Abstract
In natural produced bacteria, β-carotene hydroxylase (CrtZ) and β-carotene ketolase (CrtW) convert β-carotene into astaxanthin. To increase astaxanthin production in heterologous strain, simple and effective strategies based on the co-expression of CrtZ and CrtW were applied in E. coli. First, nine artificial operons containing crtZ and crtW genes from different sources were constructed and, respectively, introduced into E. coli ZF237T, a β-carotene producing host. Among the nine resulting strains, five accumulated detectable amounts of astaxanthin ranging from 0.49 to 8.07 mg/L. Subsequently, the protein fusion CrtZ to CrtW using optimized peptide linkers further increased the astaxanthin production. Strains expressing fusion proteins with CrtZ rather than CrtW attached to the N-terminus accumulated much more astaxanthin. The astaxanthin production of the best strain ZF237T/CrtZAs-(GS)1-WBs was 127.6% and 40.2% higher than that of strains ZF237T/crtZAsWBs and ZF237T/crtZBsWPs, respectively. The strategies depicted here also will be useful for the heterologous production of other natural products.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
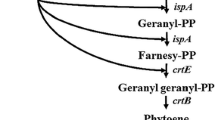
Astaxanthin (3,3′-dihydroxy-β,β-carotene-4,4′-dione) is an important natural red–orange pigment with one of the highest antioxidative activities. It is widely used in pharmaceuticals, cosmetics, health products, food and beverage due to its various health benefits [15]. Usually, the biosynthetic pathway of astaxanthin is artificially divided into two parts, the biosynthesis of β-carotene and astaxanthin formation (Fig. 1). In bacteria, the reactions from β-carotene to astaxanthin are normally catalyzed by two oxygenases, but the pathways vary. The β-carotene hydroxylase (CrtZ) is involved in hydroxyl introducing at positions C-3 and C-3′, while the ketolase (CrtW) is responsible for keto group formation at positions C-4 and C-4′ [26]. The Misawa group isolated and determined the functions of the gene cluster involved in astaxanthin production in Agrobacterium aurantiacum and identified the astaxanthin biosynthesis pathway at the level of individual genes for the first time [28]. Later they found that the astaxanthin biosynthesis enzymes from bacteria and Haematococcus pluvialis are bifunctional and have different substrates preference [10, 11]. In addition, Scaife et al. investigated 12 genes encoding β-carotene ketolases and 4 encoding β-carotene hydroxylases from 5 cyanobacterial species in vivo and reached the same conclusion [35]. In addition to the substrate preference, both CrtZ and CrtW can simultaneously catalyze the oxygenation of the respective positions on one or both of the β-ionone rings, which makes it difficult to identity the main reaction in natural astaxanthin producers that possess the two oxygenases. Consequently, conversion of β-carotene into astaxanthin is accomplished via multiple intermediate products (Fig. 1).
The biosynthetic pathway of astaxanthin in engineered E. coli. The biosynthetic pathway was artificially divided into two parts, the biosynthesis of β-carotene and astaxanthin formation. Heterologous pathways are shown in red. GAP glyceraldehyde-3P, IPP isopentenyl pyrophosphate, DMAPP dimethylallyl pyrophosphate, FPP farnesyl pyrophosphate, MEP pathway 2-C-methyl-d-erythritol-4-phosphate pathway, MVA pathway mevalonate pathway, ispA FPP synthase, crtE GGPP synthase, crtB phytoene synthase, crtI phytoene desaturase, crtY lycopene cyclase, H β-carotene hydroxylase, K β-carotene ketolase (color figure online)
In recent years, some representative microbiological systems including heterologous strains of E. coli [23, 25, 32, 38, 40], Saccharomyces cerevisiae [16, 41, 42] and Corynebacterium glutamicum [14] have been engineered to accumulate astaxanthin by various metabolic engineering strategies. The strategies which were used individually or jointly include enzyme quarrying, promoter control, optimization of ribosome binding sites and codon usage, protein engineering, discovery of new targets and gene copy number regulation. Among these studies, some engineered E. coli strains with potential for industrial applications were constructed. For example, E. coli strain E [40] and WLGB-RPP (pAX15) [32] produced astaxanthin titers of 320 mg/L and 332.23 mg/L in fed-batch fermentation, respectively. In these engineered strains, the genes involved in astaxanthin biosynthesis were introduced by a controlled arrangement of reactions or dual expression vectors. In natural producers, the genes involved in astaxanthin biosynthesis are present on the genome in form of gene cluster [30]. To our knowledge, there are only few reports about regulating astaxanthin accumulation through the combinatorial co-expression of CrtZ and CrtW within an expression cassette in engineered strains.
In this work, we used simple and effective combinatorial strategies to optimize the heterologous astaxanthin biosynthesis pathway in E. coli. The crtZ and crtW genes from different sources were combined and co-expressed. The co-expression of the crtZ gene from Brevundimonas sp. SD212 and crtW gene from Paracoccus sp. N81106 yielded the best astaxanthin accumulation. Based on the improved combinations, a series of fusion proteins were constructed and the astaxanthin production was significantly enhanced by expressing a CrtZ-W fusion protein. Finally, the astaxanthin production and content were further increased via carbon resource optimization.
Materials and methods
Strains, media and culture conditions
E. coli DH5α was used to construct recombinant plasmids and E. coli strain ZF237T was used as host for astaxanthin production [20]. LB medium [10 g tryptone, 5 g yeast extract, 10 g sodium chloride per liter with or without 1.5% (m/v) agar] containing 20 mg/L of ampicillin was used for plasmids construction. During plasmids construction, cultures were grown in a rotary shaker at 220 rpm and 37 °C or statically at 37 °C. 2 × YT medium (16 g tryptone, 10 g yeast extract, 5 g sodium chloride per liter) was used for fermentation in shake flasks. All strains are listed in Table S1.
Plasmid construction
All DNA manipulation procedures were implemented according to standard procedures [34]. Restriction enzyme digestion and overlap extension PCR [13] were performed to construct designed recombinant plasmids. All plasmids and primers used in this study are listed in Tables S1 and S2. The crtZ genes from Paracoccus sp. N81106 (A. aurantiacum), Alcaligenes sp. strain PC-1 and Brevundimonas sp. SD212, and the crtW genes from Paracoccus sp. N81106, Brevundimonas sp. SD212 and Nostoc sp. PCC7120 were codon optimized and chemically synthesized by Wuhan GeneCreate biological engineering Co. LTD (China) and cloned into plasmid pUC57. To construct the nine artificial expression operons, the crtZ gene was amplified from the plasmid pUC57-Z using the primers Z-5′-EcoRI/Z-3′-BamHI with the primer Z-5′-EcoRI containing the sequences of promoter J23119 and RBS apFAB917. Similarly, the crtW gene was amplified from the plasmid pUC57-W using the primers W-5′-BamHI/W-3′-HindIII with the primer W-5′-BamHI containing the sequences of the RBS apFAB916. The purified crtZ gene fragment was digested with EcoR I and BamH I and inserted between the corresponding sites of plasmid p5C. Subsequently, the crtW gene fragment was located at the BamH I/Hind III sites of plasmid p5C-Z after digestion with BamH I and Hind III. To construct CrtZ-W fusion proteins, the stop codon of the crtZ gene was removed and the fusion gene was inserted into plasmid p5C via enzyme digestion-ligation after splicing by overlap-extension PCR using the primers CrtZ-5′-EcoRI/CrtW-3′-HindIII. The CrtW-Z fusion proteins were constructed analogously. The linkers of different lengths, (Gly–Ser–Gly)x (x = 1, 2, 3, or 4), or (Gly–Ser)y (y = 1, 2, 3, 4, or 5), which were introduced between open reading frames, were incorporated into primers as overlap sequences. The codons of linkers for E. coli were chosen using the Codon Adaptation Tool (JCAT) (http://www.jcat.de/). All recombinant plasmids were verified through sequencing by GENEWIZ, Inc (Beijing, China).
Quantitative real-time PCR
Quantitative real-time PCR was applied to measure the transcriptional levels of genes in the engineered strains. Cells were collected after 48 h of shake-flask fermentation. Total RNA extraction and cDNA synthesis were the same as described previously [21]. Quantitative PCR was conducted using the ABI7500 Real-Time PCR kit (Applied Biosystems, USA) with SYBR® Premix Ex Taq™ II (Tli RNaseH Plus) and ROX plus (TaKaRa, Dalian, China). The primers used for qPCR are listed in Table S2. The data were quantified using the 2−ΔΔCT method as described before [22]. The rrsA gene was used as internal standard for normalization, and three biological replicates and sample repetitions were performed.
Homology modeling
The homology-based models of Alcaligenes sp. strain PC-1 CrtZ, Brevundimonas sp. SD212 CrtW, as well as the fusion proteins CrtWAs-ZBs and CrtZBs-WAs were modeled using I-TASSER (https://zhanglab.ccmb.med.umich.edu/I-TASSER/) [37]. The specific operations were described by Roy et al. [33]. To the best of our knowledge, there are no crystal structures of β-carotene hydroxylases and β-carotene ketolases in the PDB library, so we chose model 1 from each prediction output according to the C-score and TM-score, which indicate prediction reliability and alignment identity, respectively.
Shake-flask fermentation
The engineered E. coli strains were cultured in 5 mL LB medium with appropriate antibiotics at 30 °C overnight in a rotary shaker (220 rpm). The liquid cultures were then transferred to 500 mL flasks containing 100 mL of 2 × YT medium with 10 g/L glucose or 10 g/L other carbon source for fermentation optimization and appropriate antibiotics (20 mg/L for ampicillin and 10 m/L for kanamycin) to an initial OD600 of about 0.04 and cultivated under the same conditions for 96 h.
Analytical methods
Intracellular carotenoids were extracted as described previously [20]. The extract was analyzed by high-performance liquid chromatography (HITACHI Primaide, Japan) equipped with a HyPURITY C18 column (150 × 4.6 mm, 5 μm, Thermo Fisher Scientific, Inc., USA) to determine the concentration of astaxanthin and identify other carotenoids. The analyte signals were detected at 470 nm. Solvent A consisting of 90% aqueous acetonitrile (HPLC grade) and solvent B consisting of methyl alcohol-isopropyl alcohol (3:2, v/v, HPLC grade) were used as mobile phase at a flow rate of 1.0 mL/min. A 35-min gradient program was used to elute the analytes as follows: 100–10% solvent A (0–15 min), 10% solvent A (15–30 min), 10–100% solvent A (30–35 min) [42]. The authentic carotenoid standards and their retention times were as follows: astaxanthin (≥ 97%, Sigma-Aldrich, USA), 7.1 min; zeaxanthin (≥ 95%, Sigma-Aldrich, USA), 9.1 min; canthaxanthin (≥ 95%, Sigma-Aldrich, USA), 11.7 min; echinenone (≥ 95%, Sigma-Aldrich, USA), 17.6 min; lycopene (≥ 98%, Sigma-Aldrich, USA), 20.9 min; β-carotene (≥ 95%, Sigma-Aldrich, USA), 22.8 min. The OD600 was used as cell growth indicator. For E. coli, the OD600 and dry cell weight (DCW) had the following conversion relation: 1 OD600 = 0.323 g DCW/L. The concentration of glucose was determined using an SBA-40C biosensor analyzer (Institute of Microbiology, Shandong, China). The concentration of glycerol was measured by HPLC (HITACHI Primaide, Japan) equipped with an Aminex HPX-87H column (300 × 7.8 mm, Bio-Rad, Inc., USA). The mobile phase was 5 mM H2SO4 with flow rate of 0.5 mL/min, and the column temperature was 50 °C.
Results
Design of the β-carotene hydroxylase and -ketolase coexpression system
Previous studies revealed that CrtZ can efficiently convert β-carotene into zeaxanthin, as well as that the main product of zeaxanthin conversion by CrtW is astaxanthin in E. coli [7, 8]. Transcription and translation are generally coupled in bacteria, including E. coli, and the sequence of translation is related to the sequence of genes in polycistronic operons [24]. The E. coli strain ZF237T, which harbors an artificial crtEBIY operon and another 15 modified genes in its genome, is a β-carotene over-producing engineered strain [20]. To engineer it to produce astaxanthin, crtZ–crtW expression cassette was designed. The transcription of the cassette was controlled by the Anderson promoter J23119 instead of an IPTG-induced promoter, as IPTG was unfavorable for lycopene accumulation in E. coli [17]. The RBS apFAB917 was used for crtZ and apFAB916 for crtW. The combinations of the promoter and the RBSs have high mRNA and high protein levels [18]. The cassette was assembled through enzymatic digestion-ligation and inserted into the low copy number plasmid p5C (Fig. 2a) to avoid a possible severe metabolic burden on the host.
Astaxanthin production by E. coli ZF237T derivatives expressing various artificial operons and fusion proteins. a, c The designs of different operons and fusion proteins. The swallowtail arrow denotes the promoter J23119, the blue solid circle denotes the RBS apFAB917. b The production of astaxanthin by strains expressing various operons, and d CrtW-Z and CrtZ-W fusion proteins (color figure online)
Construction and expression of 9 cassettes in vivo
It has been discovered that there are many different β-carotene hydroxylases and -ketolases in nature. This facilitates isoenzyme selection to regulate astaxanthin production. Previous endeavors succeeded in increasing astaxanthin production by utilizing different hydroxylases and ketolases from various origins [23, 36, 40]. The β-carotene hydroxylases from Paracoccus sp. N81106 (CrtZPs), Alcaligenes sp. strain PC-1 (CrtZAs) and Brevundimonas sp. SD212 (CrtZBs), and the β-carotene ketolases CrtW from Paracoccus sp. N81106 (CrtWPs), Brevundimonas sp. SD212 (CrtWBs) and Nostoc sp. PCC7120 (CrtWNs) were chosen to construct expression cassettes due to their higher substrate conversion efficiencies in vivo [7, 8, 35]. The three crtZ genes were combined with the three crtW genes according to the design mentioned above so that a total of nine crtZW expression plasmids were constructed and transferred into strain ZF237T.
All the colonies harboring any of the nikne different crtZW expression plasmids showed red color after cultivation for 24 h on LB agar plates, and their carotenoid composition was analyzed. All strains accumulated lycopene, but only five accumulated detectable amounts of astaxanthin after 96 h of fermentation (Table 1). As shown in Fig. 2b, the astaxanthin production of strains ZF237T/crtZAsWPs and ZF237T/crtZPsWNs was less than 2 mg/L, while the strains ZF237T/crtZAsWBs, ZF237T/crtZBsWBs and ZF237T/crtZBsWPs produced 4.97, 7.38 and 8.07 mg/L of astaxanthin, respectively. The astaxanthin production of strain ZF237T/crtZBsWPs was 16.5 times that of strain ZF237T/crtZAsWPs. However, among the four non-astaxanthin-accumulating strains, the product distribution differed greatly (Table 1). For example, strain ZF237T/crtZPsWBs accumulated canthaxanthin without detectable amount of zeaxanthin, while strain ZF237T/crtZBsWNs accumulated zeaxanthin without detectable amounts of canthaxanthin.
Construction of fusion proteins
In Xanthophyllomyces dendrorhous, a single chromosomally encoded P450 protein, CrtS, directly converts β-carotene into astaxanthin using CrtR as electron donor [2]. However, the expression of P450 enzymes involved in terpenoids biosynthesis in E. coli is still very challenging [3, 5]. The genetic fusion of proteins was originally developed for protein expression/purification and was later adopted to enhance metabolic fluxes by facilitating substrate channeling [1, 43]. Two broad classes of fusion proteins are usually designed by changing the location of an enzyme. To produce astaxanthin from β-carotene more efficiently, three CrtZ-W and three CrtW-Z fusion proteins were designed and constructed based on the three combinations that showed higher astaxanthin production (Fig. 2b) when expressed independently, crtZAsWBs, crtZBsWBs and crtZBsWPs. The expression of the fusion proteins was controlled by the constitutive promoter J23119 and RBS apFAB917. The expression cassettes encoding these fusion proteins were also inserted into plasmid p5C after assembly (Fig. 2c).
The six fusion proteins comprised three pairs, CrtZAs-WBs and CrtWBs-ZAs, CrtZBs-WBs and CrtWBs-ZBs, CrtZBs-WPs and CrtWPs-ZBs, which were fused directly (Fig. 2c). The fusion protein expression plasmids were, respectively, introduced into strain ZF237T and the six resulting strains were cultivated in shake flasks. The astaxanthin concentrations were determined after 96 h of fermentation. Unexpectedly, the strains expressing CrtZ-W variants produced much more astaxanthin than the corresponding CrtW-Z expressing strains (Fig. 2d). The maximum difference of astaxanthin production was up to 45.9-fold within fusion protein pair (CrtZBs-WBs versus CrtWBs-ZBs). The results indicate that fusion proteins with CrtZ attached to the N-terminus are better for astaxanthin production. ZF237T/CrtZAs-WBs produced 7.89 mg/L astaxanthin, which was 58.5% higher than the corresponding non-fusion protein expressing strain ZF237T/crtZAsWBs. However, strains ZF237T/CrtZBs-WBs and ZF237T/CrtZBs-WPs produced 4.02 mg/L and 6.22 mg/L of astaxanthin, respectively, both of which were much lower than those of the corresponding non-fusion protein expressing strains (ZF237T/crtZBsWBs and ZF237T/crtZBsWPs) (Fig. 2b, d). In addition, the production of astaxanthin produced by strain ZF237T/CrtZAs-WBs was 2.2% lower than that of strain ZF237T/crtZBsWPs. We, therefore, failed to increase astaxanthin production by fusing crtZ and crtW directly, which might be due to the lack of a linker between the two folding units.
Application of peptide linkers to increase astaxanthin production
As an important component of fusion proteins, linkers can significantly affect the biological function of fusion proteins. Previous reports demonstrated that flexible linkers are well-suited for engineering the fusion proteins with expected functions, and the length of flexible linkers was found to affect the catalytic efficiency of fusion proteins [12, 19]. The fusion protein CrtZAs-WBs produced more astaxanthin than the other fusion proteins, so two types of flexible linkers with different lengths were introduced into it to further increase astaxanthin production.
The flexible linkers (Gly-Ser-Gly)x and (Gly-Ser)y are commonly used in the construction of fusion proteins [1, 6]. Here, two series of fusion proteins, CrtZAs-(Gly-Ser-Gly)x-CrtWBs (x = 1, 2, 3 and 4) and CrtZAs-(Gly-Ser)y-CrtWBs (y = 1, 2, 3, 4 and 5), were constructed, and the corresponding expression cassette was inserted in plasmid p5C, respectively. These fusion proteins were also expressed under the control of the promoter J23119 and the RBS apFAB917 (Fig. 3a). As shown in Fig. 3b, ZF237T/CrtZAs-(GS)1-WBs, ZF237T/CrtZAs-(GS)2-WBs, ZF237T/CrtZAs-(GSG)2-WBs and ZF237T/CrtZAs-(GSG)3-WBs produced 11.31 mg/L, 9.86 mg/L, 9.95 mg/L and 10.05 mg/L of astaxanthin, corresponding to an increase of 43.4%, 25.0%, 26.2% and 27.5% over the strain ZF237T/CrtZAs-WBs, respectively. Moreover, the astaxanthin productions of the four strains increased by 40.2%, 22.2%, 23.4% and 24.6% compared with the non-fusion protein expressing strain ZF237T/crtZBsWPs, respectively.
Production of astaxanthin, lycopene and β-carotene by ZF237T-derived strains expressing different CrtZAs-(GS)y-WBs and CrtZAs-(GSG)x-WBs fusion proteins. a Designs for the introduction of peptide linkers. The swallowtail arrow denotes the promoter J23119, the blue solid circle denotes the RBS apFAB917. b, c The productions of astaxanthin, lycopene and β-carotene. The horizontal axis shows the abbreviations of different fusion proteins, i.e. (GS)1 and (GSG)1 denote fusion proteins CrtZAs-(GS)1-WBs and CrtZAs-(GSG)1-WBs, respectively (color figure online)
At the same time, the analysis of by-products showed that all strains accumulated the precursors, lycopene and β-carotene (Table 2). The strains expressing the fusion proteins CrtZAs-(GS)1-WBs and CrtZAs-(GS)2-WBs produced 3.71 mg/L and 2.93 mg/L of lycopene, respectively, which was lower than the corresponding value of strain ZF237T/CrtZAs-WBs (5.78 mg/L), while strains CrtZAs-(GSG)2-WBs and CrtZAs-(GSG)3-WBs produced 6.52 mg/L and 5.30 mg/L of lycopene (Fig. 3c). The β-carotene production of the strains expressing the fusion proteins CrtZAs-(GS)y-WBs were lower than the corresponding value of strain ZF237T/CrtZAs-WBs (5.92 mg/L), while the strains expressing the fusion proteins CrtZAs-(GSG)1-WBs, CrtZAs-(GSG)2-WBs and CrtZAs-(GSG)4-WBs produced 35.69, 8.60 and 33.57 mg/L of β-carotene, which was much higher than that of strain ZF237T/CrtZAs-WBs (Fig. 3c). In addition, strains expressing the fusion proteins CrtZAs-(GS)y-WBs except for CrtZAs-(GS)5-WBs did not produce detectable amounts of canthaxanthin and echinenone, and the species of by-products increased with the increase of linker length, while strains expressing the fusion proteins CrtZAs-(GSG)x-WBs produced different intermediates with the varied length of linkers (Table 2). These results confirmed that the length of the peptide linkers (GS)x and (GSG)y can affect the catalytic efficiency of fusion proteins and thus influence by-product generation.
Optimization of the carbon source for strain ZF237T/CrtZAs-(GS)1-WBs
Glycerol has a significantly higher average degree of reduction per carbon (C3H8O3: κ = 4.67) than sugars such as glucose (C6H12O6: κ = 4) or xylose (C5H10O5: κ = 4) [9]. This implies that glycerol can provide more reducing equivalents for astaxanthin biosynthesis, where two moles of NADPH are theoretically needed to converse one mole β-carotene into an equal amount of astaxanthin [10]. It is, therefore, perhaps not surprising that Park et al. demonstrated that glycerol is better for astaxanthin accumulation than glucose [32]. We, therefore, optimized the carbon source in the fermentation medium to increase astaxanthin production in shake flasks.
As shown in Fig. 4, when 10 g/L of glycerol was added into 2 × YT medium, the astaxanthin production was 23.71 mg/L and the intracellular content reached 4.67 mg/g DCW at the end of fermentation. These values were, respectively, 109.64% and 181.33% higher than those produced in 2 × YT medium containing 10 g/L glucose (1.66 mg/g DCW). This result verified that glycerol is more favorable than glucose for astaxanthin production. We also used a mixture of glycerol and glucose as carbon source. In host strain ZF237T, phosphotransferase system is inactivated, the expression of galP is enhanced and the genes involving acetate production are not inactivated [20]. To avoid metabolic overflow, 1% (m/v) of total carbon source was added into 2 × YT medium [27]. In view of that glycerol is better for astaxanthin production than glucose, the ratio of glycerol to glucose was set in a value no less than 1 in the mixtures. The analysis of astaxanthin production after 96 h of fermentation showed that the production decreased with the decrease of the ratio of glycerol to glucose (Fig. 4). When 8 g/L glycerol and 2 g/L glucose were added, the production of astaxanthin reached 26.16 mg/L and the content reached 5.18 mg/g DCW, which were comparable to the highest level in E. coli in shake flask reported previously [32]. In addition, the two values were 10.32% and 10.76% higher than those produced in 2 × YT medium containing 10 g/L glycerol, respectively. These results were consistent with previous report that appropriate ratio of glycerol to glucose within mixed carbon source was more beneficial for aromatic compounds’ production in PTS−Glc+E. coli strain [27]. The phenomena indicated that mixed carbon source with appropriate ratio of glycerol to glucose could supply better precursors and NADPH than single carbon source for astaxanthin biosynthesis.
The astaxanthin production of strain ZF237T/CrtZAs-(GS)1-WBs in 2 × YT media with different carbon sources. Abscissa values represent the ratios of glycerol to glucose in media except 1% represents the carbon source is single carbon source glycerol, the concentration (m/v) of glycerol equal to or greater than the glucose (color figure online)
Discussion
The regulation of heterologous biosynthetic pathways is a recurrent theme in metabolic engineering, especially for the biosynthetic pathways of natural products. Thus, a simple method for effective combinatorial expression is important. Astaxanthin is an important natural pigment with extremely strong antioxidant activity which was reported to exceed those of β-carotene and α-tocopherol [15]. Recently, several engineered E. coli strains for astaxanthin biosynthesis were constructed through metabolic engineering. Examples include E. coli A1 (pFZ153) [25], E. coli ASTA-1 [23] and E. coli E/F [40]. To obtain these engineered strains, researchers adopted different strategies to optimize the expression levels of β-carotene hydroxylase and ketolase. Here, we combined β-carotene hydroxylases and ketolases from different sources within a single expression cassette to regulate astaxanthin production in E. coli through expression of artificial operons and fusion proteins. The yield of astaxanthin produced by strain ZF237T/CrtZAs-(GS)1-WBs was 127.6% and 40.2% higher than that of strains ZF237T/crtZAsWBs and ZF237T/crtZBsWPs, respectively. The production and intracellular content of astaxanthin produced by the best strain are comparable to the highest level in E. coli reported by other researchers after optimization of carbon source in shake flask.
Previous studies revealed that CrtZ from Alcaligenes sp. strain PC-1, Paracoccus sp. N81106 and Brevundimonas sp. SD212 have high zeaxanthin production efficiency from β-carotene, while CrtW from Brevundimonas sp.SD212, Paracoccus sp. N81106 and Nostoc sp. PCC7120 showed high astaxanthin production efficiency in E. coli producing zeaxanthin [7, 8, 35]. However, the ZF237T-derived strains expressing the operons crtZAsWNs, crtZBsWNs, crtZPsWPs, and crtZPsWBs did not accumulate detectable amounts of astaxanthin (Table 1). The results of qRT-PCR indicated that the operons crtZAsWNs, crtZBsWNs, crtZPsWPs, and crtZPsWBs were expressed efficiently at the RNA level (Fig. S1), yet the strains did not accumulate detectable amount of astaxanthin and we failed to relate the relative transcription levels to astaxanthin accumulation. However, we found that the total relative transcriptional levels of genes crtZ and crtW within the four operons either too low or too high compared with the other five operons, such as crtZAsWNs to crtZAsWBs and crtZPsWPs to crtZPsWNs. We speculated this possibly affected the enzyme activities. Another possible reason is the competition for common substrates between the CrtZ and CrtW enzymes due to their broad substrate spectra and different affinity for the metabolites in vivo [29]. In addition, we noted that strains ZF237T/crtZAsWNs and ZF237T/crtZBsWNs did not accumulated detectable amounts of astaxanthin while strain ZF237T/crtZPsWNs accumulated 1.67 mg/L of astaxanthin (Table 1 and Fig. 2b). qRT-PCR showed that the total relative transcriptional of genes crtZ and crtW within operons crtZAsWNs and crtZBsWNs were lower than that of operon crtZPsWNs (Fig. S1). The exact reason for these failure merits further investigation in the future.
Park et al. fused eight tags to the N- or C-terminus of the trCrBKT membrane protein and screened two signal peptides, OmpF and TrxA, which could enhance the production of astaxanthin 2.08-fold compared to that obtained without a tag [32]. Ye et al. linked CrtW and CrtZ using a fexible eight-amino acids linker and targeted the fusion to membrane via a GlpF protein fusion in E. coli, and the production of astaxanthin was increased by 215.4% after optimal localization configuration [38]. We fused CrtZ and CrtW based on our operons directly or via peptide linkers. We found that the ZF237T-derived strains expressing fusion proteins with CrtZ attached to the N-terminus accumulated much more astaxanthin than the strains expressing fusion proteins with CrtW attached to the N-terminus, and most of the ZF237T-derived strains harboring CrtWBs-ZAs fusion proteins did not accumulate astaxanthin after introducing peptide linkers (Table S4). This phenomenon was similar to the research just reported by Nogueira et al. [31]. We inferred that the arrangement of modules within fusion proteins has a significant influence on their conformations and further affects their activities. Therefore, I-TASSER protein structure predictor was used to produce homology models of the fusion proteins CrtZAs-WBs and CrtWBs-ZAs, because the astaxanthin production of strain ZF237T/CrtZAs-WBs was 58.5% higher than that of strain ZF237T/crtZAsWBs while strain ZF237T/CrtWBs-ZAs only produced an amount equivalent to 11.67% of strain ZF237T/crtZAsWBs. Sequence analysis revealed that CrtZ from Alcaligenes sp. strain PC-1 and CrtW from Brevundimonas sp. SD212 both have several conserved histidine motifs (HX4H, HX2HH or HHXHH), four in CrtZAs and three in CrtWBs (Fig. 5a). These motifs are Fe2+ binding sites and are also characteristic of membrane-associated fatty acid desaturases and sterol desaturases [8, 10]. Functional characterization of β-carotene hydroxylase and mutational analysis of β-carotene ketolase demonstrated that these motifs are indispensable for their activities [4, 39]. Considering that topology models of conserved histidine motifs were available [4, 39], we concentrated our analysis on the changes of the locations of the histidine motifs in the two fusion proteins. First, we established the models of the proteins CrtZAs and CrtWBs and found that the simulated structures of CrtZAs and CrtWBs resemble that of a cone frustum which is composed of an α helix and an aperiodical coil. The conserved histidine motifs of the two enzymes were adjacent to each other at the bottom of the cone frustum, and the secondary structures of the histidine motifs were α-helical except for the histidine motif HDGLVH of CrtZAs, which was an aperiodical coil (Fig. 5b). The two models were consistent with the topology models and prediction of 3D structures reported previously [4, 31, 39]. Second, in view of astaxanthin production, it was conceivable that the locations of the histidine motifs were different in the structures of the fusion proteins CrtZAs-WBs and CrtWBs-ZAs. Indeed, structure modeling showed that the fusion protein CrtZAs-WBs formed an oblique cylinder while the shape of CrtWBs-ZAs was cylindrical, and the hisidine motifs were distributed in different parts of the two cylinders. The surface structure analysis showed that all the histidine motifs were on the surface of the CrtZAs-WBs fusion proteins, whereas the three histidine motifs (HHEHH of CrtZAs, HDGLVH and HRLHH of CrtWBs) were inside the fusion protein CrtWBs-ZAs (Fig. S2). Therefore, our structure analysis showed that the differences in locations between the two fusion protein formats may affect the enzyme activity and substrate preference. Certainly, we had no information about the active sites of enzymes CrtZAs and CrtWBs, molecular docking could be used to identity the active sites of the two enzymes, and investigate the changes of physical distance between the sites in fusion proteins in the future. In addition, enzyme kinetics should be adopted to investigate the activities of the fusion proteins CrtZAs-WBs and CrtWBs-ZAs.
Homology based models CrtZAs, CrtWBs, CrtZAs-WBs and CrtWBs-ZAs. a The histidine motifs of CrtZ from Alcaligenes sp. strain PC-1 and CrtW from Brevundimonas sp. SD212 were highlighted in uppercase letter with different colors. b Structure-based homology models of CrtZAs, CrtWBs and fusion proteins. The colors of histidine motifs correspond to the colors of the capital letter in a (color figure online)
Different peptide linkers are always used to create different fusion proteins because the nature of linker significantly affects the correct folding and activity of fusion protein [1]. Glycine has the simplest structure, minimum molecular weight and best flexibility compared to other 19 amino acids. Serine is the most hydrophilic polar amino acid. The incorporation of Gly and Ser can maintain the flexibility and the stability of linker in aqueous solution [6]. The production of astaxanthin was significantly different among the ZF237T-derived strains expressing CrtZAs-WBs fusion proteins, with the highest value being 21-fold higher than the lowest. In addition, we noted that the accumulated amounts of lycopene and β-carotene were different among these strains. The lycopene and β-carotene production of the ZF237T-derived strains harboring fusion proteins with GSG linkers was higher than with GS linkers (Fig. 3c). These results indicated that the length and composition of the peptide linkers not only influenced the activities of the fusion proteins but also the metabolic fluxes in carotenoid biosynthesis. This was consistent with previous reports that the linker length and composition greatly affect the production of patchoulol, resveratrol and astaxanthin [1, 12, 31]. In attempt to explore the reasons, we produced homology models of fusion proteins CrtZAs-(GS)2-WBs and CrtZAs-(GSG)2-WBs via I-TASSER protein structure predictor because the strains expressing the two fusions accumulated similar amount of astaxanthin but different amounts of β-carotene and lycopene (Fig. 3b, c). The models showed that the secondary structure of histine motif HHEHH in fusion CrtZAs-(GS)2-WBs was aperiodical while it was α helix in fusion CrtZAs-(GSG)2-WBs (Fig. S3). In addition, we found that (GS)2 linker located inside of the fusion CrtZAs-(GS)2-WBs and (GSG)2 linker located outside of the fusion CrtZAs-(GSG)2-WBs (Fig. S3). We inferred these changes caused the different activities of the two fusions and thus affected the flux for astaxanthin biosynthesis.
The biosynthetic pathways of natural products often contain bi-functional enzymes and regulatory elements. To construct cell factories for the industrial manufactory of natural products, these bio-bricks must be first assembled into one or more operons and then optimized. Here, we succeeded in regulating the production of astaxanthin in vivo by combining the expression of different oxygenases. Our observations provide a reference for the construction and regulation of other pathways comprising bi- or even multi-functional enzymes.
References
Albertsen L, Chen Y, Bach LS, Rattleff S, Maury J, Brix S, Nielsen J, Mortensen UH (2011) Diversion of flux toward sesquiterpene production in saccharomyces cerevisiae by fusion of host and heterologous enzymes. Appl Environ Microbiol 77:1033–1040. https://doi.org/10.1128/aem.01361-10
Alcaíno J, Barahona S, Carmona M, Lozano C, Marcoleta A, Niklitschek M, Sepúlveda D, Baeza M, Cifuentes V (2008) Cloning of the cytochrome p450 reductase (crtR) gene and its involvement in the astaxanthin biosynthesis of Xanthophyllomyces dendrorhous. BMC Microbiol 8:169. https://doi.org/10.1186/1471-2180-8-169
Biggs BW, Lim CG, Sagliani K, Shankar S, Stephanopoulos G, De Mey M, Ajikumar PK (2016) Overcoming heterologous protein interdependency to optimize P450-mediated taxol precursor synthesis in Escherichia coli. Proc Natl Acad Sci 113:3209
Bouvier F, Keller Y, d’Harlingue A, Camara B (1998) Xanthophyll biosynthesis: molecular and functional characterization of carotenoid hydroxylases from pepper fruits (Capsicum annuum L.). Biochim Biophys Acta 1391:320–328. https://doi.org/10.1016/S0005-2760(98)00029-0
Chang MCY, Eachus RA, Trieu W, Ro D-K, Keasling JD (2007) Engineering Escherichia coli for production of functionalized terpenoids using plant P450s. Nat Chem Biol 3:274. https://doi.org/10.1038/nchembio875
Chen X, Zaro JL, Shen W-C (2013) Fusion protein linkers: property, design and functionality. Adv Drug Deliver Rev 65:1357–1369. https://doi.org/10.1016/j.addr.2012.09.039
Choi S-K, Matsuda S, Hoshino T, Peng X, Misawa N (2006) Characterization of bacterial β-carotene 3,3′-hydroxylases, CrtZ, and P450 in astaxanthin biosynthetic pathway and adonirubin production by gene combination in Escherichia coli. Appl Microbiol Biotechnol 72:1238. https://doi.org/10.1007/s00253-006-0426-2
Choi S-k, Nishida Y, Matsuda S, Adachi K, Kasai H, Peng X, Komemushi S, Miki W, Misawa N (2005) Characterization of β-carotene ketolases, CrtW, from marine bacteria by complementation analysis in Escherichia coli. Mar Biotechnol 7:515–522. https://doi.org/10.1007/s10126-004-5100-z
Durnin G, Clomburg J, Yeates Z, Alvarez PJJ, Zygourakis K, Campbell P, Gonzalez R (2009) Understanding and harnessing the microaerobic metabolism of glycerol in Escherichia coli. Biotechnol Bioeng 103:148–161. https://doi.org/10.1002/bit.22246
Fraser PD, Miura Y, Misawa N (1997) In vitro characterization of astaxanthin biosynthetic enzymes. J Biol Chem 272:6128–6135. https://doi.org/10.1074/jbc.272.10.6128
Fraser PD, Shimada H, Misawa N (1998) Enzymic confirmation of reactions involved in routes to astaxanthin formation, elucidated using a direct substrate in vitro assay. Eur J Biochem 252:229–236. https://doi.org/10.1046/j.1432-1327.1998.2520229.x
Guo H, Yang Y, Xue F, Zhang H, Huang T, Liu W, Liu H, Zhang F, Yang M, Liu C, Lu H, Zhang Y, Ma L (2017) Effect of flexible linker length on the activity of fusion protein 4-coumaroyl-CoA ligase:stilbene synthase. Mol BioSyst 13:598–606. https://doi.org/10.1039/C6MB00563B
Heckman KL, Pease LR (2007) Gene splicing and mutagenesis by PCR-driven overlap extension. Nat Protoc 2:924. https://doi.org/10.1038/nprot.2007.132
Henke N, Heider S, Peters-Wendisch P, Wendisch V (2016) Production of the marine carotenoid astaxanthin by metabolically engineered Corynebacterium glutamicum. Mar Drugs 14:124
Higuera-Ciapara I, Félix-Valenzuela L, Goycoolea FM (2006) Astaxanthin: a review of its chemistry and applications. Crit Rev Food Sci 46:185–196. https://doi.org/10.1080/10408690590957188
Jin J, Wang Y, Yao M, Gu X, Li B, Liu H, Ding M, Xiao W, Yuan Y (2018) Astaxanthin overproduction in yeast by strain engineering and new gene target uncovering. Biotechnol Biofuels 11:230. https://doi.org/10.1186/s13068-018-1227-4
Jin W, Xu X, Jiang L, Zhang Z, Li S, Huang H (2015) Putative carotenoid genes expressed under the regulation of Shine-Dalgarno regions in Escherichia coli for efficient lycopene production. Biotechnol Lett 37:2303–2310. https://doi.org/10.1007/s10529-015-1922-1
Kosuri S, Goodman DB, Cambray G, Mutalik VK, Gao Y, Arkin AP, Endy D, Church GM (2013) Composability of regulatory sequences controlling transcription and translation in Escherichia coli. Proc Natl Acad Sci 110:14024
Li G, Huang Z, Zhang C, Dong B-J, Guo R-H, Yue H-W, Yan L-T, Xing X-H (2016) Construction of a linker library with widely controllable flexibility for fusion protein design. Appl Microbiol Biotechnol 100:215–225. https://doi.org/10.1007/s00253-015-6985-3
Li Y, Lin Z, Huang C, Zhang Y, Wang Z, Tang Y-j, Chen T, Zhao X (2015) Metabolic engineering of Escherichia coli using CRISPR–Cas9 meditated genome editing. Metab Eng 31:13–21. https://doi.org/10.1016/j.ymben.2015.06.006
Lin Z, Xu Z, Li Y, Wang Z, Chen T, Zhao X (2014) Metabolic engineering of Escherichia coli for the production of riboflavin. Microb Cell Fact 13:104. https://doi.org/10.1186/s12934-014-0104-5
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Lu Q, Bu Y-F, Liu J-Z (2017) Metabolic engineering of Escherichia coli for producing astaxanthin as the predominant carotenoid. Mar Drugs 15:296
Lv X, Xu H, Yu H (2013) Significantly enhanced production of isoprene by ordered coexpression of genes dxs, dxr, and idi in Escherichia coli. Appl Microbiol Biotechnol 97:2357–2365. https://doi.org/10.1007/s00253-012-4485-2
Ma T, Zhou Y, Li X, Zhu F, Cheng Y, Liu Y, Deng Z, Liu T (2015) Genome mining of astaxanthin biosynthetic genes from Sphingomonas sp. ATCC 55669 for heterologous overproduction in Escherichia coli. Biotechnol J 11:228–237. https://doi.org/10.1002/biot.201400827
Martín JF, Gudiña E, Barredo JL (2008) Conversion of β-carotene into astaxanthin: two separate enzymes or a bifunctional hydroxylase-ketolase protein? Microb Cell Fact 7:3. https://doi.org/10.1186/1475-2859-7-3
Martínez K, de Anda R, Hernández G, Escalante A, Gosset G, Ramírez OT, Bolívar FG (2008) Coutilization of glucose and glycerol enhances the production of aromatic compounds in an Escherichia coli strain lacking the phosphoenolpyruvate: carbohydrate phosphotransferase system. Microb Cell Fact 7:1. https://doi.org/10.1186/1475-2859-7-1
Misawa N, Satomi Y, Kondo K, Yokoyama A, Kajiwara S, Saito T, Ohtani T, Miki W (1995) Structure and functional analysis of a marine bacterial carotenoid biosynthesis gene cluster and astaxanthin biosynthetic pathway proposed at the gene level. J Bacteriol 177:6575–6584. https://doi.org/10.1128/jb.177.22.6575-6584.1995
Misawa N, Shimada H (1998) Metabolic engineering for the production of carotenoids in non-carotenogenic bacteria and yeasts. J Biotechnol 59:169–181. https://doi.org/10.1016/S0168-1656(97)00154-5
Nishida Y, Adachi K, Kasai H, Shizuri Y, Shindo K, Sawabe A, Komemushi S, Miki W, Misawa N (2005) Elucidation of a carotenoid biosynthesis gene cluster encoding a novel enzyme, 2,2′-β-hydroxylase, from Brevundimonas sp. strain SD212 and combinatorial biosynthesis of new or rare xanthophylls. Appl Environ Microbiol 71:4286
Nogueira M, Enfissi EMA, Welsch R, Beyer P, Zurbriggen MD, Fraser PD (2019) Construction of a fusion enzyme for astaxanthin formation and its characterisation in microbial and plant hosts: a new tool for engineering ketocarotenoids. Metab Eng 52:243–252. https://doi.org/10.1016/j.ymben.2018.12.006
Park SY, Binkley RM, Kim WJ, Lee MH, Lee SY (2018) Metabolic engineering of Escherichia coli for high-level astaxanthin production with high productivity. Metab Eng 49:105–115. https://doi.org/10.1016/j.ymben.2018.08.002
Roy A, Kucukural A, Zhang Y (2010) I-TASSER: a unified platform for automated protein structure and function prediction. Nat Protoc 5:725. https://doi.org/10.1038/nprot.2010.5
Russell DW, Sambrook J (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor, Cold Spring Harbor Laboratory
Scaife MA, Burja AM, Wright PC (2009) Characterization of cyanobacterial β-carotene ketolase and hydroxylase genes in Escherichia coli, and their application for astaxanthin biosynthesis. Biotechnol Bioeng 103:944–955. https://doi.org/10.1002/bit.22330
Scaife MA, Ma CA, Ninlayarn T, Wright PC, Armenta RE (2012) Comparative analysis of β-carotene hydroxylase genes for astaxanthin biosynthesis. J Nat Prod 75:1117–1124. https://doi.org/10.1021/np300136t
Yang J, Zhang Y (2015) I-TASSER server: new development for protein structure and function predictions. Nucleic Acids Res 43:W174–W181. https://doi.org/10.1093/nar/gkv342
Ye L, Zhu X, Wu T, Wang W, Zhao D, Bi C, Zhang X (2018) Optimizing the localization of astaxanthin enzymes for improved productivity. Biotechnol Biofuels 11:278. https://doi.org/10.1186/s13068-018-1270-1
Ye RW, Stead KJ, Yao H, He H (2006) Mutational and functional analysis of the β-carotene ketolase involved in the production of canthaxanthin and astaxanthin. Appl Environ Microbiol 72:5829
Zhang C, Seow VY, Chen X, Too H-P (2018) Multidimensional heuristic process for high-yield production of astaxanthin and fragrance molecules in Escherichia coli. Nat Commun 9:1858. https://doi.org/10.1038/s41467-018-04211-x
Zhou P, Xie W, Li A, Wang F, Yao Z, Bian Q, Zhu Y, Yu H, Ye L (2017) Alleviation of metabolic bottleneck by combinatorial engineering enhanced astaxanthin synthesis in Saccharomyces cerevisiae. Enzyme Microb Technol 100:28–36. https://doi.org/10.1016/j.enzmictec.2017.02.006
Zhou P, Ye L, Xie W, Lv X, Yu H (2015) Highly efficient biosynthesis of astaxanthin in Saccharomyces cerevisiae by integration and tuning of algal crtZ and bkt. Appl Microbiol Biotechnol 99:8419–8428. https://doi.org/10.1007/s00253-015-6791-y
Zhou YJ, Gao W, Rong Q, Jin G, Chu H, Liu W, Yang W, Zhu Z, Li G, Zhu G, Huang L, Zhao ZK (2012) Modular pathway engineering of diterpenoid synthases and the mevalonic acid pathway for miltiradiene production. J Am Chem Soc 134:3234–3241. https://doi.org/10.1021/ja2114486
Funding
This work was supported by the National Natural Science Foundation of China (NSFC-21621004, NSFC-21776208 and NSFC-21776209).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wu, Y., Yan, P., Liu, X. et al. Combinatorial expression of different β-carotene hydroxylases and ketolases in Escherichia coli for increased astaxanthin production. J Ind Microbiol Biotechnol 46, 1505–1516 (2019). https://doi.org/10.1007/s10295-019-02214-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-019-02214-1