in the fjord sediments (Kumar et al. 2016; Koziorowska et al. 2017). The highest values of Mn (362.0 mg/Kg), Co (13.2 mg/Kg), Cu (33.6 mg/Kg), Zn (223.9 mg/Kg), Ni (40.8 mg/Kg) and Cd (0.3 mg/Kg) were found in terrestrial sediments. Similarly, the outer fjord sediment, KNS1 exhibited the highest concentrations of Pb (7.9 mg/Kg) and Hg (90.9 µg/Kg) (Table 1). The higher concentrations of Ni, Cu, and Zn noted from the terrestrial sediments (Table 1) can be linked to the abundance of soluble rocks such as carbonates and sulfides in the glacierized areas and the contact of water with glacial debris causing the enrichment of elements (Dragon and Marciniak 2010). Łokas et al. (2016) has reported the role of atmospheric circulation transporting metals from global and local sources; marine aerosols and precipitation as major suppliers of trace metals in the Arctic glacier associated and terrestrial environments. The average values of trace metal concentrations of fjord sediments were comparable to the previously reported values from Kongsfjorden sediments (Lu et al. 2013; Grotti et al. 2017) and were lower than the baseline values reported by Lu et al. (2013). The higher concentrations of Hg and Pb towards the outer fjord were also reported by Lu et al. (2013) suggesting the influence of West Spitsbergen Current bringing relatively warm and saline waters from low and middle latitudes to the Arctic (long-range transport of metals). We could also observe significant correlations between the geochemical factors such as Ni and IC (p < 0.005), TOC and Hg (p < 0.05), Mn and Co (p < 0.005) as well as Hg and Pb (p < 0.01) (Supp Table 1). The significant correlation of TOC–Hg observed in our study may indicate the possible association of Hg to organic material (Grotti et al. 2017). Similarly, the association of Hg–Pb and Mn–Co is an indication that these metals had a similar source or may be produced by similar physicochemical processes (Bai et al. 2011).
Total microbial counts, retrievable heterotrophic bacterial counts, and phylogeny of cultivated bacterial isolates
Total microbial counts (TMC) in the sediment samples ranged between 106–8 cells/g. The lowest count (4.3 × 106 cells/g) was observed in the terrestrial sediment BR while the highest count (4.9 × 108 cells/g) was recorded in the outer fjord sediment KNS1 (Table 1). The total microbial count observed in the terrestrial sediments was two-fold lower as compared to the counts from the fjord sediments (108 cells/g) in this study. Reddy et al. (2009) has reported total bacterial count in the range 2.7–11 × 107 cells/g for the sediment samples collected from the snout of Midtre Lov´enbreen glacier to the confluence point of the melt water streams of the glacier with the fjord. Similarly, Conte et al. (2018) has reported the total microbial counts from the Kongsfjorden sediments to be in the range of 107 cells/g which is one-fold lower than the values observed in our study.
The total isolated heterotrophic bacterial count was calculated for both 4 and 20 °C incubations. Half-strength R2A media yielded the highest retrievable count for the terrestrial sediments at both the incubation temperature (4 °C: VB − 9.1 × 103 CFU/g and BR − 3.2 × 103 CFU/g and 20 °C: VB − 2.8 × 103 CFU/g and BR − 1.1 × 103 CFU/g) while quarter-strength ZMA yielded the highest number of isolates for the fjord sediments at both temperatures (4 °C: 1.6 × 105–2.5 × 105 CFU/g and 20 °C: 7.7 × 104–1.1 × 105 CFU/g) (Table 1).
106 and 64 bacterial isolates were retrieved (based on colony characteristics) from terrestrial and fjord sediments respectively from 4 °C incubation. Similarly, a total of 25 and 60 bacterial isolates were retrieved from 20 °C incubation. Pigmentation observed for more than half of the total terrestrial and fjord sediment retrievable bacterial fraction has been reported as an adaptive strategy to cold environments and for UV protection (Mueller et al. 2005). Based on the banding pattern obtained by rep-PCR, 109 bacterial isolates with unique banding patterns were selected for further identification. The results from 16S rRNA gene sequencing-based on a similarity cut-off > 98.7% revealed the presence of 27 different bacterial species in terrestrial sediments (VB and BR) and 15 in the fjord sediments (KNBR, KNS1, and KNS9). Eight bacterial sequences retrieved (7 from the terrestrial sediments and 1 from the fjord sediments) exhibited < 98.7% sequence similarity with the closest type strain in the EzBioCloud database indicating their novelty (Table 2). The dominant bacterial isolates from the terrestrial and fjord sediments were also tested to estimate their optimal temperature as well as temperature range for growth. Majority of the bacterial isolates (about 85%) were psychrotolerant (capable of growth close to 0 °C but with an optimum growth temperature of > 20 °C) while 3 isolates fit into the criteria for true psychrophiles (optimal temperature for growth at about 15 °C or lower and a maximal temperature for growth at about 20 °C). The psychrophilic isolates were closely related to the species Polaromonas glacialis, Cryobacterium psychrotolerans and Polymorphobacter fuscus, all of which were isolated from the terrestrial sediments in our study.
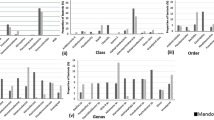
The bacterial isolates from the terrestrial and fjord sediments belonged to 4 phyla−Proteobacteria (constituted by class γ-proteobacteria and α-proteobacteria), Bacteroidetes, Actinobacteria, and Firmicutes (Fig. 2a). Phylum Actinobacteria dominated the terrestrial sediments at both 4 °C (58% of the total isolates) and 20 °C (38.5% of the total) incubation. The other dominant phyla isolated from the terrestrial sediments included Proteobacteria and Bacteroidetes. Bacteroidetes accounted for 13.4% of the total isolates from 4 °C and 33.3% of the total from 20 °C incubation. Class γ-proteobacteria was dominant at 4 °C (27.5% of the total) while class α-proteobacteria was dominant at 20 °C (25.6% of the total) (Fig. 2a).
a Percentage composition of retrievable bacteria belonging to the phylum Proteobacteria, Actinobacteria, Bacteroidetes and Firmicutes present in the terrestrial and fjord sediments from 4 °C and 20 °C incubation. Further class level abundance of the dominant phylum Proteobacteria is also shown in each pie diagram since the abundance of α-and γ-proteobacteria varied between the terrestrial and fjord sediments. b The relative abundance of bacterial OTUs at the phylum level and the abundance of major bacterial classes observed in the amplicon based culture-independent study
Actinobacterial phyla was represented by 5 different genera, of which the genus Cryobacterium was dominant with 7 different species (Supp Table 2a). The class γ-proteobacteria was represented by the genera Janthinobacterium, Herminiimonas, Polaromonas, Hydrogenophaga (order Betaproteobacteriales), and Pseudomonas (order Pseudomonadales). The genus Flavobacterium solely represented the phylum Bacteroidetes while the genus Polymorphobacter represented the class α-proteobacteria. The dominant species isolated from the terrestrial sediments from 4 °C incubation were Cryobacterium psychrotolerans (27.9% of the total species), Janthinobacterium lividum (19.5%), Cryobacterium roopkundense (19.1%), Flavobacterium degerlachei (7.6%), and Polaromonas glacialis (4.6%). Similarly, the dominant species isolated from 20 °C incubation were Polymorphobacter fuscus (25.6% of the total species), Flavobacterium degerlachei (23%), Cryobacterium luteum (17.9%), Flavobacterium omnivorum (10.3%), and Cellulomonas cellasea (7.7%).
The higher abundance of bacterial isolates affiliated to the phylum Actinobacteria in the terrestrial sediments corroborated well with the studies by Edwards et al. (2013) and Zhang et al. (2016) wherein they indicated the presence of Actinobacterial members as common inhabitants of glacier forelands and cryoconites, playing a significant role in soil development and biogeochemical cycling. The three psychrophilic bacterial species isolated in our study from the terrestrial sediments are known representatives of glacier systems of Polar and high elevation environments (Zhang et al. 2007; Margesin et al. 2012; Jia et al. 2015). Of which, the genus Cryobacterium has been previously reported as a dominant genera associated with the supraglacier systems, known to decompose a wide variety of organic carbon substrates (Poniecka et al. 2020). Liu et al. (2020) has also observed significantly higher number of genes involved in stress response, motility, and chemotaxis in the genus Cryobacterium as cold adaptive survival strategies. Similar properties of carbon substrate utilization and mechanisms to withstand unfavourable conditions—through dormancy (Darcy et al. 2011) and plasmid mediated genes (Ciok et al. 2018) has been reported in the genus Polaromonas. The third psychrophilic species belonged to the genus Polymorphobacter and dominated the bacterial fraction retrieved from 20 °C incubation. Polymorphobacter genus comprises of all psychrotolerant members isolated from low temperature environments and known for their aerobic anoxygenic photoheterotrophic nature (Jia et al. 2015; Phurbu et al. 2020). Other dominant terrestrial isolates belonged to Janthinobacterium sp. which are known to produce water-insoluble purple pigment violacein as an environmental stress responsive mechanism (Pantanella et al. 2007) and Flavobacterium sp. known for organic matter degradation in the glacier environment (Poniecka et al. 2020; Thomas et al. 2020).
γ-proteobacteria dominated the retrievable bacterial fraction in the marine fjord sediment samples at both 4 °C (65.2% of the total) and 20 °C (68.2% of the total), with the presence of 4 genera and 6 different species (Fig. 2a). Bacterial species belonging to the phylum Actinobacteria were retrieved only from the inner fjord sediment KNS9. The dominant species retrieved from the fjord sediments at 4 °C incubation were Psychrobacter fozii (26.3% of the total species), Psychrobacter cryohalolentis (17.8%), Psychrobacter glaciei (16.6%), and Flavobacterium degerlachei (16.5%). Similarly, Psychrobacter glaciei (41.9%) and Psychrobacter fozii (13.95%) dominated the retrievable fraction at 20 °C incubation.
Higher abundance of γ-proteobacterial genus Psychrobacter in the fjord sediments (Supp Table 2a and b) corroborates with the previous culture-based studies from Kongsfjorden where their potential role in carbohydrate degradation, as well as cold-active enzyme production, was demonstrated (Prasad et al. 2014; Jain and Krishnan 2017; Sinha et al. 2017). The retrieval of Actinobacterial species only from the inner fjord sediments might suggest the association of Actinobacterial taxa with the cold and less saline environment.
Flavobacterium degerlachei, belonging to the phylum Bacteroidetes, was the only common representative retrieved from both the terrestrial and fjord sediments from both incubation temperatures (Table 2) suggesting their adaptability to both the terrestrial and fjord environments and corresponds well with our previous report from the meltwaters and fjord waters (Thomas et al. 2020). The versatile nature of the Flavobacterium species could be linked to their ability to produce poly-unsaturated fatty acids, cold-adapted enzymes, and pigments along with their specialized role in uptake and degradation of high molecular weight fraction of organic matter (Liu et al. 2019).
Bacterial diversity as revealed by 16S rRNA gene amplicon sequencing
A total of 223,613 sequences and 2376 OTUs were obtained in the present study using SILVA_132 database at 99% sequence similarity. The Good’s coverage estimator of the OTUs ranged between 99.7 and 99.9% (Table 3), indicating sufficient sequence coverage in all the samples. It was found that 30,914 sequences representing 1263 OTUs were not identified based on the user database and hence have been characterized as unassigned OTUs. The unassigned OTUs accounted for about 19.98% of the total sequences identified from sediments of VB, 25% of BR, 26.8% of KNBR, 2.3% of KNS1, and 8.3% of KNS9. 1103 identified bacterial OTUs were classified into 22 phyla, 42 classes, 74 orders, 119 families, and 142 genera. In total, Proteobacteria (avg 31.2%), Epsilonbacteraeota (avg 18.9%) and Bacteroidetes (avg 16.3%) dominated the overall bacterial community. A total of 10 archaeal OTUs (1611 sequences) were identified representing the classes Bathyarchaeia, Thermoplasmata, and Nitrososphaeria. However, the number of archaeal OTUs detected in the amplicon analysis was only 0.42% of the total OTUs recovered.
The relative abundance of the major (> 1% abundance), minor (0.1–1% abundance), and rare bacterial genera (< 0.1% abundance) are summarized in Supp Fig. 2. OTUs from Proteobacteria (18.9–45.75%) and Bacteroidetes (7.5–24.2%) dominated both the terrestrial and fjord sediments. Verrucomicrobia were mostly abundant in the terrestrial sediment BR (19.4%) while Cyanobacterial OTUs were only found in the VB sediment (6.6%) Similarly, Patescibacteria and Planctomycetes were detected only in terrestrial sediments (Fig. 2b). Epsilonbacteraeota mainly resided in the outer fjord sediment KNS1 (50%) while Fusobacterial OTUs were observed only at station KNS9 (10.3%) (Fig. 2b).
The dominance of sequences affiliated to Proteobacteria, Bacteroidetes, and Verrucomicrobia from VB and BR sediments were previously reported for Svalbard lake sediments (Wang et al. 2016), although a comparatively lesser abundance of Acidobacterial phylum was found in the present study. The abundance and exclusive presence of cyanobacterial OTUs in the snout sediment (VB) might indicate their export from the supra-glacier systems to the snout region as cyanobacterial members are known to dominate the supraglacial ecosystems such as cryoconites playing a significant role in microbial community interactions and involved in cryoconite granule formation (Anesio et al. 2017; Garcia-Lopez and Cid 2017). The dominance of Verrucomicrobial OTUs in the forefield sediment BR is consistent with the results of Bendia et al. (2018), wherein they suggested the role of Verrucomicrobia in methane cycling in the glacier environments. Significant correlations were observed (Supp Table 1) between the bacterial classes Oxyphotobacteria (Cyanobacteria), VadinHA49 (Planctomycetes), ABY1 (Patescibacteria), and α-proteobacteria, which are dominant in terrestrial sediments suggesting their possible interactions with each other in the terrestrial system, contributing to ecosystem functioning. Such links were reported by Kosek et al. (2019) from Revelva catchment waters, wherein the abundance of Planctomycetes and α-proteobacteria in areas covered by cyanobacterial mats was related to their role in the degradation of algal matter. Further, the glacier snout sediment VB harbored highly diverse rare taxa (bacterial taxa having abundance < 0.1%) as compared to the other samples (64.8% of total rare taxa) (Supp Fig. 2), which in turn points out the importance of snout ecosystem acting as a 'seed bank' with less abundant highly diverse dormant taxa having unknown functional potential that could become numerically dominant and active with favorable environmental conditions (Dawson et al. 2017). This observation was further supported by the presence of 5 potential novel bacterial species from the retrievable fraction from the VB sediment (Table 2) also indicating the snout sediment to be a hotspot of rare taxa. The culture-dependent and culture-independent analysis of the snout and forefield sediments clearly indicated that their community comprise of both cosmopolitan taxa reported from the overlying meltwaters (Thomas et al. 2020), the supra-and subglacial systems (Stibal et al. 2012; Poniecka et al. 2020) and glacier-forefield soils (Nash et al. 2018; Malard et al. 2019) and rare taxa specific to the sediments.
The dominance of Proteobacteria (γ-proteobacteria), Epsilonbacteraeota, and Bacteroidetes observed in the Kongsfjorden sediments corresponds to the results of Zeng et al. (2017). An increased abundance of OTUs belonging to Epsilonbacteraeota was noted from the inner to the outer fjord sediment (Fig. 2b). This could be due to the influence of Atlantic water (Hamdan et al. 2013), which is more prevalent in the outer fjord (Cottier et al. 2005). Similarly, the dominance of OTUs belonging to class γ-proteobacteria (30.5–44.5%) and Bacteroidia (7.5–20.6%) in all fjord sediments is substantiated by our retrievable data, suggesting a vital linkage of the dominant bacterial taxa in the fjord sediments with organic matter degradation (Zeng et al. 2013).
The Shannon diversity index was highest for VB sediment (8.1) while the lowest value was noted for KNS9 sediment (4.5). We could observe a reduction in the bacterial diversity and richness from the terrestrial to the fjord sediments (Table 3). The higher Shannon diversity values for the terrestrial sediments are comparable to the values reported from soils and sediments from an Arctic lake in Svalbard (Wang et al. 2016). Despite the lower temperature and oligotrophic conditions prevailing in the glacier-influenced terrestrial environment, the higher diversity noted could be maintained by the mechanism of transient dormancy of the microbiota (Jones and Lennon 2010). The Shannon diversity values for fjord sediments are comparable to those reported by Zeng et al. (2017).
While comparing the amplicon sequencing results with the cultivable isolates, we could observe that all the species isolated in our study had their closest sequence variants among the OTUs from the amplicon results. Although cultivation-dependent techniques has the limitation of the selectivity of isolation media and culture conditions, it still stands significant in studying the physiological and metabolic potential of bacterial isolates from cold Polar environments which is otherwise not possible with the 16SrRNA amplicon sequencing approach. The combination of both these approaches has clearly described differences between the terrestrial and fjord sediments in terms of its community structure.
Bacterial community structure and their correlation with the sediment geochemical properties between the sampling sites
Canonical Correspondence Analysis showed that the total eigenvalue obtained as the summation of Axis 1 to 4 was 0.79. Axis 1 and Axis 2 accounted for 71.9% of the cumulative percentage variance observed in the relationship between species and the various environmental factors tested (Fig. 3). Among the various environmental factors tested in CCA, pore water pH showed significant influence on the species distribution (p-value < 0.005) explaining about 39% of the total variance by all variables. The significant influence of pH on the bacterial community structuring and their significant correlation with Acidobacterial and Actinobacterial OTUs (Supp Table1) was comparable to the results from the pan-Arctic survey of bacterial communities in Arctic soils by Malard et al. (2019). We could also observe a close association of IC (Inorganic Carbon) and trace metals such as Ni and Cd with the terrestrial VB bacterial community while Zn and sediment pore water pH exhibited a positive correlation with the BR community. Similarly, the close association of Pb, Hg, TOC, and water content with the fjord sediment communities was noted (Fig. 3). Therefore, our results establish a linkage between the environmental factors and bacterial community structuring associated with the terrestrial and fjord system.
Canonical correspondence analysis showing associations between the bacterial communities and geochemical properties of the terrestrial and fjord sediments. The red arrows represent environmental factors measured. The length of the arrows and the angle between each arrow and the nearest axis indicate the relative importance and closeness of that environmental factor in explaining the variation in the bacterial communities. The terrestrial sediments VB and BR are marked in orange stars and the fjord sediments KNBR, KNS1 and KNS9 are marked in blue stars
Metabolic functional analysis of sediment communities and isolates
It was well indicated in our culture-dependent and culture-independent bacterial diversity analysis that the dominant members of the terrestrial and fjord sediments are known for their capability for organic carbon degradation. We could note significant differences in the community metabolic profiles between the terrestrial and fjord sediments at both 4 °C and 20 °C incubations (p < 0.005) (Fig. 4a and b). However, at both temperatures, the terrestrial sediment community had shown a higher affinity to utilize amino acids, amines and amides (24.8–35%) as compared to the fjord community (19–25.5%). Similarly, a higher affinity for utilization of carbohydrates and polymers was noted for the fjord sediment community irrespective of temperature incubations. It was found that all 5 sediment communities were positive for the utilization of D-mannitol, Tween 80, Tween 40, and N-acetyl-D-glucosamine (GlcNAc) at both 4 and 20 °C incubations (Fig. 4a and b). Mannitol being low-molecular mass organic osmolytes, are accumulated in the cytoplasm by cold-adapted bacteria, to lower the cytoplasmic freezing point and avoid desiccation by counteracting water loss and cell shrinkage during freezing (Collins and Margesin 2019). Feltracco et al. (2020) reported the prevalence of alcohol sugar mannitol in coarse particles in Arctic aerosol denoting it as a local biogenic source. GlcNAc is a low-molecular weight amino sugar that is used for the synthesis of cell surface structures in bacteria and plays an important role in supplying carbon and energy by entering the glycolytic pathway after it is converted into fructose-6-phosphate (Álvarez-Añorve et al. 2005). GlcNAc uptake is reported as a widespread phenotype among marine bacteria by Cottrell and Kirchman (2000). Similarly, the Tween compounds, having an oleic acid moiety in their structure are commonly used by polar bacteria (Sala et al. 2008) and known to protect bacterial cells from adverse environmental conditions (Reitermayer et al. 2018). This is an indication that the terrestrial and fjord sediment communities utilize certain specific carbon substrates which could boost their survival mechanisms in Arctic environments.
Polymer substrates such as α-cyclodextrin and glycogen and carbohydrate substrate D-cellobiose (product of cellulose degradation), were utilized by all 3 fjord sediment communities (OD > 0.5) at both temperature incubations (Fig. 4a and b). Higher affinity for carbohydrates and complex polymer substrates over others observed in the fjord sediments suggests the presence of complex carbohydrate degrading bacterial groups in the fjord sediments (Teske et al. 2011; Jain and Krishnan 2017). Similarly, Pyruvic acid methyl ester, D-glucosaminic acid, D-galactonic acid-gamma lactone, L-arginine, L-asparagine, and L-serine were utilized efficiently by the terrestrial sediment communities (OD > 0.5) over other substrates. Significant correlations were noted between the ecoplate carbon substrates, bacterial community and different geochemical properties (Supp Table 3a and b). We could also observe that the linkages of community and geochemical properties with that of the ecoplate carbon substrates varied with the Ecoplate incubations at two different temperatures (Supp Table 3a and b). Shannon–Wiener (H) index and Richness index (Rs) were higher for fjord sediments as compared to terrestrial sediments at 4 °C while the values were found to be lower than terrestrial sediments at 20 °C incubation (Table 4). At these two temperatures, terrestrial and fjord communities showed variation in their metabolic responses, which may have significant implications on the substrate availability in the natural environment.
The total community metabolic response, however, might vary from the metabolic response of single bacteria representing each system. To test this hypothesis, the dominant cultivable bacterial fraction from both the terrestrial and fjord sediments was tested for their ability to grow in the presence of different carbon substrates. Some of the retrievable isolates chosen represented genera like Flavobacterium, Polaromonas and Photobacterium which accounted for > 5% relative abundance in the NGS results. For the experimental study, incubations at 4 °C and 20 °C were taken into account to determine the influence of varying temperatures on bacterial growth response. It was found that 50% of the terrestrial isolates and 70% of the fjord isolates could utilize at least 1 of the carbon substrates provided, at both temperature incubations (Table 5). Among the terrestrial isolates, the highest number of substrates were utilized by the species Cryobacterium roopkundense and Arthrobacter ginsengisoli (100% substrates utilized at 4 °C and 42% at 20 °C) belonging to Actinobacteria and Janthinobacterium lividum (37% and 21% of substrates utilized at 4 °C and 20 °C) belonging to Proteobacteria. Similarly, among the fjord isolates, maximum number of substrates were utilized by the species Psychrobacter glaciei (94.7% substrates utilized at 4 °C and 31.5% at 20 °C), Photobacterium frigidiphilum (47% at 4 °C and 26% at 20 °C) and Psychrobacter nivimaris (26% at 4 °C, 42% at 20 °C) belonging to Proteobacteria respectively (Table 5). PCA also showed that members of the bacterial phyla Proteobacteria and Actinobacteria have more affinity for utilization of different carbon substrates (Supp Fig. 3a and b).
We have also looked into the metabolic profiles of the psychrophilic species isolated in our study. Among the isolates, Polymorphobacter fuscus utilized carbohydrate (N-acetyl-D-glucosamine), polymer (Tween 80) and amino acid (L-serine) substrates at 4 °C while Cryobacterium psychrotolerans could utilize only carbohydrate substrates-D-mannitol and N-acetyl-D-glucosamine at 4 °C. Similarly, Polaromonas glacialis could utilize only one amino acid substrate-L-serine at 4 °C. There was no substrate utilization observed at 20 °C except for the isolate Polymorphobacter fuscus which could utilize Tween 80 at 20 °C (Supp Table 4a and b). Therefore, among the three psychrophilic species, Polymorphobacter sp. had a higher metabolic potential as compared to the other 2, although their metabolic capabilities were lower as compared to the psychrotolerant members isolated from the terrestrial system (Table 5). Comparison between the psychrophilic and psychrotolerant representatives of the genus Cryobacterium (C. psychrotolerans, C. roopkundense, C. luteum) indicated distinct metabolic capabilities for different species belonging to the same genera isolated from the same environment.
We could also observe that at 4 °C incubation, maximum number of bacterial isolates have utilized N-acetyl-D-glucosamine > Sucrose > Tween 80 > D-mannitol = D-galacturonic acid = L-serine = L-glutamic acid > Cellulose = D-cellobiose. Similarly, at 20 °C, we found maximum utilization of Tween80 > D-mannitol > D-galacturonic acid > N-acetyl-D-glucosamine > D-cellobiose > D-glucose = D-lactose. This corroborates well with the Ecoplate results wherein substrates such as N-acetyl-D-glucosamine, D-mannitol, and Tween 80 were utilized by all the sediment communities irrespective of 4 °C and 20 °C incubations suggesting the ecological importance of these substrates in the Arctic sedimentary environments.
Similar to the community metabolic response, we could note significant differences in the bacterial growth yields (in terms of total cell count) towards different carbon substrates between the terrestrial and fjord isolates at both 4 °C and 20 °C incubations (p value < 0.05). Figure 5a and b indicate a decline in bacterial growth in terms of total cell count (*107 cells/ml) towards different carbon sources at 20 °C incubation as compared to 4 °C incubation. An exception was noted for the isolates F4 (Planococcus halocryophilus) and F7 (Paeniglutamicibacter antarcticus), wherein they yielded higher cell counts at 20 °C in the presence of different carbohydrates, polymers, and amino acid substrates. There was a noticeable temperature selection particularly for the terrestrial isolates i.e. 4 °C incubation found to have higher growth yields for different substrates than 20 °C incubation. Production of cold-adapted enzymes having high level of specific activity at low temperatures might have enhanced their affinity to utilize different substrates at lower temperature (Struvay and Feller 2012). This needs further evidence through experiments involving varying temperatures and varying substrate concentrations as well as whole-genome based studies on the psychrotolerant isolates which can shed light on the different mechanisms involved in high metabolic activity at temperatures much below their temperature optima for growth.
a Bacterial growth yield results expressed in terms of bacterial cell count (*107cells/mL) for both terrestrial and fjord isolates grown in the presence of selected carbon substrates at 4 °C incubation observed at 12th day of growth, b Bacterial growth yield results for terrestrial and fjord isolates grown in the presence of selected carbon substrates at 20 °C incubation observed at 12th day of growth. Terrestrial isolates are marked as T1–T10 while fjord isolates are marked as F1–F10 along with the respective phylum details described in brackets as Bactero-Bacteroidetes, Actino-Actinobacteria, Proteo-Proteobacteria and Firmi-Firmicutes. Details of the isolates: T1-Flavobacterium degerlachei (4VBSedT3), T2-Arthrobacter ginsegisoli (4VBSedA2), T3-Janthinobacterium lividum (4BR1SedT2), T4-Polymorphobacter fuscus (20BRSedZ4), T5-Cryobacterium roopkundense (4VBSedR1), T6-Cryobacterium luteum (20VBSedR2), T7-Cryobacterium psychrotolerans (4BR2SedZ2), T8-Flavobacterium omnivorum (20VBSedR5), T9-Polaromonas glacialis (4VBSedT8) and T10-Cellulomonas cellasea (4BR1SedT3). F1-Flavobacterium degerlachei (4KNS1SedZ1), F2-Psychrobacter glaciei (4KNS1SedA4), F3-Paracoccus sediminilitoris (20KNBRSedZ3), F4-Planococcus halocryophilus (4KNS9SedZ1), F5-Psychrobacter fozii (4KNS1SedAA3), F6-Photobacterium frigidiphilum (4KNS9SedR2), F7-Paeniglutamicibacter antarcticus (4KNS9SedA1), F8-Psychrobacter nivimaris (20KNBRSedR2), F9-Psychrobacter cryohalolentis (4KNS9SedA2), F10-Flavobacterium frigoris (4KNS1SedA3)
Another aspect of our study was to check whether phylogenetically related bacterial members exhibit similar growth response to varying substrates and varying temperature incubations or whether they are influenced by the sediment-type. For this, isolates belonging to the species Flavobacterium degerlachei, one retrieved from the terrestrial sediment and one from the fjord sediment were included in the study. Similarly, different bacterial species belonging to the same genera were also considered for the experiment (Cryobacterium roopkundense, Cryobacterium luteum, and Cryobacterium psychrotolerans from the terrestrial sediments, Psychrobacter glaciei, Psychrobacter fozii, Psychrobacter nivimaris and Psychrobacter cryohalolentis from the fjord sediments). Our results indicated a clear distinction in the growth response of different isolates irrespective of their phylogenetic relatedness (Table 5 and Supp Table 4a, b). Similar results were reported by Poniecka et al. (2020) wherein they observed striking differences in the metabolic capabilities of closely related isolates assigned to the same OTU. Since bacterial strains/species can adjust their substrate utilization profiles according to condition changes, the phylogenetic origin seems to be a less important structuring component of sediment bacterial communities (Freese et al. 2010). Detailed whole-genome level studies could further give insights into the metabolic pathways in phylogenetically similar bacterial species from terrestrial and fjord environments.
Conclusion
Significant differences in the carbon substrate utilization profiles between the terrestrial and fjord sediments at both 4 °C and 20 °C incubations (p < 0.005) implies the adaptive responses of bacterial members to utilize the available substrates in the varying natural environment. The dominance of class γ-proteobacteria observed in the fjord sediments along with the preferential utilization of carbohydrates and complex polymers in the metabolic profile of bulk sediments indicated their potential role in complex organic matter degradation in the fjord sediments. Higher taxonomic diversity with the presence of diverse rare taxa noted in the terrestrial sediment VB along with the higher affinity to utilize amino acids, amines and amides need to be emphasized further for a better understanding of their ecological role in the glacier-influenced terrestrial environment. Utilization of D-mannitol, N-acetyl-D-glucosamine, and Tween 80 by both terrestrial and fjord sediment communities as well as by the cultivated bacterial fraction indicates the readily available nature of such carbon substrates in the Arctic sedimentary environments, promoting survivability of bacteria in the extremely cold environments. The significant influence of temperature on the growth response of terrestrial and fjord isolates towards different carbon substrates was indicated in our study, with phylogeny found to be a less important structuring component of bacterial metabolic potential. Thus, our polyphasic approach contributes to a better understanding of the taxonomic and metabolic diversity of bacteria associated with the terrestrial and fjord sedimentary system, which could be a base to understand bacterial-environmental interactions and underlying ecological processes.
Data availability
The raw nucleotide sequences generated for this study can be found in the NCBI Sequence Read Archive (SRA) database under the SRA accession number: PRJNA475645. The 16S rRNA gene sequences (104 sequences) obtained from the cultivation-dependent study are available in GenBank under the accession numbers MN080149-MN080222, MT309496-MT309525.
References
Álvarez-Añorve LI, Calcagno ML, Plumbridge J (2005) Why does Escherichia coli grow more slowly on glucosamine than on N-acetylglucosamine? Effects of enzyme levels and allosteric activation of GlcN6P deaminase (NagB) on growth rates. J Bacteriol 187:2974–2982. https://doi.org/10.1128/JB.187.9.2974-2982.2005
Anesio AM, Lutz S, Chrismas NAM, Benning LG (2017) The microbiome of glaciers and ice sheets. npj Biofilms Microbiomes 3:1–11. https://doi.org/10.1038/s41522-017-0019-0
Bai J, Cui B, Chen B et al (2011) Spatial distribution and ecological risk assessment of heavy metals in surface sediments from a typical plateau lake wetland, China. Ecol Model 222:301–306. https://doi.org/10.1016/j.ecolmodel.2009.12.002
Bendia AG, Signori CN, Franco DC et al (2018) A mosaic of geothermal and marine features shapes microbial community structure on deception Island Volcano, Antarctica. Front Microbiol 9:1–13. https://doi.org/10.3389/fmicb.2018.00899
Bolyen E, Rideout JR, Dillon MR et al (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857. https://doi.org/10.1038/s41587-019-0209-9
Buongiorno J, Herbert LC, Wehrmann LM et al (2019) Complex microbial communities drive iron and sulfur cycling in Arctic Fjord sediments. Appl Environ Microbiol 85:1–16. https://doi.org/10.1128/AEM.00949-19
Ciok A, Budzik K, Zdanowski MK et al (2018) Plasmids of psychrotolerant Polaromonas spp. isolated from arctic and Antarctic glaciers—diversity and role in adaptation to polar environments. Front Microbiol 9:1–17. https://doi.org/10.3389/fmicb.2018.01285
Collins T, Margesin R (2019) Psychrophilic lifestyles: mechanisms of adaptation and biotechnological tools. Appl Microbiol Biotechnol 103:2857–2871. https://doi.org/10.1007/s00253-019-09659-5
Conte A, Papale M, Amal S et al (2018) Bacterial community structure along the subtidal sandy sediment belt of a high Arctic fjord (Kongsfjorden, Svalbard Islands). Sci Total Environ. 620:203–211. https://doi.org/10.1016/j.scitotenv.2017.11.077
Cottier F, Tverberg V, Inall M et al (2005) Water mass modification in an Arctic fjord through cross-shelf exchange: The seasonal hydrography of Kongsfjorden, Svalbard. J Geophys Res Ocean 110:1–18. https://doi.org/10.1029/2004JC002757
Cottrell MT, Kirchman DL (2000) Natural assemblages of marine Proteobacteria and members of the Cytophaga-flavobacter cluster consuming low-and high-molecular-weight dissolved organic matter. Appl Environ Microbiol 66:1692–1697. https://doi.org/10.1128/AEM.66.4.1692-1697.2000
Darcy JL, Lynch RC, King AJ et al (2011) Global distribution of Polaromonas phylotypes—evidence for a highly successful dispersal capacity. PLoS ONE. https://doi.org/10.1371/journal.pone.0023742
Dawson W, Hör J, Egert M et al (2017) A small number of low-abundance bacteria dominate plant species-specific responses during rhizosphere colonization. Front Microbiol 8:1–13. https://doi.org/10.3389/fmicb.2017.00975
Descamps S, Aars J, Fuglei E et al (2017) Climate change impacts on wildlife in a high Arctic archipelago—Svalbard, Norway. Glob Chang Biol 23:490–502. https://doi.org/10.1111/gcb.13381
Dragon K, Marciniak M (2010) Chemical composition of groundwater and surface water in the Arctic environment (Petuniabukta region, central Spitsbergen). J Hydrol 386:160–172. https://doi.org/10.1016/j.jhydrol.2010.03.017
Edwards A, Anesio AM, Rassner SM et al (2011) Possible interactions between bacterial diversity, microbial activity and supraglacial hydrology of cryoconite holes in Svalbard. ISME J 5:150–160. https://doi.org/10.1038/ismej.2010.100
Edwards A, Rassner SME, Anesio AM, et al (2013) Contrasts between the cryoconite and ice-marginal bacterial communities of Svalbard glaciers. Polar Res 32:19468. https://doi.org/10.3402/polar.v32i0.19468
Epstein SS, Alexander D, Cosman K et al (1997) Enumeration of sandy sediment bacteria: Are the counts quantitative or relative? Mar Ecol Prog Ser 151:11–16. https://doi.org/10.3354/meps151011
Fang XM, Zhang T, Li J et al (2019) Bacterial community pattern along the sediment seafloor of the Arctic fjorden (Kongsfjorden, Svalbard). Antonie Van Leeuwenhoek Int J Gen Mol Microbiol 112:1121–1136. https://doi.org/10.1007/s10482-019-01245-z
Feltracco M, Barbaro E, Tedeschi S et al (2020) Interannual variability of sugars in Arctic aerosol: biomass burning and biogenic inputs. Sci Total Environ. https://doi.org/10.1016/j.scitotenv.2019.136089
Freese HM, Eggert A, Garland JL, Schumann R (2010) Substrate utilization profiles of bacterial strains in plankton from the River Warnow, a humic and eutrophic river in North Germany. Microb Ecol 59:59–75. https://doi.org/10.1007/s00248-009-9608-7
Garcia-Lopez E, Cid C (2017) The role of microbial ecology in glacier retreat. In: Glaciers evolution in a changing world. InTech
Garland JL, Mills AL (1991) Classification and characterization of heterotrophic microbial communities on the basis of patterns of community-level sole-carbon-source utilization. Appl Environ Microbiol 57:2351–2359. https://doi.org/10.1128/aem.57.8.2351-2359.1991
Gomez E, Ferreras L, Toresani S (2006) Soil bacterial functional diversity as influenced by organic amendment application. Bioresour Technol 97:1484–1489. https://doi.org/10.1016/j.biortech.2005.06.021
Grotti M, Soggia F, Ardini F et al (2017) Trace elements in surface sediments from Kongsfjorden, Svalbard: occurrence, sources and bioavailability. Int J Environ Anal Chem 97:401–418. https://doi.org/10.1080/03067319.2017.1317762
Hamdan LJ, Coffin RB, Sikaroodi M et al (2013) Ocean currents shape the microbiome of Arctic marine sediments. ISME J 7:685–696. https://doi.org/10.1038/ismej.2012.143
Hop H, Pearson T, Hegseth EN et al (2002) The marine ecosystem of Kongsfjorden, Svalbard. Polar Res 21:167–208. https://doi.org/10.1111/j.1751-8369.2002.tb00073.x
Jain A, Krishnan KP (2017) A glimpse of the diversity of complex polysaccharide-degrading culturable bacteria from Kongsfjorden, Arctic Ocean. Ann Microbiol 67:203–214
Jia L, Feng X, Zheng Z et al (2015) Polymorphobacter Fuscus sp. nov., isolated from permafrost soil, and emended description of the genus polymorphobacter. Int J Syst Evol Microbiol 65:3920–3925. https://doi.org/10.1099/ijsem.0.000514
Jones SE, Lennon JT (2010) Dormancy contributes to the maintenance of microbial diversity. Proc Natl Acad Sci U S A 107:5881–5886. https://doi.org/10.1073/pnas.0912765107
Kim M, Jung JY, Laffly D et al (2017) Shifts in bacterial community structure during succession in a glacier foreland of the high Arctic. FEMS Microbiol Ecol 93:1–9. https://doi.org/10.1093/femsec/fiw213
Kosek K, Luczkiewicz A, Kozioł K et al (2019) Environmental characteristics of a tundra river system in Svalbard. Part 1: bacterial abundance, community structure and nutrient levels. Sci Total Environ 653:1571–1584. https://doi.org/10.1016/j.scitotenv.2018.11.378
Koziorowska K, Kuliński K, Pempkowiak J (2017) Distribution and origin of inorganic and organic carbon in the sediments of Kongsfjorden, Northwest Spitsbergen, European Arctic. Cont Shelf Res 150:27–35. https://doi.org/10.1016/j.csr.2017.08.023
Kritzberg ES, Duarte CM, Wassmann P (2010) Changes in Arctic marine bacterial carbon metabolism in response to increasing temperature. Polar Biol 33:1673–1682. https://doi.org/10.1007/s00300-010-0799-7
Kumar V, Tiwari M, Nagoji S, Tripathi S (2016) Evidence of anomalously low δ13C of marine organic matter in an Arctic Fjord. Sci Rep 6:1–9. https://doi.org/10.1038/srep36192
Lin X, Zhang L, Liu Y, Li Y (2017) Bacterial and archaeal community structure of pan-Arctic Ocean sediments revealed by pyrosequencing. Acta Oceanol Sin 36:146–152. https://doi.org/10.1007/s13131-017-1030-2
Liu Q, Liu HC, Zhou YG, Xin YH (2019) Microevolution and adaptive strategy of Psychrophilic species flavobacterium bomense sp. Nov Isol Glaciers Front Microbiol. https://doi.org/10.3389/fmicb.2019.01069
Liu Y, Shen L, Zeng Y et al (2020) Genomic insights of Cryobacterium isolated from ice core reveal genome dynamics for adaptation in Glacier. Front Microbiol 11:1–15. https://doi.org/10.3389/fmicb.2020.01530
Łokas E, Zaborska A, Kolicka M et al (2016) Accumulation of atmospheric radionuclides and heavy metals in cryoconite holes on an Arctic Glacier. Chemosphere 160:162–172. https://doi.org/10.1016/j.chemosphere.2016.06.051
Lu Z, Cai M, Wang J et al (2013) Levels and distribution of trace metals in surface sediments from Kongsfjorden, Svalbard, Norwegian Arctic. Environ Geochem Health 35:257–269. https://doi.org/10.1007/s10653-012-9481-z
Lutz S, Anesio AM, Edwards A, Benning LG (2017) Linking microbial diversity and functionality of arctic glacial surface habitats. Environ Microbiol 19:551–565
Malard LA, Anwar MZ, Jacobsen CS, Pearce DA (2019) Biogeographical patterns in soil bacterial communities across the Arctic region. FEMS Microbiol Ecol 95:1–28. https://doi.org/10.1093/femsec/fiz128
Margesin R, Spröer C, Zhang DC, Busse HJ (2012) Polaromonas glacialis sp. nov. and Polaromonas cryoconiti sp. nov., isolated from alpine glacier cryoconite. Int J Syst Evol Microbiol 62:2662–2668. https://doi.org/10.1099/ijs.0.037556-0
Mueller DR, Vincent WF, Bonilla S, Laurion I (2005) Extremotrophs, extremophiles and broadband pigmentation strategies in a high arctic ice shelf ecosystem. FEMS Microbiol Ecol 53:73–87. https://doi.org/10.1016/j.femsec.2004.11.001
Nash MV, Anesio AM, Barker G et al (2018) Metagenomic insights into diazotrophic communities across Arctic glacier forefields. FEMS Microbiol Ecol 94:1–12. https://doi.org/10.1093/femsec/fiy114
Nuth C, Kohler J, König M et al (2013) Decadal changes from a multi-temporal glacier inventory of Svalbard. Cryosphere 7:1603–1621. https://doi.org/10.5194/tc-7-1603-2013
Overland J, Dunlea E, Box JE et al (2019) The urgency of Arctic change. Polar Sci 21:6–13. https://doi.org/10.1016/j.polar.2018.11.008
Pantanella F, Berlutti F, Passariello C et al (2007) Violacein and biofilm production in Janthinobacterium lividum. J Appl Microbiol 102:992–999. https://doi.org/10.1111/j.1365-2672.2006.03155.x
Pfennig N (1974) Rhodopseudomonas globiformis, sp. n., a new species of the Rhodospirillaceae. Arch Microbiol 100:197–206. https://doi.org/10.1007/BF00446317
Phurbu D, Liu ZX, Liu HC et al (2020) Polymorphobacter arshaanensis sp. Nov., containing the photosynthetic gene pufml, isolated from a volcanic lake. Int J Syst Evol Microbiol 70:1093–1098. https://doi.org/10.1099/ijsem.0.003880
Poniecka EA, Bagshaw EA, Sass H et al (2020) Physiological capabilities of cryoconite hole microorganisms. Front Microbiol 11:1783. https://doi.org/10.3389/fmicb.2020.01783
Prasad S, Manasa P, Buddhi S et al (2014) Diversity and bioprospective potential (cold-active enzymes) of cultivable marine bacteria from the subarctic glacial fjord, Kongsfjorden. Curr Microbiol 68:233–238. https://doi.org/10.1007/s00284-013-0467-6
Quast C, Pruesse E, Yilmaz P et al (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:D590–D596. https://doi.org/10.1093/nar/gks1219
Reddy PVV, Rao SSSN, Pratibha MS et al (2009) Bacterial diversity and bioprospecting for cold-active enzymes from culturable bacteria associated with sediment from a melt water stream of Midtre Lov’enbreen glacier, an Arctic glacier. Res Microbiol 160:538–546
Reitermayer D, Kafka TA, Lenz CA, Vogel RF (2018) Interrelation between tween and the membrane properties and high pressure tolerance of Lactobacillus plantarum. BMC Microbiol 18:1–14. https://doi.org/10.1186/s12866-018-1203-y
Sala MM, Terrado R, Lovejoy C et al (2008) Metabolic diversity of heterotrophic bacterioplankton over winter and spring in the coastal Arctic Ocean. Environ Microbiol 10:942–949. https://doi.org/10.1111/j.1462-2920.2007.01513.x
Sanders ER (2012) Aseptic laboratory techniques: plating methods. J vis Exp. https://doi.org/10.3791/3064
Sinha RK, Krishnan KP, Hatha AAM et al (2017) Diversity of retrievable heterotrophic bacteria in Kongsfjorden, an Arctic fjord. Braz J Microbiol 48:51–61. https://doi.org/10.1016/j.bjm.2016.09.011
Sinha RK, Krishnan KP, Thomas FA et al (2019) Polyphasic approach revealed complex bacterial community structure and function in deep sea sediment of ultra-slow spreading Southwest Indian Ridge. Ecol Indic. https://doi.org/10.1016/j.ecolind.2018.08.063
Skidmore M, Anderson SP, Sharp M et al (2005) Comparison of microbial community compositions of two subglacial environments reveals a possible role for microbes in chemical weathering processes. Appl Environ Microbiol 71:6986–6997
Stibal M, Hasan F, Wadham JL et al (2012) Prokaryotic diversity in sediments beneath two polar glaciers with contrasting organic carbon substrates. Extremophiles 16:255–265. https://doi.org/10.1007/s00792-011-0426-8
Struvay C, Feller G (2012) Optimization to low temperature activity in psychrophilic enzymes. Int J Mol Sci 13:11643–11665. https://doi.org/10.3390/ijms130911643
Sułowicz S, Bondarczuk K, Ignatiuk D et al (2020) Microbial communities from subglacial water of naled ice bodies in the forefield of Werenskioldbreen. Svalbard Sci Total Environ. https://doi.org/10.1016/j.scitotenv.2020.138025
Takahashi S, Tomita J, Nishioka K et al (2014) Development of a prokaryotic universal primer for simultaneous analysis of bacteria and archaea using next-generation sequencing. PLoS One 9:e105592
Teske A, Durbin A, Ziervogel K et al (2011) Microbial community composition and function in permanently cold seawater and sediments from an Arctic fjord of Svalbard. Appl Environ Microbiol 77:2008–2018. https://doi.org/10.1128/AEM.01507-10
Thomas FA, Sinha RK, Krishnan KP (2020) Bacterial community structure of a glacio-marine system in the Arctic (Ny-Ålesund, Svalbard). Sci Total Environ. https://doi.org/10.1016/j.scitotenv.2019.135264
Wang NF, Zhang T, Yang X et al (2016) Diversity and composition of bacterial community in soils and lake sediments from an arctic lake area. Front Microbiol 7:1–9. https://doi.org/10.3389/fmicb.2016.01170
Winkelmann D, Knies J (2005) Recent distribution and accumulation of organic carbon on the continental margin west off Spitsbergen. Geochem Geophys Geosys. https://doi.org/10.1029/2005GC000916
Yang A, Yen C (2012) PCR Optimization of BOX-A1R PCR for microbial source tracking of Escherichia coli in waterways. J Exp Microbiol Immunol 16:85–89
Yoon S-H, Ha S-M, Kwon S et al (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol 67:1613
Zeng Y-X, Zhang F, He J-F et al (2013) Bacterioplankton community structure in the Arctic waters as revealed by pyrosequencing of 16S rRNA genes. Antonie Van Leeuwenhoek 103:1309–1319
Zeng YX, Yu Y, Li HR, Luo W (2017) Prokaryotic community composition in arctic Kongsfjorden and sub-arctic northern Bering sea sediments as revealed by 454 pyrosequencing. Front Microbiol. https://doi.org/10.3389/fmicb.2017.02498
Zhang B, Wu X, Zhang W et al (2016) Diversity and succession of Actinobacteria in the Forelands of the Tianshan Glacier, China. Geomicrobiol J 33:716–723. https://doi.org/10.1080/01490451.2015.1085468
Zhang DC, Wang HX, Cui HL et al (2007) Cryobacterium psychrotolerans sp. nov., a novel psychrotolerant bacterium isolated from the China No. 1 glacier. Int J Syst Evol Microbiol 57:866–869. https://doi.org/10.1099/ijs.0.64750-0
Acknowledgements
The authors wish to express their gratitude to the Director, National Centre for Polar and Ocean Research, Ministry of Earth Sciences, India for his support and interest in this work. Thanks are due to colleagues at the Cryobiology laboratory at NCPOR, School of Environmental Sciences at M.G.University, and Kerala University of Fisheries and Ocean Studies for their support. This is NCPOR contribution number- J-13/2021-22.
Funding
Ministry of Earth Sciences, India.
Author information
Authors and Affiliations
Contributions
FAT and KPK designed the research plan and methodology. The formal analysis was carried out by FAT and MM. Bioinformatics analysis and statistical analysis of data were done by FAT. FAT wrote the manuscript under the supervision of KPK. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Thomas, F.A., Mohan, M. & Krishnan, K.P. Bacterial diversity and their metabolic profiles in the sedimentary environments of Ny-Ålesund, Arctic. Antonie van Leeuwenhoek 114, 1339–1360 (2021). https://doi.org/10.1007/s10482-021-01604-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-021-01604-9