Abstract
A total of 60 offspring obtained from controlled reciprocal crosses of ‘Mamaei’ and ‘Marcona’ cultivars of almond along with parents were evaluated using 31 morphological and biochemical traits of the nut and kernel. According to the results, attribute values such as shell weight (1.39–5.68 g), shell retention (0–5 codes), nut weight (2.16–7.60 g), kernel weight (0.35–2.51 g), and double kernel percentage (0–60%) were variable. These traits showed a high coefficient of variation. Also, the values of biochemical characteristics and compounds of kernel include protein (14.54–25.16%), ash (1.79–4.69%), oil percentage (47.93–62.94%), soluble carbohydrates (0.64–4.65%), insoluble carbohydrate (3.30–7.13%), and vitamin E content (71–220 mg · 100 g−1 FW) were variable in the offspring and parents. Simple correlation coefficients between traits showed a significant positive or negative correlation between some measured characteristics related to the nut, kernel, and biochemical characteristics. The principal component analysis (PCA) reduced the evaluated traits to 10 main factors that explained 74.45% of the total variance. The nut and kernel traits greatly affected the differentiation of cultivars and genotypes. Cluster analysis at 25 Euclidean distances divided cultivars and offspring into two main groups. Attributes such as nut length, width, and shape, as well as protein content and kernel taste, effectively differentiated the parents and progenies. By reducing the Euclidean distance, cultivars and offspring were divided into four main groups. Among the important factors for separating the main clusters were traits such as the length, width, and thickness of nut, as well as the weight, length, width, size, and taste of kernel and shell thickness. Finally, 16 offspring, including ‘MarM2’, ‘MarM4’, ‘MarM5’, ‘MarM6’, ‘MarM8’, ‘MarM10’, ‘MarM17’, ‘MMar2’, ‘MMar6’, ‘MMar7’, ‘MMar15’, ‘MMar24’, ‘MMar25’, ‘MMar27’, ‘MMar28’, and ‘MMar33’ with parents ‘Mamaei’ and ‘Marcona’ cultivars, showed relative superiority over other offspring in terms of morphological and biochemical traits. The identified superior offspring, which were even superior to the parents in some important almond breeding traits, are good candidates for use in almond breeding programs to achieve improved cultivars.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Almond (Prunus dulcis L.) is a valuable nut grown in many temperate and subtropical regions for domestic consumption and export (Yada et al. 2011). P. dulcis L. belongs to the genus Prunus, the subspecies Amygdalus of the family Rosaceae (Ayaz et al. 2020; Zahedi et al. 2020). The almond kernel contains significant content of protein, amino acids, fatty acids, carbohydrates, vitamins, and minerals (Barreca et al. 2020). Iran, with a cultivated area of 79392 ha, is considered one of the most important producers of almonds, and with an annual production of about 164,348 tons, it ranks third in the world after the United States and Spain (FAO 2021).
In successful breeding programs, the identification and evaluation of superior germplasm are essential. They are a necessary step in preserving hereditary reserves, which is the fundamental basis for genetic research and breeding programs. The breed and production of cultivars depend on the careful selection between plants, which depends on the identification of cultivars and their diversity (Hajnajari et al. 2019; Hajivand et al. 2020; Valaei and Bernousi 2020).
In almond breeding programs, the relationships between the traits and their correlations should be determined. The significant correlation between characteristics indirectly helps select important attributes, facilitating and accelerating breeding programs (Hansche et al. 1972). Studies have shown that the genetic diversity in the germplasm resulting from almond crosses in different parts of Italy will lead to valuable sources of genes (Rigoldi et al. 2015).
A total of 137 accessions from 18 wild almond species native to Iran showed that the weight and width of the nut as well as the weight of the kernel had a higher coefficient of variation (45.8%) than the other studied traits and rich sources of new germplasm of almond help improve crop yield (Sorkheh et al. 2009). A reduction in the fruit size, pollen-ovary self-incompatibility, and bitter taste of the kernel are the most common barriers to germplasm use in wild almond species (Sorkheh et al. 2009).
The quantitative and qualitative characteristics of fruit in the 50 superior almond genotypes and ‘Tuono’ cultivar were investigated (Estaji et al. 2013, 2014). The results showed that the studied genotypes had significant differences with each other in terms of all studied traits and characteristics, such as nut size, shriveling of kernel, length, and thickness of the kernel, softness of shell, and marking of outer shell had the largest share of difference between genotypes (Estaji et al. 2013, 2014). Study of 94 almond offspring and their parents showed that traits such as fruit length and dry weight most change and dry kernel weight and nut thickness showed the least differences among other characteristics. On the other hand, estimating the percentage of heritability for oil content was the parents’ average (Turkmen et al. 2020).
The ‘Mamaei’ is an almond variety native to Iran. In terms of growth habit, it has a spreading, and fruiting habit is on a 1-year branch. Flowering time is early to medium (compared to ‘Ferragnes’ as very late flowering cultivar), and it has big flowers, white color and is self-incompatible (Imani and Shamili 2018b). The softness of shell in this variety is hard (difficult to break, need hammer), the size of fruit is large, shape of fruit and kernel is elongated. Also, in this cultivar, the color of kernel is bright yellow, percentage of kernel is 40–45%, double kernel is more than 50%, and quality and vitamin E of kernel are high and better than other Iranian almond cultivars. ‘Mamaei’ yields are medium to high, about 2–2.2 tons per hectare.
The ‘Marcona’ variety originated in Spain. The growth habit of this cultivar is semi-upright, strong vigor, flowering time is intermediate to late (compared to ‘Tardy nonpareil’ an extremely late flowering cultivar), with small flowers, self-incompatible, and mixed fruiting habit (1-year branch and spur) (Imani and Shamili 2018b; Pérez-Sánchez and Morales-Corts 2021). The softness of shell in this variety is hard (difficult to break, need hammer), fruit shape is round, kernel percentage is 32–35%, the percentage of double is 0–2%, and kernel color is brownish yellow. The average yield of ‘Marcona’ is 2.5–3 tons per hectare (Imani and Shamili 2018b; Pérez-Sánchez and Morales-Corts 2021). Based on Mendelian inheritance, it is possible to create progeny with superior fruit and kernel traits in the mutual crossing of two cultivars, ‘Mamaei’ and ‘Marcona’.
Also, many studies addressing the morphological and biochemical characterization of almond cultivars have been undertaken in Iran and different countries mainly located around the Mediterranean basin by Sepahvand et al. (2014), Imani and Shamili (2018b), Rasouli et al. (2019), Khojand et al. (2022), Montero-Riquelme (1993), Felipe (2000), Cordeiro et al. (2001), Kodad (2008), Kodad and Socias i Company (2008), Vargas et al. (2009), Kodad et al. (2013), and Pérez-Sánchez and Morales-Corts (2021).
Multivariate statistical methods are needed to evaluate cultivars and genotypes based on quantitative and qualitative properties and relationships. Multivariate statistical method is a statistical technique that simultaneously analyzes data with more than one statistical variable (Todde et al. 2016). Among these methods, factor and cluster analysis are more valuable. Factor analysis method can put the number of traits evaluated in effective groups. In cluster analysis, individuals in a cluster have many similarities in terms of the studied features, and individuals in separate clusters are more heterogeneous in terms of traits (Estaji et al. 2013).
This study aimed to evaluate 60 almond offspring resulting from the controlled reciprocal crossing of ‘Mamaei’ and ‘Marcona’ cultivars with parents and the relationships between their important morphological and biochemical traits. Also, the selection of superior offspring compared to other progeny and parents, based on the evaluated features, to be used in almond breeding programs was another target of this research.
Materials and Methods
Plant Material
This research was carried out in the Kamal Abad collection of the Seed and Plant Breeding Research Institute (SPBRI), affiliated to the Ministry of Agriculture Jihad of Iran, during 2019–2020. The study investigated 60 almond offspring obtained from controlled reciprocal crosses between ‘Mamaei’ and ‘Marcona’ cultivars (Table 1) with parents. The method of naming the offspring (5 years old) was conducted based on the first letters of the maternal and paternal parents (Mamaei: M and Marcona: Mar).
Measurement of Traits
In all, 31 morphological, biochemical, and qualitative characteristics of the nut and kernel were evaluated (Table 2). A total of 20 fruits were randomly harvested from different directions of the tree and used to study the traits. Some traits were coded based on the descriptor of almond (Gulcan 1985; UPOV 2011) (Table 2). We measured ash by burning organic matter up to 575 °C (Aktas et al. 2015). After moisture measurements, about 2.5 g of dry samples were placed in an electric furnace at 250 °C (with 10 °C/min intervals). The temperature held constant at 250 °C for 30 min and then increased to 575 °C (10 °C/min intervals) for 3 h and then cooled down to 100 °C. The ash was immediately moved to a desiccator while cooling to room temperature before measurements. After weighing, samples were kept in air-tight bags under desiccation to avoid the re-absorption of moisture (Aktas et al. 2015).
Total protein was determined according to Kjeldahl’s (V50 model, Bakhshi Company, Iran) method (Okay 2002; El Hawary et al. 2014). Samples were warmed up with sulphuric acid to decompose the organic matter via oxidation to release ammonium sulfate. The solution is then distilled with sodium hydroxide in order to convert the ammonium to ammonia. The amount of ammonia (and therefore the nitrogen) is determined by back titration using boric acid and sodium carbonate solution using methyl orange as a pH indicator.
In order to determine the content of soluble carbohydrates, 300 g of dried almond kernel samples were poured into 25 ml tubes and the extraction of soluble sugars was performed using Kochert’s (1987) method. To remove excess deposits and other waste compounds, 5 ml of 5% ZnSO4 and 4.7 ml of 0.3 N barium hydroxide were added to the tubes and vortexed again. Then the tubes were centrifuged at 3000 rpm at room temperature for 20 min. At the same time, 1 ml of illuminants and 1 ml of standard solutions were transferred to new 25 ml tubes and 0.5 ml of 5% phenol solution was added to each of them and shaken vigorously. Then, 2.5 ml of 98% sulfuric acid was added to the tubes by pressure using a syringe (dispenser). After 45 min, the absorbance of the samples was measured at a wavelength of 485 nm by a spectrophotometer (JENUS model, UV-1200, USA). Finally, the content of soluble sugars was calculated based on the absorption rate of the samples and comparing it with the standard.
In order to measure insoluble carbohydrates, after the extraction of soluble sugars, the resulting pulp was collected and dried in an oven (SHD96A, Iran) at 50 °C for 2 h. Then, 4.5 ml of distilled water and 6 ml of 51% perchloric acid were added to the samples and placed in the refrigerator for 14 h at a temperature of 4 °C and the continuation of the measurement method was carried out in the same way as the measurement of soluble sugars (Kochert 1987).
The Soxhlet method (Hamilton and Rossell 1986; Golzari et al. 2013) was used to measure oil percentage. After grinding, the almond powder was extracted using Soxhlet (SX100‑G model, Bakhshi Company, Iran) method (temperature 45 °C and solvent pure methanol and chloroform in the ratio of 50:50). The solvent in the extracted oil was separated using an oven under vacuum and the amount of oil was determined.
Vitamin E using the device high-performance liquid chromatography (HPLC) (unicam 200-crystal model, England) was measured with an array-photodiode detector (Çelik et al. 2019). The fluorescence detector was set at 295 nm wavelength and 330 nm wavelength for excitation and emission, respectively. The mobile phase consisted of methanol: distilled water (97:3 v/v). Flow rate was set at 1.05 ml min−1. Peaks were determined according to retention times compared to the known four standards (Sigma-Aldrich) (Çelik et al. 2019).
Statistical Analyses
The frequency of traits, descriptive statistics, simple correlation between characteristics, cluster analysis and factor analysis were performed using SPSS software version 25 according to Rasouli’s et al. (2013) method. To calculate the coefficient of variation, the standard deviation of each trait was calculated by dividing it by the average of that trait. Using factor rotation technique and varimax method, factor analysis was performed. In each main and independent factor, coefficients of 0.5 and above were considered significant (Rasouli et al. 2013; Imani and Shamili 2018b). Cluster analysis and grouping of offspring and cultivars were performed using Ward’s method or minimum variance based on square Euclidean distance and distances were calculated after data standardization (Rasouli et al. 2013). Also, to compare the parents and offspring data obtained from the measured traits, analyzed in the form of completely randomized design with three replications using SAS V9.1 software (SAS Institute Inc., Cary, NC, USA). Comparing the average data was conducted using Duncan’s multi-domain test. Finally, the superior offspring were identified regarding different traits (Azimi 2020, 2023).
Results and Discussion
Frequency of Traits
The changes in some of the studied traits in offspring had a normal distribution. The frequency percentages are shown in Figs. 1, 2, 3, 4, 5 and 6, respectively. The results showed that in most of the evaluated traits, such as softness of shell, marking of outer shell, nut shape, kernel pubescence, kernel size, and double kernel, the highest frequency distribution was related to the middle classes. The highest frequencies were obtained for softness of shell (code 4: semi-hard), marking of outer shell (code 5: medium), nut shape (code 3: oblong), kernel shape (code 3: low), kernel size (code 5: medium), and the percentage of double kernel (between 0–20%), which is consistent with the results of Rasouli et al.’s (2019) research in some traits. Estaji et al. (2013), to evaluate the morphological diversity of 50 genotypes of almonds, investigated 22 vegetative and reproductive characteristics, including kernel, fruit, and tree features. Their results showed that the highest diversity and coefficient of variation were reported in the traits of kernel size, marking in outer shell, shell softness, and percentage of double kernel, which was consistent with the results of this experiment.
Descriptive Statistics of Traits
The numerical average of some important traits measured in the studied offspring and cultivars is given in Table 3. According to the results, some characteristics such as shell weight, nut weight, kernel weight, and percentage of double kernel showed high diversity and coefficient of variation (Table 3). Based on the results, some traits such as shell weight (1.39–5.68 g), shell retention (0–5 codes), nut weight (2.16–7.60 g), kernel weight (0.35–2.51 g), and double kernel percentage (0–60%) were variable and these traits showed a high coefficient of variation. These results are coordinated with the findings of Sorkheh et al. (2009), who reported that the weight and width of the nut as well as the weight of the kernel had a higher coefficient of variation in almond species native to Iran. Regarding fruit traits, Socias i Company et al. (2017) also observed that the almond parameters are highly variable depending on the cultivar.
The range of changes in shell weight was 1.39–5.68% and the progenies of ‘MarM4’, ‘MMar13’, and ‘MMar14’ had the highest shell weight and the progenies of ‘MarM17’, ‘MarM9’, and ‘MMar24’ had the lowest shell weight, respectively (Table 4). The shell of almond is a function of the ratio of cellulose, hemicellulose, and lignin as the main components of the shell and is inversely related to the percentage of kernel. The higher kernel percentage, the lower shell softness and vice versa (Comas et al. 2019). The range of changes in nut weight varied from 2.16 to 7.60 g and the range of the kernel weight varied from 0.35 to 2.51 g (Tables 3 and 4). The diversity index obtained from both traits was 28.92 and 36.25%, respectively, which shows that changes in the weight of the nut and kernel were high among the progenies (Table 3). The mean kernel weight was 1.19 g, with offspring including ‘MMar24’, ‘MMar28’, and ‘MMar27’ having the highest kernel weight, while ‘MarM4’, ‘MarM16’, and ‘MMar6’ had the lowest kernel weight. The nut and kernel weight during packaging, transportation, and marketing is estimated as the most important quality trait of the product. Studies have shown that the most variables that directly and indirectly affect the weight of the nut include weight, thickness, length, and width of the kernel, respectively (Imani and Shamili 2018a; Pérez-Sánchez and Morales-Corts 2021).
The mean percent of double kernel was estimated to be 24.84% (Table 3), with the progenies of ‘MMar41’, ‘MMar33’, and ‘MMar27’ having the highest rate of percent (60%) of double kernel (Table 4). Double kernels are caused by fertilization and development of both ovules that are present in the ovary. Typically, one of the ovules is aborted. The presence of double kernel depends more on the genetics of the cultivar. The percentage of double kernel is considered an undesired trait of almond breeding programs, since the deformation of the kernels complicates the screening and grading process (Egea and Burgos 1995; Rasouli and Imani 2016; Imani and Shamili 2018a; Pérez-Sánchez and Morales-Corts 2021). Mosavi et al. (2009), to evaluate the morphological diversity of 55 cultivars and genotypes of almond, investigated 29 quantitative and qualitative traits of the nut and kernel. Their results showed that traits such as nut weight, kernel percentage, percentage of double kernels, softness, and shell thickness had the highest variation among the traits that were consistent with the results of this experiment in some cases. Due to the diversity of the studied traits, selecting different values for a trait is possible. Also, high-diversity traits can be used to evaluate progenies for more accurate statistical analysis.
Protein is one of the major chemical components of almond and cultivars with values above 23% has high nutrient quality (Pérez-Sánchez and Morales-Corts 2021). The range of protein value in kernel varied from 14.54% to 25.26%. Its mean content was 19.95%, with values above 23% in the ‘Marcona’ cultivars and ‘MarM2’, ‘MarM5’, ‘MarM6’, ‘MarM16’, ‘MMar15’, and ‘MMar32’ offspring (Table 5). This average result agrees with that reported by Kodad (2017) for Spanish almond cultivars (15.7–21.1% protein content of the kernel dry weight). Also, these results were consistent with Pérez-Sánchez and Morales-Corts (2021) findings that reported average protein content was 21.13% in almond cultivars grown in the central-western Iberian Peninsula.
The average soluble-carbohydrate and non-soluble carbohydrate content were 3.45% and 5.05%, respectively (Table 5). These results conformed to Pérez-Sánchez and Morales-Corts (2021), who reported that the average carbohydrate content was 3.89% in almond cultivars grown in the central-western Iberian Peninsula. Also, these results for kernel carbohydrate composition agree with those reported by Kodad (2017) for Spanish almond cultivars (1.8–7.6% carbohydrates content of the kernel fresh weight). In this sense, it is also important to point out that sugars, starch, and some sugar alcohols are the only carbohydrate forms present in the almond kernels that can be digested, absorbed, and metabolized by humans to provide a source of energy (Kodad 2017; Pérez-Sánchez and Morales-Corts 2021). Also, Schirra (1997) reported that soluble sugars, while present in relatively low amounts, are sufficient to make kernels sweet-tasting in almond.
Furthermore, the range of kernel ash varied from 1.79% to 4.69% and its average content was 3.32% (Table 5). Also, this result is in agreement with Romojaro et al. (1977), Saura-Calixto et al. (1981), and Pérez-Sánchez and Morales-Corts (2021), who reported low variability for this parameter in Spanish almond cultivars (3.05–3.60%). The almond kernel is considered a good source of mineral elements, playing an important role in human health (Kodad 2017).
The range of kernel oil percentage from 47.93% (‘MarM10’) to 62.94% (‘MMar17’) (Table 5). These results agree with those reported by Kodad et al. (2013) that the total almond kernel oil content varied from 48.7% to 64.59% in Spanish almond cultivars. Saura-Calixto et al. (1981) observed that the average kernel oil content was 53.37%. Martínez-Gómez et al. (2002) reported that 12 varieties of almonds contained between 30% and 51% oil. Khojand et al. (2022) stated that the kernel oil content changed from 53.67% to 54.26%. In some other studies, the quantity of P. dulcis oil was determined in the range of 45.9% to 61.7% (Mehran and Filssof 1974; Khojand et al. 2022).
Also, in this study, the highest content of kernel vitamin E was related to ‘MarM4’ (220 mg · 100 g−1 FW) (Tables 3 and 5). Furthermore, the range of kernel vitamin E varied from 71 to 220 mg · 100 g−1 FW and its average content was 125.82 mg · 100 g−1 FW (Tables 3 and 5). Studies show that the concentration of tocopherol as a precursor of vitamin E in almond oil depends on the genotype and climatic conditions of the year under investigation, as well as the environmental conditions of the region (Kodad 2008; Kodad and Socias i Company 2008; Vargas et al. 2009; Kodad et al. 2013). Different results have been obtained based on different cultivars and genotypes in other countries (Kodad et al. 2018; Pérez-Sánchez and Morales-Corts 2021).
Simple Correlation Coefficients of Traits
The correlation coefficients between the measured traits indicated positive and negative correlations between some characteristics. The results of this analysis are shown in Tables 6, 7 and 8. The weight of the nut showed a significant relationship with length, and size of the nut, which is consistent with the results of other researchers (Rasouli et al. 2013; Khadivi-Khub and Etemadi-Khah 2015; Gouta et al. 2019). The length of the nut had a significant correlation with the thickness of the nut (r = 0.6), kernel length (r = 0.65), and kernel width (r = 0.36). Also, the thickness of the nut showed a positive correlation with the mentioned properties of the kernel, which was consistent with the results of Hajivand (2020).
The traits of the kernel showed a positive and significant correlation with each other. The length of the kernel showed a positive and significant correlation with the width (r = 0.57), thickness (r = 0.27), as well as weight of the kernel (r = 0.59), which was consistent with Khadivi et al. (2019b) experiment (Tables 6, 7 and 8). As expected, a significant positive correlation between the nut weight and shell thickness (r = 0.48) and the nut size (r = 0.41) (Tables 6, 7 and 8). Also, a positive and significant correlation was observed between the nut size and kernel size (r = 0.48), which was consistent with the results of Mosavi et al. (2009). In general, the morphological traits of nut and kernel are significantly related to each other, while these traits showed no significant relationship with biochemical parameters. The studies showed no significant correlation between kernel weight and shell softness; these two factors act independently (Khadivi et al. 2019b).
The results of Tables 6, 7 and 8 show that the thicker the shell of the nut, the hardness of the shell increases. Dicenta and Garcia (1992) related the shell of the fruit to the content of lignin deposited in the endocarp during fruit development. They stated that a dominant gene controls softness of the shell with two alleles (D: very hard shell, d: paper-like shell). Also, a significant correlation was between the thickness of the shell and kernel pubescence, kernel color, and the percentage of double kernel. These results are consistent with the results of Rasouli et al. (2019) and Mosavi et al. (2009). The results showed significant correlation between the percentage of double kernel, nut size (r = 0.31), nut shape (r = 0.20), and kernel shape (r = 0.30). A significant positive correlation was between shriveling kernel and shell retention (r = 0.26) of the kernel. Also, kernel pubescence significantly correlated with kernel shriveling (r = 0.32) (Tables 6, 7 and 8).
On the other hand, there was a significant negative correlation between softness of shell with nut weight (r = −0.42), shell weight (r = −0.48), shell thickness (r = −0.32), and protein percentage (r = −0.28). In other words, the offspring with hardness shell, nut weight, shell weight, and thickness were less. Also, the percentage of double kernel had a significant negative correlation with the shell thickness (r = −0.35) (Tables 6, 7 and 8). Therefore, the offspring with a higher double kernel percentage had less shell thickness. These results are somewhat aligned with the findings of Mosavi et al. (2009), Rasouli et al. (2019), and Khadivi et al. (2019a).
The high correlation coefficients between morphological and important traits from the perspective of fruit growing (such as fruit, kernel, leaf size, and phenology) have been reported in other species of the genus Prunus such as plums, peaches, and cherries. Accordingly, it can be concluded that these traits have a similar effect in determining crop potential as well as germplasm differentiation. In addition, these results can be used in breeding programs or identification of ecotypes of almonds in field studies (Khadivi et al. 2019a).
Factor Analysis
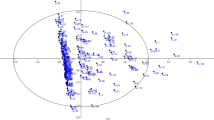
Tables 9 and 10 shows the results of factor analysis, among which the first, second, and third factors showed the highest contribution to the explanation of variance. The relative variance of each factor indicates the importance of that factor in the total variance of the studied traits and is expressed as a percentage. In factor analysis, a total of 10 main and independent factors, whose eigenvalues were more than 1, were able to explain 74.45% of the total variance (Tables 9 and 10). Some evaluated traits such as weight, length, width, thickness and size of nut and weight, length, width, kernel dry weight, and shell weight, were in the first-factor group, which explained 21.73% of the variance (Tables 9 and 10). The traits such as shell thickness kernel, and double kernel were in the second-factor group, which explained 10.75% of the variance (Tables 9 and 10). Some of the nut and kernel properties, which were divided into the first and second factor groups, played the most significant role in differentiating the progenies from each other. These two factors together explained 32.49% of the total variance (Tables 9 and 10). The traits, such as kernel humidity percentage, were in the third-factor group and explained 8.61% of the total variance (Tables 9 and 10). In the fourth factor group, the trait of nut shape and kernel shape were included, which explained 7.34% of the variance, and characteristics such as kernel pubescence, shriveling of kernel, and shell retention were in the fifth-factor group and explained 5.67% of the total variance (Tables 9 and 10). The sixth-factor group (PC6) included no soluble carbohydrates and making of outer shell, which explained 4.71% of the variance. In the seventh factor group (PC7), protein and oil percentages were included which explained 4.59% of the total variance. The eighth-factor group (PC8) included vitamin E, which explained 4.26% of the variance (Tables 9 and 10). In the ninth-factor group (PC9), soluble carbohydrate was included, which explained 3.64% of the total variance in the tenth-factor (PC10), the traits of kernel color were positioned with 3.12% variance (Tables 9 and 10).
Rasouli et al. (2019) reported that the nut and kernel traits played an important role in grouping of almond genotypes and cultivars and were effective traits in factor analysis. Khadivi et al. (2019b) used factor analysis to evaluate the morphological diversity of different cultivars of almonds. They reported the nut and kernel traits as important and effective features in this analysis, which is consistent with the study results. Khadivi-Khub and Etemadi-Khah (2015), in a study investigating the phenotypic diversity in the germplasm of selected cultivars of almonds, reported that factor analysis reduced the evaluated traits to three main factors, including the nut and kernel properties including length, width, and dry weight of the nut and kernel as well as the percentage of double kernel were reported as effective factors on the differences between the studied progenies. The results of the present study also show that the traits of the nut and kernel played the most role in factor analysis (Imani and Shamili 2018a).
Cluster Analysis
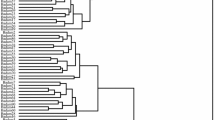
This study performed cluster analysis based on all measured traits (Table 2) by Ward’s method (Fig. 7). At 25 Euclidean distance, the progenies were divided into two main groups, which were important factors in differentiating cultivars from each other in this distance, such as the nut length, width, shape, protein content and protein content, and kernel taste. By reducing the distance from 25 to eight progenies were divided into four main groups. The important factors for differentiation in this distance were traits such as the nut length, width, and thickness, kernel weight, length, width, dry weight, shell thickness, size, taste, protein content, oil percentage, and vitamin E.
Group 1:
This included the highest number of progenies, i.e., 28 progenies out of 60 progenies studied are in this group, including: ‘MMar19’, MMar32, MarM12, MarM14, ‘MMar32’, ‘MMar40’, ‘MarM1’, ‘MMar36’, ‘MMar3’, ‘MMar26’, ‘MMar1’, ‘MMar4’, MMar22, MMar5, ‘MMar29’, ‘MMar42’, ‘MarM4’, ‘MarM7’, ‘MarM16’, ‘MMar6’, ‘MMar10’, ‘MMar12’, ‘MMar39’, ‘MMar43’, ‘MarM9’, ‘MMar16’, and ‘MarM3’. In general, these progenies were similar in terms of most of the measured traits, especially the traits related to nut and kernel, and had semi-hard shell, medium to small size of nut and kernel, light color of shell, medium percentage of protein, and oil, as well as and low vitamin E content.
Group 2:
Nine progenies out of 60 progenies studied are in this group, including: ‘MMar18’, ‘MMar20’, ‘MMar35’, ‘MMar34’, ‘MarM13’, ‘MMar14’, ‘MMar8’, ‘MMar9’, and ‘MMar30’. This group had longer nut and kernel length, medium nut and kernel size, less shriveling, non-shell retention, high percentage of humidity and oil. Also, MMar9, which had the most soluble carbohydrates, was included in this group.
Group 3:
This included progenies ‘MMar11’, ‘MMar38’, ‘MMar2’, ‘MMar25’, ‘MarM5’, ‘MarM15’, ‘MMar13’, and ‘Marcona’ cultivar. The progeny of this group had elongated nut shape, sweeter taste, medium kernel color and high ash and oil percentage. ‘MarM15’ progeny with the highest amount of vitamin E (220 mg · 100 g−1 FW) and ‘Marcona’ parent with the highest amount of protein (25.16%) were included in this group.
Group 4:
Offspring ‘MMar33’, ‘MMar41’, ‘MMar17’, ‘MarM17’, ‘MarM2’, ‘MMar31, ‘MMar37’, ‘MMar21’, ‘MarM8’, ‘MMar24’, ‘MMar7’, ‘MarM6’, ‘MMar28’, ‘MarM10’, ‘MMar15’, ‘MMar27’, and ‘Mamaei’ cultivar were included in this group. The progeny of this group had the largest nut and kernel size, round to elongated nut shape, light to dark kernel color, as well as the highest amount of protein and insoluble carbohydrates. ‘MMar24’ progeny with the highest kernel weight (2.51 g) and ‘MarM17’ with paper skin were included in this group. ‘MarM6’ and ‘MMar28’ progenies, which had more similar characteristics, were put together.
In cluster analysis, individuals in a cluster have many similarities in terms of the studied traits but fewer similarities with individuals in separate clusters in terms of that trait (Dejampour et al. 2018). In a study, phenotypic diversity of 100 Iranian seed genotypes of almonds was investigated (Rasouli et al. 2019). The study results showed that the properties of the nut and kernel, including the nut shape, shell softness, and percentage of double kernel were among the important traits in distinguishing cultivars and genotypes from each other (Rasouli et al. 2019). In another study, the morphological characteristics of 55 Iranian and foreign almond cultivars were studied in terms of 29 quantitative and qualitative traits of the nut and kernel. The cluster analysis results divided the studied traits, genotypes, and cultivars into six main groups. The important factors in separating the main clusters, such as the present study, were traits such as fruit length and shape, thickness, and softness of shell (Mosavi et al. 2009). Although the measured biochemical characteristics were also effective in grouping the progeny, and in the meantime, vitamin E content, oil percentage, and proteins were more effective traits. Pérez-Sánchez and Morales-Corts (2021) reported in the central-western Iberian Peninsula the almond cultivars ‘Gorda José’ and ‘Marcelina’ fruits were quite heavy (nuts: > 9.1 g; kernels: > 1.9 g), with very low percentages of double kernels (< 3%) and high nutritional value (> 50% lipids; > 21% proteins). These reports showed the importance of fruit and kernel traits and their biochemical compounds in introducing superior cultivars, which aligns with the present research findings. Furthermore, the PCA and cluster analysis results showed that agromorphological and chemical analysis could provide reliable information on the variability in almond genotypes (Pérez-Sánchez and Morales-Corts 2021), which was mainly following the findings of this research in cluster analysis.
Finally, in Table 11, according to investigated important traits, 16 superior offspring were selected along with their parents from among the 62 almond cultivars and progeny. The range of nut weight in the 16 selected superior progenies varied between 3.8 g (‘MarM12’) to 7.60 g (‘MMar27’) (Table 11). Moreover, the range of kernel weight ranged from 1.51 g (‘MarM5’) to 2.51 g (‘MMar24’), double kernel percentage from 0 to 60%, oil percentage from 47.93% (‘MarM10’) to 62.08% (‘MMar33’), and vitamin E content varied from 74 mg · 100 g−1 FW (‘MarM12’) to 210 mg · 100 g−1 FW (‘MMar27’). The results of mean comparison showed that in the width (30.32 mm) and nut thickness (24.41 mm), offspring of ‘MarM17’ and in the traits of nut length (39.54 mm) and kernel weight (2.51 g), ‘MMar7’ and ‘MMar24’ were higher than the parents and other offspring, respectively (Tables 3 and 11, Fig. 8). While the ‘Mamaei’ parent had longer kernel (26.79 mm) and ‘MarM5’ offspring had wider kernel (16.86 mm). Also, ‘MarM15’ offspring had more vitamin E (220 mg · 100 g−1 FW), ‘MMar33’ offspring had more oil (62.08%) and ‘MarM5’ offspring had more ash among selected superior progenies (Table 11, Fig. 8). Based on the results, the change of place between the maternal and paternal parents did not greatly affect the almond offspring’s important nut and kernel traits that resulted from controlled reciprocal crosses between ‘Mamaei’ and ‘Marcona’ cultivars. However, some offspring were superior to parents in nut and kernel traits that can be relative to hybrid vigor. This work constitutes an important step in using superior progeny in future almond breeding programs to obtain cultivars with superior nut and kernel quality.
Conclusion
In this research, 60 offspring resulting from controlled reciprocal crosses between ‘Mamaei’ and ‘Marcona’ as two commercial almond cultivars, along with their parents, were evaluated in terms of quantitative, qualitative, and biochemical traits of nut and kernel, so that the superior offspring identified be exploited in future almond breeding programs. The studied progenies showed high phenotypic diversity and, therefore, can be used as a potential source of germplasm in almond breeding. According to some important and commercial traits of almonds such as nut and kernel weight, nut and kernel size, percentage of shell, oil content, average carbohydrates content, vitamin E content, and percentage of double kernel, 16 offspring, including ‘MarM2’, ‘MarM4’, ‘MarM5’, ‘MarM6’, ‘MarM8’, ‘MarM10’, ‘MarM17’, ‘MMar2’, ‘MMar6’, ‘MMar7’, ‘MMar15’, ‘MMar24’, ‘MMar25’, ‘MMar27’, ‘MMar28’, and ‘MMar33’ with parents ‘Mamaei’ and ‘Marcona’ cultivars, were relatively superior to other offspring. Therefore, it is recommended to use them as parents in breeding programs, as well as after regional evaluations and observing the stability of traits in selected superior offspring, for the use of breeders and almond producers.
References
Aktas T, Thy P, Williams RB, Caffrey Z, Khatami R, Jenkins BM (2015) Characterization of almond processing residues from the Central Valley of California for thermal conversion. J Fuel Process Technol 140:132–147
Ayaz Z, Zainab B, Khan S, Abbasi AM, Elshikh MS, Munir A, Al-Ghamdi AA, Alajmi AH, Alsubaie QD, Mustafa AE (2020) In Silico authentication of amygdalin as a potent anticancer compound in the bitter kernels of family Rosaceae. Saudi J Biol Sci 27(9):1–15
Azimi MH (2020) Evaluation yield and genetically factors in different cultivars of gladiolus. Ornam Hortic 26:8–17
Azimi MH (2023) Intervarietal hybridization and observation of high-quality offspring of cut Freesia flowers. Int J Hortic Sci Technol 10(1):11–22
Barreca D, Nabavi SM, Sureda A, Rasekhian M, Raciti R, Silva AS, Annunziata G, Arnone A, Tenore GC, Süntar İ (2020) Almonds (Prunus dulcis Mill. DA webb): A source of nutrients and health-promoting compounds. Nutrients 12:672
Çelik F, Balta MF, Ercişli S, Gündoğdu M, Karakaya O, Yaviç A (2019) Tocopherol contents of almond genetic resources from Eastern and Western Turkey. Erwerbs-Obstbau 61(3):257–262
Comas JF, i Company RS, Segura JM (2019) Shell hardness in almond: Cracking load and kernel percentage. Sci Hortic 245:7–11
Cordeiro V, Monteiro A, Oliveira M, Ventura J (2001) Study of some physical characters and nutritive composition of the Portuguese’s (local) almond varieties. Options Méditerranéennes 56:333–337
Dejampour J, Zarrinbal M, Fathi H, Mousavizadeh S (2018) Fruit characteristics of some almond cultivars and genotypes of northwest of Iran. Seed Plant Improv J 33(2):195–213
Dicenta F, Garcia JE (1992) Phenotypical correlation among some traits in almond. J Genet Breed 46:241–246
Egea J, Burgos L (1995) Double kerneled fruits in almond (Prunus dulcis Mill.) as related to pre-blossom temperatures. Ann Appl Biol 126(1):163–168
El Hawary SS, Sokkar NM, El Halawany AM, Mokbel HA (2014) Study of nutritional contents of Prunus Amygdalus Batsch Seeds. Egypt J Hosp Med 57:437–443
Estaji A, Ebadi A, Fattahi Moghadam MR, Alifar M (2013) Evaluation of fifty almond genotypes characteristics obtained from crossing between some superior Iranian genotypes and ‘Tuono’ cultivar. J Plant Prod 20(2):253–270
Estaji A, Ebadi A, Fatahi Moghadam M, Aali Far M (2014) Evaluation of offspring obtained from crossing between superior local genotypes of almond and ‘Tuono’ cultivar from the viewpoint of self-compatible quantitative and qualitative characteristics. Fruit 6(1):43–57
FAO (2021) FAOSTAT database results. http:// Fao.stat.org/stat.org/stat/almond. Accessed 17 Feb 2021
Felipe AJ (2000) El Almendro. El Material Vegetal, vol I. Mira Editores, Zaragoz
Golzari M, Rahemi M, Hassani D, Vahdati K, Mohammadi N (2013) Protein content, fat and fatty acids of kernel in some persian walnut (Juglans regia L.) cultivars affected by kind of pollen. J Foot Sci Technol 38(10):21–31
Gouta H, Ksia E, Martinez-Gomez P (2019) Agronomical evaluation of local Tunisian almond cultivar and their breeding prospects. Eur J Hortic Sci 84(2):73–84
Gulcan R (1985) Descriptor list for almond (Prunus amygdalus)
Hajivand S (2020) Identification and initial evaluation of superior almond genotypes of traditional orchards from Qazvin based on some quantitative traits. Iran J Hortic Sci Technol 21(2):161–170
Hajivand S, Moghaddasi R, Zeraatkish Y, Mohammadinejad A (2020) An application of stochastic frontier gravity approach (the case of Iran’s potential agricultural exports). Int J Anal Appl 18:482–492
Hajnajari H, Kohneshine Leily H, Bakhshi D (2019) Selection of promising early ripening progenies and assessment of earliness heritability in the breeding program of apple. Agric Conspectus Sci 84(3):245–256
Hamilton RJ, Rossell JB (1986) Analysis of oils and fats. Elsevier, pp 18–336
Hansche PE, Beres V, Forde HT (1972) Estimates of quantitative genetic properties of walnut and their implication for cultivar improvement. Sci Hortic 97:279–285
Imani A, Shamili M (2018a) Almond nut weight assessment by stepwise and path analysis. Int J Fruit Sci 18(3):338–343
Imani A, Shamili M (2018b) Phenology and pomology of almond’s cultivars and genotypes using multivariate analysis. Adv Hortic Sci 32(1):27–32
IPGRI (1985) Descriptors list for almond (Prunus amygdalus). International Board for Plant Genetic Resources, Rome (Revised)
Khadivi A, Mohammadi M, Asgari K (2019a) Morphological and pomological characterizations of sweet cherry (Prunus avium L.), sour cherry (Prunus cerasus L.) and duke cherry (Prunus × gondouinii Rehd.) to choose the promising selections. Sci Hortic 257:1–10
Khadivi A, Safdari L, Hajian MH, Safari F (2019b) Selection of the promising almond (Prunus amygdalus L.) genotypes among seedling origin trees. Sci Hortic 256:1–10
Khadivi-Khub A, Etemadi-Khah A (2015) Phenotypic diversity and relationships between morphological traits in selected almond (Prunus amygdalus) germplasm. Agrofor Syst 89:205–216
Khojand S, Zeinalabedini M, Azizinezhad R, Imani A, Ghaffari MR (2022) Diversity of nut and kernel weight, oil content, and the main fatty acids of some almond cultivars and genotypes. J Nuts 14(1):33–44
Kochert G (1987) Carbohydrate determination by the phenolsulfuric acid method. In: Handbook of physiological methods, physiological and biochemical methods, p 95
Kodad O (2008) Fruit quality in almond as related to the type of pollination in self-compatible genotypes. J Am Soc Hortic Sci 133(3):320–326
Kodad O (2017) Chemical composition of almond nuts. In: Socias i Company R, Gradziel TM (eds) Almonds. Botany, production and uses. CABI, Wallingford, pp 428–448
Kodad O, Socias i Company R (2008) Variability of oil content and of major fatty acid composition in almond (Prunus amygdalus Batsch) and its relationship with kernel quality. J Agric Food Chem 56:4096–4101
Kodad O, Estopañán G, Juan T, Socias i Company R (2013) Genetic diversity in an almond germplasm collection: Application of a chemometric approach. Acta Hortic 976:237–242
Kodad O, Socias i Company IR, Alonso JM (2018) Genotypic and environmental effects on tocopherol content in almond. Antioxidants (basel) 7:1–9
Martínez-Gómez P, Gradziel TM, Ortega E, Dicenta F (2002) Low temperature storage of almond pollen. HortScience 37:691–692
Mehran M, Filssof M (1974) Characteristics of Iranian almond nuts and oils. J Am Oil Chem Soc 51:433–434
Montero-Riquelme FJ (1993) Caracterización Morfológica del Almendro. Servicio de Publicaciones de la Universidad de Castilla-LaMancha, Murcia
Mosavi A, Fatahi R, Zamani Z, Imani A (2009) Evaluation of quantitative and qualitative characteristics of some almond cultivars and genotypes. Iran J Agric Sci 41(20):119–131
Okay Y (2002) The comparison of some pistachio cultivars regarding their fat, fatty acids and protein content. J Gartenbauwiss 67(3):107–113
Pérez-Sánchez R, Morales-Corts MR (2021) Agromorphological characterization and nutritional value of traditional almond cultivars grown in the Central-Western Iberian Peninsula. Agronomy 11(6):1–19
Rasouli M, Imani A (2016) Effect of supplementary pollination by different pollinizers on fruit set and nut physicochemical traits of Supernova a self-compatible almond. Fruits J 71(5):229–306
Rasouli M, Fattahi R, Zamani Z, Imani A, Ebadi A (2013) A study of the phenotypic diversity of some almond cultivars and genotypes, using morphological traits. Iran J Hortic Sci 43(4):357–370
Rasouli M, Jafari Taeme A, Rahmati Joneidabad M (2019) Evaluation of genetic variation of some almond genotypes using morphological markers. Res Pomol 4(1):106–120
Rigoldi MP, Rapposelli E, De Giorgio D, Resta P, Porceddu A (2015) Genetic diversity in two Italian almond collections. Electron J Biotechnol 18:40–45
Romojaro F, García JE, López-Andreu FJ (1977) Estudio sobre la composición química de variedades de almendra del sureste español. An Edafol Agrobiol 36:121–131
Saura-Calixto F, Bauzá M, Martínez F, Argamentería A (1981) Amino acids, sugars, and inorganic elements in the sweet almond. J Agric Food Chem 29:509–511
Schirra M (1997) Postharvest technology and utilization of almonds. Hortic Rev 20:267–292
Sepahvand R, Delfan B, Ghanbarzadeh S, Rashidipour M, Veiskarami GH, Ghasemian-Yadegari J (2014) Chemical composition, antioxidant activity and antibacterial effect of essential oil of the aerial parts of Salvia sclareoides. Asian Pac J Trop Med 7:S491–S496
Socias i Company R, Ansón JM, Espiau MT (2017) Taxonomy, botany and physiology. In: Socias i Company R, Gradziel TM (eds) Almonds. Botany, production and uses. CABI, Wallingford, pp 1–42
Sorkheh K, Shiran B, Rouhi V, Asadi E, Jahanbazi H, Moradi H, Gradziel TM, Martínez-Gómez P (2009) Phenotypic diversity within native Iranian almond (Prunus spp.) species and their breeding potential. Genet Resour Crop Evol 56(7):947–961
Todde G, Murgia L, Caria M, Pazzona A (2016) multivariate statistical analysis approach to characterize mechanization, structural and energy profile in Italian dairy farms. Energy Rep 2:129–134
Turkmen S, Zamorano MJ, Xu H, Fernández-Palacios H, Robaina L, Kaushik S, Izquierdo M (2020) Parental LC-PUFA biosynthesis capacity and nutritional intervention with alpha-linolenic acid affect performance of Sparus aurata progeny. J Exp Biol 223:jeb214999
UPOV (2011) Guidelines for the Conduct of Test for Distinctness, Homogeneity and Stability. Almond (Prunus amygdalus Batsch). International Union for the Protection of New Varieties of Plants, Geneva
Valaei B, Bernousi I (2020) Phenotypic diversity for fruit yield and some related traits in tomato (Solanum lycopersicum L.) local populations of Kalaybar region. J Plant Prod 43(3):387–396
Vargas FJ, Romero M, Clavé J, Alegre S, Miarnau X (2009) Variedades de Almendro IRTA. Ibercaja, Madrid
Yada S, Lapsley K, Huang G (2011) A review of composition studies of cultivated almonds: Macronutrients and micronutrients. J Food Compos Anal 24(4):469–480
Zahedi SM, Abdelrahman M, Sadat Hosseini M, Yousefi R, Phan Tran L (2020) Physical and biochemical properties of 10 wild almond (Amygdalus scoparia) accessions naturally grown in Iran. J Food Biosci 37:129–134
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
M. Zahedi, M. Rasouli, A. Imani, O. Khademi and S.K. Jari declare that they have no competing interests.
Rights and permissions
Springer Nature oder sein Lizenzgeber (z.B. eine Gesellschaft oder ein*e andere*r Vertragspartner*in) hält die ausschließlichen Nutzungsrechte an diesem Artikel kraft eines Verlagsvertrags mit dem/den Autor*in(nen) oder anderen Rechteinhaber*in(nen); die Selbstarchivierung der akzeptierten Manuskriptversion dieses Artikels durch Autor*in(nen) unterliegt ausschließlich den Bedingungen dieses Verlagsvertrags und dem geltenden Recht.
About this article
Cite this article
Zahedi, M., Rasouli, M., Imani, A. et al. Evaluation of Quantitative, Qualitative, and Biochemical Traits of Almond Offspring from Controlled Reciprocal Crosses Between ‘Mamaei’ and ‘Marcona’ Cultivars. Erwerbs-Obstbau 65, 1525–1543 (2023). https://doi.org/10.1007/s10341-023-00900-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10341-023-00900-0