Abstract
In this study, 48 almond (Prunus dulcis Miller [D.A. Webb]) genotypes were analyzed in terms of detailed morphological parameters. On the other hand, the incompatibility S genotypes in these genotypes were determined using a polymerase chain reaction (PCR) approach with allele-specific primers. High morphological diversity among the genotypes was observed. Most of the total variation (71.94%) in the seven phenological and five morphological traits consisted of the first three main principal components (PCs). The all-important traits ensured a positive value, but leaf colour had a negative value for the first three PCs. On the other hand, foliation time and petiole length showed low variation among the studied genotypes. According to the results of PCR using the AS1II- and AmyC5R-specific primers in a single reaction, the amplification was successful. The results showed amplification of nine different self-incompatibility alleles (S1, S2, S3, S5, S6, S10, S11, S12 and S13) and of the self-compatibility allele Sf. The PCR-amplified fragments ranged from 600 to 1600 bp. The self-compatibility allele Sf and S3 allele had the same band size at 1200 bp. The number of self-compatible genotypes was 12, including ‘Marta’, ‘F. Barese’, ‘Tuono’ and ‘Super Nova’ cultivars. S1, S2, S5 and S6 were the most common alleles, as each was found in almond genotypes assayed here. The least common alleles were S10, S11, S12 and S13 alleles, and especially S10 was determined only in ‘Dokuzoguz’ cultivar. The PCR approach is an easy, low-cost tool for early identification of self-compatible progeny seedlings. From these results, it could be concluded that these local genotypes might be considered as potential candidates to be used in breeding programs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The almond (Prunus dulcis Miller [D.A. Webb]) belongs to the Rosaceae family, genus Prunus. Over 30 species have been described by botanists. The genus almond is one of the oldest tree nut crops and occupies a very peculiar place among fruit trees (Sorkheh et al. 2009). Because of almond’s tolerance to cold, drought and salinity, it is considered an important tree crop and is cultivated in different climatic regions of world. Almond was reported to originate in Central Asia, but almond species grown under a subtropical Mediterranean climate of mild, wet winters and warm, dry summers (Kester and Gradiziel 1996). Since the climate in Türkiye changes from subtropical to terrestrial, most plant species such as almond grow naturally and economically throughout the country (Ercişli 2004). Almond tree is one of the popular nut trees worldwide and produces fruits with high economic value. Türkiye is the main producer of this crop (FAO 2022).
There has been a genetically rich almond population since it has been mostly grown with seed for many years in Türkiye (Aslantas and Guleryuz 1999). In this regard, Türkiye, which has very valuable almond types, has an advantage that is not found in many other countries in terms of the breeding material. This large genetic diversity provides great benefits in the selection of superior genotypes especially in breeding research (Sumbul and Bayazit 2019). It is very significant in terms of almond cultivation that the variation of genetic material is revealed. The morphological and phenological properties are usually used for cultivar identification in almond (De Giorgio and Polignano 2001). Morphological characterization of the almond accessions is the credentials for plant breeders and is also essential to study the genetic diversity within the collection (Van Hintum et al. 2000; Sepahvand et al. 2015). Revealing the diversity of genetic sources, traditional multivariate statistical analyses such as principal component analysis (PCA), hierarchical cluster analysis, and multidimensional scaling analysis are used (Odong et al. 2011).
Although almond is generally a self-incompatible species, some self-compatible cultivars have been identified (Halasz et al. 2006). Almond is a gametophytic species in terms of self-incompatibility. Self-incompatibility is controlled by a single S locus with multiple codominant alleles. It is expressed within the styles of flowers as S‑RNase glycoproteins, which are responsible for the inactivation of the self-pollen tube growth (Wirthensohn et al. 2011). It was reported that these glycoproteins displayed a similar effect in most species of genus Prunus, including almond, apricot, sweet cherry and plum (Yamane et al. 1999; Wunsch and Hormoze 2004; Alonso and Socias I Company 2006; Hajilou et al. 2006). In total, 23 S-alleles of incompatibility (from S1 to S23, and also Sf) have been identified in the cultivated almond species (Ortega et al. 2005). For determining the self-compatible cultivars, two methods have mainly been used to: controlled pollination in the field and S‑RNase analysis in the laboratory (Boskovic et al. 2003). However, new technologies based on DNA analysis have lately been developed (Mousavi et al. 2010, 2014).
Identification of a self-incompatible genotype by PCR consists of the amplification of target DNA by specific primers designed for DNA sequences coding for the S alleles. To determine the different S‑alleles, researchers have designed specific primers. In these studies, it was determined the S1 and S3 alleles by Ma and Oliveira (2001); S7, S8, and Sf alleles by Channuntapipat et al. (2001); S2, S5, S9, S23, and Sf alleles by Channuntapipat et al. (2003). Sanchez-Perez et al. (2004) reported that 10 different S‑incompatibility alleles and Sf-compatibility alleles could be determined by the PCR method, but they could not be differentiated because the S3 allele and Sf allele had the same band size. Sheikh-Alian et al. (2010) used AS1II and AmyC5R primers to determine the self-compatibility of almond cultivars and genotypes. In the study, S1, S2, S3, S5, S7, S9, S10, S11, S12 and S13 alleles were identified, and foreign cultivars showed the same band size S‑alleles. The Iranian genotypes showed alleles as follows: S1 and S7 alleles in ‘Yalda’; S3 or Sf allles in ‘Shahroodi’, ‘Azar’ and ‘Shokoofeh’; S7 and S12 alleles in ‘Talkh-13’; S1 and S13 alleles in ‘Genotype-5’; S13 allele in ‘Genotype-11’. Rasouli (2017) examined some morphological traits and self-incompatibility of 71 almond genotypes. In the study, simple and multiplex specific PCR analyses were performed to identify self- incompatibility alleles in genotypes. As a result of the PCR method performed with the combination of AS1II and AmyC5R primers, the amplifications of 10 self-incompatibility alleles (S1, S2, S3, S5, S6, S7, S8, S10, S12 and S unknown allele) and Sf alleles were determined.
The identification of self-compatible almond cultivars is important for orchard planning and for designing crosses and selection parents in breeding programs. The aim of this study was to evaluate the morphological traits and determine self-incompatibility alleles in 48 foreign and Turkish almond cultivars and genotypes.
Materials and Methods
Plant Material
The 48 almond genotypes (nine foreign cultivars and 39 Turkish cultivars and selected genotypes) (Table 1) were located the ex-situ germplasm collection on the Mediterranean coast of Mersin, Türkiye. Turkish genotypes were selected based on some important morphological and phenological traits from different regions of Türkiye.
Morphological Traits
The morphological trait values can be found in previous studies (Pinar et al. 2016). The seven phenological and five morphological characteristics were used to perform statistical analysis. The times of bud swell, bud burst, pink bud, start of blooming, full blooming, end of blooming and foliation of all almond genotypes was used as phenological characterization data. The almond cultivars and genotypes had considerable differences in blooming time. The minimum and maximum values in genotypes were as follows: 5 January–14 February for bud swell time, 10 January–18 February for bud burst time, 16 January–26 February for pink bud time, 20 January–1 March for start of blooming time, 25 January–13 March for full blooming time, 29 January–20 March for end of blooming time and 2 February–28 March for foliation time. Genotypes were then divided into three groups based on flowering date early, middle, and late within this period, indicating differences in the chilling requirements of the studied genotypes. Most of the genotypes were middle flowering according to full blooming time. ‘Genotypes 6/12’ (TA-22) and ‘2902’ (TA-24) were the earliest to bloom, flowering on 20 January. The latest genotypes to bloom were ‘42/2’ (TA-8) and ‘47/2’ (TA-26) that was very-late flowering (1 March). In addition, leaf length (mm), leaf width (mm), leaf colour, leaf edge serration and petiole length (mm) were used for morphological data analysis. The leaf length, leaf width and petiole length of almond genotypes ranged from 14.86 mm to 84.91 mm, from 2.11 mm to 30.79 mm, and from 1.80 mm to 35.90 mm, respectively. The genotypes were categorized as light green, green and dark green in leaf colour, and serrated and non-serrated in leaf edge serration.
Data Analysis
Morphological data analysis was performed by NTSYS-pc software (Version 2.2. Exeter Software. Setauket, NY, USA) (Rohlf 2004). Morphological data matrices were standardized using the STAN option. Then SIMINT option was used to compute population distance matrix. The distance matrix was used to set up a dendrogram based on the unweighted pair group method with arithmetic mean (UPGMA) method using the SHAN option. To determine the efficiency of clustering, the cophenetic correlation coefficient (CPCC) was calculated with the Mantel method (Mantel 1967).
PCA was used to measured morphometric distances between trees of all the populations. The PAST program was used to follow the distribution of genotypes and morphological traits according to the first two PCs in the scatter plot.
Identification of S-alleles
DNA Extraction
The leaf tissue was ground in a muller under liquid nitrogen, then DNA was extracted using the CTAB method according to the protocol of Doyle and Doyle (1990). DNA concentration was determined by spectrophotometric measurement. The DNA solution was brought to a concentration of 10 ng/μL and stored at −20 °C.
PCR Amplification
In this study, specific PCR primers [AS1II (forward; 5′-TATTTTCAATTTGTGCAACAATGG-3′) and AmyC5 (reverse;5′-CAAAATACCACTTCATTGTAACAAC-3′)] were used to determine of S‑genotypes (Tamura et al. 2000; Sanchez-Perez et al. 2004). PCR was carried out 15 μL of reaction volume containing 2 μl DNA (30 ng), 1.5 μl 10xPCR Buffer, 0.2 μl Taq DNA polymerase (5 u/μL), 1 μl dNTP (2.5 mM), 1.5 μl MgCl2 (25 mM), 2 μl 10 mM each primer and 6.8 μl H2O. Reactions were performed in accordance with the procedure: initial denaturation at 95 °C for 3 min, 30 cycles with denaturation at 95 °C for 30 s, annealing at 54 °C for 45 s (depending on the primer), extension at 72 °C for 1 min 15 s, and a final extension step at 72 °C for 10 min (GeneAmp_PCR System 9700, Perkin–Elmer Corp, California). PCR products were run in 1.5% agarose gel at 110 volts for 2 h. Resultant bands were visualized under UV light after staining with ethidium bromide. The band widths determined by using 100 bp DNA ladder (Vilanova et al. 2005).
Furthermore, the deduced amino acid sequences of S1–S29 almond alleles were compared with sequences of other Prunus S‑RNases, displaying 97% similarity or higher, as deduced using the FASTA program in the European Bioinformatics Institute toolkit (http://www.ebi.ac.uk).
Results and Discussion
Principal Components (PCs) and Cluster Analyses
PCA was applied based on morphological data of almond genotypes with the PAST software. The major components were described by variables and their relationship was mirrored on principal components scores (Table 2). Most of the total variation (71.94%) in the seven phenological and five morphological traits occurred from the first three PCs with Eigenvalues > 1. The PC‑I, PC-II and PC-III represented 45.21%, 16.63% and 10.13% of total variability, respectively. In each PC, a coefficient greater than 0.3 was determined as the border to define the cut-off limit for the coefficients of the correct vectors (Hanci 2019). PC‑I showed six variables with higher scores (over 0.369 absolute value) related to all phenological characters, except foliation. The highest value of PC-II corresponded to leaf length and width. The separation in PC-III was due to changes in leaf colour and leaf edge serration. The all-important characteristics provided a positive value, but leaf colour had a negative value for the first three PCs in the PCA. On the other hand, foliation time and petiole length showed low variation among the studied genotypes.
Our finding, for the first three PCs, was similar with the values obtained (68%) by Nikoumanesh et al. (2011) for 62 almond genotypes from Iran, but was higher than those found by Zeinalabedini et al. (2012) for 71 almond genotypes from Iran, by Colic et al. (2012) for 19 almond genotypes from Serbia, and by Khadivi-Khub and Etemadi-Khah (2015) for 90 almond genotypes from Iran. The highest first three PCs in our study show that the properties observed are excellent for the classification of almond genotypes. Sakar et al. (2019) reported the first two principal components explained 95% of total variance in five phenological traits of almond from 12 local genotypes. Similarly, in a study conducted in Iran, it was reported that 13 phenotypic traits successfully explained genetic diversity in 33 almond cultivars/genotypes (Imani et al. 2021). Iranian almond germplasm, a large proportion of the variation was explained by the traits related to leaf and tree (Nikoumanesh et al. 2011). In the study by Sepahvand et al. (2015), the variables contributing to nut and kernel size were more important than leaf traits in different almond collections.
The distribution of the 48 almond accessions on the two-dimensional graph is shown in Fig. 1, according to the first two major PCs. Almond genotypes showed generally distribution on the graph, but not grouped. According to this analysis, accessions TA-22 and TA-24, accessions TA-25 and TA-32, accession TA‑8 and accession TA-40 are located separately from others as in the cluster analysis. The distribution of the measured morphological traits is shown in Fig. 2. According to this graph, the leaf edge serration of the genotypes is located in a different region compared to the other traits. On the other hand, the foliation character had the lowest effect on the distinction of accessions.
The cluster analysis grouped the 48 almond accessions into two main clusters, and they were further divided into two sub-clusters (Fig. 3). Five genotypes were in the first cluster (I), and the 42/2 genotype (TA‑8 accession) was grouped separately from the others in the first sub-cluster (I-A). In all, 43 of the 48 genotypes were included in the second cluster (II). There were 10 genotypes in the 1st sub-cluster (II-A), and 33 genotypes in the 2nd sub-cluster (II-B). On the other hand, genotypes in the 2nd sub-cluster (II-B) were grouped into two sub-groups with a distance value more than 15.0. The UPGMA dendrogram and the scatter plot approved the high variation between almond genotypes. The relative harmony between the results of PCs and cluster analyzes showed that morphological traits can ensure credible information on the variability in genotypes. Similar results were observed in almond genotypes in Lebanon (Talhouk et al. 2000), Serbia (Colic et al. 2012) and Iran (Sorkheh et al. 2009; Zeinalabedini et al. 2012; Khadivi-Khub and Etemadi-Khah 2015).
Identification of Self-Incompatibility Alleles
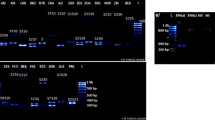
In this study, the primers developed by Tamura et al. (2000) were used for identification of S‑alleles in some almond genotypes. According to the results of PCR using the AS1II- and AmyC5R-specific primers in a single reaction, the amplification was successful. In the study by Sanchez-Perez et al. (2004) was used for identification S‑alleles the two primers (AS1II/AmyC5R). In the same study, it was proved that the lengths of amplified fragments in both single PCR and multiplex PCR were similar. Martínez-Gómez et al. (2003) reported that the eight S‑alleles in almond cultivars identified in single PCR reactions using these PCR primers. The primers (AS1II and AmyC5R) used in this study were quite efficient in the identification of several S‑alleles. The results showed amplification of nine different self-incompatibility alleles (S1, S2, S3, S5, S6, S10, S11, S12 and S13) and of the self-compatibility allele Sf. The PCR-amplified fragments were size from 600 to 1600 bp. The amplified fragments of Sf self-compatibility allele and S3 self-incompatibility allele had the same band size in 1200 bp. The findings of Sanchez-Perez et al. (2004) was similar with findings in this study. DNA fragment sizes corresponding to each S‑allele agreed in most status with the available data. Thus, S‑genotypes of the new tested almond selections were determined (Table 3).
The number of self-compatible genotypes was 12, including ‘Marta’, ‘F. Barese’, ‘Tuono’ and ‘Super Nova’ cultivars. For 16 genotypes, two alleles were confirmed; and for other genotypes, only one allele was determined. S1, S2, S5 and S6 were the most common alleles, as each was found in almond genotypes assayed here. The least common alleles were S10, S11, S12 and S13 alleles, and especially S10 was determined only in ‘Dokuzoguz’ cultivar. S‑allele frequencies were probably related to geographical origin. The S‑allele frequencies results were a little different than those reported by Lopez et al. (2006), who found that the alleles S1, S5, S7, and S8 in 133 almond cultivars were observed more frequently than others. Valizadeh and Ershadi (2009) reported that the alleles S2, S7 and S13 were the most common alleles in 16 Iranian almond cultivars. On the other hand, the S‑genotypes of Iranian and foreign almond genotypes have been reviewed by Rasouli (2017). The study found that the alleles S1 and S2 were observed more frequently than others.
Conclusions
In this study, a wide phenotypic and morphologic diversity in all evaluated traits was detected. The presence of such diversity and the maintenance of genetic diversity are important for future breeding of almond. But it may be beneficial to use molecular data to determine genetic diversity and relatedness. On the other hand, the determination of eight local genotypes with Sf allele in the study is promising to consider the self-inefficiency in almond species. According to the results, the selection of the appropriate variety and pollinator in the new orchards will contribute to production and cultivation. However, in parallel with the research and goals in the world, especially in almond type, the process of obtaining a national variety of almond type that can grow naturally in Anatolia for thousands of years will be initiated by contributing to the selection of the right parents to determine a late and self-efficient variety.
References
Alonso JM, Socias-Company R (2006) Almond S‑genotype identification by PCR using specific and non-specific allele primers. Acta Hortic 726:67–70
Aslantas R, Guleryuz M (1999) Almond selection in microclimate areas of northeast Anatolia. XI Grempa Seminar on Pistacios and Almonds, Sanliurfa, 01–04 September 1999, pp 339–342
Boskovic R, Tobutt KR, Batlle I, Duval H, Martinez-Gomez P, Gradziel TM (2003) Stylar ribonucleases in almond: Correlation with, and prediction of incompatibility genotypes. Plant Breed 122:70–76
Channuntapipat C, Sedgley M, Collins G (2001) Sequences of the cDNAs and genomic DNAs encoding the S1, S7, S8, and Sf alleles from almond, Prunus dulcis. Theor Appl Genet 103:1115–1122
Channuntapipat C, Wirthensohn M, Ramesh SA, Battle I, Arus P, Sedgley M, Collins G (2003) Identification of incompatibility genotypes in almond (Prunus dulcis Mill.) using specific primers basen on the introns of the S‑alleles. Plant Breed 122:164–168
Colic S, Rakonjac V, Zec G, Nikolic D, Aksic MF (2012) Morphological and biochemical evaluation of selected almond [Prunus dulcis (Mill.) D.A.Webb] genotypes in northern Serbia. Turk J Agric For 36:429–438
De Giorgio D, Polignano GB (2001) Evaluating the biodiversity of almond cultivars from germplasm collection field in Southern Italy. Sustain Glob Farm 56:305–311
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Ercisli S (2004) A short review of the fruit germplasm resources of Turkey. Genet Resour Crop Evol 51:419–435
FAO (2021) Food and Agriculture Organization of the United Nations. “Crops data”. https://www.fao.org/faostat/en/#data/QCL
Hajilou J, Grigorian V, Mohammadi SA, Nazemieh A, Romero C, Vilanova S, Burgos L (2006) Self- and Cross- (in) compatibility between important apricot cultivars in northwest Iran. J Hortic Sci Biotechnol 81:513–517
Halasz J, Hegedus A, Pedryc A (2006) Review of the molecular background of selfincompatibility in rosaceous fruit trees. Int J Hortic Sci 12(2):7–18
Hanci F (2019) Genetic variability in peas (Pisum sativum L.) from Turkey asssessed with molecular and morphological markers. Folia Hort 31(1):101–116
Imani A, Amani G, Shamili M, Mousavi A, Rezai H, Rasouli M, Martínez-García PJ (2021) Diversity and broad sense heritability of phenotypic characteristic in almond cultivars and genotypes. Int J Hortic Sci Technol 8(3):281–289
Kester DE, Gradiziel TM (1996) Almonds. In: Janick J, Moore JN (eds) Nuts. Fruit Breeding, vol III. John Wiley & Sons, New York, pp 1–97
Khadivi-Khub A, Etemadi-Khah A (2015) Phenotypic diversity and relationships between morphological traits in selected almond (Prunus amygdalus) germplasm. Agroforest Syst 89:205–216
Lopez M, Vargas FJ, Batlle I (2006) Self-(in)compatibility almond genotypes: a review. Euphytica 150:1–16
Ma R, Oliveira MM (2001) Molecular cloning of the self-incompatibility genes S1 and S3 from almond (Prunus dulcis cv. Ferragnès). Sex Plant Reprod 14:163–167
Mantel N (1967) The detection of disease clustering and generalized regression approach. Cancer Res 27:209–220
Martínez-Gómez P, López M, Alonso JM, Ortega E, Batlle I, Socías-Company R, Dicenta F, Dandekar AM, Gradziel TM (2003) Identification of self-incompatibility alleles in almond and related Prunus species using PCR. Acta Hortic 622:397–401
Mousavi A, Fatahi Moghadam MR, Zamani Z, Imani A (2010) Evaluation of quantitative and qualitative characteristics of some almond cultivars and genotypes. Iran J Hortic Sci 41(2):119–131
Mousavi A, Fatahi R, Zamani Z, Imani A, Dicenta F, Ortega E (2014) Genetic variation and frequency of S‑alleles in Iranian almond cultivars. Acta Hortic 1028:45–48
Nikoumanesh K, Ebadi A, Zeinalabedini M, Gogorcena Y (2011) Morphological and molecular variability in some Iranian almond genotypes and related Prunus species and their potentials for rootstock breeding. Sci Hortic 129:108–118
Odong TL, Van Heerwaarden J, Jansen J, Van Hintum TJL, Van Eeuwijk FA (2011) Determination of genetic structure of germplasm collections: are traditional hierarchical clustering methods appropriate for molecular marker data? Theor Appl Genet 123:195–205
Ortega E, Sutherland BG, Dicenta F, Boskovic R, Tobutt KR (2005) Determination of incompatibility genotypes in almond using first and second intron consensus primers: detection of new S alleles and correction of reported S genotypes. Plant Breed 124:188–196
Pinar H, Bircan M, Uzun A, Coskun OF, Yaman M (2016) Evulation of phenological and pomological characters of some almond genotypes and cultivars in Turkey. International Conference on Sustainable Development, Skopje, 19–23 September 2016, pp 385–388
Rasouli M (2017) The study of morphological traits and identification of self-incompatibility alleles in almond cultivars and genotypes. J Nuts 8(2):137–150
Rohlf FJ (2004) NTSYS-pc numerical taxonomy and multivariate analysis system. Version 2.11V. Exeter software, Setauket, New York
Sakar H, El-Yamani M, Boussakouran A, Rharrabti Y (2019) Reproductive phenology of some local almond [Prunus dulcis (Mill.) D.A. Webb] genotypes from northern Morocco. J Anal Sci Appl Biotechnol 1(2):49–54
Sanchez-Perez R, Dicenta F, Martinez-Gomez P (2004) Identification of S‑alleles in almond using multiplex PCR. Euphytica 138:263–269
Sepahvand E, Khadivi-Khub A, Momenpour A, Fallahi E (2015) Evaluation of an almond collection using morphological variables to choose superior trees. Fruits 70(1):53–59
Sheikh-Alian A, Vezvaei A, Ebadi A, Fatahi-Moghadam MR, Sarkhosh A (2010) Determination and identification of selfincompatibility alleles in selective Iranian and foreign almond (Prunus dulcis M.) cultivars by PCR method. Iran J Hortic Sci 41(3):247–252
Sorkheh K, Shiran B, Rouhi V, Asadi E, Jahanbazi H, Moradi H, Gradziel TM, Martinez-Gómez P (2009) Phenotypic diversity within native Iranian almond (Prunus spp.) species and their breeding potential. Genet Resour Crop Evol 56:947–961
Sumbul A, Bayazit S (2019) Pomological and chemical attributes of almond genotypes from Hatay province (in Turkish). Int J Agric Wildl Sci 5(1):1–10
Talhouk SN, Lubani RT, Baalbaki R, Zurayk R, Al-Khatib A, Parmaksizian L, Jaradat AA (2000) Phenotypic diversity and morphological characterization of Amygdalus species in Lebanon. Genet Resour Crop Evol 47:93–104
Tamura M, Ushijima K, Sassa H, Hirano H, Tao R, Gradziel TM, Dandekar AM (2000) Identification of self-incompatibility genotypes of almond by allele-specific PCR analysis. Theor Appl Genet 101:344–349
Valizadeh B, Ershadi A (2009) Identification of self-incompatibility alleles in Iranian almond cultivars by PCR using consensus and allele-specific primers. J Hortic Sci Biotechnol 84(3):285–290
Van Hintum TJL, Brown AHD, Spillane C, Hodgkin T (2000) Core collections of plant genetic resources. IPGRI Technical Bulletin No. 3,. nternational Plant Genetic Resources Institute, Rome
Vilanova S, Romero C, Llacer G, Badenes ML (2005) Identification of self incompatibility aleles in apricot by PCR and sequence analysis. J Am Soc Hortic Sci 130(6):893–898
Wirthensohn M, Rahemi M, Fernández-Martí A (2011) Identification of self-incompatibility genotypes and DNA fingerpriting of some Australian almond cultivars. Acta Hortic 912:561–566
Wunsch A, Hormoze JI (2004) S‑allele identification by PCR analysis in sweet cherry cultivars. Plant Breed 123:1–6
Yamane H, Tao R, Sugiora A (1999) Identification and cDNA cloning for S‑RNases in selfincompatible Japanese plum (Prunus salisina cv, sordum). Plant Biotechnol 16(5):389–396
Zeinalabedini M, Khayam-Nekoui M, Imani A, Majidian P (2012) Identification of selfcompatibility and self-incompatibility genotypes in almond and some Prunus species using molecular markers. Seed Plant Improv J 28(2):227–238
Funding
This work has been supported by Erciyes University Scientific Research Projects Coordination Unit under Grant number of FBA-2019-8708.
Author information
Authors and Affiliations
Contributions
Hasan Pınar contributed 35%, Ercan Yıldız 30%, Aydın Uzun 25%, Mustafa Bircan 10%.
Corresponding author
Ethics declarations
Conflict of interest
H. Pınar, E. Yıldız, M. Bircan and A. Uzun declare that they have no competing interests.
Additional information
Availability of data and material
Available
Rights and permissions
Springer Nature oder sein Lizenzgeber (z.B. eine Gesellschaft oder ein*e andere*r Vertragspartner*in) hält die ausschließlichen Nutzungsrechte an diesem Artikel kraft eines Verlagsvertrags mit dem/den Autor*in(nen) oder anderen Rechteinhaber*in(nen); die Selbstarchivierung der akzeptierten Manuskriptversion dieses Artikels durch Autor*in(nen) unterliegt ausschließlich den Bedingungen dieses Verlagsvertrags und dem geltenden Recht.
About this article
Cite this article
Pınar, H., Yıldız, E., Bircan, M. et al. Identification of Genetic Diversity Using Morphological Properties and Self-Incompatibility Alleles in Selected Prunus dulcis Miller (D.A. Webb) Genotypes. Erwerbs-Obstbau 65, 1595–1602 (2023). https://doi.org/10.1007/s10341-023-00874-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10341-023-00874-z