Abstract
Planctomycetes occur in almost all aquatic ecosystems on earth. They have a remarkable cell biology, and members of the orders Planctomycetales and Pirellulales feature cell division by polar budding, perform a lifestyle switch from sessile to motile cells and have an enlarged periplasmic space. Here, we characterise a novel planctomycetal strain, Pla110T, isolated from the surface of polystyrene particles incubated in the Baltic Sea. After phylogenetic analysis, the strain could be placed in the family Planctomycetaceae. Strain Pla110T performs cell division by budding, has crateriform structures and grows in aggregates or rosettes. The strain is a chemoheterotroph, grows under mesophilic and neutrophilic conditions, and exhibited a doubling time of 21 h. Based on our phylogenetic and morphological characterisation, strain Pla110T (DSM 103387T = LMG 29693T) is concluded to represent a novel species belonging to a novel genus, for which we propose the name Polystyrenella longa gen. nov., sp. nov.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Planctomycetes are bacteria with a Gram-negative cell envelope architecture belonging to the PVC superphylum, along with Verrucomicrobia, Lentisphaerae, Kiritimatiellaeota, Candidatus Omnitrophica and Chlamydiae. Many representatives of this superphylum have major medical and biotechnological relevance (Wagner and Horn 2006), and can play key roles in global biogeochemical cycles (Peeters and van Niftrik 2018; Strous et al. 1999; Wiegand et al. 2018).
Planctomycetes are found in many environments, e.g. on (marine) algae (Bengtsson and Øvreås 2010; Bengtsson et al. 2012; Bondoso et al. 2014, 2015, 2017; Lage and Bondoso 2014; Vollmers et al. 2017), in peat bogs in northern wetlands (Kulichevskaya et al. 2012) or in hot springs (Slobodkina et al. 2015). They are assumed to metabolise complex carbon substrates released by biotic surfaces (Frank et al. 2014; Jeske et al. 2013; Lachnit et al. 2013; Wiegand et al. 2018). In the past, Planctomycetes were thought to have a number of exceptional traits (Devos et al. 2013; Devos and Reynaud 2010; Fuerst and Sagulenko 2011; Fuerst and Webb 1991; König et al. 1984; Lindsay et al. 1997; Lonhienne et al. 2010) that would place them at the base of the eukaryotes, but these properties have been re-interpreted in the recent years (Acehan et al. 2013; Boedeker et al. 2017; Jeske et al. 2015; Jogler 2014; Jogler et al. 2011; Jogler and Jogler 2013; Neumann et al. 2014; Rast et al. 2017; Rivas-Marin et al. 2016; Santarella-Mellwig et al. 2013). The cell envelope architecture of Planctomycetes was classified as Gram-negative (Boedeker et al. 2017; Devos 2014; van Teeseling et al. 2015).
Nevertheless, Planctomycetes are non-canonical as they have many remarkable properties. They perform cell division by budding or binary fission (or even a combination) while lacking most of the canonical cell division proteins, including FtsZ (Jogler et al. 2012; Pilhofer et al. 2008; Wiegand et al. 2020). Many Planctomycetes can perform a lifestyle switch between sessile mother cells and motile daughter cells (Jogler et al. 2011; Wiegand et al. 2020). They can also have unusual crateriform structures on their cell surfaces, which are visible by electron microscopy. Their periplasm is enlarged and forms large invaginations into the cytoplasm. Several planctomycetal strains are regarded as potential talented producers of as yet uncharacterised small bioactive molecules (Graça et al. 2016; Jeske et al. 2016; Wiegand et al. 2018). Many Planctomycetes possess large numbers of giant genes (Kohn et al. 2016; Reva et al. 2008) and are amongst the bacterial phyla with the highest number of predicted genes of unknown function (Overmann et al. 2017; Wiegand et al. 2018).
In this study, we describe a novel strain, Pla110T, which was isolated from polystyrene particles incubated in the Baltic Sea. The strain is part of the family Planctomycetaceae, which includes the limnic model planctomycete Planctopirus limnophila. Most of the other known members of this clade are marine bacteria.
Materials and methods
Isolation and cultivation
Strain Pla110T was collected during a sampling campaign, in which polystyrene particles were stored in small incubators, submerged in water and incubated at the given location for 14 days. The strain characterised here was isolated from the Baltic Sea 120 m off the shore at Heiligendamm, Germany (54.146 N 11.843 E). Isolation was performed as previously described (Wiegand et al. 2020).
The strain was cultivated and analysed in M1H medium supplemented with N-acetyl-glucosamine (NAG) and artificial seawater (ASW) (M1H NAG ASW medium) as described earlier (Wiegand et al. 2020) and incubated in a shaking incubator at 28 °C and 110 rpm. Medium supplemented with 1.5% (w/v) washed agar was used for plates.
Light microscopy and electron microscopy
Phase contrast micrographs were taken with a Nikon Eclipse Ti inverted microscope and a Nikon DS-Ri2 camera. To ensure good image quality, specimens were immobilised using a 1% (w/v) agarose cushion and MatTek glass bottom dishes (35 mm, No. 1.5) (Boedeker et al. 2017). ImageJ (Rueden et al. 2017) was used to examine cell size by sequentially applying an Otsu threshold, a watershed, and count particles.
Field emission scanning electron microscopy was performed as described (Boersma et al. 2019). Briefly, bacteria were fixed in formaldehyde, washed, and placed on cover slips coated with poly-l-lysine solution. Cover slips were then fixed in 1% (v/v) glutaraldehyde and washed twice before dehydrating in a graded series of acetone (10, 30, 50, 70, 90, 100% (v/v)) on ice. Samples from the 100% acetone step were brought to room temperature before placing them in fresh 100% acetone. Samples were then subjected to critical-point drying with liquid CO2 (CPD 300, Leica). Dried samples were covered with a gold/palladium (80/20) film by sputter coating (SCD 500, Bal-Tec) before examination in a field emission scanning electron microscope (Zeiss Merlin) using the Everhart Thornley HESE2 detector and the inlens SE detector in a 25:75 ratio at an acceleration voltage of 5 kV.
Physiological analyses
Determination of the pH optimum for growth was performed at 28 °C, with buffering agents 100 mM 2-(N-morpholino)ethanesulfonic acid (MES) at pH 5 and 6, 100 mM (4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid) (HEPES) at pH 7, 7.5 and 8, or 100 mM N-cyclohexyl-2-aminoethanesulfonic acid (CHES) at pH 9 and 10. Temperature optimum for growth determination was performed at pH 7.5 with temperatures ranging from 10 to 40 °C. Cell densities were inferred from optical density measurements (OD600).
Genome information
The genome and 16S rRNA gene sequence of strain Pla110T are available from GenBank under accession numbers CP036281 and MK554533, respectively. Information on genome sequencing and assembly was previously published (Wiegand et al. 2020).
Phylogenetic analysis
16S rRNA gene-based phylogeny was computed for strain Pla110T, the type strains of all described planctomycetal species (assessed in January 2020), all isolates recently described (Boersma et al. 2019; Kallscheuer et al. 2019a, b, c, d, 2020; Kohn et al. 2019; Peeters et al. 2019; Rensink et al. 2020; Wiegand et al. 2020) and with an outgroup of strains from outside the Planctomycetes, but part of the PVC superphylum. The alignment of 16S rRNA genes was made with SINA (Pruesse et al. 2012). Phylogenetic analysis was performed employing a maximum likelihood approach with 1,000 bootstraps, the nucleotide substitution model GTR, gamma distribution, and estimation of proportion of invariable sites using GTRGAMMAI (Stamatakis 2014).
The genomes for the genome-based analyses were gathered from GenBank, including the sequence for strain Pla110T. Completeness and contamination of the genome was determined using CheckM v1.0.131 (Parks et al. 2015). The average nucleotide identity (ANI) was calculated using OrthoANI (Lee et al. 2016), the average amino acid identity (AAI) was computed with the aai.rb script from the enveomics collection (Rodriguez-R and Konstantinidis 2016) and the percentage of conserved proteins (POCP) was determined as previously described (Qin et al. 2014). The rpoB nucleotide sequences (encoding the β-subunit of the DNA-dependent RNA polymerase) were taken from the genome annotations and the sequence identities were determined as described before (Bondoso et al. 2013). Upon extracting only those parts of the sequences that would have been sequenced with the described primer set, the alignment and matrix calculation was done with Clustal Omega (Sievers et al. 2011). The genus thresholds for rpoB were taken from a previous study (Kallscheuer et al. 2019d).
For the multi-locus sequence analysis (MLSA), the unique single-copy core genome of all analysed genomes was determined with proteinortho5 (Lechner et al. 2011) with the ‘selfblast’ option enabled. The protein sequences of the resulting orthologous groups were aligned using MUSCLE v.3.8.31 (Edgar 2004). After clipping, partially aligned C- and N-terminal regions and poorly aligned internal regions were filtered using Gblocks (Castresana 2000). The final alignment was concatenated and clustered using the maximum likelihood method implemented by RaxML (Stamatakis 2014) with the ‘rapid bootstrap’ method and 500 bootstrap replicates. The outgroup consisted of concatenated gene sets of strains from the order Pirellulales.
Results and discussion
Morphological and physiological analyses
Strain Pla110T forms white colonies, while colony colours of related strains range from white to ochre (Table 1). Mature cells of strain Pla110T are attached to each other by loose polar fimbriae, enabling the cells to grow in aggregates or rosettes (Fig. 1a). Fibres originate from crateriform structures, which were also observed on one of the two poles of Pla110T cells.
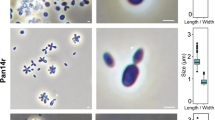
Morphological and physiological characterisation of strain Pla110T.
a SEM and phase contrast microscopy images showing elongated cells that divide by polar budding with the division site always at the wider side of the cell. Asterisks (*) indicate the nascent bud of a dividing cell. Scale bars represent 1 µm. b Boxplot of cell size, and growth rate of strain Pla110T at various pH values and temperatures
When examined with phase contrast microscopy, the cells appeared to be pear-shaped and on average 1.4 ± 0.2 by 0.7 ± 0.1 µm in size (Fig. 1b), which is relatively elongated for a planctomycetal cell (Table 1). This characteristic makes them distinguishable from their close relatives. Typical for members of Planctomycetaceae, the cells perform cell division by polar budding. The wider side of the cell was always observed to be the origin of the nascent bud during both SEM and phase contrast microscopy (Fig. 1a).
Strain Pla110T was isolated from the surface of plastic (polystyrene), whereas many of its close relatives were found on plant surfaces and can subsist on sugars. The strain was found to grow chemoorganotrophically, to be strictly aerobic and to grow at temperatures ranging from 10 to 27 °C, with the optimum at 24 °C (Fig. 1b). The pH values that permit growth of the strain range from 6.5 to 8.5, with the optimum at 7.5 (Fig. 1b). The growth rate of this strain was calculated to be 0.033 h− 1, which corresponds to a doubling time of 21 h.
Genomic characteristics
Compared to other Planctomycetes, the genome of the novel strain features an average number of transposable elements per Mb (0.82). Proteins per Mb (752), genes per Mb (762) and tRNAs per Mb (7.67) are all at the lower end of the planctomycete spectrum (Table 1). The genome is 6,125,480 bp in length, features two 16S rRNA genes and has a coding density of 84.5%. The G + C content of strain Pla110T is 55.3%, which is in the lower range compared to other members of the family Planctomycetaceae (50.4–70.7%, Table 1). The number of transposable elements per Mb (0.82) is higher than for most of its close relatives.
Phylogenetic analysis
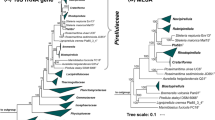
Based on 16S rRNA gene comparison and MLSA, strain Pla110T clusters within the family Planctomycetaceae according to the recent re-definition (Dedysh et al. 2019) (Fig. 2). However, it remains elusive from these analyses which genus is the current closest neighbour, although both trees showed a relationship with the type strain of Gimesia maris, to which high pairwise 16S rRNA homology was found (90.5%). The 16S rRNA sequence similarities between strain Pla110T and the described genera within the family Planctomycetaceae range from 81.7–90.6% (Fig. 3). These identity values are below the suggested threshold for genera of 94.5% (Yarza et al. 2014), indicating that strain Pla110T is not part of any established genus, but instead represents a novel genus.
Phylogenetic inference of strain Pla110T
a 16S rRNA gene-based phylogenetic tree of described Planctomycetes and strain Pla110T indicated in blue. Bootstrap values are indicated as a proportion of 1,000 re-samplings (in %). The outgroup consists of three 16S rRNA genes from the PVC superphylum. b Whole genome-based MLSA phylogeny, with bootstrap values based on 500 re-samplings at the nodes (in %). The outgroup consists of several representatives of the order Pirellulales. Planctomyces bekefii (Dedysh et al. 2020) was included in the 16S rRNA gene-based comparison, but not in the MLSA comparison as only metagenome-assembled genomes are available
Comparison of phylogenetic markers to separate species and genera
To determine strain Pla110T as a novel genus and species, the strain was compared to several related strains. Methods used: 16S rRNA gene sequence identity (16S), average amino acid identity (AAI), rpoB gene identity, average nucleotide identity (ANI), and percentage of conserved proteins (POCP). Planctomyces bekefii (Dedysh et al. 2020) was included in the 16S rRNA gene comparisons, but not in the others as only metagenomic bins are available
Other phylogenetic markers, such as rpoB similarity (Bondoso et al. 2013), AAI (Konstantinidis and Tiedje 2005) and POCP (Qin et al. 2014) provide additional means to determine if strain Pla110T is indeed part of a new genus (Fig. 3). Comparison of AAI values of strain Pla110T and other members of the family Planctomycetaceae result in an identity range between 48.3% and 52.8% (Fig. 3). This range falls below the threshold of 60%, supporting the conclusion that this strain is indeed part of a novel genus (Luo et al. 2014). Comparison of the POCP yielded values of 35.7–44.3% (Fig. 3), which are below the 50% cut-off value, again indicating a separate genus. The similarity of a 1200 bp region of the rpoB gene was used previously to infer phylogeny in genera belonging to the family Planctomycetaceae (Kallscheuer et al. 2019d). For the novel strain, we observed an identity of the mentioned partial sequence of the rpoB gene of 64.9–72.4% to other Planctomycetaceae members (Fig. 3). Again, these values are below the proposed threshold range for genera of 75.5–78% (Kallscheuer et al. 2019d), which is in line with the delineation of Pla110T from existing genera.
Taken together, based on results obtained for phylogeny and morphology we conclude that strain Pla110T belongs to a novel genus. We propose the name Polystyrenella gen. nov., with Polystyrenella longa sp. nov. as type species. Pla110T is the type strain of Polystyrenella longa.
Polystyrenella gen. nov.
Polystyrenella (Po.ly.sty.re.nel’la. N.L. neut. n. polystyrenum polystyrene; N.L. fem. dim. n. Polystyrenella an organism isolated from polystyrene).
The members of this genus have a Gram-negative cell envelope architecture, are aerobic, mesophilic, neutrophilic and heterotrophic. Cells are pear-shaped and divide by polar budding. The genus is part of the family Planctomycetaceae, order Planctomycetales, class Planctomycetia, phylum Planctomycetes.
Polystyrenella longa sp. nov.
Polystyrenella longa (lon’ga. L. fem. adj. longa long; corresponding to the longish appearance of the cells).
The members of this species are pear-shaped cells that grow in aggregates or rosettes. The cells are between 1.4 ± 0.2 by 0.7 ± 0.1 µm in size and divide by polar budding. The cells produce fimbriae originating from one of the cell poles, and have crateriform structures on this fibre pole. The type strain grows at 10–27 °C (optimum 24 °C) and pH 6.5–8.5 (optimum 7.5). Forms white colonies on M1H NAG ASW agar. The G + C content of the type strain genome is 50.3%.
The type strain, Pla110T (DSM 103387T = LMG 29693T), was isolated from polystyrene particles submerged near Heiligendamm in the Baltic Sea. The genome (accession no. CP036281) and 16S rRNA gene sequence (accession no. MK554533) are available from GenBank.
References
Acehan D, Santarella-Mellwig R, Devos DP (2013) A bacterial tubulovesicular network. J Cell Sci 127:277–280
Bengtsson MM, Øvreås L (2010) Planctomycetes dominate biofilms on surfaces of the kelp Laminaria hyperborea. BMC Microbiol 10:261
Bengtsson MM, Sjøtun K, Lanzén A, Øvreås L (2012) Bacterial diversity in relation to secondary production and succession on surfaces of the kelp Laminaria hyperborea. ISME J 6:2188–2198
Boedeker C, Schuler M, Reintjes G, Jeske O, van Teeseling MC, Jogler M, Rast P, Borchert D, Devos DP, Kucklick M, Schaffer M, Kolter R, van Niftrik L, Engelmann S, Amann R, Rohde M, Engelhardt H, Jogler C (2017) Determining the bacterial cell biology of Planctomycetes. Nat Commun 8:14853
Boersma A, Kallscheuer N, Wiegand S, Rast R, Peeters S, Mesman R, Heuer A, Boedeker C, Jetten M, Rohde M, Jogler M, Jogler C (2019) Alienimonas californiensis gen. nov. sp. nov., a novel Planctomycete isolated from the kelp forest in Monterey Bay. Antonie van Leeuwenhoek. https://doi.org/10.1007/s10482-019-01367-4
Bondoso J, Harder J, Lage OM (2013) rpoB gene as a novel molecular marker to infer phylogeny in Planctomycetales. Antonie van Leeuwenhoek 104:477–488
Bondoso J, Balague V, Gasol JM, Lage OM (2014) Community composition of the Planctomycetes associated with different macroalgae. FEMS Microbiol Ecol 88:445–456
Bondoso J, Albuquerque L, Nobre MF, Lobo-da-Cunha A, da Costa MS, Lage OM (2015) Roseimaritima ulvae gen. nov., sp. nov. and Rubripirellula obstinata gen. nov., sp. nov. two novel planctomycetes isolated from the epiphytic community of macroalgae. Syst Appl Microbiol 38:8–15
Bondoso J, Godoy-Vitorino F, Balague V, Gasol JM, Harder J, Lage OM (2017) Epiphytic Planctomycetes communities associated with three main groups of macroalgae. FEMS Microbiol Ecol 93:fiw255
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552
Dedysh SN, Kulichevskaya IS, Beletsky AV, Ivanova AA, Rijpstra WIC, Damsté JSS, Mardanov AV, Ravin NV (2019) Lacipirellula parvula gen. nov., sp. nov., representing a lineage of planctomycetes widespread in low-oxygen habitats, description of the family Lacipirellulaceae fam. nov. and proposal of the orders Pirellulales ord. nov., Gemmatales ord. nov. and Isosphaerales ord. nov. Syst Appl Microbiol 43: 126050
Dedysh SN, Henke P, Ivanova AA, Kulichevskaya IS, Philippov DA, Meier-Kolthoff JP, Goker M, Huang S, Overmann J (2020) 100-year-old enigma solved: identification, genomic characterization and biogeography of the yet uncultured Planctomyces bekefii. Environ Microbiol 22:198–211
Devos DP, Reynaud EG (2010) Evolution. Intermediate steps. Science 330:1187–1188
Devos DP, Jogler C, Fuerst JA (2013) The 1st EMBO workshop on PVC bacteria-Planctomycetes-Verrucomicrobia-Chlamydiae superphylum: exceptions to the bacterial definition? Antonie Van Leeuwenhoek 104:443–449
Devos DP (2014) PVC bacteria: variation of, but not exception to, the Gram-negative cell plan. Trends Microbiol 22:14–20
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Frank O, Michael V, Pauker O, Boedeker C, Jogler C, Rohde M, Petersen J (2014) Plasmid curing and the loss of grip - The 65-kb replicon of Phaeobacter inhibens DSM 17395 is required for biofilm formation, motility and the colonization of marine algae. Syst Appl Microbiol 38:120–127
Fuerst JA, Sagulenko E (2011) Beyond the bacterium: Planctomycetes challenge our concepts of microbial structure and function. Nat Rev Microbiol 9:403–413
Fuerst JA, Webb RI (1991) Membrane-bounded nucleoid in the eubacterium Gemmata obscuriglobus. Proc Natl Acad Sci USA 88:8184–8188
Graça AP, Calisto R, Lage OM (2016) Planctomycetes as Novel Source of Bioactive Molecules. Front Microbiol 7:1241
Jeske O, Jogler M, Petersen J, Sikorski J, Jogler C (2013) From genome mining to phenotypic microarrays: Planctomycetes as source for novel bioactive molecules. Antonie Van Leeuwenhoek 104:551–567
Jeske O, Schüler M, Schumann P, Schneider A, Boedeker C, Jogler M, Bollschweiler D, Rohde M, Mayer C, Engelhardt H, Spring S, Jogler C (2015) Planctomycetes do possess a peptidoglycan cell wall. Nat Commun 6:7116
Jeske O, Surup F, Ketteniß M, Rast P, Förster B, Jogler M, Wink J, Jogler C (2016) Developing techniques for the utilization of Planctomycetes as producers of bioactive molecules. Front Microbiol 7:1242
Jogler C, Glöckner FO, Kolter R (2011) Characterization of Planctomyces limnophilus and development of genetic tools for its manipulation establish it as a model species for the phylum Planctomycetes. Appl Environ Microbiol 77:5826–5829
Jogler C, Waldmann J, Huang X, Jogler M, Glöckner FO, Mascher T, Kolter R (2012) Identification of proteins likely to be involved in morphogenesis, cell division, and signal transduction in Planctomycetes by comparative genomics. J Bacteriol 194:6419–6430
Jogler M, Jogler C (2013) Towards the development of genetic tools for Planctomycetes. In: Fuerst JA (ed) Planctomycetes: cell structure, origins and biology. Springer, Berlin, pp 141–164
Jogler C (2014) The bacterial ‘mitochondrium’. Mol Microbiol 94:751–755
Kallscheuer N, Jogler M, Wiegand S, Peeters SH, Heuer A, Boedeker C, Jetten MS, Rohde M, Jogler C (2019a) Rubinisphaera italica sp. nov. isolated from a hydrothermal area in the Tyrrhenian Sea close to the volcanic island Panarea. Antonie van Leeuwenhoek, https://doi.org/10.1007/s10482-019-01329-w
Kallscheuer N, Jogler M, Wiegand S, Peeters SH, Heuer A, Boedeker C, Jetten MS, Rohde M, Jogler C (2019b) Three novel Rubripirellula species isolated from plastic particles submerged in the Baltic Sea and the estuary of the river Warnow in northern Germany. Antonie van Leeuwenhoek. https://doi.org/10.1007/s10482-019-01368-3
Kallscheuer N, Wiegand S, Jogler M, Boedeker C, Peeters SH, Rast P, Heuer A, Jetten MSM, Rohde M, Jogler C (2019c) Rhodopirellula heiligendammensis sp. nov., Rhodopirellula pilleata sp. nov., and Rhodopirellula solitaria sp. nov. isolated from natural or artificial marine surfaces in Northern Germany and California, USA, and emended description of the genus Rhodopirellula. Antonie van Leeuwenhoek, https://doi.org/10.1007/s10482-019-01366-5
Kallscheuer N, Wiegand S, Peeters SH, Jogler M, Boedeker C, Heuer A, Rast P, Jetten MSM, Rohde M, Jogler C (2019d) Description of three bacterial strains belonging to the new genus Novipirellula gen. nov., reclassificiation of Rhodopirellula rosea and Rhodopirellula caenicola and readjustment of the genus threshold of the phylogenetic marker rpoB for Planctomycetaceae. Antonie van Leeuwenhoek, https://doi.org/10.1007/s10482-019-01374-5
Kallscheuer N, Wiegand S, Heuer A, Rensink S, Boersma AS, Jogler M, Boedeker C, Peeters SH, Rast P, Jetten MS, Rohde M, Jogler C (2020) Blastopirellula retiformator sp. nov. isolated from the shallow-sea hydrothermal vent system close to Panarea Island. Antonie van Leeuwenhoek. https://doi.org/10.1007/s10482-019-01377-2
Kohn T, Heuer A, Jogler M, Vollmers J, Boedeker C, Bunk B, Rast P, Borchert D, Glöckner I, Freese HM, Klenk HP, Overmann J, Kaster AK, Wiegand S, Rohde M, Jogler C (2016) Fuerstia marisgermanicae gen. nov., sp. nov., an unusual member of the phylum Planctomycetes from the German Wadden Sea. Front Microbiol 7: 2079
Kohn T, Wiegand S, Boedeker C, Rast P, Heuer A, Jetten M, Schüler M, Becker S, Rohde C, Müller R-W, Rohde M, Engelhardt H, Jogler M, Jogler C (2019) Planctopirus ephydatiae, a novel Planctomycete isolated from a freshwater sponge. Syst Appl Microbiol 43:126022
König E, Schlesner H, Hirsch P (1984) Cell wall studies on budding bacteria of the Planctomyces/Pasteuria group and on a Prosthecomicrobium sp. Arch Microbiol 138:200–205
Konstantinidis KT, Tiedje JM (2005) Genomic insights that advance the species definition for prokaryotes. Proc Natl Acad Sci USA 102:2567–2572
Konstantinidis Rodriguez-R,LM, KT (2016) The enveomics collection: a toolbox for specialized analyses of microbial genomes and metagenomes. PeerJ Preprints 4: e1900v1
Kulichevskaya IS, Serkebaeva YM, Kim Y, Rijpstra IC, Sinninghe Damste JS, Liesack W, Dedysh SN (2012) Telmatocola sphagniphila gen. nov., sp. nov., a novel dendriform planctomycete from northern wetlands. Front Microbiol 3:146
Lachnit T, Fischer M, Kunzel S, Baines JF, Harder T (2013) Compounds associated with algal surfaces mediate epiphytic colonization of the marine macroalga Fucus vesiculosus. FEMS Microbiol Ecol 84:411–420
Lage OM, Bondoso J (2014) Planctomycetes and macroalgae, a striking association. Front Microbiol 5:267
Lechner M, Findeiss S, Müller L, Marz M, Stadler P, Prohaska S (2011) Proteinortho: Detection of (Co)Orthologs in Large-Scale Analysis. BMC Bioinformatics 12:124
Lee I, Ouk Kim Y, Park SC, Chun J (2016) OrthoANI: An improved algorithm and software for calculating average nucleotide identity. Int J Syst Evol Microbiol 66:1100–1103
Lindsay MR, Webb RI, Fuerst JA (1997) Pirellulosomes: A new type of membrane-bounded cell compartment in planctomycete bacteria of the genus Pirellula. Microbiology-UK 143:739–748
Lonhienne TG, Sagulenko E, Webb RI, Lee KC, Franke J, Devos DP, Nouwens A, Carroll BJ, Fuerst JA (2010) Endocytosis-like protein uptake in the bacterium Gemmata obscuriglobus. Proc Natl Acad Sci USA 107:12883–12888
Luo C, Rodriguez RL, Konstantinidis KT (2014) MyTaxa: an advanced taxonomic classifier for genomic and metagenomic sequences. Nucleic Acids Res 42:e73
Neumann S, Wessels HJ, Rijpstra WI, Sinninghe Damste JS, Kartal B, Jetten MS, van Niftrik L (2014) Isolation and characterization of a prokaryotic cell organelle from the anammox bacterium Kuenenia stuttgartiensis. Mol Microbiol 94:794–802
Overmann J, Abt B, Sikorski J (2017) Present and Future of Culturing Bacteria. Annu Rev Microbiol 71:711–730
Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW (2015) CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res 25:1043–1055
Peeters SH, van Niftrik L (2018) Trending topics and open questions in anaerobic ammonium oxidation. Curr Opin Chem Biol 49:45–52
Peeters SH, Wiegand S, Kallscheuer N, Jogler M, Heuer A, Jetten MSM, Rast P, Boedeker C, Rohde M, Jogler C (2019) Three marine strains constitute the novel genus and species Crateriforma conspicua in the phylum Planctomycetes. Antonie van Leeuwenhoek. https://doi.org/10.1007/s10482-019-01375-4
Pilhofer M, Rappl K, Eckl C, Bauer AP, Ludwig W, Schleifer KH, Petroni G (2008) Characterization and evolution of cell division and cell wall synthesis genes in the bacterial phyla Verrucomicrobia, Lentisphaerae, Chlamydiae, and Planctomycetes and phylogenetic comparison with rRNA genes. J Bacteriol 190:3192–3202
Pruesse E, Peplies J, Glöckner FO (2012) SINA: accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 28:1823–1829
Qin QL, Xie BB, Zhang XY, Chen XL, Zhou BC, Zhou J, Oren A, Zhang YZ (2014) A proposed genus boundary for the prokaryotes based on genomic insights. J Bacteriol 196:2210–2215
Rast P, Glockner I, Boedeker C, Jeske O, Wiegand S, Reinhardt R, Schumann P, Rohde M, Spring S, Glockner FO, Jogler C, Jogler M (2017) Three Novel Species with Peptidoglycan Cell Walls form the New Genus Lacunisphaera gen. nov. in the Family Opitutaceae of the Verrucomicrobial Subdivision 4. Front Microbiol 8:202
Rensink S, Wiegand S, Kallscheuer N, Rast P, Peeters SH, Heuer A, Boedeker C, Jetten MS, Rohde M, Jogler M, Jogler C (2020) Description of the novel planctomycetal genus Bremerella, containing Bremerella volcania sp. nov., isolated from an active volcanic site, and reclassification of Blastopirellula cremea as Bremerella cremea comb. nov. Antonie van Leeuwenhoek, https://doi.org/10.1007/s10482-019-01378-1
Reva O, Tümmler B (2008) Think big–giant genes in bacteria. Environ Microbiol 10:768–777
Rivas-Marin E, Canosa I, Santero E, Devos DP (2016) Development of Genetic Tools for the Manipulation of the Planctomycetes. Front Microbiol 7:914
Rueden CT, Schindelin J, Hiner MC, DeZonia BE, Walter AE, Arena ET, Eliceiri KW (2017) ImageJ2: ImageJ for the next generation of scientific image data. BMC Bioinformatics 18:529
Santarella-Mellwig R, Pruggnaller S, Roos N, Mattaj IW, Devos DP (2013) Three-dimensional reconstruction of bacteria with a complex endomembrane system. PLoS Biol 11:e1001565
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7:539
Slobodkina GB, Kovaleva OL, Miroshnichenko ML, Slobodkin AI, Kolganova TV, Novikov AA, van Heerden E, Bonch-Osmolovskaya EA (2015) Thermogutta terrifontis gen. nov., sp. nov. and Thermogutta hypogea sp. nov., thermophilic anaerobic representatives of the phylum Planctomycetes. Int J Syst Evol Microbiol 65:760–765
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313
Strous M, Fuerst JA, Kramer EH, Logemann S, Muyzer G, van de Pas-Schoonen KT, Webb R, Kuenen JG, Jetten MS (1999) Missing lithotroph identified as new planctomycete. Nature 400:446–449
van Teeseling MC, Mesman RJ, Kuru E, Espaillat A, Cava F, Brun YV, VanNieuwenhze MS, Kartal B, van Niftrik L (2015) Anammox Planctomycetes have a peptidoglycan cell wall. Nat Commun 6:6878
Vollmers J, Frentrup M, Rast P, Jogler C, Kaster AK (2017) Untangling Genomes of Novel Planctomycetal and Verrucomicrobial Species from Monterey Bay Kelp Forest Metagenomes by Refined Binning. Front Microbiol 8:472
Wagner M, Horn M (2006) The Planctomycetes, Verrucomicrobia, Chlamydiae and sister phyla comprise a superphylum with biotechnological and medical relevance. Curr Opin Biotechnol 17:241–249
Wiegand S, Jogler M, Jogler C (2018) On the maverick Planctomycetes. FEMS Microbiol Rev 42:739–760
Wiegand S, Jogler M, Boedeker C, Pinto D, Vollmers J, Rivas-Marín E, Kohn T, Peeters SH, Heuer A, Rast P, Oberbeckmann S, Bunk B, Jeske O, Meyerdierks A, Storesund JE, Kallscheuer N, Lücker S, Lage OM, Pohl T, Merkel BJ, Hornburger P, Müller R-W, Brümmer F, Labrenz M, Spormann AM, Op den Camp HJM, Overmann J, Amann R, Jetten MSM, Mascher T, Medema MH, Devos DP, Kaster A-K, Øvreås, L, Rohde, M, Galperin, MY, Jogler, C, 2019. Cultivation and functional characterization of 79 planctomycetes uncovers their unique biology. Nature Microbiology.Wiegand, Jogler S, M, Boedeker, C, Pinto, D, Vollmers, J, Rivas-Marín, E, Kohn, T, Peeters, SH, Heuer, A, Rast, P (2020) Cultivation and functional characterization of 79 Planctomycetes uncovers their unique biology. Nat Microbiology 5: 126–140
Yarza P, Yilmaz P, Pruesse E, Glöckner FO, Ludwig W, Schleifer K-H, Whitman WB, Euzéby J, Amann R, Rosselló-Móra R (2014) Uniting the classification of cultured and uncultured bacteria and archaea using 16S rRNA gene sequences. Nat Rev Microbiol 12:635
Acknowledgements
Open Access funding provided by Projekt DEAL. Part of this research was funded by the Deutsche Forschungsgemeinschaft grants KA 4967/1–1 and JO 893/4 − 1, Grant ALWOP.308 of the Nederlandse Organisatie voor Wetenschappelijk Onderzoek (NWO), SIAM (Soehngen Institute for Anaerobic Microbiology) Grant No. 024002002 and the Radboud Excellence fellowship. We thank Ina Schleicher for skillful technical assistance. Brian Tindall and Regine Fähnrich from the DSMZ as well as the BCCM/LMG Bacteria collection we thank for support during strain deposition. We thank our collaborators Sonja Oberbeckmann and Matthias Labrenz (IOW Warnemünde, Germany) for sampling support.
Author information
Authors and Affiliations
Contributions
S.H.P. and S.W. wrote the manuscript, analysed the data and prepared the figures, S.W. performed the genomic and phylogenetic analysis, A.H. and M.J. isolated the strains and performed the initial cultivation and strain deposition, S.H.P. and C.B. performed the light microscopic analysis, N.K. and M.S.M.J. contributed to text preparation and revised the manuscript, M.R. performed the electron microscopic analysis. C.J. supervised A.H. and the study. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethicals statement
This article does not contain any studies with animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Peeters, S.H., Wiegand, S., Kallscheuer, N. et al. Description of Polystyrenella longa gen. nov., sp. nov., isolated from polystyrene particles incubated in the Baltic Sea. Antonie van Leeuwenhoek 113, 1851–1862 (2020). https://doi.org/10.1007/s10482-020-01406-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-020-01406-5