Abstract
A pleomorphic Gram-negative, motile coccobacillus was isolated from the gills of a wild-caught bluespotted ribbontail ray after its sudden death during quarantine. Strain 141012304 was observed to grow aerobically, to be clearly positive for cytochrome oxidase, catalase, urease and was initially identified as “Brucella melitensis” or “Ochrobactrum anthropi” by Matrix-assisted laser desorption/ionization-time of flight mass spectrometry and VITEK2-compact®, respectively. Affiliation to the genus Brucella was confirmed by bcsp31 and IS711 PCR as well as by Brucella species-specific multiplex PCR, therein displaying a characteristic banding pattern recently described for Brucella strains obtained from amphibian hosts. Likewise, based on recA sequencing, strain 141012304 was found to form a separate lineage, within the so called ‘atypical’ Brucella, consisting of genetically more distantly related strains. The closest similarity was detected to brucellae, which have recently been isolated from edible bull frogs. Subsequent next generation genome sequencing and phylogenetic analysis confirmed that the ray strain represents a novel Brucella lineage within the atypical group of Brucella and in vicinity to Brucella inopinata and Brucella strain BO2, both isolated from human patients. This is the first report of a natural Brucella infection in a saltwater fish extending the host range of this medically important genus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Brucellosis is a significant public health threat and an estimated number of approximately 500,000 novel human infections per year mark it as one of the most important bacterial zoonoses worldwide (Godfroid et al. 2005). Historically, the genus comprises six so called ‘classical’ Brucella species (the type species Brucella melitensis and Brucella abortus, Brucella canis, Brucella ovis, Brucella neotomae and Brucella suis) albeit novel members have been added to this group in recent years including Brucella pinnipedialis and Brucella ceti from marine ecosystems and Brucella papionis from baboons (Foster et al. 2007; Whatmore et al. 2014). All classical Brucella species are genetically highly related to each other with average core genome nucleotide identities (ANI) of >99 % and harbour identical 16S rRNA and recA gene sequences (Scholz et al. 2016b).
In contrast, the recently described members Brucella microti, Brucella inopinata and Brucella vulpis belong to the informally named ‘atypical’ group of the genus, exhibiting either atypical phenotypic traits (B. inopinata, B. microti) and/or represent genetically more distantly related species (B. inopinata, B. vulpis). B. microti exhibits a unique position as it is phenotypically clearly different from the classical Brucella species but genetically nearly identical to B. suis (Audic et al. 2009). All ‘atypical’ Brucella species still share average nucleotide identities values of 98 % compared with the type species, B. melitensis, but, with the exception of B. microti, differ by 2–5 nucleotides in their 16S rRNA and recA gene sequences (Scholz et al. 2010; Scholz et al. 2016b). Furthermore, their genomes commonly carry additional genetic material not found in ‘classical’ Brucella species but present in other soil associated bacteria of the Alphaproteobacteria. Most of the accessory genes with known function encode for additional metabolic proteins, ABC transporters or represent bacteriophages and mobile genetic elements, which indicates a different ecology in comparison to the classical host-adapted Brucella species (Scholz et al. 2016a). In addition to the validly named species, some candidates are awaiting species description, which were recently isolated from different homoeothermic (McDonald et al. 2006; Tiller et al. 2010a, b) as well as poikilothermic host species (Eisenberg et al. 2012; Fischer et al. 2012; Whatmore et al. 2015; Soler-Lloréns et al. 2016). Based on their biochemical and molecular profiles, these potential novel Brucella species are also members of the ‘atypical’ Brucella group (Wattam et al. 2012; Mühldorfer et al. 2016; Scholz et al. 2016a). Two members of the atypical group, B. inopinata and Brucella sp. strain BO2 were isolated from diseased human patients. An overview of the ‘atypical’ Brucella species is given in Table 1.
Human brucellosis has always been attributed to an animal reservoir (Godfroid et al. 2005), although data for the reservoir hosts and zoonotic potential of the novel members of the genus is missing.
The present report focusses on the characterisation of another novel member of the ‘atypical’ Brucella group, recently isolated from the gills of a wild-caught bluespotted ribbontail ray. To the best of our knowledge this represents the first identification of a distinct Brucella sp. from a saltwater fish, thereby furthering the knowledge on Brucella in cold-blooded hosts.
Materials and methods
Case description
A recently introduced group of five wild-caught adult bluespotted ribbontail rays (Taeniura lymma) was housed in an aquarium enclosure for quarantine reasons in a German zoo. The rays had been caught and imported from Bali and were at first maintained together with two bluestreak cleaner wrasses (Labroides dimidiatus) and a yellow tang (Zebrasoma flavescens) in a nearby German university’s animal facility. At the zoo, the rays were kept in a 500 L quarantine aquarium with 35 g/L sodium chloride at 25.5–26.0 °C water temperature and external metal-vapour discharge lamp illumination. The aquarium was equipped with a separate filter system and some hiding tubes as the only furnishing. The rays had never been housed together with or near to amphibians nor came into contact with amphibian waste waters. Neither in the breeding facility nor during quarantine were zookeepers at the same time simultaneously responsible for fish and amphibian care. From six months after arrival at the zoo, all animals died over a period of four months, but because of advanced decay, necropsy was performed only from two individuals. Animal husbandry fulfilled ethical standard guidelines according to the code of ethics and animal welfare of the World Association of Zoos and Aquariums (http://www.waza.org/files/webcontent/1.public_site/5.conservation/code_of_ethics_and_animal_welfare/Code%20of%20Ethics_EN.pdf).
Phenotypic characterisation
Bacterial isolation and biochemical identification
Bacterial isolates were obtained during post mortem examination. Native tissue samples were processed for bacterial culture. Briefly, organ samples were flame sterilised and the surface of a fresh cut was directly inoculated onto culture media. Agar plates were incubated for up to 48 h at 20 °C using aerobic conditions (Columbia agar with 5 % sheep blood [SBA; Oxoid, Wesel, Germany] and Gassner agar; [VWR, Darmstadt, Germany]). Phenotypic characterisation was performed by standard microbiological procedures: bacterial colonies were tested for haemolytic properties on SBA, catalase activity with 3 % H2O2 on microscopic slides and for presence of cytochrome oxidase with the BBL DrySlide® Oxidase system (Becton–Dickinson, Heidelberg, Germany). Urease, hydrogen sulfide, indole, motility and oxidative and fermentative glucose assimilation were tested on Christensen agar, SIM and OF medium in slant agar tubes, respectively (all Merck, Darmstadt, Germany). Microscopic examinations of fixed smears were performed using Gram’s stain. For further identification the VITEK2-compact® with GN card system for Gram-negative bacteria (bioMérieux, Nürtingen, Germany) was used. Subcultures were also incubated on Brucella-agar with and without crystal violet (Oxoid) under aerobic and capnophilic conditions at different incubation temperatures (20 and 37 °C).
Identification by matrix-assisted laser desorption/ionization-time of flight mass spectrometry (MALDI-TOF MS)
Bacterial isolates were selected from the culture plates and then applied to steel-targets according to manufacturer’s instructions (BrukerBiotyper, BrukerDaltonics, Bremen, Germany). Isolates were prepared using both a direct transfer and an acetonitrile-formic acid extraction protocol provided by the manufacturer and analysed on a Bruker Microflex LT system by MALDI-TOF MS using Biotyper Version V3.3.1.0. The standard and the ‘Security Relevant’ (SR) database used (DB 5627, BrukerDaltonics) comprised spectra from 6 different B. melitensis strains. The MALDI Biotyper real-time classification (RTC) software considers MALDI scores >2.3 and >2.0 as secure species and genus identification levels, respectively. The identification was repeated three times to verify the original findings. Furthermore, identification of the concomitant bacterial flora was also carried out by MALDI-TOF MS.
Cluster analysis of infrared spectra of strains obtained by Fourier transform-infrared spectroscopy (FT-IR)
The ray strain from this study, as well as the strains used for comparison, was cultivated independently in three replicates at 37 °C for 24 h on SBA. Harvesting of cells, preparation of bacteria films on zinc selenide plates, drying and handling were performed as described previously (Kuhm et al. 2009). The dried bacterial films were used directly for examination by FT-IR spectroscopy. Infrared (IR) spectra were recorded for each sample in a transmission mode from 500 to 4000 cm−1 with an FT-IR spectrometer (Tensor27 with HTS-XT-module, Bruker Optics, Ettlingen, Germany). Acquisition and first analysis of data were carried out using OPUS Software (version 4.2, Bruker Optics). IR spectra from the ray strain from this study as well as from reference strains of ‘atypical’ Brucella strains, B. suis and Ochrobactrum intermedium were compared by cluster analysis (Helm et al. 1991), a comparative method based on species or even strain specific spectral differences in their patterns of biomolecules. For cluster analysis, the second derivation of the vector normalised spectra in the wave number range of 870–1760 and 2780–3100 cm−1 were used for calculation with Ward’s algorithm [OPUS 4.2; (Ward 1963)].
Genotypic characterisation
PCR analysis
The Brucella genus specific bscp31 gene was amplified according to Baily et al. (1992). Amplification of the Brucella specific insertion sequence 711 (IS711) was done as described by Bricker and Halling (1994). AMOS-PCR was carried out following the methodology of Bricker and Halling (1994). For identification at the species level, a muliplex PCR assay for the identification and differentiation of all Brucella species was performed according to the method described by Garcia-Yoldi et al. (2006).
Genome sequencing and phylogenetic reconstruction
The genome sequence of strain 141012304 was determined using the PacBio (Pacific Biosiences, Menio Park, USA) sequencing platform. Briefly, DNA was extracted using a Qiagen spin column kit (Qiagen, Hilden, Germany). Genome sequencing was carried out by a sequencing company (GATC, Konstanz, Germany). De novo assembly of the PacBio generated reads (1 SMRT cell) was done using the freely available SMRT analysis software (v. 2.3.0) (http://www.pacb.com/devnet/) and the HGAP3 algorithm, with a read length minimum of 2500 bp. Genes that have no orthologues (singletons) in classical Brucella species were calculated using the singletons option of the EDGAR platform (available at http://edgar.computational.bio) and an in-house Brucella genome database consisting of all known classical Brucella species. Singletons were further processed and analysed by MG-RAST metagenomics analysis server (metagenomics.anl.gov) to detect similarities to other microorganisms and evidence for their potential origin and function.
A core genome-based phylogenetic tree was constructed as described previously using the EDGAR platform (Blom et al. 2016). Briefly, the core genome with B. melitensis 16MT as the reference genome and the type strains and biovars of various Brucella species was calculated using the implemented function of EDGAR. The core genome consisted of 2316 coding sequences (CDS) per genome. Multiple alignments of the nucleotide coding sequences or their translated products were created for all core genes using MUSCLE (Edgar 2004). The gene set alignments were concatenated to one large multiple alignment. Finally, a phylogenetic tree (amino acid-based) was generated using the F84 (DNA) or Kimura (AA) distance matrix and the neighbour joining method with 200 repetitions as implemented in PHYLIP. Genome sequences used in this project were retrieved from either http://www.ebi.ac.uk/genomes/ or https://www.patricbrc.org.
RecA gene analysis
The recA gene sequence of the ray strain was extracted from the genome sequence and compared with the consensus recA gene sequences of all classical species and members of the ‘atypical’ Brucella group, comprising a set of 36 frog strains, two strains from Australian rodents (83–13 and NF 2653), B. inopinata BO1T, strain BO2 and B. vulpis F60T. The recA gene sequences of B. inopinata BO1T, strain BO2 and the Australian rodent strains were extracted from the genome sequences available at http://patric.vbi.vt.edu/. The recA sequences of the 36 frog strains were obtained from Scholz et al. (2016a). Briefly, partial recA sequences (628 bp) were aligned using MUSCLE implemented in MEGA v6 (Tamura et al. 2013). Phylogenetic reconstructions were performed using the neighbor joining (Kimura 2-parameter substitution model) and maximum likelihood (Jukes-Cantor) methods of MEGA with 1000 repetitions. The type strains of Ochrobactrum anthropi and O. intermedium served as outgroup. Trees were exported in newick format and further processed using the software figtree, available from http://tree.bio.ed.ac.uk/software/figtree/.
Calculation and analysis of singletons unique for the ray isolate
Genes unique in strain 141012304 from this study but not present in the ‘classical’ Brucella species were calculated using the singleton function of EDGAR (Suppl. Table S1). A second calculation was done that also included genome sequences of B. inopinata BO1T, strain BO2, B. vulpis and members of Ochrobactrum in order to determine genes unique to strain 141012304 only (Suppl. Table S2).
Identified singletons were further compared with the RefSeq database using the BLAST+ implementation of BLAST P with an initial e-value cutoff of 1e−10 (Camacho et al. 2009; Tatusova et al. 2014). Results were filtered for the best two hits of every query and the annotation and source organism for these best hits were extracted from the database.
Results
Phenotypic characterisation
Presumptive and biochemical identification
The ray sampled in this study was highly decomposed and tissue samples yielded a heavy growth of Vibrio harveyi. From the gills, a pleomorphic Gram-negative coccobacillus (strain 141012304) could be cultivated on SBA as regular, slightly moist, whitish, non-haemolytic colonies, measuring approximately 1–2 mm in diameter after 48 h. The strain was found to be motile, positive for cytochrome oxidase and catalase and displayed a prompt and strong urease activity (within 20–30 min). Oxidative and fermentative glucose metabolism and H2S production were found to be inactive and negative, respectively. No growth on Gassner’s agar was observed after 24–72 h, but the isolate was observed to grow well on Brucella-agar plates with and without crystal violet, both at 20 and 37 °C within 24 h. In general, better growth was observed in an aerobic atmosphere as compared with CO2-incubation. Biochemical identification using the VITEK2-compact® system yielded ‘Ochrobactrum anthropi’ with an accordance of 97 % (bioprofile number 1001205350701001).
MALDI-TOF MS
Using MALDI-TOF mass spectrometry and the BioTyper S/R database strain 141012304 was unequivocally identified to the species level as B. melitensis with score levels of 2.30 to 2.35 using both a direct transfer and an acetonitrile-formic acid extraction protocol in sample preparation. Following the manual addition of respective spectra from the ray strain as well as from other ‘atypical’ Brucella strains to the database, all these strains could be clearly differentiated from database entries of B. melitensis and O. intermedium (Fig. 1). Spectra from the ray strain were indistinguishable from spectra from other ‘atypical’ Brucella strains obtained from amphibians.
Dendrogram including main spectra peak lists (msp) of Ochrobactrum intermedium LMG 3301T HAM and Brucella melitensis 5520 available in the Bruker Taxonomy database; spectra of ‘atypical’ brucellae from a bluespotted ribbontail ray from this study (in bold) as well as from various amphibians [(Mühldorfer et al. 2016); see Table 1] were measured using an extraction protocol provided by the manufacturer. The dendrogram was generated using the BioTyper msp Dendrogram Creation Standard Method (version 1.4) of the MALDI Biotyper OC Software (version 3.1, build 66). The database used (DB 5627, Bruker Daltonics) comprised 24 spectra from B. melitensis 5520; Ttype strain, HAM Harmsen strain collection, Uni Münster, Germany
Characterisation of Brucella spp. by FT-IR
Comparison of the IR spectra from the ray strain, as well as from other ‘atypical’ Brucella strains, showed a clear separation of the strains from cold-blooded animals from B. suis and O. intermedium (Suppl. Fig. S1). Again, the spectrum from the ray strain was indistinguishable from the spectra from other ‘atypical’ Brucella strains obtained from amphibians.
Molecular characterisation
PCR analysis
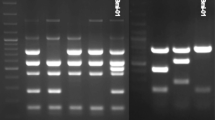
Strain 141012304 was unequivocally identified as a member of the genus Brucella by amplification of the Brucella-specific markers bcsp31 and IS711. A species differentiating multiplex-PCR (Bruceladder-PCR) revealed a pattern consisting of five fragments (152, 272, 450, 587 and 794 bp) also found among Brucella isolated from frogs (Eisenberg et al. 2012). No PCR product was amplified using the AMOS-PCR.
Analysis of the ray isolate genome
De novo assembly of the PacBio sequence reads obtained from strain 141012304 resulted in a high quality genome with an average coverage of 120x, comprising two chromosomes as single contigs of 2,145,953 bp and 1,296,428 bp, respectively. No plasmid was detected. The GC content was determined to be 57.1 %, which is in agreement with other Brucella species. The calculated average nucleotide and amino acid identities compared with the type species B. melitensis 16MT were 97.5 and 98.45 %, respectively (not shown), confirming affiliation to the genus Brucella.
The entire virB operon (Vir1-11) is present on chromosome 2 of strain 141012304 (peg 1100–peg 1110). In classical Brucella species, this operon encodes for a type IV secretion system which is induced intracellularly in macrophages and therefore is essential for lethality of Brucella. As in B. inopinata and strain BO2, a gene cluster encoding a system for L-rhamnose uptake and catabolism (L-rhamnose mutarotase, dTDP-4-dehydrorhamnose 3,5-epimerase) is present in close proximity to flagellar genes (Wattam et al. 2012). The wbk operon, encoding the perosamine-based O-antigen in classical Brucella species, is absent and therefore suggests an altered LPS pathway as described for strain BO2 (Wattam et al. 2012).
Genes present in strain 141012304 but absent in ‘classical’ Brucella species
The singleton calculation revealed a total of 269 genes (~10 % of the entire genome) shared among the ray strain and the ‘atypical’ Brucella group including members of Ochrobactrum which were absent from the ‘classical’ Brucella species (see Suppl. Table S1). This showed that genes not found in the ‘classical’ Brucella species are frequently shared among the ‘atypical’ Brucella group and Ochrobactrum genomes. Among these genes, 132 were unique for the ray isolate (see Suppl. Table S2). As determined by a subsequent BLAST search, most of the singletons were of hypothetical function. From genes with known functions, the majority coded for ABC transporters and proteins of metabolic function. Several phage-related genes and transposases not found in the classical species were also present. High similarities, with identities of up to 100 %, were detected with corresponding sequences of B. inopinata, strain BO2 and members of Ochrobactrum (see Suppl. Table S1). High sequence similarities were also detected with members of Rhizobium, Mesorhizobium and Agrobacterium in particular, with several mobile genetic elements (insertion sequences, transposases).
The BLAST search also revealed that various genes were organised in gene clusters. Major clusters consisted of phage related genes. One cluster consisting of nine genes (peg752-peg761) was located on a 12.7 kb fragment on chromosome one and encodes an ectoine uptake system (via ABC transporter) with high similarity (>95 %) to that of O. intermedium (see Suppl. Table S1, marked in gray) (Soler-Lloréns et al. 2016). Ectoine and hydroxyectoine are used by various bacteria as protectants against osmotic and cold stress (Kuhlmann et al. 2011).
Phylogenetic analysis
The core genome calculated for strain 141012304 together with the reference strains of all Brucella species and selected ‘atypical’ strains consists of 2316 coding sequences. The phylogenetic reconstruction based on these genes confirmed that strain 141012304 belongs to the ‘atypical’ group of Brucella species with closest placement to B. inopinata and strain BO2, both of which were isolated from human patients (Fig. 2). This clade also comprised two isolates from Australian rodents (83–13 and NF 2653) as well as B. vulpis, the genetically most diverse Brucella species. B. microti was confirmed as being a member of the ‘classical’ species with a basal position, despite its distinctive phenotype.
Core-genome-based phylogenetic neighbor-joining tree with 200 repetitions. Bar 0.002 substitutions per site. The bluespotted ribbontail ray strain 141012304 from this study is indicated in bold letters. Accession numbers are given in brackets. For details on ‘atypical’ brucellae from various amphibians we refer to (Mühldorfer et al. 2016; Scholz et al. 2016a) and Table 1
RecA gene analysis
In contrast to the ‘classical’ species harbouring identical recA gene sequences, analysis of the recA genes of a comprehensive collection of ‘atypical’ Brucella strains revealed an unexpected high degree of heterogeneity (Fig. 3). Strains from cold-blooded animals group into at least nine different clades, of which two share higher sequence similarities with the ‘classical’ Brucella group. The recA sequence of the ray isolate from this study is distantly related to ‘classical’ members and shows higher similarities to some isolates obtained from edible bull frogs.
Phylogenetic tree from maximum likelihood analysis of the recA gene alignment of various classical and ‘atypical’ Brucella strains including the one from a bluespotted ribbontail ray from this study (bold) and from various amphibians (see Table 1). The tree was calculated with 100 bootstrap repetitions. Ochrobactrum anthropi and O. intermedium served as outgroup. Bar 0.01 substitutions per site. Accession numbers are given in brackets
Nucleotide sequence accession numbers and deposition
The genome sequences of isolate 141012304 have been submitted to the GenBank database under accession nos. LT605585 and LT605586 (study ID PRJEB15074), respectively. Strain 141012304 has been deposited at DSM 103976.
Discussion
In contrast to ‘classical’ or ‘core’ Brucella species, which are genetically and phenotypically highly related to each other with average nucleotide identities of >99 %, an ‘atypical’ Brucella group was recently proposed (Wattam et al. 2012). This expanding group consists of genetically more distantly related members of the genus from various host species and geographical origins, together with the two validly named species B. inopinata and B. vulpis (Scholz et al. 2010; Scholz et al. 2016b). This group of ‘atypical’ brucellae comprises strains from Australian rodents (Tiller et al. 2010a), a set of 36 strains obtained from various frog species (Eisenberg et al. 2012; Fischer et al. 2012; Whatmore et al. 2015), and the B. inopinata-related strain BO2 which was isolated from a human patient with chronic destructive pneumonia (Tiller et al. 2010b).
Noteworthy, with the exception of B. microti, all members of the ‘atypical’ group studied to date are equipped with additional genetic information not found in ‘classical’ Brucella species, but present in other soil-associated bacteria such as members of Ochrobactrum, Paracoccus and Agrobacterium. Most of the accessory genes encode for metabolic functions, membrane associated proteins or ABC transporters, suggesting a reservoir in the environment. This also implies a different ecology compared with the group of the host adapted ‘classical’ species, most of which are highly pathogenic for humans (Soler-Lloréns et al. 2016). It seems likely that members of the ‘atypical’ group are widely distributed in the environment, infecting various animal species and occasionally humans. The wide distribution among amphibians from different continents and in the ray from this study is indicative of a water-born transmission route, an assumption that has to be proven by further studies. B. microti, despite its different phenotype, does not carry additional genetic information but is genetically nearly identical to B. suis (Audic et al. 2009). In this case the altered phenotype presumably relies on differential gene expression and not on a changed gene content.
Recently it was shown that isolates from amphibians are much more common than previously thought and that they form different clusters inside the overall Brucella phylogeny (Mühldorfer et al. 2016; Scholz et al. 2016a). Despite their close relationship to B. inopinata BO1T and strain BO2, both isolated from human patients with brucellosis, there is presently no direct evidence that isolates from frogs are infectious for humans or animals other than amphibians. On the other hand, pathologies due to Brucella infections have also been described in uncompromised amphibians (Shilton et al. 2008; Eisenberg et al. 2012; Fischer et al. 2012; Mühldorfer et al. 2016; Scholz et al. 2016a). In this study we identified another B. inopinata-like member of the atypical Brucella group, isolated from a bluespotted ribbontail ray, suggesting an even broader distribution of B. inopinata-related strains among various animal species worldwide.
Like other ‘atypical’ fast-growing brucellae, strain 141012304 displayed a biochemical profile similar to those of Ochrobactrum species (Shilton et al. 2008; Eisenberg et al. 2012; Whatmore et al. 2015; Mühldorfer et al. 2016). Consequently fast growing brucellae such as B. inopinata, strain BO2, the frog strains, and the strain from the present study are readily misidentified as Ochrobactrum sp. using commercially available biochemical identification systems (Mühldorfer et al. 2016). This diagnostic problem has been reported also for the ‘classical’ Brucella spp. (Elsaghir and James 2003) and may result in a risk of missing these medically important pathogens.
As with the amphibian strains, MALDI-TOF MS, which identifies bacteria based on their highly conserved ribosomal proteins, was able to identify the ray strain with high score levels to the correct genus and thus proved very suitable as an initial screening method, which may prevent misidentification of such isolates during diagnostic workflows. However, as a prerequisite the database used needs to contain the appropriate spectra of the genus Brucella (Cunningham and Patel 2013). An extended “security relevant” database version provided by the manufacturer helped to identify the ray strain as B. melitensis, which presently represents the only entry in the database. By introducing a representative set of spectra from different frog Brucella it became possible to display ‘atypical’ isolates in a separate cluster to the ‘core’ Brucella species represented by B. melitensis (Fig. 1). Furthermore, an affiliation of the ray strain to amphibian Brucella strains was also possible by FT-IR spectroscopy, as this technique derives signals from an even larger range of biomolecules (Suppl. Fig. S1).
The isolation of a novel Brucella strain from a marine cold-blooded vertebrate host again significantly extends the ecologic range of the genus and suggests that all vertebrate classes including reptiles may be suitable hosts for members of Brucella. With respect to fish, primary susceptibility including clinical disease has been already proven in Nile catfish (Clarias gariepinus) (Salem and Mohsen 1997), although infection could be linked to B. melitensis biovar 3 contamination during illegal carcass disposal (El-Tras et al. 2010). Raw seafood could also be a possible source for marine Brucella infections in humans (Hernandez-Mora et al. 2013) and fish might, furthermore, also serve as vectors for Brucella infections of marine mammals (Nymo et al. 2016). Importantly, novel members of the ‘atypical’ Brucella group not only behave like classical species in the murine infection model (Jiménez de Bagüés et al. 2014) but are even capable of inducing disease in humans (Tiller et al. 2010b), and, unless otherwise proven, should be considered as potential zoonotic pathogens.
Where and how the infection of the rays in this study took place remains elusive and if other individuals from the same group also suffered from Brucella infection. All the other individuals were reported to have died but were too putrid for pathological examination. It remains therefore speculative if the fish from this report really suffered from Brucella infection and if it already acquired these bacteria in the wild or came in contact with this microorganism during the first or final aquaculture. IS711 PCR testing of separate tissues revealed that only the gills were colonised with this Brucella strain (data not shown). Because of the timely detection within the zoo’s quarantine unit no further spread of infection was observed.
Given the relatively high incidence of Brucella isolations from amphibians in recent years (Shilton et al. 2008; Eisenberg et al. 2012; Fischer et al. 2012; Whatmore et al. 2015; Mühldorfer et al. 2016; Scholz et al. 2016a) it is tempting to speculate that Brucella in fish might also be more common in professional and hobbyist aquaculture than previously expected. Their role as a potential pathogen (or member of the normal flora with the ability to act as opportunistic pathogen at times of lowered immunity) awaits clarification. As can be concluded from amphibians, brucellae may be found in apparently healthy individuals, but can also cause a variety of different pathologies ranging from individual, localised disease manifestations (e.g. subcutaneous abscess, skin lesions, swollen paravertebral ganglia, panopthalmitis) to systemic bacterial infections with high mortalities (Mühldorfer et al. 2016). In Nile catfish, skin lesions were found to be associated with B. melitensis infection (El-Tras et al. 2010).
Presently, we do not know enough about specific host-bacteria relationships or possible adaptations. As recently reported, Brucella isolated from frogs also contain numerous genetic elements presumably acquired from environmental bacteria, possibly indicating an evolutionary connection link between a soil saprophyte and a mammalian pathogen (Scholz et al. 2016a). Fish are—like amphibians—primitive vertebrates, that fit well into this concept and might represent both asymptomatically infected as well as facultative susceptible reservoir hosts for Brucella spp.
In summary, we have isolated a novel Brucella strain from a bluespotted ribbontail ray and, hence, demonstrate occurrence of indigenous Brucella bacteria in a second class of cold-blooded vertebrates. The isolate is motile, fast growing and phenotypically very similar to strains that were recently isolated from amphibian hosts. Notably motility was recently found to be a unique feature among other ‘atypical’ Brucella from frogs (Eisenberg et al. 2012; Mühldorfer et al. 2016) and has also been demonstrated for another frog strain B13-0095 as well as for B. inopinata BO1T and strain BO2 (Soler-Lloréns et al. 2016). In contrast, the ‘atypical’ Australian rodent strains represented by NF2653 are non-motile which can possibly be explained by flagellar pseudogenes (Soler-Lloréns et al. 2016). As determined by recA and complete genome analysis the ray strain from this study belongs to the ‘atypical’ Brucella group. Based on recA gene sequences, and in contrast to the ‘classical’ members of the genus, the ‘atypical’ brucellae are characterised by a remarkable heterogeneity. The ray strain forms a separate lineage closely related to B. inopinata and Brucella strain BO2 and—with respect to recA gene sequence—also to some of the amphibian strains. The presence of additional genetic material from various other soil-living bacteria suggests extensive horizontal gene transfer and indicates a different ecology compared to classical Brucella species. As with frog brucellae, these strains might represent an evolutionary connection between a soil-associated ancestor and the highly host-adapted classical Brucella species infecting humans and animals. Our findings significantly enhance the understanding of the ecology and pathogenic potential of members of the genus Brucella.
References
Audic S, Lescot M, Claverie JM, Scholz HC (2009) Brucella microti: the genome sequence of an emerging pathogen. BMC Genom 10:352. doi:10.1186/1471-2164-10-352
Baily GG, Krahn JB, Drasar BS, Stoker NG (1992) Detection of Brucella melitensis and Brucella abortus by DNA amplification. J Trop Med Hyg 95:271–275
Blom J, Kreis J, Spanig S, Juhre T, Bertelli C, Ernst C, Goesmann A (2016) EDGAR 2.0: an enhanced software platform for comparative gene content analyses. Nucleic Acids Res 44:W22–28. doi:10.1093/nar/gkw255
Bricker BJ, Halling SM (1994) Differentiation of Brucella abortus bv. 1, 2, and 4, Brucella melitensis, Brucella ovis, and Brucella suis bv. 1 by PCR. J Clin Microbiol 32:2660–2666
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST+: architecture and applications. BMC Bioinform 10:421. doi:10.1186/1471-2105-10-421
Cunningham SA, Patel R (2013) Importance of using Bruker’s security-relevant library for Biotyper identification of Burkholderia pseudomallei, Brucella species, and Francisella tularensis. J Clin Microbiol 51:1639–1640. doi:10.1128/JCM.00267-13
De BK et al (2008) Novel Brucella strain (BO1) associated with a prosthetic breast implant infection. J Clin Microbiol 46:43–49. doi:10.1128/JCM.01494-07
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797. doi:10.1093/nar/gkh340
Eisenberg T et al (2012) Isolation of potentially novel Brucella spp. from frogs. Appl Environ Microbiol 78:3753–3755. doi:10.1128/AEM.07509-11
Elsaghir AA, James EA (2003) Misidentification of Brucella melitensis as Ochrobactrum anthropi by API 20NE. J Med Microbiol 52:441–442. doi:10.1099/jmm.0.05153-0
El-Tras WF, Tayel AA, Eltholth MM, Guitian J (2010) Brucella infection in fresh water fish: evidence for natural infection of Nile catfish, Clarias gariepinus, with Brucella melitensis. Vet Microbiol 141:321–325. doi:10.1016/j.vetmic.2009.09.017
Fischer D, Lorenz N, Heuser W, Kampfer P, Scholz HC, Lierz M (2012) Abscesses associated with a Brucella inopinata-like bacterium in a big-eyed tree frog (Leptopelis vermiculatus). J Zoo Wildl Med 43:625–628. doi:10.1638/2011-0005R2.1
Foster G, Osterman BS, Godfroid J, Jacques I, Cloeckaert A (2007) Brucella ceti sp. nov. and Brucella pinnipedialis sp. nov. for Brucella strains with cetaceans and seals as their preferred hosts. Int J Syst Evol Microbiol 57:2688–2693. doi:10.1099/ijs.0.65269-0
Garcia-Yoldi D, Marin CM, de Miguel MJ, Munoz PM, Vizmanos JL, Lopez-Goni I (2006) Multiplex PCR assay for the identification and differentiation of all Brucella species and the vaccine strains Brucella abortus S19 and RB51 and Brucella melitensis. Rev1 Clin Chem 52:779–781. doi:10.1373/clinchem.2005.062596
Godfroid J et al (2005) From the discovery of the Malta fever’s agent to the discovery of a marine mammal reservoir, brucellosis has continuously been a re-emerging zoonosis. Vet Res 36:313–326. doi:10.1051/vetres:2005003
Helm D, Labischinski H, Schallehn G, Naumann D (1991) Classification and identification of bacteria by Fourier-transform infrared spectroscopy. J Gen Microbiol 137:69–79
Hernandez-Mora G, Palacios-Alfaro JD, Gonzalez-Barrientos R (2013) Wildlife reservoirs of brucellosis: brucella in aquatic environments. Revue scientifique et technique (International Office of Epizootics) 32:89–103
Jiménez de Bagüés MP, Iturralde M, Arias MA, Pardo J, Cloeckaert A, Zygmunt MS (2014) The new strains Brucella inopinata BO1 and Brucella species 83-210 behave biologically like classic infectious Brucella species and cause death in murine models of infection. J Infect Dis 210:467–472. doi:10.1093/infdis/jiu102
Kuhlmann AU, Hoffmann T, Bursy J, Jebbar M, Bremer E (2011) Ectoine and hydroxyectoine as protectants against osmotic and cold stress: uptake through the SigB-controlled betaine-choline- carnitine transporter-type carrier EctT from Virgibacillus pantothenticus. J Bacteriol 193:4699–4708. doi:10.1128/JB.05270-11
Kuhm AE, Suter D, Felleisen R, Rau J (2009) Identification of Yersinia enterocolitica at the species and subspecies levels by Fourier transform infrared spectroscopy. Appl Environ Microbiol 75:5809–5813. doi:10.1128/AEM.00206-09
McDonald WL et al (2006) Characterization of a Brucella sp. strain as a marine-mammal type despite isolation from a patient with spinal osteomyelitis in New Zealand. J Clin Microbiol 44:4363–4370. doi:10.1128/JCM.00680-06
Mühldorfer K, Wibbelt G, Szentiks CA, Fischer D, Scholz HC, Zschöck M, Eisenberg T (2016) The role of ‘atypical’ Brucella in amphibians: Are we facing a novel emerging pathogen? J Appl Microbiol
Nymo IH, Seppola M, Al Dahouk S, Bakkemo KR, Jimenez de Bagues MP, Godfroid J, Larsen AK (2016) Experimental challenge of Atlantic cod (Gadus morhua) with a Brucella pinnipedialis strain from hooded seal (Cystophora cristata). PLoS One 11:e0159272. doi:10.1371/journal.pone.0159272
Salem SF, Mohsen A (1997) Brucellosis in fish. Vet Med 42:5–7
Scholz HC et al (2010) Brucella inopinata sp. nov., isolated from a breast implant infection. Int J Syst Evol Microbiol 60:801–808. doi:10.1099/ijs.0.011148-0
Scholz HC, Mühldorfer K, Shilton C, Suresh B, Whatmore AM, Blom J, Eisenberg T (2016a) Worldwide emergence of genetically diverse Brucella isolates in exotic frogs provides insights into Brucella evolution. PloS One under revision
Scholz HC et al (2016b) Brucella vulpis sp. nov., a novel Brucella species isolated from mandibular lymph nodes of red foxes (Vulpes vulpes) in Austria. Int J Syst Evol Microbiol. doi:10.1099/ijsem.0.000998
Shilton CM, Brown GP, Benedict S, Shine R (2008) Spinal arthropathy associated with Ochrobactrum anthropi in free-ranging cane toads (Chaunus [Bufo] marinus) in Australia. Vet Pathol 45:85–94. doi:10.1354/vp.45-1-85
Soler-Lloréns PF et al (2016) A Brucella spp isolate from a Pac-Man frog (Ceratophrys ornata) reveals characteristics departing from classical brucellae. Front Cell Infect Microbiol. doi:10.3389/fcimb.2016.00116
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. doi:10.1093/molbev/mst197
Tatusova T, Ciufo S, Fedorov B, O’Neill K, Tolstoy I (2014) RefSeq microbial genomes database: new representation and annotation strategy. Nucleic Acids Res 42:D553–559. doi:10.1093/nar/gkt1274
Tiller RV, Gee JE, Frace MA, Taylor TK, Setubal JC, Hoffmaster AR, De BK (2010a) Characterization of novel Brucella strains originating from wild native rodent species in North Queensland, Australia. Appl Environ Microbiol 76:5837–5845. doi:10.1128/AEM.00620-10
Tiller RV et al (2010b) Identification of an unusual Brucella strain (BO2) from a lung biopsy in a 52 year-old patient with chronic destructive pneumonia. BMC Microbiol 10:23. doi:10.1186/1471-2180-10-23
Ward JH (1963) Hierarchical grouping to optimize an objective function. J Amer Statist Assoc 58:236–244
Wattam AR et al (2012) Comparative genomics of early-diverging Brucella strains reveals a novel lipopolysaccharide biosynthesis pathway. MBio 3:e00212–e00246. doi:10.1128/mBio.00246-12
Whatmore AM et al (2014) Brucella papionis sp. nov., isolated from baboons (Papio spp.). Int J Syst Evol Microbiol 64:4120–4128. doi:10.1099/ijs.0.065482-0
Whatmore AM et al. (2015) Isolation of Brucella from a White’s tree frog (Litoria caerulea) JMM Case Reports:1–5. doi: 10.1099/jmmcr.0.000017
Acknowledgments
The authors like to thank Anna Mohr for excellent technical support. The Hessian State Laboratory is supported by the Hessian Ministry for the Environment, Climate Change, Agriculture and Consumer Protection.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed.
Additional information
Tobias Eisenberg and Holger C. Scholz have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Suppl. Fig. S1
Cluster analysis of respective spectra obtained by Fourier transform-infrared spectroscopy (FT-IR) using OPUS software (version 4.2, Bruker Optics). Representative IR-spectra of Brucella suis 161003792, Ochrobactrum intermedium 09-7-D-05279-1 as well as brucellae from a bluespotted ribbontail ray from this study (in bold) and from various amphibians [(Mühldorfer et al. 2016); see Table 1] were used for calculation with the Ward’s algorithm. The dendrogram depicts the arrangement of isolates in groups according to their spectral differences. Supplementary material 1 (PDF 157 kb)
Suppl. Table S1
Detail analysis of protein coding genes unique to the bluespotted ribbontail ray strain 141012304 in comparison to ‘classical’ Brucella species. Singletons identified for strain 141012304 in comparison to ‘classical’ Brucella species were compared to the RefSeq database (Tatusova et al. 2014) using the BLAST+ implementation (Camacho et al. 2009) of BLASTP with an initial evalue cutoff of 1e-10. Results were filtered for the best two hits of every query and the annotation and source organism for these best hits were extracted from the database. Supplementary material 2 (XLSX 44 kb)
Suppl. Table S2
Detail analysis of protein coding genes unique to the bluespotted ribbontail ray strain 141012304 in comparison to all Brucella species used in this study. Supplementary material 3 (TXT 29 kb)
Rights and permissions
About this article
Cite this article
Eisenberg, T., Riße, K., Schauerte, N. et al. Isolation of a novel ‘atypical’ Brucella strain from a bluespotted ribbontail ray (Taeniura lymma). Antonie van Leeuwenhoek 110, 221–234 (2017). https://doi.org/10.1007/s10482-016-0792-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-016-0792-4