Abstract
Traditionally, detection of epileptic seizures based on the visual inspection of neurologists is tedious, laborious and subjective. To overcome such disadvantages, numerous seizure detection techniques involving signal processing and machine learning tools have been developed. However, there still remain the problems of automatic detection with high efficiency and accuracy in distinguishing normal, interictal and ictal electroencephalogram (EEG) signals. In this study we propose a novel method for automatic identification of epileptic seizures in singe-channel EEG signals based upon time-scale decomposition (ITD), discrete wavelet transform (DWT), phase space reconstruction (PSR) and neural networks. First, EEG signals are decomposed into a series of proper rotation components (PRCs) and a baseline signal by using the ITD method. The first two PRCs of the EEG signals are extracted, which contain most of the EEG signals’ energy and are considered to be the predominant PRCs. Second, four levels DWT is employed to decompose the predominant PRCs into different frequency bands, in which third-order Daubechies (db3) wavelet function is selected for analysis. Third, phase space of the PRCs is reconstructed based on db3, in which the properties associated with the nonlinear EEG system dynamics are preserved. Three-dimensional (3D) PSR together with Euclidean distance (ED) has been utilized to derive features, which demonstrate significant difference in EEG system dynamics between normal, interictal and ictal EEG signals. Fourth, neural networks are then used to model, identify and classify EEG system dynamics between normal (healthy), interictal and ictal EEG signals. Finally, experiments are carried out on the University of Bonn’s widely used and publicly available epilepsy dataset to assess the effectiveness of the proposed method. By using the 10-fold cross-validation style, the achieved average classification accuracy for eleven cases is reported to be 98.15%. Compared with many state-of-the-art methods, the results demonstrate superior performance and the proposed method can serve as a potential candidate for the automatic detection of seizure EEG signals in the clinical application.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Epilepsy is a chronic neurological disorder caused due to abnormal and excessive brain neuronal activity, in which Electroencephalogram (EEG) signal is the most commonly used and efficient clinical technique to assess epilepsy due to its inexpensiveness and availability (Zhang et al. 2017). Traditionally, detection of epileptic seizures based on the visual inspection of neurologists is tedious, laborious and subjective (Martis et al. 2015). In addition, it requires expertise in the analysis of long-duration EEG signals (Scheuer and Wilson 2004). In those application scenarios absence of experts, for example, in emergency, computer-aided automatic detection of epileptic seizure becomes significant. To overcome above-mentioned disadvantages, numerous seizure detection techniques involving signal processing and machine learning tools have been developed, such as support vector machine (SVM), extreme learning machine (ELM), random forest (RF) and deep learning (Zhang and Chen 2016; Song et al. 2016; Mursalin et al. 2017; Acharya et al. 2018; Ullah et al. 2018; Li et al. 2019; Subasi et al. 2019; Sharma et al. 2018; Sharma and Pachori 2017a, b; Bhati et al. 2017a, b; Bhattacharyya and Pachori 2017; Tiwari et al. 2016; Sharma et al. 2017; Bhattacharyya et al. 2017; Sharma and Pachori 2015; Kumar et al. 2015; Pachori and Patidar 2014; Bajaj and Pachori 2012, 2013; Pachori and Bajaj 2011; Pachori 2008; Pachori et al. 2015). However, it still remains an open problem of automatic detection with high efficiency and accuracy in distinguishing normal, interictal and ictal EEG signals (Djemili et al. 2016). In attempt to sovle the problem, various algorithms have been developed. Since EEG signals are the redundant discrete-time sequences, numerous methods with combination of time-domain, frequency-domain, time-frequency-domain and nonlinear analysis have been proposed (Acharya et al. 2013). For the time-domain analysis, representative techniques such as linear prediction (Sheintuch et al. 2014), fractional linear prediction (Joshi et al. 2014), principal component analysis (PCA) based radial basis function neural network (Kafashan et al. 2017), etc, have been proposed for seizure detection and EEG classification. For the frequency-domain analysis, with an assumption that EEG signals are stationary, Fourier transform is usually employed to extract features for epileptic seizure detection. Samiee et al. (2015) applied the rational Discrete Short Time Fourier Transform (DSTFT) to extract features for the separation of seizure epochs from seizure-free epochs using a Multilayer Perceptron (MLP) classifier. Considering the non-stationary nature of EEG signals (Subasi and Gursoy 2010), for the time-frequency-domain analysis, a wavelet transform tool together with certain classifier has usually been used for the epileptic seizure detection. Hassan et al. (2016) decomposed the EEG signal segments into sub-bands using Tunable-Q factor wavelet transform (TQWT) and several spectral features were extracted. Then bootstrap aggregating was employed for epileptic seizure classification. For the nonlinear analysis, various nonlinear parameters extracted through different types of entropies (Acharya et al. 2015), Lyapunov exponent (Shayegh et al. 2014), fractal dimension (Zhang et al. 2015), correlation dimension (Sato et al. 2015), recurrence quantification analysis (RQA) (Timothy et al. 2017) and Hurst exponent (Lahmiri 2018) methods have been used for automatic detection of epileptic EEG signals. Aarabi and He (2017) developed a method on the fusion of features extracted from correlation dimension, correlation entropy, noise level, Lempel–Ziv complexity, largest Lyapunov exponent, and nonlinear interdependence for the detection of focal EEG signals.
Despite the fact that these previous approaches have demonstrated respectable classification accuracy, the potential of nonlinear methods has not been thoroughly investigated. The EEG signal is highly random, nonlinear, nonstationary and non-Gaussian in nature (Acharya et al. 2013), for which nonlinear features characterize the EEG more accurately than linear models (Wang et al. 2017). Considering this characteristics, several self-adaptive signal processing methods, such as empirical mode decomposition (EMD) (Huang et al. 1998; Huang and Kunoth 2013), local mean decomposition (LMD) (Park et al. 2011) and intrinsic time-scale decomposition (ITD) (Frei and Osorio 2007), can be employed to extract effective and predominant features from EEG signals (Li et al. 2013; Zahra et al. 2017). EMD decomposes a multi-component signal into a series of single components and a residual signal while LMD decomposes any complicated signal into a series of product functions. However, there exist some drawbacks in these methods, in which the EMD method contains over envelope, mode mixing, end effects and unexplainable negative frequency caused by Hilbert transformation (Chen et al. 2011), while the LMD method has distorted components, mode mixing and time-consuming decomposition (Li et al. 2015). To address these problems, recently, a new technical tool named ITD, has been introduced by Frei and Osorio (2007) for analyzing data from nonstationary and nonlinear processes. Compared with EMD, more local characteristic information of the signal can be utilized in ITD method. In addition, the negative frequency caused by Hilbert transform has been completely eliminated (Feng et al. 2016). Furthermore, the computational efficiency has been significantly improved. With high decomposition efficiency and frequency resolution, ITD can help decompose a complex signal into several proper rotation components (PRCs) and a baseline signal, which leads to the accurate extraction of the dynamic features of nonlinear signals. Meanwhile, there is no spline interpolation and screening process in ITD method which contains low edge effect (An et al. 2012; Xing et al. 2017). ITD can better preserve and extract the EEG system dynamics which is effective for the classification of normal, interictal and ictal EEG signals. Phase space reconstruction (PSR) is another popular nonlinear tool for analyzing composite, nonlinear and nonstationary signals (Takens 1980; Xu et al. 2013; Lee et al. 2014; Chen et al. 2014; Jia et al. 2017). The principle of PSR is to transform the properties of a time series into topological properties of a geometrical object which is embedded in a space, wherein all possible states of the system are represented. Each state corresponds to a unique point, and this reconstructed space is sharing the same topological properties as the original space. The dynamics in the reconstructed state space is equivalent to the original dynamics. Hence reconstructed phase space is a very useful tool to extract nonlinear dynamics of the signal (Takens 1980; Xu et al. 2013; Lee et al. 2014; Chen et al. 2014; Jia et al. 2017). It is hypothesized that EEG system dynamics between normal, interictal and ictal EEG signals are significantly different, which implies that PSR offers the potential to compute the difference and classify these EEG signals.
The novelty of this work lies in four aspects: (1) ITD method is employed to measure the variability of EEG signals and the first and second proper rotation components (PRCs) are extracted as predominant PRCs which contain most of the EEG signals’ energy; (2) discrete wavelet transform (DWT) decomposes the predominant PRCs into different frequency bands, which are used to construct the reference variables. (3) 3D phase space of the two PRCs components is reconstructed, in which the properties associated with the EEG system dynamics are preserved; (3) EEG system dynamics can be modeled and identified using neural networks, which employ the ED of 3D PSR of the reference variables as features; (4) the difference of EEG system dynamics between normal, interictal and ictal EEG signals is computed and used for the discrimination between the three groups based on a bank of estimators. Detailed description is illustrated as follows. In the present study we propose a combined and computational method from the area of nonlinear method and machine learning for the classification of normal, interictal and ictal EEG signals. To explore the underlying motor strategies in the three groups, neural networks together with ITD, discrete wavelet transform (DWT) and PSR are implemented for this purpose. The complete algorithm encompasses four principal stages: (1) EEG signals are decomposed into a series of proper rotation components (PRCs) and a baseline signal by using the ITD method. The first two PRCs of the EEG signals are extracted, which contain most of the EEG signals’ energy and are considered to be the predominant PRCs. (2) four levels DWT is employed to decompose the predominant PRCs into different frequency bands, in which third-order Daubechies (db3) wavelet function is selected for analysis. (3) Phase space of the PRCs is reconstructed based on db3, in which the properties associated with the nonlinear EEG system dynamics are preserved. Three-dimensional (3D) PSR together with Euclidean distance (ED) has been utilized to derive features, which demonstrate significant difference in EEG system dynamics between normal, interictal and ictal EEG signals. (4) Neural networks are then used to model, identify and classify EEG system dynamics between normal (healthy), interictal and ictal EEG signals.
The rest of the paper is organized as follows. Section 2 introduces the details of the proposed method, including the Bonn dataset, data description, ITD, DWT, PSR, ED, feature extraction and selection, learning and classification mechanisms. Section 3 presents experimental results. Sections 4 and 5 give some discussions and conclusions, respectively.
2 Method
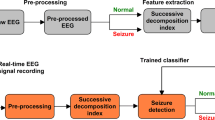
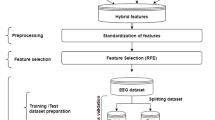
In this section, we propose a method to discriminate between normal, interictal and ictal EEG signals using the information obtained from nonlinear EEG dynamics. It is divided into the training stage and the classification stage and follows the following steps. In the first step, ITD is applied to decompose EEG signals into several PRCs to extract predominant components. In the second step, DWT is employed to decompose the predominant PRCs into different frequency bands. In the third step, PSR is applied to extract nonlinear dynamics of EEG signals and Euclidean distances are computed. Finally, feature vectors are fed into the neural networks for the modeling and identification of EEG system dynamics. The difference of dynamics between normal (healthy), interictal and ictal EEG signals will be applied for the classification task. The flowchart of the proposed algorithm is illustrated in Fig. 1.
2.1 EEG database
In the present study we use the open and publicly available Bonn University database (Andrzejak et al. 2001) consisting of five different sets (Z, O, N, F and S), each of which contains 100 single-channel EEG segments of 23.6-s duration. All EEG signals were recorded at a sampling rate of 173.61 Hz using a 128-channel amplifier system with an average common reference. Band-pass filter was set with the frequency 0.53–40 Hz. Hence each signal has 4097 recordings, which means the data length of each signal is 4097. Set Z and O contain surface EEG recordings that were carried out on five healthy subjects in relaxing state. Set Z was recorded when subjects’ eyes were open while set O was recorded when subjects’ eyes were closed. Sets N, F, and S contain intracranial recordings from depth and strip electrodes collected from five epileptic patients. Set N contains seizure-free intervals collected from the hippocampal formation of opposite hemisphere, set F contains seizure-free intervals collected from epileptogenic zone, and set S contains epileptic seizure segments originated from all channels. EEG recordings from Z–O, N–F and S datasets were defined as normal (healthy), interictal and ictal signals, respectively.
2.2 Intrinsic time-scale decomposition (ITD)
Intrinsic time-scale decomposition (ITD) is suitable for analyzing nonstationary and nonlinear signals such as the EEG signals. Without resorting to the spline interpolation to signal extrema and sifting in mono-component separation, it decomposes a signal into proper rotation components (PRCs) that are suitable to calculate the instantaneous frequency and amplitude, based on the baseline defined via linear transform. The obtained decomposition result precisely preserves the temporal information of each component regarding signal critical points and riding waves, with a time resolution equal to the time scale of the occurrence of extrema in the raw signal (Feng et al. 2016). Based on the single wave analysis, it extracts accurately the inherent instantaneous amplitude and frequency/phase information and other relevant morphological features (Frei and Osorio 2007).
For a time series signal I(t), define the operator L to extract the baseline signal from I(t) and the residual signal is called the proper rotation component (PRC). The decomposed signal I(t) can be expressed as
where B(t) is the baseline signal and H(t) is the proper rotation.
The decomposition procedure of a nonlinear signal can be summarized by the following steps:
Step 1 Find the local extrema of the signal I(t), denoted by \(I_k\), and the corresponding occurrence time instant \(\tau _k, k=0,1,2,\ldots \). For convenience \(\tau _0=0\).
Step 2 Suppose the operators B(t) and H(t) are given over the interval \([0, \tau _k]\), and I(t) is set on the interval \(t\in [0, \tau _{k+2}]\). Then on the interval \([\tau _k, \tau _{k+1}]\) between two adjacent extrema \(I_k\) and \(I_{k+1}\), the piecewise baseline extraction operator is defined as
$$\begin{aligned} LI(t)=B(t)=B_k+(\frac{B_{k+1}-B_k}{I_{k+1}-I_k})\times (I(t)-I_k), \quad t\in [\tau _k, \tau _{k+1}], \end{aligned}$$(2)where
$$\begin{aligned} B_{k+1}=\beta [I_k+(\frac{\tau _{k+1}-\tau _{k}}{\tau _{k+2}-\tau _{k}})(I_{k+2}-I_k)]+(1-\beta )I_{k+1}, \end{aligned}$$(3)and \(0<\beta <1\), typically \(\beta =0.5\).
Step 3 After extracting the baseline signal, the operator \(\Theta \) for extracting the residual signal as PRCs is defined as
$$\begin{aligned} \Theta I(t)\equiv (1-L)I(t)=I(t)-B(t) \end{aligned}$$(4)
According to the definition, the PRC is a riding wave with the highest frequency on the baseline. Therefore, ITD separates the PRC in a frequency order from high to low. In addition, the PRC is obtained directly by subtracting the baseline from the input signal, without resorting to any sifting within each iterative decomposition. Thus, ITD has low computational complexity, and more importantly, avoids the smoothing of transients and time-scale smearing due to repetitive sifting (Feng et al. 2016).
Take the baseline B(t) as the input signal I(t), and repeat steps (1)–(3), until the baseline becomes a monotonic function or a constant. Eventually, the raw signal will be decomposed into PRCs and a trend (Feng et al. 2016)
where \(\rho \) is the decomposition level.
Samples of the ITD of EEG signals from the five sets are demonstrated in Fig. 2.
2.3 Discrete wavelet transform (DWT)
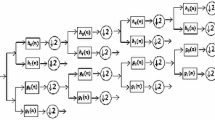
Wavelet transform is an effective time-frequency tool for the analysis of non-stationary signals. Discrete Wavelet Transform (DWT) is a procedure for the decomposition of input signal H(t) (H(t) is the PRC of the EEG signal in this work) into sets of function, called wavelets, by scaling and shifting of mother wavelet function. Consequently, the decomposition i.e. set of wavelet coefficients are formed.
To accomplish this, the signal H(t) can be reconstructed as linear combination of wavelets and weighting wavelet coefficients. The setting of appropriate wavelet function and the number of decomposition levels is of great importance for correctly reconstructing the signal H(t). In order to extract five physiological EEG bands, four levels DWT with third-order Daubechies (db3) wavelet function have been used (Table 1 represents the frequency distribution of the DWT-based coefficients of the PRCs of the EEG signals at 173.6 Hz), from which the choice of the mother wavelet is supported by many works in literature (Vavadi et al. 2010; Tawfik 2016; Li et al. 2017). Figure 3 shows samples of EEG channel of five sets and their decomposed frequency bands of predominant PRCs. Since the frequency components above 40 Hz is lack of use in epilepsy analysis, in order to reduce the feature dimension, the advisable sub-bands (D4 and A4) are selected for feature acquisition.
2.4 Phase space reconstruction (PSR)
It is sometimes necessary to search for patterns in a time series and in a higher dimensional transformation of the time series (Sun et al. 2015). Phase space reconstruction is a method used to reconstruct the so-called phase space. The concept of phase space is a useful tool for characterizing any low-dimensional or high-dimensional dynamic system. A dynamic system can be described using a phase space diagram, which essentially provides a coordinate system where the coordinates are all the variables comprising mathematical formulation of the system. A point in the phase space represents the state of the system at any given time (Lee et al. 2014; Sivakumar 2002). Every db3 wavelet function of the PRC of the EEG signals can be written as the time series vector \(V=\{v_1,v_2,v_3, \ldots ,v_K\}\), where K is the total number of data points. The phase space can be reconstructed according to Lee et al. 2014:
where \(j=1,2, \ldots ,K-(d-1)\tau \), d is the embedding dimension of the phase space and \(\tau \) is a time lag. It is worthwhile to mention that the properties associated with the EEG dynamics are preserved in the reconstructed phase space.
The behaviour of the signal over time can be visualized using PSR (especially when \(d=2\) or 3). In this work, we have confined our discussion to the value of embedding dimension \(d=3\), because of their visualization simplicity. In addition, different studies have found this value to best represent the attractor for human movement (Venkataraman and Turaga 2016; Som et al. 2016). For \(\tau \), we either use the first-zero crossing of the autocorrelation function for each time series or the average \(\tau \) value obtained from all the time series in the training dataset using the method proposed in Michael (2005). In this study, we consider the values of time lag \(\tau =1\) to test the classification performance. PSR for \(d=3\) has been referred to as 3D PSR.
Reconstructed phase spaces have been proven to be topologically equivalent to the original system and therefore are capable of recovering the nonlinear dynamics of the generating system (Takens 1980; Xu et al. 2013). This implies that the full dynamics of the EEG system are accessible in this space, and for this reason, features extracted from it can potentially contain more and/or different information than the common features extraction method (Chen et al. 2014).
3D PSR is the plot of three delayed vectors \(V_j,V_{j+1}\) and \(V_{j+2}\) to visualize the dynamics of human EEG system. Euclidian distance (ED) of a point \((V_j,V_{j+1},V_{j+2})\), which is the distance of the point from origin in 3D PSR and can be defined as Lee et al. 2014
ED measures can be used in features extraction and have been studied and applied in many fields, such as clustering algorithms and induced aggregation operators (Merigó and Casanovas 2011).
2.5 Feature extraction and selection
In order to obtain more efficient features, this paper proposes the following extraction scheme.
(1) ITD of the EEG signals and derivation of predominant PRCs. The signals obtained by ITD method, which are a series of decomposing signals, cannot be directly used to classify because of the high feature dimension. To solve this problem, the Pearson’s correlation coefficient is calculated to measure the correlation between the first four PRCs and the original EEG signals. The PRC with higher correlation coefficient is more highly correlated to the original signal, which means the signal energy is mostly concentrated in this PRC as well. In the present study most of the energy is concentrated in PRC1 and PRC2 components, which have the most important information from the EEG signals and are considered to be the predominant PRCs (seen from Table 2).
(2) Four levels DWT is employed to decompose the predominant PRCs into different frequency bands, in which third-order Daubechies (db3) wavelet function is selected for analysis. D4 and A4 of the PRC1 and PRC2 EEG signals are regarded as reference variables \([PRC1^{D4},PRC1^{A4},PRC2^{D4},PRC2^{A4}]^T\) and are used for feature derivation.
(3) Reconstruct the phase space of the reference variables with selected values of d and \(\tau \);
(4) Compute ED of 3D PSR of the reference variables. Concatenate them to form a feature vector \([ED_{j}^{PRC1^{D4}},ED_j^{PRC1^{A4}},ED_j^{PRC2^{D4}},ED_j^{PRC2^{A4}}]^T\).
For the Bonn epileptic database, EEG signals are analyzed and signal dynamics are extracted by using ITD, DWT and 3D PSR. First, ITD of the EEG signals are exhibited in Fig. 2. Four levels DWT of the PRC1 and PRC2 of EEG signals from the five sets are demonstrated in Fig. 3. The db3 of the first two PRCs are utilized to form the reference variables \([PRC1^{D4},PRC1^{A4},PRC2^{D4},PRC2^{A4}]^T\). Samples of the 3D PSR of the reference variables are exhibited in Fig. 4. After 3D PSR, features of \([ED_j^{PRC1^{D4}},ED_j^{PRC1^{A4}},ED_j^{PRC2^{D4}},ED_j^{PRC2^{A4}}]^T\) for EEG signals of the five sets are derived through ED computation, as demonstrated in Fig. 5. As we have analyzed before, significant difference in EEG system dynamics have been reported between EEG signals of five sets, which can also be seen obviously from Fig. 4.


2.6 Training and modeling mechanism based on selected features
In this section, we present a scheme for modeling and deriving of EEG system dynamics from normal, interictal and ictal EEG signals based on the above mentioned features.
Consider a general nonlinear EEG system dynamics in the following form:
where \(x=[x_1,\ldots ,x_n]^T\in R^n\) are the system states which represent the features \([ED_j^{PRC1^{D4}},ED_j^{PRC1^{A4}},ED_j^{PRC2^{D4}},ED_j^{PRC2^{A4}}]^T\), p is a constant vector of system parameters. \(F(x;p)=[f_1(x;p),\ldots ,f_n(x;p)]^T\) is a smooth but unknown nonlinear vector representing the EEG system dynamics, v(x; p) is the modeling uncertainty. Since the modeling uncertainty v(x; p) and the EEG system dynamics F(x; p) cannot be decoupled from each other, we consider the two terms together as an undivided term, and define \(\phi (x;p):=F(x;p)+v(x;p)\) as the general EEG system dynamics. Then, the following steps are taken to model and derive the EEG system dynamics via deterministic learning theory (Wang and Hill 2006, 2007, 2009).
In the first step, standard RBF neural networks are constructed in the following form
where Z is the input vector, \(W=[w_1,\ldots ,w_N]^T\in R^N\) is the weight vector, N is the node number of the neural networks, and \(S(Z)=[s_1(\parallel Z-\mu _1\parallel ),..., s_N(\parallel Z-\mu _N\parallel )]^T\), with \(s_i(\parallel Z-\mu _i\parallel )=\exp [\frac{-(Z-\mu _i)^T(Z-\mu _i)}{\eta _i^2}]\) being a Gaussian function, \(\mu _i(i=1,...,N)\) being distinct points in state space, and \(\eta _i\) being the width of the receptive field.
In the second step, the following dynamical RBF neural networks are employed to model and derive the EEG system dynamics \(\phi (x;p)\):
where \(\hat{x}=[\hat{x}_1,\ldots ,\hat{x}_n]\) is the state vector of the dynamical RBF neural networks, \(A=diag[a_1,\ldots ,a_n]\) is a diagonal matrix, with \(a_i>0\) being design constants, localized RBF neural networks \(\hat{W}_j^TS_j(x)=\sum \nolimits _{i=1}^N w_{ij}s_{ij}(x)\) are used to approximate the unknown \(\phi (x;p)\), where \(\hat{W}_j=[w_{1j},\ldots ,w_{Nj}]^T\), \(S_j=[s_{1j},\ldots ,s_{Nj}]^T\), for \(j=1,\ldots ,n\).
The following law is used to update the neural weights
where \(\tilde{x}_i=\hat{x}_i-x_i, \tilde{W}_i=\hat{W}_i-W_i^*\), \(W_i^*\) is the ideal constant weight vector such that \(\phi _i(x;p)={W_i^*}^TS_i(x)+\epsilon _i(x)\), \(\epsilon _i(x)<\epsilon ^*\) represents the neural network modeling error, \(\Gamma _i=\Gamma _i^T>0\), and \(\sigma _i>0\) is a small value.
With Eqs. (8)–(10), the derivative of the state estimation error \(\tilde{x}_i\) satisfies
In the third step, by using the local approximation property of RBF neural networks, the overall system consisting of dynamical model (12) and the neural weight updating law (11) can be summarized into the following form in the region \(\Omega _\zeta \)
and
where \(\epsilon _{\zeta i}=\epsilon _i-\tilde{W}_{\bar{\zeta }i}^TS_{\bar{\zeta }}(x)\). The subscripts \((\cdot )_\zeta \) and \((\cdot )_{\bar{\zeta }}\) are used to stand for terms related to the regions close to and far away from the trajectory \(\varphi _\zeta (x_0)\). The region close to the trajectory is defined as \(\Omega _\zeta :=\{Z|\mathrm {dist}(Z,\varphi _\zeta )\le d_{\iota }\}\), where \(Z=x, d_\iota >0\) is a constant satisfying \(s(d_\iota )>\iota \), \(s(\cdot )\) is the RBF used in the network, \(\iota \) is a small positive constant. The related subvectors are given as: \(S_{\zeta i}(x)=[s_{j1}(x),\ldots ,s_{j\zeta }(x)]^T\in R^{N_\zeta }\), with the neurons centered in the local region \(\Omega _\zeta \), and \(W_\zeta ^*=[w_{j1}^*,\ldots ,w_{j\zeta }^*]^T\in R^{N_\zeta }\) is the corresponding weight subvector, with \(N_\zeta <N\). For localized RBF neural networks, \(|\tilde{W}_{\bar{\zeta }i}^TS_{\bar{\zeta i}}(x)|\) is small, so \(\epsilon _{\zeta i}=O(\epsilon _i)\).
By the convergence result, we can obtain a constant vector of neural weights according to
where \(t_b>t_a>0\) represent a time segment after the transient process. Therefore, we conclude that accurate identification of the function \(\phi _i(x;p)\) is obtained along the trajectory \(\varphi _\zeta (x_0)\) by using \(\bar{W}_i^TS_i(x)\), i.e.,
where \(\epsilon _{i2}=O(\epsilon _{i1})\) and subsequently \(\epsilon _{i2}=O(\epsilon ^*)\).
2.7 Classification mechanism
In this section, we present a scheme to classify normal, interictal and ictal EEG signals.
Consider a training dataset consisting of EEG signal patterns \(\varphi _\zeta ^k\), \(k=1,\ldots ,M\), with the kth training pattern \(\varphi _\zeta ^k\) generated from
where \(F^k(x;p^k)\) denotes the EEG system dynamics, \(v^k(x;p^k)\) denotes the modeling uncertainty, \(p^k\) is the system parameter vector.
As demonstrated in Sect. 2.6, the general EEG system dynamics \(\phi ^k(x;p^k):=F^k(x;p^k)+v^k(x;p^k)\) can be accurately derived and preserved in constant RBF neural networks \(\bar{W}^{k^T}S(x)\). By utilizing the learned knowledge obtained in the training stage, a bank of M estimators is constructed for the training EEG signal patterns as follows:
where \(k=1,\ldots ,M\) is used to stand for the kth estimator, \(\bar{\chi }^k=[\bar{\chi }_1^k,\ldots ,\bar{\chi }_n^k]^T\) is the state of the estimator, \(B=diag[b_1, \ldots , b_n]\) is a diagonal matrix which is kept the same for all estimators, x is the state of an input test EEG signal pattern generated from Eq. (8).
In the classification phase, by comparing the test EEG signal pattern (standing for a normal, interictal or ictal EEG signal pattern) generated from EEG system (8) with the set of M estimators (18), we obtain the following test error systems:
where \(\tilde{\chi }_i^k=\bar{\chi }_i^k-x_i\) is the state estimation (or synchronization) error. We compute the average \(L_1\) norm of the error \(\tilde{\chi }_i^k(t)\)
where \(\mathrm {T}_c\) is the cycle of EEG signals.
The fundamental idea of the classification between normal, interictal and ictal EEG signals is that if a test EEG signal pattern is similar to the trained EEG signal pattern \(s~(s\in \{1,\ldots ,k\})\), the constant RBF network \(\bar{W}_i^{s^T}S_i(x)\) embedded in the matched estimator s will quickly recall the learned knowledge by providing accurate approximation to EEG system dynamics. Thus, the corresponding error \(\Vert \tilde{\chi }_i^s(t)\Vert _1\) will become the smallest among all the errors \(\Vert \tilde{\chi }_i^k(t)\Vert _1\). Based on the smallest error principle, the appearing test EEG signal pattern can be classified. We have the following classification scheme.
Classification scheme If there exists some finite time \(t^s,s\in \{1,\ldots ,k\}\) and some \(i\in \{1,\ldots ,n\}\) such that \(\Vert \tilde{\chi }_i^s(t)\Vert _1<\Vert \tilde{\chi }_i^k(t)\Vert _1\) for all \(t>t^s\), then the appearing EEG signal pattern can be classified.
3 Experimental results
Experiments are implemented using matlab software and tested on an Intel Core i7 6700K 3.5 GHz computer with 32 GB RAM. We assign feature vector sequences for all the EEG signals in the Bonn database. Based on the method described in Sect. 2.5, we extract features through EEG signal time series which means input of the RBF neural networks \(x=[ED_j^{PRC1^{D4}},ED_j^{PRC1^{A4}},ED_j^{PRC2^{D4}},ED_j^{PRC2^{A4}}]^T\). In order to eliminate data difference between different features, all feature data are normalized to \([-1, 1]\).
Several experiments are carried out to verify the effectiveness of the proposed method. The classification results will be evaluated with the 10-fold and leave-one-out cross-validation styles. The data are divided into the training and test subsets. For the 10-fold cross-validation, the data set is divided into ten subsets. Each time, one of the ten subsets is used as the test set and the other night subsets are put together to form a training set. For the leave-one-out cross-validation style, each time we select one EEG signal pattern for classification, the rest of the EEG signal patterns for training. This process is repeated K (representing the number of EEG signal patterns) times and the leave-one-out classification accuracy is calculated as the average of the classification accuracy of all of the individually left-out patterns.
For the evaluation, six performance parameters are used including the Sensitivity (SEN), the Specificity (SPF), the Accuracy (ACC), the Positive Predictive Value (PPV), the Negative Predictive Value (NPV) and the Matthews Correlation Coefficient (MCC) (Azar and El-Said 2014). To be accurate, a classifier must have a high classification accuracy, a high sensitivity, as well as a high specificity (Chu 1999). For a larger value of MCC, the classifier performance will be better (Azar and El-Said 2014; Yuan et al. 2007).
In the past literature, various approaches focused on the classification of EEG signals between Sets A and E. However the effectiveness of the classification between different groups of datasets was not investigated thoroughly. It is therefore more desirable to figure out the ability of the proposed method to classify EEG signals containing different combinations of datasets (Z, O, N, F and S). To address this issue, 11 different classification problems are made from aforementioned datasets. All experiments described in Table 3 focus on distinguishing normal, interictal and ictal EEG signals. Cases 1 to 8 deal with the binary classification while cases 9 to 11 accomplish multi-class classification.
The classification results on different cases have been illustrated in Tables 4, 5, 6, 7, 8, 9, 10, 11, 12, 13 and 14 with 10-fold and leave-one-out cross-validation styles. Our study demonstrates the accuracy improvements to differentiate between normal, interictal and ictal EEG signals. Overall, our classification approach achieves good performance, which indicates that the proposed pattern recognition system can effectively differentiate between different classes of EEG signals by using nonlinear features and neural network based classification tools.
4 Discussion
The experimental results of this study demonstrate that normal, interictal and ictal EEG signals could be detected automatically by means of hybrid feature extraction methods and neural networks. The proposed scheme focuses not only on providing evidence to support the claim that interictal and ictal EEG signals demonstrate altered dynamics compared to normal EEG signals, but also on providing an automatic and objective method to distinguish between the three groups of EEG signals.
Recently, different methods reported in the literature have been proposed to automatically detect the normal, interictal and ictal EEG signals. It should be noted that all of the recent methods demonstrated in Table 15 were evaluated using 10-fold cross-validation.
For case 1 (A–E), Isik and Sezer (2012) used tools including Wavelet Transform (WT), Multilayer Perceptron (MLP) and Elman artificial neural networks (ANN), and the achieved classification accuracy was 96%. Du et al. (2012) extracted principal component features using principal components analysis (PCA) on 15 high-order spectra (HOS) features. Then eight different classifier including ANN, MLP, RBF network, random forest, rotation forest, logistic regression, model trees, simple logistic regression, and bagging were employed to evaluate the classification performance of the proposed features, in which the simple logistic regression achieved the highest accuracy of 94.5%. Zhang et al. (2018) combined fuzzy distribution entropy with wavelet packet decomposition, Kruskal–Wallis nonparametric one-way analysis of variance and k-nearest neighbor (KNN) classifier to classify the EEG signals and the achieved best accuracy was 100%. In our proposed method, the achieved accuracy is 99%.
For cases 2–4, the classification accuracy achieved by our proposed method is 99.25%, 99.02%, and 98.18% respectively. Tawfik (2016) achieved the classification accuracy of 85%, 93.5%, and 96.5%, respectively, for these experimental cases with the combination of weighted permutation entropy and SVM. Recently, with the development of deep learning method, Ahmedt-Aristizabal et al. (2018) used recurrent Neural Networks (RNNs) via the use of Long-Short Term Memory (LSTM) networks and achieved classification accuracy of 94.75%, 97.25%, and 96.5% for these experimental cases, respectively. However, there are efficient formulas behind deep learning success (Goceri 2018), parameters such as batch size should be chosen carefully (Goceri and Gooya 2018). In comparison, the achieved accuracy for cased 2–4 in our proposed method is 99.5%, 98.5% and 99.5%, respectively.
For cases 5–7, our proposed method achieved the classification accuracy of 98%, 99.5% and 98%, respectively. For case 5 (NF-S), Joshi et al. (2014) utilized the fractional linear prediction technique together with the SVM classifier and the reported classification accuracy was 95.33%. For the same case, Diykh et al. (2017) used complex networks approach and reported the classification accuracy of 97.8%. For case 6 (Z–F), Jaiswal and Banka (2017) employed the local neighbor descriptive pattern (LNDP) and one-dimensional local gradient pattern (1D-LGP) together with ANN for the classification and reported the accuracy of 99.90%. Kaya et al. (2014) used one-dimensional local binary pattern (1D-LBP) to extract the histogram features and fed them into the BayesNet classifier for the classification. The reported accuracy was 99.50%. For case 7 (ZONF-S), Kumar et al. (2014) used the DWT-based fuzzy approximate entropy to extract features and fed them into the SVM classifier to achieve the classification accuracy of 97.38%. Mursalin et al. (2017) used the improved correlation-based feature selection method (ICFS) together with Random Forest classifier and reported the classification accuracy of 97.4%.
For case 8 (ZO–NFS), Kaya et al. (2014) reported accuracy of 93%. Acharya et al. (2018) used the 13-layer deep convolutional neural network (CNN) algorithm and reported the accuracy of 88.7%. In comparison, the achieved accuracy in our proposed method is 95.2%.
Case 9 (Z–F–S) is a multi-class classification problem including three classes. Jaiswal and Banka (2017) reported the accuracy of 98.22% and 97.06% by using LNDP and 1D-LGP, respectively. Kaya et al. (2014) reported the classification accuracy of 95.67% with 1D-LBP. Li et al. (2017) used the dual-tree complex wavelet transform (DT-CWT) to decompose EEG signals into five constituent sub-bands, which were associated with the nonlinear features of Hurst exponent (H), Fractal Dimension (FD) and Permutation Entropy (PE). Then four classifiers including SVM, KNN, random forest and rotation forest were employed and the reported classification accuracy was 98.87%. In comparison, the achieved accuracy in our proposed method is 99%.
Case 10 (ZO–NF–S) is another multi-class classification problem including three classes. Wang et al. (2011) reported the accuracy of 97.13% by using wavelet packet entropy features together with a classifier of artificial neural network. Acharya et al. (2012) reported the classification accuracy of 99% with Wavelet packet decomposition and Gaussian mixture model. In comparison, the achieved accuracy in our proposed method is 99.4%.
Case 11 (Z–O–N–F–S) is a five-class classification problem. Zahra et al. (2017) decomposed the EEG signal to its multiple intrinsic scales by using the multivariate empirical mode decomposition algorithm. After removing the intrinsic mode functions (IMFs) belonging to noise and other unnecessary artifacts, classification on the remainder of IMFs has been performed by employing a feature vector via artificial neural network framework. The reported accuracy was 87.2%. In comparison, the achieved accuracy in our proposed method is 94%.
Different from the above discussed methods, this study proposes a hybrid method to extract effective features based on ITD, DWT, PSR and ED. These features are fed into dynamical estimators which are consisting of RBF neural networks to classify different classes of EEG signals. Comparison of the classification performance to other state-of-the-art methods on the same database is demonstrated in Table 15. The proposed method provides an average classification accuracy of 98.15% for the eleven cases through 10-fold cross-validation. Due to the use of 10-fold cross-validation, the classification performance is robust. The method studied in this paper has the potential to serve as a supportive technical means to other approaches such as fMRI for the diagnosis of epilepsy.
Because the dimension of the features and the number of the neurons used in the study is 4 and 83,521, respectively, the computational load is relatively high. It is also time-consuming to carry out the ITD and DWT computation which may increase the complexity. However, with the development of computer technology, more powerful workstations and high-performance computers have been used to improve the computational capacity and reduce the computing time. This makes it easier to carry out ITD and DWT computation and become applicable in real-time applications, which significantly reduces the complexity. Hence it is acceptable to implement the experiments on an Intel Core i7 6700K 3.5 GHz computer with 32 GB RAM in the present study. In future work the authors will try to optimize the algorithm structure and adopt new computing technology and equipment to improve the computational performance and further reduce the complexity.
In general, the experimental results have shown that the proposed method can acquire high accuracy in epilepsy detecting on two-class, three-class and five-class classification problems. This demonstrates that our scheme is appropriate in solving problems with multiple classes. Automated analysis of epileptic seizure activity has a strong clinical potential. Also, it can be more important to produce mobile health technologies about the disease (Goceri and Songul 2018). Another important property is its computational simplicity after employing high-performance computers, which reduces the complex and makes it possible to be deployed in clinical applications. Consequently, this new approach can better meet clinical demands in the aspects of efficiency, functionality, universality and simplicity with satisfactory accuracy. These characteristics would make this method become an attractive alternative offer for actual clinical diagnosis. There are many factors in the proposed method that work together to improve the classification performance, which includes the following advantages. ITD could extract most important information of the EEG signals through predominant PRCs. DWT decomposes the predominant PRCs into different frequency bands, which are used to construct the reference variables. PSR plots EEG system dynamics along the advisable db3 sub-bands (D4 and A4) of PRCs trajectory in a 3D phase space diagram and visualizes the EEG system dynamics. ED measures and derives features, which are fed into RBF neural networks for the modeling, identification and classification of EEG system dynamics between normal, interictal and ictal EEG signals. However, some limitations such as the regulation principle of the embedding dimension and time lag, the relationship between the classification performance and the PSR parameters, still need to be improved and overcome. It would be of interest to develop strategy for adaptive selection of PSR parameters which could create best classification performance.
5 Conclusions
In this study, effective feature extraction techniques including ITD, DWT, PSR and ED have been introduced for epileptic EEG signal classification. All the techniques extract informative features for classification, which are computationally simple and easy to implement. The results of this study indicate that the pattern classification of EEG signals can offer an objective method to assess the disparity of EEG system dynamics between normal, interictal and ictal EEG signals. However, some limitations such as the relatively small size of the database, the regulation principle of the embedding dimension and time lag, still need to be improved and overcome. In future research, features introduced in other methods such as complete ensemble empirical mode decomposition (CEEMD), various entropies, Hurst exponent, mean-frequency (MF) and root-mean-square (RMS) bandwidth, Lempel–Ziv complexity, largest Lyapunov exponent, fractal dimension and other nonlinear features, can also be explored in the proposed framework to evaluate its classification performance. The results of the present study can be improved further by using wide database with more patients and various features. In addition, the future scope of this research will include identification of seizure stages besides the seizure detection part.
References
Aarabi A, He B (2017) Seizure prediction in patients with focal hippocampal epilepsy. Clin Neurophysiol 128(7):1299–1307
Acharya UR, Sree SV, Alvin APC, Suri JS (2012) Use of principal component analysis for automatic classification of epileptic EEG activities in wavelet framework. Expert Syst Appl 39(10):9072–9078
Acharya UR, Sree SV, Swapna G, Martis RJ, Suri JS (2013) Automated EEG analysis of epilepsy: a review. Knowl Based Syst 45:147–165
Acharya UR, Fujita H, Sudarshan VK, Bhat S, Koh JE (2015) Application of entropies for automated diagnosis of epilepsy using EEG signals: a review. Knowl Based Syst 88:85–96
Acharya UR, Oh SL, Hagiwara Y, Tan JH, Adeli H (2018) Deep convolutional neural network for the automated detection and diagnosis of seizure using EEG signals. Comput Biol Med 100:270–278
Ahmedt-Aristizabal D, Fookes C, Nguyen K, Sridharan S (2018) Deep classification of epileptic signals. arXiv preprint arXiv:1801.03610
An X, Jiang D, Chen J, Liu C (2012) Application of the intrinsic time-scale decomposition method to fault diagnosis of wind turbine bearing. J Vib Control 18(2):240–245
Andrzejak RG, Lehnertz K, Mormann F, Rieke C, David P, Elger CE (2001) Indications of nonlinear deterministic and finite-dimensional structures in time series of brain electrical activity: dependence on recording region and brain state. Phys Rev E 64(6):061907
Azar AT, El-Said SA (2014) Performance analysis of support vector machines classifiers in breast cancer mammography recognition. Neural Comput Appl 24:1163–1177
Bajaj V, Pachori RB (2012) Classification of seizure and nonseizure EEG signals using empirical mode decomposition. IEEE Trans Inf Technol Biomed 16(6):1135–1142
Bajaj V, Pachori RB (2013) Epileptic seizure detection based on the instantaneous area of analytic intrinsic mode functions of EEG signals. Biomed Eng Lett 3(1):17–21
Bhati D, Pachori RB, Gadre VM (2017a) A novel approach for time-frequency localization of scaling functions and design of three-band biorthogonal linear phase wavelet filter banks. Digit Signal Process 69:309–322
Bhati D, Sharma M, Pachori RB, Gadre VM (2017b) Time-frequency localized three-band biorthogonal wavelet filter bank using semidefinite relaxation and nonlinear least squares with epileptic seizure EEG signal classification. Digit Signal Process 62:259–273
Bhattacharyya A, Pachori RB (2017) A multivariate approach for patient-specific EEG seizure detection using empirical wavelet transform. IEEE Trans Biomed Eng 64(9):2003–2015
Bhattacharyya A, Pachori R, Upadhyay A, Acharya U (2017) Tunable-Q wavelet transform based multiscale entropy measure for automated classification of epileptic EEG signals. Appl Sci 7(4):385
Chen B, He Z, Chen X, Cao H, Cai G, Zi Y (2011) A demodulating approach based on local mean decomposition and its applications in mechanical fault diagnosis. Meas Sci Technol 22(5):055704
Chen M, Fang Y, Zheng X (2014) Phase space reconstruction for improving the classification of single trial EEG. Biomed Signal Process Control 11:10–16
Chu K (1999) An introduction to sensitivity, specificity, predictive values and likelihood ratios. Emerg Med Australas 11(3):175–181
Diykh M, Li Y, Wen P (2017) Classify epileptic EEG signals using weighted complex networks based community structure detection. Expert Syst Appl 90:87–100
Djemili R, Bourouba H, Korba MA (2016) Application of empirical mode decomposition and artificial neural network for the classification of normal and epileptic EEG signals. Biocybern Biomed Eng 36(1):285–291
Du X, Dua S, Acharya RU, Chua CK (2012) Classification of epilepsy using high-order spectra features and principle component analysis. J Med Syst 36(3):1731–1743
Feng Z, Lin X, Zuo MJ (2016) Joint amplitude and frequency demodulation analysis based on intrinsic time-scale decomposition for planetary gearbox fault diagnosis. Mech Syst Signal Process 72:223–240
Frei MG, Osorio I (2007) Intrinsic time-scale decomposition: time-frequency-energy analysis and real-time filtering of non-stationary signals. In: Proceedings of the royal society of London A: mathematical, physical and engineering sciences, vol 463, no 2078. The Royal Society, pp 321–342
Goceri E (2018) Formulas behind deep learning success. In: International conference on applied analysis and mathematical modeling, p 156
Goceri E, Gooya A (2018) On the importance of batch size for deep learning. Paper presented at the international conference on mathematics. Istanbul, Turkey
Goceri E, Songul C (2018) Mobile health technologies for patients with mental illness. In: International conference on advanced technologies, Antalya, Turkey, p 146
Hassan AR, Siuly S, Zhang Y (2016) Epileptic seizure detection in EEG signals using tunable-Q factor wavelet transform and bootstrap aggregating. Comput Methods Programs Biomed 137:247–259
Huang B, Kunoth A (2013) An optimization based empirical mode decomposition scheme. J Comput Appl Math 240:174–183
Huang NE, Shen Z, Long SR, Wu MC, Shih HH, Zheng Q, Liu HH (1998) The empirical mode decomposition and Hilbert spectrum for nonlinear and non-stationary time series analysis. Proc R Soc Lond A Math Phys Eng Sci R Soc 454(1971):903–995
Isik H, Sezer E (2012) Diagnosis of epilepsy from electroencephalography signals using multilayer perceptron and Elman artificial neural networks and wavelet transform. J Med Syst 36(1):1–13
Jaiswal AK, Banka H (2017) Local pattern transformation based feature extraction techniques for classification of epileptic EEG signals. Biomed Signal Process Control 34:81–92
Jia J, Goparaju B, Song J, Zhang R, Westover MB (2017) Automated identification of epileptic seizures in EEG signals based on phase space representation and statistical features in the CEEMD domain. Biomed Signal Process Control 38:148–157
Joshi V, Pachori RB, Vijesh A (2014) Classification of ictal and seizure-free EEG signals using fractional linear prediction. Biomed Signal Process Control 9:1–5
Kafashan M, Ryu S, Hargis MJ, Laurido-Soto O, Roberts DE, Thontakudi A, Ching S (2017) EEG dynamical correlates of focal and diffuse causes of coma. BMC Neurol 17(1):197
Kaya Y, Uyar M, Tekin R, Yildirim S (2014) 1D-local binary pattern based feature extraction for classification of epileptic EEG signals. Appl Math Comput 243:209–219
Kumar Y, Dewal ML, Anand RS (2014) Epileptic seizure detection using DWT based fuzzy approximate entropy and support vector machine. Neurocomputing 133:271–279
Kumar TS, Kanhangad V, Pachori RB (2015) Classification of seizure and seizure-free EEG signals using local binary patterns. Biomed Signal Process Control 15:33–40
Lahmiri S (2018) Generalized Hurst exponent estimates differentiate EEG signals of healthy and epileptic patients. Physica A Stat Mech Appl 490:378–385
Lee SH, Lim JS, Kim JK, Yang J, Lee Y (2014) Classification of normal and epileptic seizure EEG signals using wavelet transform, phase-space reconstruction, and Euclidean distance. Comput Methods Programs Biomed 116(1):10–25
Li S, Zhou W, Yuan Q, Geng S, Cai D (2013) Feature extraction and recognition of ictal EEG using EMD and SVM. Comput Biol Med 43(7):807–816
Li Y, Xu M, Wei Y, Huang W (2015) Rotating machine fault diagnosis based on intrinsic characteristic-scale decomposition. Mech Mach Theory 94:9–27
Li M, Chen W, Zhang T (2017a) Classification of epilepsy EEG signals using DWT-based envelope analysis and neural network ensemble. Biomed Signal Process Control 31:357–365
Li M, Chen W, Zhang T (2017b) Automatic epileptic EEG detection using DT-CWT-based non-linear features. Biomed Signal Process Control 34:114–125
Li Y, Cui WG, Huang H, Guo YZ, Li K, Tan T (2019) Epileptic seizure detection in EEG signals using sparse multiscale radial basis function networks and the Fisher vector approach. Knowl Based Syst 164:96–106
Martis RJ, Tan JH, Chua CK, Loon TC, Yeo SWJ, Tong L (2015) Epileptic EEG classification using nonlinear parameters on different frequency bands. J Mech Med Biol 15(03):1550040
Merigó JM, Casanovas M (2011) Induced aggregation operators in the Euclidean distance and its application in financial decision making. Expert Syst Appl 38:7603–7608
Michael S (2005) Applied nonlinear time series analysis: applications in physics, physiology and finance, vol 52. World Scientific, Singapore
Mursalin M, Zhang Y, Chen Y, Chawla NV (2017) Automated epileptic seizure detection using improved correlation-based feature selection with random forest classifier. Neurocomputing 241:204–214
Pachori RB (2008) Discrimination between ictal and seizure-free EEG signals using empirical mode decomposition. Res Lett Signal Process 2008:14
Pachori RB, Bajaj V (2011) Analysis of normal and epileptic seizure EEG signals using empirical mode decomposition. Comput Methods Programs Biomed 104(3):373–381
Pachori RB, Patidar S (2014) Epileptic seizure classification in EEG signals using second-order difference plot of intrinsic mode functions. Comput Methods Programs Biomed 113(2):494–502
Pachori RB, Sharma R, Patidar S (2015) Classification of normal and epileptic seizure EEG signals based on empirical mode decomposition. In: Zhu Q, Azar AT (eds) Complex system modelling and control through intelligent soft computations, studies in fuzziness and soft computing. Springer, Switzerland, pp 367–388
Park C, Looney D, Van Hulle MM, Mandic DP (2011) The complex local mean decomposition. Neurocomputing 74(6):867–875
Samiee K, Kovacs P, Gabbouj M (2015) Epileptic seizure classification of EEG time-series using rational discrete short-time Fourier transform. IEEE Trans Biomed Eng 62(2):541–552
Sato Y, Doesburg SM, Wong SM, Ochi A, Otsubo H (2015) Dynamic preictal relations in FCD type II: potential for early seizure detection in focal epilepsy. Epilepsy Res 110:26–31
Scheuer ML, Wilson SB (2004) Data analysis for continuous EEG monitoring in the ICU: seeing the forest and the trees. J Clin Neurophysiol 21(5):353–378
Sharma R, Pachori RB (2015) Classification of epileptic seizures in EEG signals based on phase space representation of intrinsic mode functions. Expert Syst Appl 42(3):1106–1117
Sharma RR, Pachori RB (2017a) Time-frequency representation using IEVDHM-HT with application to classification of epileptic EEG signals. IET Sci Meas Technol 12(1):72–82
Sharma M, Pachori RB (2017b) A novel approach to detect epileptic seizures using a combination of tunable-Q wavelet transform and fractal dimension. J Mech Med Biol 17(07):1740003
Sharma M, Pachori RB, Acharya UR (2017) A new approach to characterize epileptic seizures using analytic time–frequency flexible wavelet transform and fractal dimension. Pattern Recognit Lett 94:172–179
Sharma RR, Varshney P, Pachori RB, Vishvakarma SK (2018) Automated system for epileptic EEG detection using iterative filtering. IEEE Sens Lett 2(4):1–4
Shayegh F, Sadri S, Amirfattahi R, Ansari-Asl K (2014) A model-based method for computation of correlation dimension, Lyapunov exponents and synchronization from depth-EEG signals. Comput Methods Programs Biomed 113(1):323–337
Sheintuch L, Friedman A, Efrat N, Tifeeret C, Shorer Z, Neuman I, Shallom I (2014) O16: detection of epileptiform activity using multi-channel linear prediction coefficients and localization of epileptic foci based on EEG-fMRI data. Clin Neurophysiol 125:S33
Sivakumar B (2002) A phase-space reconstruction approach to prediction of suspended sediment concentration in rivers. J Hydrol 258(1–4):149–162
Som A, Krishnamurthi N, Venkataraman V, Turaga P (2016) Attractor-shape descriptors for balance impairment assessment in Parkinson’s disease. In: IEEE conference on engineering in medicine and biology society, pp 3096–3100
Song JL, Hu W, Zhang R (2016) Automated detection of epileptic EEGs using a novel fusion feature and extreme learning machine. Neurocomputing 175:383–391
Subasi A, Gursoy MI (2010) EEG signal classification using PCA, ICA, LDA and support vector machines. Expert Syst Appl 37(12):8659–8666
Subasi A, Kevric J, Canbaz MA (2019) Epileptic seizure detection using hybrid machine learning methods. Neural Comput Appl 31(1):317–325
Sun Y, Li J, Liu J, Chow C, Sun B, Wang R (2015) Using causal discovery for feature selection in multivariate numerical time series. Mach Learn 101(1–3):377–395
Takens F (1980) Detecting strange attractors in turbulence. In: Dynamical systems and turbulence, Warwick 1980, Springer, Berlin, 1981, pp 366–381
Tawfik NS, Youssef SM, Kholief M (2016) A hybrid automated detection of epileptic seizures in EEG records. Comput Electr Eng 53:177–190
Timothy LT, Krishna BM, Nair U (2017) Classification of mild cognitive impairment EEG using combined recurrence and cross recurrence quantification analysis. Int J Psychophysiol 120:86–95
Tiwari AK, Pachori RB, Kanhangad V, Panigrahi BK (2016) Automated diagnosis of epilepsy using key-point-based local binary pattern of EEG signals. IEEE J Biomed Health Inform 21(4):888–896
Ullah I, Hussain M, Aboalsamh H (2018) An automated system for epilepsy detection using EEG brain signals based on deep learning approach. Expert Syst Appl 107:61–71
Vavadi H, Ayatollahi A, Mirzaei A (2010) A wavelet-approximate entropy method for epileptic activity detection from EEG and its sub-bands. J Biomed Sci Eng 3(12):1182
Venkataraman V, Turaga P (2016) Shape distributions of nonlinear dynamical systems for video-based inference. IEEE Trans Pattern Anal Mach Intell 38(12):2531–2543
Wang C, Hill DJ (2006) Learning from neural control. IEEE Trans Neural Netw 17(1):130–146
Wang C, Hill DJ (2007) Deterministic learning and rapid dynamical pattern recognition. IEEE Trans Neural Netw 18(3):617–630
Wang C, Hill DJ (2009) Deterministic learning theory for identification, recognition and control. CRC Press, Boca Raton
Wang D, Miao D, Xie C (2011) Best basis-based wavelet packet entropy feature extraction and hierarchical EEG classification for epileptic detection. Expert Syst Appl 38(11):14314–14320
Wang L, Xue W, Li Y, Luo M, Huang J, Cui W, Huang C (2017) Automatic epileptic seizure detection in EEG signals using multi-domain feature extraction and nonlinear analysis. Entropy 19(6):222
Xing Z, Qu J, Chai Y, Tang Q, Zhou Y (2017) Gear fault diagnosis under variable conditions with intrinsic time-scale decomposition-singular value decomposition and support vector machine. J Mech Sci Technol 31(2):545–553
Xu B, Jacquir S, Laurent G, Bilbault JM, Binczak S (2013) Phase space reconstruction of an experimental model of cardiac field potential in normal and arrhythmic conditions. In: 35th annual international conference of the IEEE engineering in medicine and biology society, pp 3274–3277
Yuan Q, Cai C, Xiao H, Liu X, Wen Y (2007) Diagnosis of breast tumours and evaluation of prognostic risk by using machine learning approaches. In: Huang DS, Heutte L, Loog M (eds) Advanced intelligent computing theories and applications. With aspects of contemporary intelligent computing techniques. Springer, Berlin, pp 1250–1260
Zahra A, Kanwal N, ur Rehman N, Ehsan S, McDonald-Maier KD (2017) Seizure detection from EEG signals using multivariate empirical mode decomposition. Comput Biol Med 88:132–141
Zhang T, Chen W (2016) LMD based features for the automatic seizure detection of EEG signals using SVM. IEEE Trans Neural Syst Rehabilit Eng 25(8):1100–1108
Zhang Y, Zhou W, Yuan S, Yuan Q (2015) Seizure detection method based on fractal dimension and gradient boosting. Epilepsy Behav 43:30–38
Zhang T, Chen W, Li M (2017) AR based quadratic feature extraction in the VMD domain for the automated seizure detection of EEG using random forest classifier. Biomed Signal Process Control 31:550–559
Zhang T, Chen W, Li M (2018) Fuzzy distribution entropy and its application in automated seizure detection technique. Biomed Signal Process Control 39:360–377
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant Nos. 61773194, 61304084), by the Natural Science Foundation of Fujian Province (Grant No. 2018J01542), by Fujian Provincial Training Foundation For “Bai-Qian-Wan Talents Engineering”, by the Program for New Century Excellent Talents in Fujian Province University and by the Training Program of Innovation and Entrepreneurship for Undergraduates (Grant No. 201811312002).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zeng, W., Li, M., Yuan, C. et al. Identification of epileptic seizures in EEG signals using time-scale decomposition (ITD), discrete wavelet transform (DWT), phase space reconstruction (PSR) and neural networks. Artif Intell Rev 53, 3059–3088 (2020). https://doi.org/10.1007/s10462-019-09755-y
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10462-019-09755-y