Abstract
Marine sponges are abundant and ecologically important components of coral reefs and have been shown to harbour exceptionally high microbial densities, which can differ substantially among sponge species. However, this dichotomy between high and low microbial abundance (HMA, LMA) sponges is still not fully understood, particularly as concerns the archaeal community. This study aims to fill this gap by analysing (using 454-pyrosequencing of the 16S rRNA gene) how the archaeal community varies among known LMA (Stylissa carteri, and Stylissa massa), known HMA (Hyrtios erectus and Xestospongia testudinaria) and unknown HMA/LMA status sponge species (Ectyoplasia coccinea, Paratetilla bacca and Petrosia aff. spheroida) collected in a remote location in which very few sponge microbial composition studies have been previously performed (Mayotte, Comores archipelago, France) and comparing the results with those reported in four other geographical areas. Based on archaeal community composition, the known LMA sponges formed a distinct cluster together with Paratetilla bacca, Ectyoplasia coccinea and seawater while the known HMA sponge X. testudinaria formed a cluster with Petrosia aff. spheroida. The known HMA sponge H. erectus, in turn, had an intermediate archaeal community between HMA sponges and sediment samples. In addition to the above, we also showed significant compositional congruence between archaeal and bacterial communities sampled from the same sponge individuals. HMA sponges were mainly dominated by members assigned to the genus Nitrosopumilus while LMA sponges were mainly dominated by members assigned to the genus Cenarchaeum. In general, there was no clear difference in richness between HMA and LMA sponges. Evenness, however, was higher in HMA than LMA sponges. Whilst the present study corroborates some of the traits commonly associated with the HMA–LMA dichotomy (higher evenness in Mayotte HMA sponges), this was not consistent across geographical areas showing that more research is needed to fully understand the HMA/LMA dichotomy as concerns Archaea.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Marine sponges are sessile, filter-feeding organisms, which harbour exceptionally high microbial densities in their mesohyl (Diaz and Rützler 2001; Moitinho-Silva et al. 2014a, b) with a wide variety of functions. For example, sponge microbial communities are involved in the biosynthesis of secondary metabolites, antibiotics, cofactors and vitamins, in nutrient transport and utilization (ammonium oxidation, carbon fixation, denitrification and nitrification), in redox sensing and response, and in the degradation of benzoic compounds (Thomas et al. 2010; Fan et al. 2012; Hentschel et al. 2012). This functional repertoire influences sponge health and the surrounding environment; by filtering large volumes of seawater, sponges, furthermore, contribute to water column composition (Bell 2008; Hoffmann et al. 2009; McMurray et al. 2014) and thus play important ecological roles in the ecosystem (e.g., Yahel et al. 2003; Hoffmann et al. 2009; de Goeij et al. 2013). Sponges are among the most studied of invertebrate microbial hosts and their microbial communities have been described as distinct when compared to other non-sponge hosts or non-host biotopes such as sediment or water (Hentschel et al. 2002). Sponges have also been grouped according to the density and diversity of their prokaryotic symbionts (Kamke et al. 2010). High microbial abundance (HMA) sponges have microbial concentrations varying from 108 to 1010 bacteria/g of tissue while low microbial abundance sponges (LMA) have microbial concentrations close to that of seawater, i.e., varying from 105–106 bacteria/g of tissue (Hentschel et al. 2006). The fact that HMA sponges have lower pumping rates than LMA suggests that they rely more on their higher microbial densities to acquire energy than their LMA counterparts (Weisz et al. 2007). This dichotomy however goes beyond microbial abundances and seems to extend to morphology, physiology, microbial diversity, microbial composition and microbial specificity (Erwin et al. 2015 and references therein). LMA sponges tend to host low diversity microbial communities, which are compositionally similar to those found in seawater (Moitinho-Silva et al. 2014b; Thacker and Freeman 2012; Bayer et al. 2014a, b; Blanquer et al. 2013; Polónia et al. 2013, 2015). Additionally, HMA and LMA sponges also seem to be dominated by distinct microbial members. These indicator taxa include the phyla Chloroflexi, Acidobacteria, Actinobacteria, PAUC34f, Gemmatimonadetes, SBR1093, Poribacteria, AncK6, Nitrospirae and Spirochaetae for HMA sponges and Proteobacteria, Bacteroidetes, Planctomycetes and Firmicutes for LMA sponges (Schmitt et al. 2011; Bayer et al. 2014a, b; Moitinho-Silva et al. 2017). The known LMA and HMA indicators, however, all belong to the bacterial domain (Bayer et al. 2014a, b; Moitinho-Silva et al. 2017) and very few studies have specifically analysed this dichotomy in terms of the archaeal community (Bayer et al. 2014b; Chaib De Mares et al. 2017; Turon and Uriz 2020).
The archaeal lineages reported in sponges (Euryarchaeota, Thaumarchaeota and Parvarchaeota; Preston et al. 1996; Polónia et al. 2013, 2015; Jackson et al. 2013; Kennedy et al. 2014; Thomas et al. 2016) can, in some sponge species (e.g., Dragmacidon mexicanum and Inflatella pellicula), dominate the microbial community (Jackson et al. 2013; Preston et al. 1996). Although this denotes an important role for archaeal members in these hosts, the exact functions of these microbial members in sponges are still poorly understood. Thaumarchaeota members, more specifically members assigned to the genus Candidatus Nitrosopumilus and Cenarchaeum, have been frequently associated with sponges and are able to convert ammonium to nitrite and fix CO2 through ammonia oxidation. Archaeal community members may thus be important drivers of nitrification in these hosts (Hallam et al. 2006a, b; Bayer et al. 2008; Radax et al. 2012) and consequently of the sponge nitrogen metabolism and nitrogen cycle (Zhang et al. 2014).
Here, using 454-pyrosequencing of the 16S rRNA gene, the archaeal communities detected in nine distinct biotopes, namely, sediment, seawater and seven different sponge species were assessed and a comparison between the LMA and HMA sponges among geographical areas was performed with the aim of ascertaining to what extent the HMA/LMA dichotomy can be extended to Archaea. The studied sponge species included two known LMA sponges: Stylissa carteri and Stylissa massa, two known HMA sponges: Hyrtios erectus and Xestospongia testudinaria and the species Ectyoplasia coccinea, Paratetilla bacca, and Petrosia aff. spheroida with unknown HMA/LMA status. In a previous study, de Voogd et al. (2019) found that the bacterial communities of E. coccinea and P. bacca resembled those of LMA sponges while the bacterial community of Petrosia aff. spheroida resembled that of HMA sponges. The main objectives of the present study were to (1) compare archaeal richness, evenness and composition among biotopes (sponge host species, seawater and sediment), (2) assess if there are significant differences in the relative abundances of selected higher taxa among biotopes, (3) identify abundant OTUs that significantly discriminate between pairs of biotopes, (4) test for concordance in composition between archaeal and bacterial communities in the studied biotopes, (5) assign HMA or LMA status to sponge species and (6) compare the compositional characteristics of HMA and LMA sponge species in Mayotte with similar species sampled in other areas.
Materials and methods
Study area
Mayotte is part of France and of the Comores archipelago (Indian Ocean), which is considered a biodiversity hotspot (Agnarsson and Kuntner 2012; Said Hassane et al. 2020). The coastal areas of Mayotte are considered a priority region in the framework of National biodiversity strategy by the French government. Increasing human population numbers, however, pose a threat to the marine ecosystems of the region (Kiszka et al. 2007). The archipelago is located to the northwest of Madagascar in the Mozambique channel (Supplementary Fig. 1) and consists of two main islands of volcanic origin (Grande Terre and Petite Terre) and some other smaller islands. Surrounding the main island, there is an almost continuous barrier reef forming a lagoon, with an area of 1500 km2 and 3–15 km wide making it one of the world's largest lagoons.
Sampling
Using SCUBA diving and snorkeling, two to three replicates of 7 sponge species, seawater and sediment were collected at different sites inside and outside the lagoon at the western side of Grande Terre (between 12° 56.470′ S 45° 04.305′ E and 13° 00.375′ S 45° 08.250′ E) and at depths ranging from 3 to 25 m from May the 4th to the 11th, 2013 (Supplementary Table 1). Sponge species were identified using classical morphological characters by NJ de Voogd; voucher specimens have been deposited in the sponge collection of Naturalis Biodiversity Center. Among the studied species, two have been previously identified as, or have been shown to, house bacterial communities very similar to LMA sponges [Stylissa carteri (2 samples), and Stylissa massa (3 samples); family Scopalinidae, order Scopalinida]. Two other species have been designated as HMA sponges [Hyrtios erectus (3 samples, family Thorectidae, order Dictyoceratida) and Xestospongia testudinaria (3 samples, family Petrosiidae, order Haplosclerida)] (Moitinho‐Silva et al. 2014b, 2017; Gloeckner et al. 2014; Cleary et al. 2015, 2018; Polónia et al. 2018). Three sponge species had unknown HMA/LMA status [Ectyoplasia coccinea (2 samples, family Raspailiidae, order Axinellida), Paratetilla bacca (2 samples, family Tetillidae, order Tetractinellida), and Petrosia aff. spheroida (3 samples, family Petrosiidae order Haplosclerida)]. A more in depth description of these species can be found in de Voogd et al. (2019).
Small sponge fragments (enough for identification and DNA extraction) were sampled including segments of surface and interior in order to sample, as much as possible, the whole archaeal community. The sediment samples (3 replicates) were taken using the mini-core method (Polónia et al. 2016) while water samples (3 replicates) were collected with a 1.5 L bottle from which 1 L of water was filtered through a Millipore® White Isopore Membrane Filter (0.22 µm pore size) to obtain the archaeoplankton. All samples were stored in absolute alcohol and kept at temperatures lower than 4 ºC immediately after collection. Once in the laboratory, samples were stored at − 80 ºC until DNA extraction.
DNA extraction and pyrosequencing
We isolated PCR-ready total community DNA (TC-DNA) from sediment, seawater and sponge samples using the FastDNA® SPIN Kit for soil (MP Biomedicals) following the manufacturer's instructions. Briefly, we prepared sediment samples by centrifuging each one for 30 min at 4400 rpm and 4 ºC and the membrane filter (seawater sample) and sponge samples by cutting them into small pieces. The whole membrane filter and 500 mg of sediment or sponge were transferred to Lysing Matrix E tubes containing a mixture of ceramic and silica particles. The microbial cell lysis was performed in the FastPrep® Instrument (Q Biogene) for 80 s at a speed of 6.0 m s−1. Extracted DNA was eluted into DNase/Pyrogen-Free Water to a final volume of 50 μl and stored at − 20 °C until use. Prior to 454 pyrosequencing, the amplicons of the archaeal 16S rRNA gene were obtained using the Archaea specific primers ARC344f-mod and Arch958R-mod (Pires et al. 2012). After a denaturation step at 94 °C during 5 min, 35 thermal cycles of 1 min at 94 °C, 1 min at 56 °C and 1 min at 72 °C were carried out followed by an extension step at 72 °C for 7 min (first amplification; Pires et al. 2012). Using the amplicons of the archaeal 16S rRNA gene as template, the V3-V4 regions were amplified using barcoded fusion primers, 524F-10-ext and Arch958R-mod (second amplification; Pires et al. 2012). After 4 min denaturation at 94 °C, 30 cycles of 94 °C for 30 s, 50 °C for 45 s and 68 °C for 60 s and a final extension at 68 °C for 10 min were carried out.
Following previous studies (Cleary et al. 2015; de Voogd et al. 2015), barcoded pyrosequencing libraries were analysed using the QIIME (Quantitative Insights Into Microbial Ecology; Caporaso et al. 2010) software package. In QIIME, separate fasta and qual files were used as input for the split_libraries.py script. Default arguments were used except for the minimum sequence length, which was set at 218 bps after removal of forward primers and barcodes; backward primers were removed using the 'truncate only' argument and a sliding window test of quality scores was enabled with a value of 50 as suggested in the QIIME description for the script. In addition to user-defined cut-offs, the split_libraries.py script performs several quality filtering steps (http://qiime.org/scripts/split_libraries.html). OTUs were selected using UPARSE with usearch7 using a sequence similarity threshold of 97% (Edgar 2013). The UPARSE sequence analysis tool (Edgar 2013) provides clustering, chimera checking and quality filtering on de-multiplexed sequences. Chimera checking was performed using the UCHIME algorithm (Edgar et al. 2011). The quality filtering, as implemented in usearch7, filters noisy reads and preliminary results suggest it gives results comparable to other denoisers such as AmpliconNoise but is much less computationally expensive (http://drive5.com/usearch/features.html; last checked 2014-01-20). First, reads were filtered with the -fastq_filter command and the following arguments -fastq_trunclen 250 -fastq_maxee 0.5 -fastq_truncqual 15. Sequences were then dereplicated and sorted using the -derep_fulllength and -sortbysize commands. OTU clustering was performed using the -cluster_otus command. Representative sequences were selected using the pick_rep_set.py script in QIIME using the 'most_abundant' method. Taxonomy was assigned to reference sequences of OTUs using default arguments in the assign_taxonomy.py script in QIIME with the SILVA_132_QIIME_release database and the uclust classifier method (Quast et al. 2013). The make_otu_table.py script in QIIME was used to generate a square matrix of OTUs × SAMPLES and subsequently rarefied to 3485 sequences per sample with the single_rarefaction.py script in QIIME. This was used as input for further analyses using the R package (R Core Team 2013). Sequence identifiers of closely related taxa of numerically abundant OTUs (≥ 130 sequence reads) were downloaded using the NCBI Basic Local Alignment Search Tool (BLAST) command line 'blastn' tool with the -db argument set to nt (Zhang et al. 2000). BLAST identifies locally similar regions between sequences, compares sequences to extant databases and assesses the significance of matches; functional and evolutionary relationships can subsequently be inferred. Each run produces a list of hits based on significant similarity between pairs of sequences, i.e., the target sequence and taxa present in the database (or no hits if no significantly similar sequences are found). A discussion of how significance is determined can be found at http://www.ncbi.nlm.nih.gov/BLAST/tutorial/Altschul-1.html.
For the OTU table, OTUs not classified as Archaea (e.g., bacteria or unclassified at the level of domain) were removed prior to statistical analysis. For the statistical analyses, only biotopes/species with three replicates were included. This table was then loge (x + 1) transformed and distance matrices constructed using the Bray–Curtis index with the vegdist() function in the vegan package (Oksanen et al. 2009) in R. Principal Coordinates Analysis (PCO) was performed using the cmdscale() function in R with the Bray–Curtis distance matrix as input to assess the variation in OTU composition among biotopes. We tested for significant variation in composition among all biotopes (sponge species, sediment and water) using the adonis() function in vegan. In the adonis analysis, the Bray–Curtis distance matrix of species composition was the response variable with biotope as independent variable. The number of permutations was set at 999; all other arguments used the default values set in the function. Weighted averages scores were computed for OTUs on the first two PCO axes using the wascores() function in the vegan package.
We tested for significant differences in the relative abundance of the most abundant phyla among biotopes and among sponge species with an analysis of deviance using the glm() function in R. In the GLM analysis, we set the family argument to ‘tweedie’ (Tweedie 1984) with var.power = 1.5 and link.power = 0 (a compound Poisson– gamma distribution). Using the GLM models, we tested for significant variation among biotopes and among sponge species using the anova() function in R with the F test. We subsequently used the emmeans() function in the emmeans library (Lenth 2017) to perform multiple comparisons of mean abundance among biotopes using the false discovery rate (fdr) method in the adjust argument. The simper() function in vegan was used to identify significantly discriminating OTUs between pairs of biotopes based on a square root transformed OTU table and 999 permutations. The discriminating OTUs contribute the most to differences between pairs of biotopes. In addition to the above, we also identified dominant and possible core OTUs of each biotope and their relative abundance in those biotopes and calculated rarefied species richness (Gotelli and Colwell 2001), Shannon's (H′) diversity index (Shannon and Weaver 1949) and evenness using the diversity() and specnumber() functions of the Vegan package (Oksanen et al. 2009).
We used results of the present study, namely, relative richness and evenness, the number of core OTUs, the mean abundance of these core OTUs, the mean abundance of the single most abundant OTU and the mean abundances of the main Phyla (Thaumarchaeota, Euryarchaeota) of LMA (S. massa, S. carteri) and HMA (X. testudinaria, H. erectus) species from Mayotte and compared these with the same species sampled from other areas (Spermonde Archipelago, Kepulauan Seribu, Berau and Misool coral reef systems, Indonesia; hereafter Makassar, Jakarta, Berau and Papua respectively; Polónia et al. 2014, 2015, 2016, 2017, 2018). We also performed this comparison using sediment and seawater. Note, however, that core status in the present study was based on very small sample sizes and more individuals of a species should be sampled before reliably determining whether or not they are core members. Closely related organisms (identified using BLAST) of the most abundant OTUs of all these studies were also compared along with data from Polónia and Cleary (2019). All these studies used the same sampling, preservation, extraction, amplification and pyrosequencing methodology. In the data analysis the only difference resided in the use of the SILVA_132_QIIME_release database for taxonomy assignment and the rarefaction of the square matrix of OTUs x SAMPLES in the present study compared with the previous studies that used the Greengenes 13_5 release and no rarefaction.
Finally, we used Procustes analysis with the procrustes function in the vegan package in R to test for significant congruence between archaeal and bacterial communities inhabiting sponges. Variation in bacterial composition from the same sponge specimens was previously published by de Voogd et al. (2019) and this data was used in the present study. Procrustes analysis entails scaling, which adjusts one configuration 'Y' to maximum similarity with another configuration 'X'. The scaling is non-symmetric given that Y is scaled to fit X. In addition to the procrustes() function, the protest() function in vegan was used to estimate the significance of the Procrustes statistic. The latter function uses a statistic (r = sqrt(1-ss)) derived from the symmetric Procrustes sum of squares 'ss' and calls the procrustes() function a given number of times (1000 permutations in the present case). Detailed descriptions of the functions used here can be found in R and online in reference manuals (http://cran.r-project.org/web/packages/vegan/index.html; Accessed 26-9-2011).
Results
Sequencing yielded 83,601 sequences, binned to 455 archaeal OTUs, which were assigned to 9 phyla, 18 classes and 21 orders after rarefaction, quality control, OTU picking (OTU cut-off 97%), chimera removal and removal of OTUs not assigned to Archaea.
Higher taxon abundance
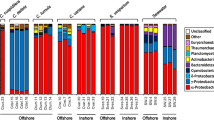
The relative abundance of all higher archaeal taxa differed significantly among biotopes (Fig. 1; Supplementary Fig. 2). All the sponge species (≥ 94.5%) and sediment (90.42% ± 8.96) samples were dominated by members of the Thaumarchaeota phylum (Fig. 1) with no significant difference among sponge species (Fig. 2; Supplementary Table 2). In contrast, the relative abundance of Euryarchaeota was significantly higher in seawater (99.57% ± 0.57) than all other biotopes (Figs. 1, 3 and Supplementary Table 2). In addition to Thaumarchaeota and Euryarchaeota, sediment samples also included members assigned to the Nanoarchaeaeota, Crenarchaeota, Asgardaeota, Hydrothermarchaeota, and Diapherotrites. Hydrothermarchaeota and Diapherotrites were only found in sediment. The relative abundance of Nanoarchaeaeota was, however, highest in the sponge Petrosia aff. spheroida but this was only significantly higher than H. erectus (Fig. 1; Supplementary Table 2).
Mean relative abundance of the most abundant archaeal phyla, classes, orders, evenness and richness for Paratetilla bacca (Pb), Stylissa carteri (Sc), Stylissa massa (Sm), Ectyoplasia coccinea (Ec), Hyrtios erectus (He), Petrosia aff. spheroida (Ps), Xestospongia testudinaria (Xt), sediment (Sd), and seawater (Wt). Error bars represent a single standard deviation. Results of the GLM analyses are shown in the top‐right corner of each subfigure
Mean relative abundance of the most abundant archaeal phyla, classes, orders, evenness and richness for Paratetilla bacca (Pb), Stylissa carteri (Sc), Stylissa massa (Sm), Ectyoplasia coccinea (Ec), Hyrtios erectus (He) and Petrosia aff. spheroida (Ps), Xestospongia testudinaria (Xt). Error bars represent a single standard deviation. Results of the GLM analyses are shown in the top‐right corner of each subfigure
Relative abundance of the most abundant OTUs and, in the online version, colour-coded according to archaeal classes for the demosponges: Paratetilla bacca (Pb), Stylissa carteri (Sc), Stylissa massa (Sm), Ectyoplasia coccinea (Ec), Hyrtios erectus (He), Petrosia aff. spheroida (Ps), Xestospongia testudinaria (Xt) and the environmental biotopes: sediment (Sd), and seawater (Wt). The size of the symbol is proportional to the percentage of sequences represented by a given OTU. The y-axis shows the OTU-id numbers. The y-axis numbers have been colour coded red for the Cenarchaeum genus OTUs and Green for Candidatus Nitrosopumilus genus OTUs. The x-axis biotope identifications have been colour coded orange for known LMA sponges; blue for known HMA sponges and purple for sponge species of unknown LMA-HMA status
Among sponges, E. coccinea (5.48 ± 2.76%), P. bacca (1.84 ± 2.31%) and the known LMA S. massa (1.94 ± 1.19%) and S. carteri (1.41 ± 0.77%) had the highest relative abundances of Euryarchaeota (Fig. 2). The relative abundance of this phylum was also significantly different between the LMA S. massa and the HMA sponge species H. erectus and X. testudinaria and the sponge Petrosia aff. spheroida, but there was no significant difference among the HMA sponge species and Petrosia aff. spheroida (Supplementary Table 2).
Dominant and core OTUs
Only 2 OTUs were present in all sponge species (OTUs 1 and 766); both were assigned to the Nitrosopumilaceae family. The possible core community of each biotope, in turn, varied from 5 in X. testudinaria to 53 in sediment (Supplementary Table 3). Among sponges, the highest number of possible core OTUs (17) was observed in H. erectus and E. coccinea. The mean abundance of the single most abundant OTU varied between 88 and 98% for P. bacca, E. coccinea and the known LMA sponge species S. carteri and S. massa and between 56 and 66% for Petrosia aff. spheroida, and the known HMA sponge species H. erectus and X. testudinaria. In the environmental biotopes, these values varied from 12% in sediment to 23% in seawater (Supplementary Table 3). The cumulative mean abundance of the possible core OTUs made up 86.02% of sediment samples and 99.3% of seawater samples (Supplementary Table 3). In sponges, these values varied between 98.98 and 99.63% for P. bacca, E. coccinea, and the known LMA sponge species S. carteri and S. massa, and between 92.58 and 95.87% for Petrosia aff. spheroida, and the known HMA sponge species H. erectus and X. testudinaria.
Dominance was much less pronounced in seawater and sediment and both water and sediment housed distinct subsets of OTUs, which were rare in the sponge biotopes. The most abundant OTUs in sponges were also usually rare in sediment and seawater.
Paratetilla bacca and E. coccinea each housed a single, highly abundant OTU namely OTU-3 (assigned to the genus Cenarchaeum) and OTU-6 (assigned to the genus Candidatus Nitrosopumilus), respectively (Fig. 3). Both Stylissa species housed the same highly abundant OTUs namely OTU-1 and OTU-357 (both assigned to the genus Cenarchaeum) (Fig. 3). Among the remaining species, OTU dominance was less pronounced. OTU-2 (assigned to the family Nitrosopumilaceae) was the most abundant OTU of H erectus; OTU-4 (assigned to the genus Candidatus Nitrosopumilus) was the most abundant OTU of X. testudinaria and OTU-1009 (assigned to the genus Candidatus Nitrosopumilus) was the most abundant OTU of Petrosia aff. spheroida.
Some of these highly abundant OTUs significantly discriminated between pairs of biotopes. (Supplementary Table 4). For example, OTU-5 significantly discriminated X. testudinaria from seawater and the sponge species S. massa and H. erectus. OTU-4 significantly discriminated X. testudinaria and Petrosia aff. spheroida from seawater (in the case of X. testudinaria) and the sponge species S. massa and H. erectus. OTU-4 and OTU-5 were particularly abundant (albeit not host specific) in X. testudinaria and were closely related (100%, 97.6%) to organisms obtained from the known HMA species X. muta and Luffariella sp., respectively (Supplementary Table 5). OTU-1 and OTU-357 significantly discriminated the sponge species Petrosia aff. spheroida, X. testudinaria, H. erectus and seawater from the S. massa. These OTUs were related to an organism obtained from the sponge species Phakellia fusca (Supplementary Table 5). OTU-1 was highly abundant in both Stylissa species (15418 sequences) but was also found in all other sponge species albeit in very low numbers (15 sequences distributed among the other sponge samples). OTU-357 was also abundant in both Stylissa species (1659 sequences) with only two sequences found in two other sponge species (Petrosia aff. Spheroida and X. testudinaria). OTU-1009 significantly discriminated Petrosia aff. spheroida from S. massa, X. testudinaria and seawater. This OTU was particularly abundant in Petrosia aff. spheroida (5197 sequences) but was not restricted to this sponge species, with 597 sequences found in other sponge species (mainly in H. erectus and X. testudinaria). This OTU was also closely related (100%) to an organism obtained from the sponge species Theonella swinhoei (Supplementary Table 5). OTU-2 and OTU-7 were abundant in, but not restricted to H. erectus (with sequences also found in other sponge species and in environmental samples) and were closely related (> 99%) to organisms obtained from the sponge species Aplysina aerophoba and Cinachyra sp., respectively. OTU-3, which dominated the archaeal community of P. bacca (6834 sequences; with only 20 sequences recorded in other sponge species), was related (97.2%) to an organism obtained from tropical marine sediment collected in Karwar, Arabian Sea coast, India. OTU-6, the dominant OTU of E. coccinea (6497 sequences; with only 191 sequences recorded in other biotopes), was related (98.2%) to an organism previously obtained from sediment samples collected in the Oujiang river, Zhejiang province of China (Supplementary Table 5).
Diversity patterns
Richness was significantly higher in sediment than all other biotopes and significantly higher in H. erectus than S. massa, Petrosia aff. spheroida and X. testudinaria (Supplementary Table 2). Evenness was significantly higher in sediment and seawater than all other biotopes, but there was no significant difference between sediment and seawater. Evenness was also higher in Petrosia aff. spheroida and the known HMA sponge species H. erectus and X. testudinaria than all remaining sponge biotopes. Evenness was lower in P. bacca than all other biotopes (Supplementary Table 2).
Community composition
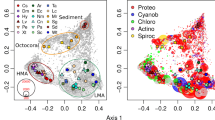
There was a highly significant difference in composition among biotopes (adonis: F5,17 = 15.46, p = 0.001, R2 = 0.870; Fig. 4). Variation among biotopes, thus, explained 87% of the variation in composition. The first PCO axis (Fig. 4), separated samples of seawater, S. massa, S. carteri, E. coccinea and P. bacca from samples of sediment, Petrosia aff. spheroida and the known HMA sponge species X. testudinaria and H. erectus. The second PCO axis (Fig. 4) separated sediment samples from Petrosia aff. spheroida and X. testudinaria samples (cluster 2) with samples of H. erectus in-between. The patterns observed in the relative abundances (Fig. 1) and in the clusters seen in Fig. 4 are also evident in the bubbleplot of the most abundant OTUs (≥ 130 sequences; Fig. 3) with the sponges S. massa, S. carteri and P. bacca (cluster 1 of Fig. 4) having higher percentages of OTUs assigned to the genus Cenarchaeum and to the Class Thermoplasmata (together with E. coccinea) when compared to the sponge species Petrosia aff. spheroida and the known HMA sponge species X. testudinaria and H. erectus. The most abundant seawater OTUs were more abundant in the sponge species P. bacca, E. coccinea and the known LMA sponge species S. massa and S. carteri than in Petrosia aff. spheroida and the known HMA sponge species H. erectus and X. testudinaria (Fig. 3).
a Ordination showing the first two axes of the PCO analysis. Light grey symbols represent operational taxonomic unit (OTU) scores with the symbol size representing their abundance (number of sequence reads). b Procrustes analysis comparing archaeal and bacterial OTU composition (arrowbase indicates the corresponding positions of the samples in the bacterial ordination (de Voogd et al. 2019) while arrowheads indicate the corresponding positions of the samples in the archaeal ordination). Symbols represent samples (colour-coded in the online version) of Paratetilla bacca (Pb), Stylissa carteri (Sc), Stylissa massa (Sm), Ectyoplasia coccinea (Ec), Hyrtios erectus (He), Petrosia aff. spheroida (Ps), Xestospongia testudinaria (Xt), sediment (Sd), and seawater (Wt)
The Procrustes analysis revealed significant congruence between archaeal and bacterial communities (R = 0.831, P < 0.001; Fig. 4b). Hyrtios erectus samples had the highest residual distances (0.33) when compared to all the other samples and thus, lowest congruence between their archaeal and bacterial communities.
The main results concerning to sponge biotopes and comparison between HMA–LMA status are summarized in Fig. 5. Note, however that, the HMA–LMA comparison is between two closely related LMA sponge species (S massa, S. carteri) and two distantly related HMA sponge species (X. testudinaria, H. erectus).
Mean relative abundance of the Dominant OTU (a) and the Euryarchaeota phylum (b), diversity (c), richness (d) and evenness (e) indices and ordination showing the first two axes of the PCO analysis (f) for Paratetilla bacca (Pb), Stylissa carteri (Sc), Stylissa massa (Sm), Ectyoplasia coccinea (Ec), Hyrtios erectus (He) and Petrosia aff. spheroida (Ps), Xestospongia testudinaria (Xt). Bars, symbols and x-axis biotope/category identifications have been colour coded in the online version orange for known LMA sponges and for LMA category; blue for known HMA sponges and for the HMA category and purple for sponge species of unknown LMA–HMA status. In the PCO (f) very small circles represent OTUs ≤ 100 sequence reads
LMA and HMA across geographical areas
The comparison between closely related organisms (identified using BLAST) to the most abundant OTUs, diversity indices, mean abundance of the main Phyla (Thaumarchaeota, Euryarchaeota), number of core OTUs, mean abundance of core OTUs and mean abundance of the single most abundant OTU of each LMA (S. massa, S. carteri) and HMA (X. testudinaria, H. erectus) species and environmental biotopes in Mayotte and 4 other geographical areas are shown in Supplementary Tables 6, 7, 8, 9.
Some of the most abundant OTUs in the present and previously published studies were found to be closely related to the same set of organisms in GenBank (Supplementary Table 6). For example, in the present study, OTUs 1 and 357, assigned to the genus Cenarchaeum and abundant in S. massa and S. carteri, were closely related (99.77–100%) to an organism (Accession number: HQ877742) obtained from the sponge Phakellia fusca (Supplementary Table 6). The most abundant OTUs recorded from the same sponge species collected in Berau, Papua and Jakarta were also found to be related to this organism. OTU-4 in the present study was abundant in X. testudinaria and was closed related (100%) to an organism (Accession number: GQ485797) obtained from the sponge X. muta; abundant OTUs recorded from X. testudinaria, but collected in Papua and Jakarta were also closely related to this organism. OTU-2 in the present study was abundant in the sponge species H. erectus and closely related (99.11%) to an organism (Accession number: EF529650) obtained from the sponge Aplysina aerophoba; the most abundant OTU recorded from the same sponge species in Makassar was also closely related to this organism.
The diversity indices of LMA species, HMA species and environmental biotopes for various Indopacific coral reef systems are shown in Supplementary Table 7. The HMA sponge species X. testudinaria had higher diversity and evenness than sympatric LMA sponge species in Mayotte, Papua and Jakarta and higher richness in Makassar and Jakarta. Thaumarchaeota members were more abundant in sponge and sediment biotopes and Euryarchaeota members more abundant in seawater in all research areas. Among sponges, Euryarchaeota members were more abundant in LMA sponges (S. carteri and S. massa) across all studied coral reef systems (Supplementary Table 8). The mean abundances of the single most abundant OTU and core OTUs were higher in S. carteri and S. massa than in X. testudinaria and H. erectus (Supplementary Table 9).
Discussion
Higher taxon abundance
Most of the sponge and sediment archaeal sequences were assigned to the Thaumarchaeota phylum (> 94% in sponges and 90% in sediment). Albeit in very low abundances, Euryarchaeota was also found in all sponge species and dominated the seawater archaeal community (99.6%). These results are in accordance with other studies, which have also found low abundances of euryarchaeotal members in sponge species (Holmes and Blanch 2007; Lee et al. 2011; Dupont et al. 2013; Kennedy et al. 2014) and seawater samples almost totally consisting of archaeal reads classified as Euryarchaeota (Lee et al. 2011). In contrast and in line with previous studies, sediment (Gaidos et al. 2011; Polónia et al. 2013) and sponge samples (Polónia et al. 2014, 2015; Turon and Uriz 2020) were mostly dominated by thaumarchaeotal members. Most of the 17 tropical sponge species analyzed by Turon and Uriz (2020) were also dominated by different genera of the Thaumarchaeota phylum. Other authors, however, have reported seawater samples with Thaumarchaeota percentages ranging from 17 to 68% (Kennedy et al. 2014) and an absence of sequences assigned to the Euryarchaeota phylum in sponges (Turque et al. 2010; Alex and Antunes 2015) suggesting that this trait may be related to certain environmental conditions.
Dominant and core OTUs
Only 2 core OTUs were observed in every sponge specimen. Similar to other studies, we found the sponge archaeal community to consist of a small number of abundant core OTUs (Turon and Uriz 2020). At the species level, the highest number of possible core OTUs was 17 in E. coccinea and H. erectus. Note, however, that core status in the present study was based on very small sample sizes and more individuals of a species should be sampled before reliably determining whether or not they are core members. That being said, 17 possible core OTUs is a low number when compared to those found by Turon and Uriz (2020) in the sponge species Dysidea sp. (77). The relative abundance of these core OTUs varied from 86.02 in sediment to 99.63% in S. carteri. In sponges, the lowest abundance was observed in H. erectus (92.58%). In the 17 tropical sponge species from Nha Trang Bay (Vietnam), Turon and Uriz (2020) reported relative abundances of core OTUs higher than 60% in most of their species.
Of the 39 abundant (≥ 130 sequences) OTUs, all had similarities > 97% to previously published sequences; 12 were closely related to organisms in GenBank, which were closely related to OTUs of the same host species from different areas (Polónia et al. 2014, 2015, 2016, 2017, 2018; Polónia and Cleary 2019) and only one had similarity < 97% to previously published sequences. OTU-16 (only found in one sample of Petrosia aff. spheroida; 235 sequences) had a similarity of 88.24% with organisms obtained from an ocean drilling core sample from Shimokita Penninsula (Japan).
The mean abundance of the single most abundant OTU, was relatively high in the sponge species S. massa, S. carteri, P. bacca and E. coccinea. The affiliation of the most abundant OTU in these sponges, however, differed between both Stylissa species and P. bacca and E. coccinea. OTU-1 was the most abundant OTU in both Stylissa species, while OTU-3 was the most abundant OTU in P. bacca. Both of these OTUs were assigned to the genus Cenarchaeum. OTU-6, assigned to the genus Candidatus Nitrosopumilus, in contrast was the most abundant OTU in E. coccinea. This group of sponges shared several lesser abundant OTUs with seawater.
The presence of Cenarchaeum and Candidatus Nitrosopumilus members in sponges has been widely reported (Preston et al. 1996; Turque et al. 2010; Jackson et al. 2013; Kennedy et al. 2014; Zhang et al. 2014; Lopes et al. 2019; Turon and Uriz 2020). Members of these two genera are important chemolithoautotrophic, ammonia-oxidizing archaeons (AOA). The Nitrosopumilus species type (N. maritimus) uses bicarbonate as the sole carbon source and has its growth inhibited by organic carbon (Könneke et al. 2005; Francis et al. 2007). Some species, which is not the case of the type species, also may use urea as a source of ammonia for growth and energy production (Qin et al. 2017). The Cenarchaeum species type (Cenarchaeum symbiosum) has been reported, based on genomic predictions, as either a strict autotroph, or a mixotroph using both carbon dioxide and organic material as carbon sources (Hallam et al. 2006a, b) and having the ability to use urea as an alternative source of ammonia (Kerou and Schleper 2015; Hallam et al. 2006a, b). This may empower this taxon with the ability of chemolithoautotrophic growth in low pH environments (Burton and Prosser 2001).
Diversity patterns
As expected, richness and evenness were highest in sediment samples. Among sponges, OTU richness was highest in H. erectus and E. coccinea followed by Petrosia aff. spheroida, S. carteri, S. massa, X. testudinaria and lowest in P. bacca. Evenness, in turn, was higher in X. testudinaria, H. erectus (HMA) and Petrosia aff. spheroida than the remaining sponge species and was lowest in P. bacca. The Shannon diversity index varied from 0.025 (P. bacca) to 1.16 (Petrosia aff. Spheroida), which contrasts with Turon and Uriz (2020) who found most of the archaeal Shannon diversity indices ranging between 1 and 2. In de Voogd et al. (2019), pooled rarefied bacterial richness was lowest in P. bacca and highest in S. carteri. Still, in bacterial communities, Erwin et al. (2015) reported richness and evenness values higher in the sponge species Agelas oroides, Chondrosia reniformis and Petrosia ficiformis than in the sponge species Axinella damicornis, Dysidea avara and Spirastrella cunctatrix and intermediate values in the seawater samples. Here, seawater had higher evenness than all the sponge samples and richness was higher than all sponges with the exceptions of E. coccinea and H. erectus. In the prokaryotic community, Cleary et al. (2019) also reported higher evenness values in the HMA sponge species Aaptos lobata, H. erectus and X. testudinaria.
Community composition
In line with previous studies (Polónia 2014, 2015; Turon and Uriz 2020), the biotope/species identity was an important structuring factor explaining 87% of the variation in archaeal composition. Additionally, our results showed significant congruence between the OTU ordination patterns of archaeal and bacterial communities. In line with the bacterial OTU ordination analysed in de Voogd et al. (2019), samples of Petrosia aff. spheroida clustered together with X. testudinaria, a well-known HMA species. Likewise, samples of E. coccinea and P. bacca clustered together with the two known LMA sponges (S. massa and S. carteri). Previous studies have, however, designated sponges of the genus Ectyoplasia as HMA species (Gloeckner et al. 2014; Schmitt et al. 2008) in contrast to our results and those of de Voogd et al. (2019).
In contrast to the OTU ordination analysed in Voogd et al. 2019, the HMA sponge H. erectus had an archaeal community, which was intermediate in composition between X. testudinaria and Petrosia aff. spheroida and sediment. This may be related to the fact that this sponge species, lives cryptically embedded in the sediment and covered in coral sand where it uses exogenous materials such as sediment grains to build-up its skeleton (Polónia et al. 2015). The bacterial community of these samples, however, did not show the same pattern and clustered closer together with the aforementioned HMA sponges, suggesting that, in the case of H. erectus, there is a relatively lower congruence between archaeal and bacterial communities as shown by the higher residual distances of these samples (Fig. 4b). These Procrustes results were consistent with those reported by Chaib De Mares et al. (2017) who, comparing the congruence between bacterial and archaeal communities of the sponge species Aplysina aerophoba, Aplysina cauliformis, Dysidea avara, Dysidea etheria and seawater, also found a significant congruence between both data sets.
HMA versus LMA
The higher percentage of sequences assigned to the Euryarchaeota in LMA sponges (S. carteri and S. massa) when compared to HMA sponges (H. erectus, X. testudinaria) observed in the present study was consistent with results obtained in other geographical areas (Polónia et al. 2014, 2015, 2016, 2017, 2018; Moitinho-Silva et al. 2017). Our results are thus consistent with the concept that LMA sponges have a microbial community closer to that of seawater than their HMA counterparts. Based on this apparently consistent trait, we thus suggest that the sponge species P. bacca and E. coccinea are LMA species.
The most abundant OTUs (1 and 3) of both Stylissa species and P. bacca were assigned to the Cenarchaeum genus. In contrast, the abundant OTUs of X. testudinaria, H. erectus, Petrosia aff. spheroida and E. coccinea (4, 5, 6, 339 and 1009) were assigned to the genus Candidatus Nitrosopumilus. In a previous study (Polónia et al. 2018), we showed that 82.50% of the archaeal sequences of S. carteri (LMA) were assigned to the Cenarchaeum genus while 97.78% and 96.48% of the archaeal sequences of X. testudinaria (HMA) and Aaptos lobata (putative HMA), respectively, were assigned to the genus Candidatus Nitrosopumilus.
These results suggest an association of Cenarchaeum spp. with LMA sponges and Candidatus Nitrosopumilus spp. with HMA sponges. However, these results differ from other studies that reported Nitrosopumilus as abundant in LMA sponges such as, Mycale laxissima (Zhang et al. 2014), Amphimedon sp. (Turon and Uriz 2020), Paraleucilla magna (Lopes et al. 2019) and more abundant in the LMA sponges Dysidea fragilis, Halichondria panicea, Dysidea avara and Dysidea etheria than in the HMA sponges Erylus formosus, Ircinia variabilis, Petrosia ficiformis, Rhopaloeides odorabile, Aplysina aerophoba and Aplysina cauliformis (Moitinho-Silva et al. 2017; Chaib De Mares et al. 2017). Additionally, a study of 17 sponge species (some of them LMA; Turon and Uriz 2020) only found a preponderance (> 50%) of OTUs assigned to the genus Cenarchaeaum in a single sponge species—Haliclona sp.—recently predicted as LMA by Moitinho-Silva et al. (2017) and Turon et al. (2018). Although our results point to a link between LMA status and the genus Cenarchaeum and between HMA status and the genus Candidatus Nitrosopumilus, the comparison to other studies shows that this is an inconsistent trait. In line with this, Moitinho-Silva et al. (2017) suggested that the HMA–LMA dichotomy is best resolved at phylum and class levels.
The mean abundance of the single most abundant OTU and the mean abundance of the core OTUs was in general higher in the LMA sponge species S. carteri and S. massa than in the HMA sponge species X. testudinaria and H. erectus. This result also confirms previous studies in other geographical areas (Polónia et al. 2014, 2015, 2016, 2017, 2018). There are several reports of LMA sponge species mostly dominated by a single OTU. Crambe crambe, a LMA sponge, was reported to be dominated by a single bacterial species assigned to the Betaproteobacteria (Croué et al. 2013) while 60% of the Niphates digitalis reads (another LMA sponge) belonged to a single OTU assigned to the Alphaproteobacteria (Gantt et al. 2019). Additionally, Knobloch et al. (2019) reported that a single core OTU dominated the bacterial community of Halichondria panicea (LMA) and in a study of bacterial community profiles in LMA sponges, Giles et al. (2013) reported the dominance of a highly abundant OTU in each of the studied sponge species with values ranging from 15% in S. carteri to 81% in N. digitalis.
Other studies have reported the dominance of a single archaeal OTU in other sponge species. Jackson et al. (2013), for example, reported the presence of a single OTU (assigned to the Cenarcheaceae) dominating (55–69%) the archaeal community of Inflatella pellicula, which was found at very low abundances in seawater (0.07%). Here, only 3 (0.02%) sequences of OTU-1 were found in a sediment sample and no sequences of OTU-3 were found in environmental samples. According to Kennedy et al. (2014), the microbial communities (archaea and bacteria) of Inflatella pellicula and Lissodendoryx diversichela were typical of LMA sponges and were dominated by a relatively small number of OTUs. Turon and Uriz (2020), however, reported Dysidea sp. (a genus with some species classified as LMA; Moitinho-Silva et al. 2017) as having equal contributions of several OTUs to the overall archaeal community. Based on this mostly consistent trait, the sponge species P. bacca and E. coccinea would be considered LMA (with mean abundance of the single most abundant OTU > 93%) while Petrosia aff. spheroida would be considered HMA (with mean abundance of the single most abundant OTU ~ 66%).
In Mayotte, OTU richness was higher in H. erectus and E. coccinea than all other sponge species. Evenness, in turn, was higher in both HMA species (X. testudinaria and H. erectus) than the LMA species (S. carteri and S. massa). In the other indopacific coral reefs, the HMA species X. testudinaria had higher diversity and evenness than sympatric LMA sponge species in Papua and Jakarta, but not in Makassar and Berau and higher richness in Makassar and Jakarta, but not in the remaining areas (Polónia et al. 2014, 2015, 2016, 2017, 2018). The present and previous results, thus, do not fully corroborate other studies showing HMA sponges hosting richer and more diverse (even) microbial communities than their LMA counterparts (Poppell et al. 2014; Erwin et al. 2015; Moitinho-Silva et al. 2017; Cleary et al. 2019).
Conclusions
Our study showed that known LMA sponges had higher abundances of the phylum Euryarchaeota, higher mean abundances of the single most abundant OTU and Core OTUs and relatively low evenness compared to known HMA sponge species. The higher microbial richness generally associated with HMA sponges was not found in the present study. Based on our results, we predict that P. bacca is a LMA species and Petrosia aff. spheroida is a HMA sponge species. The status of E. coccinea was not clear and it displayed intermediate traits. Finally, archaeal and bacterial community composition was significantly congruent.
Availability of data and materials
The DNA sequences generated in this study can be downloaded from the National Center for Biotechnology Information (NCBI) Sequence Read Archive (SRA): SRP071901: PRJNA315454.
References
Agnarsson I, Kuntner M (2012) The generation of a biodiversity hotspot: biogeography and phylogeography of the western Indian Ocean islands. Curr Topics Phylogenet Phylogeogr Terrest Aquat Syst 33:82
Alex A, Antunes A (2015) Pyrosequencing characterization of the microbiota from Atlantic intertidal marine sponges reveals high microbial diversity and the lack of co-occurrence patterns. PLoS ONE 10:e0127455
Bayer K, Schmitt S, Hentschel U (2008) Physiology, phylogeny and in situ evidence for bacterial and archaeal nitrifiers in the marine sponge Aplysina aerophoba. Environ Microbiol 10:2942–2955
Bayer K, Moitinho-Silva L, Brümmer F, Cannistraci CV, Ravasi T et al (2014a) GeoChip-based insights into the microbial functional gene repertoire of marine sponges (high microbial abundance, low microbial abundance) and seawater. FEMS Microbiol Ecol 90:832–843
Bayer K, Kamke J, Hentschel U (2014b) Quantification of bacterial and archaeal symbionts in high and low microbial abundance sponges using real-time PCR. FEMS Microbiol Ecol 89:679–690
Bell JJ (2008) The functional roles of marine sponges. Estuarine Coastal Shelf Sci 79:341–353. https://doi.org/10.1016/j.ecss.2008.05.002
Blanquer A, Uriz MJ, Galand PE (2013) Removing environmental sources of variation to gain insight on symbionts vs. transient microbes in high and low microbial abundance sponges. Environ Microbiol 15:3008–3019
Burton SA, Prosser JI (2001) Autotrophic ammonia oxidation at low pH through urea hydrolysis. Appl Environ Microbiol 67:2952–2957
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335
Chaib De Mares M, Sipkema D, Huang S, Bunk B, Overmann J, Van Elsas JD (2017) Host specificity for bacterial, archaeal and fungal communities determined for high-and low-microbial abundance sponge species in two genera. Front Microbiol 8:2560
Cleary DFR, de Voogd NJ, Polónia ARM, Freitas R, Gomes NCM (2015) Composition and predictive functional analysis of bacterial communities in seawater, sediment and sponges in an Indonesian coral reef environment. Microb Ecol 70:889–903. https://doi.org/10.1007/s00248-015-0632-5
Cleary DFR, Polónia ARM, Becking LE, de Voogd NJ, Purwanto GH, Gomes NCM (2018) Compositional analysis of bacterial communities in seawater, sediment and high and low microbial abundance sponges in the Misool coral reef system, Indonesia. Mar Biodivers 48:1889–1901. https://doi.org/10.1007/s12526-017-0697-0
Cleary DFR, Swierts T, Coelho FJ, Polónia ARM, Huang YM, Ferreira MR et al (2019) The sponge microbiome within the greater coral reef microbial metacommunity. Nat Commun 10:1644
Croué J, West NJ, Escande ML, Intertaglia L, Lebaron P, Suzuki MT (2013) A single betaproteobacterium dominates the microbial community of the crambescidine-containing sponge Crambe crambe. Sci Rep 3:2583
de Goeij JM, Van Oevelen D, Vermeij MJA, Osinga R, Middelburg JJ, de Goeij AFPM, Admiraal W (2013) Surviving in a marine desert: the sponge loop retains resources within coral reefs. Science 342:108–110. https://doi.org/10.1126/science.1241981
de Voogd NJ, Cleary DFR, Polónia ARM, Gomes NC (2015) Bacterial community composition and predicted functional ecology of sponges, sediment and seawater from the thousand islands reef complex, West Java, Indonesia. FEMS Microbiol Ecol 91:fiv019
de Voogd NJ, Gauvin-Bialecki A, Polónia ARM, Cleary DFR (2019) Assessing the bacterial communities of sponges inhabiting the remote western Indian Ocean island of Mayotte. Mar Ecol 39:e12517
Diaz MC, Rützler K (2001) Sponges: an essential component of Caribbean coral reefs. Bull Mar Sci 69:535–546
Dupont S, Corre E, Li Y, Vacelet J, Bourguet-Kondracki ML (2013) First insights into the microbiome of a carnivorous sponge. FEMS Microbiol Ecol 86:520–531
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10:996
Edgar R, Haas B, Clemente J, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200
Erwin PM, Coma R, López-Sendino P, Serrano E, Ribes M (2015) Stable symbionts across the HMA–LMA dichotomy: low seasonal and interannual variation in sponge-associated bacteria from taxonomically diverse hosts. FEMS Microbiol Ecol 91:fiv115
Fan L, Reynolds D, Liu M, Stark M, Kjelleberg S, Webster NS, Thomas T (2012) Functional equivalence and evolutionary convergence in complex communities of microbial sponge symbionts. Proc Natl Acad Sci USA 109:E1878–E1887. https://doi.org/10.1073/pnas.1203287109
Francis CA, Beman JM, Kuypers MM (2007) New processes and players in the nitrogen cycle: the microbial ecology of anaerobic and archaeal ammonia oxidation. ISME J 1:19
Gantt SE, McMurray SE, Stubler AD, Finelli CM, Pawlik JR, Erwin PM (2019) Testing the relationship between microbiome composition and flux of carbon and nutrients in Caribbean coral reef sponges. Microbiome 7:124
Giles EC, Kamke J, Moitinho-Silva L, Taylor MW, Hentschel U, Ravasi T, Schmitt S (2013) Bacterial community profiles in low microbial abundance sponges. FEMS Microbiol Ecol 83:232–241. https://doi.org/10.1111/j.1574-6941.2012.01467.x
Gloeckner V, Wehrl M, Moitinho-Silva L, Gernert C, Schupp P, Pawlik JR et al (2014) The HMA–LMA dichotomy revisited: an electron microscopical survey of 56 sponge species. Biol Bull 227:78–88
Gotelli NJ, Colwell RK (2001) Quantifying biodiversity: procedures and pitfalls in the measurement and comparison of species richness. Ecol Lett 4:379–391
Hallam SJ, Konstantinidis KT, Putnam N, Schleper C, Watanabe YI, Sugahara J et al (2006a) Genomic analysis of the uncultivated marine crenarchaeote Cenarchaeum symbiosum. Proc Natl Acad Sci USA 103:18296–18301
Hallam SJ, Mincer TJ, Schleper C, Preston CM, Roberts K, Richardson PM, DeLong EF (2006b) Pathways of carbon assimilation and ammonia oxidation suggested by environmental genomic analyses of marine Crenarchaeota. PLoS Biol 4:e95
Hentschel U, Hopke J, Horn M, Friedrich AB, Wagner M, Hacker J, Moore BS (2002) Molecular evidence for a uniform microbial community in sponges from different oceans. Appl Environ Microbiol 68:4431–4440
Hentschel U, Usher KM, Taylor MW (2006) Marine sponges as microbial fermenters. FEMS Microbiol Ecol 55:167–177
Hentschel U, Piel J, Degnan SM, Taylor MW (2012) Genomic insights into the marine sponge microbiome. Nat Rev Microbiol 10:641–654. https://doi.org/10.1038/nrmicro2839
Hoffmann F, Radax R, Woebken D, Holtappels M, Lavik G, Rapp HT, Schläppy M-L, Schleper C, Kuypers MMM (2009) Complex nitrogen cycling in the sponge Geodia barretti. Environ Microbiol 11:2228–2243. https://doi.org/10.1111/j.1462-2920.2009.01944.x
Holmes B, Blanch H (2007) Genus-specific associations of marine sponges with Group I Crenarchaeotes. Mar Biol 150:759–772. https://doi.org/10.1007/s00227-006-0361-x
Jackson SA, Flemer B, McCann A, Kennedy J, Morrissey JP, O’Gara F, Dobson AD (2013) Archaea appear to dominate the microbiome of Inflatella pellicula deep sea sponges. PLoS ONE 8:e84438
Kamke J, Taylor MW, Schmitt S (2010) Activity profiles for marine sponge-associated bacteria obtained by 16S rRNA vs 16S rRNA gene comparisons. ISME J 4:498
Kennedy J, Flemer B, Jackson SA, Morrissey JP, O’Gara F, Dobson AD (2014) Evidence of a putative deep sea specific microbiome in marine sponges. PLoS ONE 9:e91092
Kerou M, Schleper C (2015) Candidatus Cenarchaeum. Bergey's Manual of Systematics of Archaea and Bacteria, pp 1–4
Kiszka J, Ersts PJ, Ridoux V (2007) Cetacean diversity around the Mozambique Channel island of Mayotte (Comoros archipelago). J Cetacean Res Manag 9:105
Knobloch S, Jóhannsson R, Marteinsson V (2019) Bacterial diversity in the marine sponge Halichondria panicea from Icelandic waters and host-specificity of its dominant symbiont “Candidatus Halichondribacter symbioticus”. FEMS Microbiol Ecol 95:fiy220
Könneke M, Bernhard AE, José R, Walker CB, Waterbury JB, Stahl DA (2005) Isolation of an autotrophic ammonia-oxidizing marine archaeon. Nature 437:543
Lee OO, Wang Y, Yang J, Lafi FF, Al-Suwailem A, Qian PY (2011) Pyrosequencing reveals highly diverse and species-specific microbial communities in sponges from the Red Sea. ISME J 5:650–664
Lenth R (2017) emmeans: estimated marginal means, aka least-squares means. https://CRAN.Rproject.org/package=emmeans
Lopes MF, Mágna B, Klautau M, Esteves EL, Albano RM (2019) Microbiota of the alien species Paraleucilla magna (Porifera, Calcarea) from the Southwestern Atlantic, and a comparison with that of other calcareous sponges. BioRxiv, 626192.
McMurray SE, Pawlik JR, Finelli CM (2014) Trait-mediated ecosystem impacts: how morphology and size affect pumping rates of the Caribbean giant barrel sponge. Aquat Biol 23:1–13. https://doi.org/10.3354/ab00612
Moitinho-Silva L, Seridi L, Ryu T, Voolstra CR, Ravasi T, Hentschel U (2014a) Revealing microbial functional activities in the Red Sea sponge Stylissa carteri by metatranscriptomics. Environ Microbiol 16:3683–3698
Moitinho-Silva L, Steinert G, Nielsen S, Hardoim CC, Wu YC, McCormack GP et al (2017) Predicting the HMA–LMA status in marine sponges by machine learning. Front Microbiol 8:752
Moitinho-Silva L, Bayer K, Cannistraci CV, Giles E, Ryu T, Seridi L et al (2014b) Specificity and transcriptional activity of microbiota associated with low and high microbial abundance sponges from the Red Sea. Mol Ecol 23:1348–1363
Oksanen J, Kindt R, Legendre P, O’Hara B, Simpson GL, Solymos P, Wagner H. 2009. Vegan: community ecology package. R Package Version. 1:15–14. http://www.cran.rproject.org/package=vegan
Pires AC, Cleary DFR, Almeida A, Cunha Â, Dealtry S, Mendonça-Hagler LC et al (2012) Denaturing gradient gel electrophoresis and barcoded pyrosequencing reveal unprecedented archaeal diversity in mangrove sediment and rhizosphere samples. Appl Environ Microbiol 78:5520–5528
Polónia ARM, Cleary DFR, Duarte LN, de Voogd NJ, Gomes NC (2014) Composition of Archaea in seawater, sediment, and sponges in the Kepulauan seribu reef system, Indonesia. Microb Ecol 67:553–567
Polónia ARM, Cleary DFR, Freitas R, de Voogd NJ, Gomes NC (2015) The putative functional ecology and distribution of archaeal communities in sponges, sediment and seawater in a coral reef environment. Mol Ecol 24:409–423
Polónia ARM, Cleary DFR, Freitas R, Coelho FJRC, de Voogd NJ, Gomes NCM (2016) Comparison of archaeal and bacterial communities in two sponge species and seawater from an Indonesian coral reef environment. Mar Genomics 29:69–80
Polónia ARM, Cleary DFR, Freitas R, Gomes NCM, de Voogd NJ (2017) Archaeal and bacterial communities of Xestospongia testudinaria and sediment differ in diversity, composition and predicted function in an Indonesian coral reef environment. J Sea Res 119:37–53
Polónia ARM, Cleary DFR, Coelho FJRC, Becking LE, de Voogd NJ, Toha AHA, Gomes NCM (2018) Compositional analysis of archaeal communities in high and low microbial abundance sponges in the Misool coral reef system, Indonesia. Mar Biol Res 14:537–550
Polónia ARM, Cleary DFR (2019) Archaeal communities in sponge, sediment and water from marine lakes and open water habitats. Mar Biol Res 15:259–274
Poppell E, Weisz J, Spicer L, Massaro A, Hill A, Hill M (2014) Sponge heterotrophic capacity and bacterial community structure in high-and low-microbial abundance sponges. Mar Ecol 35:414–424
Preston CM, Wu KY, Molinski TF, DeLong EF (1996) A psychrophilic crenarchaeon inhabits a marine sponge: Cenarchaeum symbiosum gen. nov., sp. nov. P Nat Acad Sci 93:6241–6246
Qin W, Heal KR, Ramdasi R, Kobelt JN, Martens-Habbena W, Bertagnolli AD et al (2017) Nitrosopumilus maritimus gen. nov., sp. nov., Nitrosopumilus cobalaminigenes sp. nov., Nitrosopumilus oxyclinae sp. nov., and Nitrosopumilus ureiphilus sp. nov., four marine ammonia-oxidizing archaea of the phylum Thaumarchaeota. Int J Syst Evol Microbiol 67:5067–5079
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:D590–D596
Radax R, Hoffmann F, Rapp HT, Leininger S, Schleper C (2012) Ammonia-oxidizing archaea as main drivers of nitrification in cold-water sponges. Environ Microbiol 14:909–923. https://doi.org/10.1111/j.1462-2920.2011.02661.x
R Core Team (2013) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria ISBN 3-900051-07-0. http://www.R-project.orghttp://www.R-project.org
Said Hassane C, Fouillaud M, Le Goff G, Sklirou AD, Boyer JB, Trougakos IP et al (2020) Microorganisms associated with the marine sponge Scopalina hapalia: a reservoir of bioactive molecules to slow down the aging process. Microorganisms 8:1262
Schmitt S, Deines P, Behnam F, Wagner M, Taylor MW (2011) Chloroflexi bacteria are more diverse, abundant, and similar in high than in low microbial abundance sponges. FEMS Microbiol Ecol 78:497–510
Schmitt S, Angermeier H, Schiller R, Lindquist N, Hentschel U (2008) Molecular microbial diversity survey of sponge reproductive stages and mechanistic insights into vertical transmission of microbial symbionts. Appl Environ Microbiol 74:7694–7708
Shannon E, Weaver W (1949) The Mathematical Theory of Communication. University of Illinois Press, Urbana
Thacker RW, Freeman CJ (2012) Sponge–microbe symbioses: recent advances and new directions. Adv Mar Biol 62:57–111
Thomas T, Rusch D, DeMaere MZ, Yung PY, Lewis M, Halpern A, Heidelberg KB, Egan S, Steinberg PD, Kjelleberg S (2010) Functional genomic signatures of sponge bacteria reveal unique and shared features of symbiosis. ISME J 4:1557–1567. https://doi.org/10.1038/ismej.2010.74
Thomas T, Moitinho-Silva L, Lurgi M, Björk JR, Easson C, Astudillo-Gárcia C, Olson JB, Erwin PM, Lopez-Legentil S, Luter H, Chaves-Fonnegra A, Costa R, Schupp PJ, Steindler L, Erpenbeck D, Gilbert J, Knight R, Ackermann G, Lopez JV, Taylor MW, Thacker RW, Montoya JM, Hentschel U, Webster NS (2016) Diversity, structure and convergent evolution of the global sponge microbiome. Nat Commun 7:1–12
Turon M, Uriz MJ (2020) New insights into the archaeal consortium of tropical sponges. Front Mar Sci 6:789. https://doi.org/10.3389/fmars.2019.00789
Turon M, Cáliz J, Garate L, Casamayor EO, Uriz MJ (2018) Showcasing the role of seawater in bacteria recruitment and microbiome stability in sponges. Sci Rep 8:1–10
Turque AS, Batista D, Silveira CB, Cardoso AM, Vieira RP, Moraes FC et al (2010) Environmental shaping of sponge associated archaeal communities. PLoS ONE 5:e15774
Tweedie MCK (1984) An index which distinguishes between some important exponential families. In: Ghosh JK, Roy J (eds) Statistics: applications and new directions—proceedings of the Indian Statistical Institute Golden Jubilee international conference. Indian Statistical Institute, Calcutta, pp 579–604
Weisz JB, Hentschel U, Lindquist N, Martens CS (2007) Linking abundance and diversity of sponge-associated microbial communities to metabolic differences in host sponges. Mar Biol 152:475–483
Yahel G, Sharp JH, Marie D, Häse C, Genin A (2003) In situ feeding and element removal in the symbiont-bearing sponge Theonella swinhoei: bulk DOC is the major source for carbon. Limnol Oceanogr 48:141–149. https://doi.org/10.4319/lo.2003.48.1.0141
Zhang Z, Schwartz S, Wagner L, Miller W (2000) A greedy algorithm for aligning DNA sequences. J Comput Biol 7:203–214
Zhang F, Pita L, Erwin PM, Abaid S, López-Legentil S, Hill RT (2014) Symbiotic archaea in marine sponges show stability and host specificity in community structure and ammonia oxidation functionality. FEMS Microbiol Ecol 90:699–707. https://doi.org/10.1111/1574-6941.12427
Acknowledgements
Research permits were issued via the Prefecture of Mayotte. We thank Cécile Debitus, Bruno Fichou, Stephan Aubert, Philippe Prost, and Jean‐Pierre Bellanger for their support.
Funding
This study was financed through the ANR‐Netbiome under grant No ANR‐11‐EBIM‐0006 and a contribution to the project LESS CORAL [PTDC/AAC-AMB/115304/2009] funded by FEDER, through COMPETE2020 - Programa Operacional Competitividade e Internacionalização (POCI), and by national funds (OE), through the Portuguese Foundation for Science and Technology (FCT)/MCTES. Thanks are also due, to FCT/MCTES for the financial support to CESAM (UIDP/50017/2020 + UIDB/50017/2020), through national funds. Ana R.M. Polónia was supported by a postdoctoral scholarship (SFRH/BPD/117563/2016) funded by FCT/MCTES and by the European Social Fund (ESF).
Author information
Authors and Affiliations
Contributions
N.J.d.V. and D.F.R.C. designed the study; N.J.d.V. and A.G.B. collected the samples; A.R.M.P. performed the laboratory work; D.F.R.C. and A.R.M.P. performed the data analysis; A.R.M.P., D.F.R.C., A.G.B., and N.J.d.V wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent for publication
All authors read and approved the final version of the manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the supplementary information.
Supplementary Fig. 1
(a) Location map with (b) inset showing the island of Mayotte (PDF 182 kb)
Supplementary Fig. 2
Relative abundance of the most abundant classes in the demosponges Paratetilla bacca (Pb), Stylissa carteri (Sc), Stylissa massa (Sm), Ectyoplasia coccinea (Ec), Hyrtios erectus (He), Petrosia aff. spheroida (Ps), Xestospongia testudinaria (Xt); sediment (Sd), and seawater (Wt) (PDF 35 kb)
Supplementary Table 1
Sample list with the sample code, sponge species, Group code, Host, high microbial abundance (HMA) or low microbial abundance (LMA) type, collection site (location), and GPS coordinates (XLS 26 kb)
Supplementary Table 2
Results of emmeans analysis showing pairwise comparisons of differences in the relative abundances of selected phyla between biotopes based on the Tukey test. Significance: * 0.01 < Pr < 0.05 ** 0.001 < Pr < 0.01; *** Pr < 0.001 (XLS 52 kb)
Supplementary Table 3
Sample list with the sample code, biotope, number of replicates, total number of OTUs, number of core OTUs mean abundance of core OTUs, mean abundance of the single most abundant OTUs, number of specific OTUs, Diversity, Richness and Evenness. (XLS 36 kb)
Supplementary Table 4
Results of simper analysis showing the contribution of OTUs to differences in similarity between pairs of samples. OTUs that contribute significantly to differences are indicated: * 0.01 < P < 0.05 ** 0.001 < P < 0.01; *** P < 0.001 (XLSX 72 kb)
Supplementary Table 5
List of abundant (≥ 130 sequence reads) OTUs and closely related organisms identified using BLAST search. OTU: OTU number; Abund: number of sequence reads; Taxonomic classification, Acc: Genbank accession numbers of closely related organisms identified using BLAST; Seq: sequence similarity of these organisms with our representative OTU sequences; Source: isolation source of organisms identified using BLAST (XLS 36 kb)
Supplementary Table 6
List of abundant Mayotte OTUs with the same closely related organisms identified using BLAST search in other geographic areas. OTU: OTU number; Study: Geographic area; Reference: reference of the study in which the OTU were previous reported; Core: biotopes where the OTUs were core OTUs; Group: biotope where the OTUs were found; Abund: number of sequence reads; Database: Database used in QIIME; Taxonomic classification; GI:GenBank GenInfo sequence identifiers; Acc: Genbank accession numbers of closely related organisms identified using BLAST; Seq: sequence similarity of these organisms with our representative OTU sequences; Source: isolation source of organisms identified using BLAST; Location: Geographic region of the organisms identified using BLAST. In the ‘Group’ category, the most dominant OTUs and the OTUs predominantly found in a given biotope (not considering OTUs ≤ 5 sequences) are indicated by one asterisk (*) and/or marked bold respectively. Sm: S. massa; Sc: S. carteri; Wt: Water; He: H. erectus; Ps: Petrosia aff. spheroida; Sed: Sediment; Xt: X. testudinaria; Ap: Aaptos lobata; Bie: Biemna fortis. (XLS 41 kb)
Supplementary Table 7
Mean and standard deviation (SD) of Diversity (H′), Richness (S) and Evenness (J) indices reported in the LMA sponge species S. massa and S. carteri and the HMA sponge species X. testudinaria and H. erectus (N: number of replicates) across several geographic regions. Makassar: Spermonde Archipelago coral reef system; Jakarta: Kepulauan Seribu Reef System, Berau: Berau reef system and Papua: Misool coral reef system, Indonesia resorting to the results published in our previous studies (Polónia et al. 2014; 2015; 2016; 2017; 2018). (XLS 42 kb)
Supplementary Table 8
Mean and standard deviation (SD) relative abundance of the Thaumarchaeota and Euryarchaeota phyla reported in the LMA sponge species S. massa and S. carteri and the HMA sponge species X. testudinaria and H. erectus (N: number of replicates) across several geographic regions. Makassar: Spermonde Archipelago coral reef system; Jakarta: Kepulauan Seribu Reef System, Berau: Berau reef system and Papua: Misool coral reef system, Indonesia resorting to the results published in our previous studies (Polónia et al. 2014; 2015; 2016; 2017; 2018). (XLS 41 kb)
Supplementary Table 9
Number of core OTUs (C.OTUs) the mean abundance of these core OTUs (C.OTUs(%)) and the mean abundance of the single most abundant OTU (D.OTU(%)) in the LMA sponge species S. massa and S. carteri and the HMA sponge species X. testudinaria and H. erectus (N: number of replicates) across several geographic regions. Makassar: Spermonde Archipelago coral reef system; Jakarta: Kepulauan Seribu Reef System, Berau: Berau reef system and Papua: Misool coral reef system, Indonesia resorting to the results published in our previous studies (Polónia et al. 2014; 2015; 2016; 2017; 2018). (XLS 39 kb)
Rights and permissions
About this article
Cite this article
Polónia, A.R.M., Cleary, D.F.R., Gauvin‐Bialecki, A. et al. Archaeal communities of low and high microbial abundance sponges inhabiting the remote western Indian Ocean island of Mayotte. Antonie van Leeuwenhoek 114, 95–112 (2021). https://doi.org/10.1007/s10482-020-01503-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-020-01503-5