Abstract
Resveratrol has been the subject of numerous scientific investigations due to its health-promoting activities against a variety of diseases. However, developing feasible and efficient microbial processes remains challenging owing to the requirement of supplementing expensive phenylpropanoic precursors. Here, various metabolic engineering strategies were developed for efficient de novo biosynthesis of resveratrol. A recombinant malonate assimilation pathway from Rhizobium trifolii was introduced to increase the supply of the key precursor malonyl-CoA and simultaneously, the clustered regularly interspaced short palindromic repeats interference system was explored to down-regulate fatty acid biosynthesis pathway to inactivate the malonyl-CoA consumption pathway. Down-regulation of fabD, fabH, fabB, fabF, fabI increased resveratrol production by 80.2, 195.6, 170.3, 216.5 and 123.7%, respectively. Furthermore, the combined effect of these genetic perturbations was investigated, which increased the resveratrol titer to 188.1 mg/L. Moreover, the efficiency of this synthetic pathway was improved by optimizing the expression level of the rate-limiting enzyme TAL based on reducing mRNA structure of 5′ region. This further increased the final resveratrol titer to 304.5 mg/L. The study described here paves the way to the development of a simple and economical process for microbial production of resveratrol.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
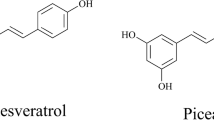
Resveratrol (3, 5, 4′-trihydroxy-trans-stilbene), a polyphenolic compound of the stilbene family, is responsible for a decreased risk of diabetes and heart diseases due to its anti-oxidative, anti-inflammatory, anti-cancer, and chemo-preventive activities [1, 9]. Currently, resveratrol is mainly obtained by extraction from grapes, peanuts, bushes and other plants. This inefficient and tedious process hampers its application as a widespread nutraceutical. Because of this, microbial production of resveratrol has attracted worldwide interest as this method is easily scaled to commercial application and avoids using toxic catalysts [5, 33].
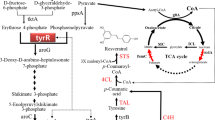
As shown in Fig. 1, three catalytic steps are required for the conversion of aromatic amino acid l-tyrosine to resveratrol [6, 18]. This process first begins with the deamination of l-tyrosine to p-coumaric acid by the action of tyrosine ammonia lyase (TAL). 4-Coumarate: CoA ligase (4CL) mediates its corresponding CoA ester, coumaroyl-CoA formation when p-coumaric acid has been generated. Coumaroyl-CoA is subsequently condensed with three malonyl-CoA units to form resveratrol by the sequential action of the type III polyketide synthase, stilbene synthase (STS).
Engineering of E. coli metabolic pathways for efficient formation of resveratrol. a The schematics of malonyl-CoA synthesis system. This included malonate assimilation pathway (matB and matC) and CRISPR interference system. The plasmids used for dCas9 and sgRNA expression and the process to repress fatty acid biosynthesis were shown. b The overall resveratrol biosynthesis pathway was shown. The repressed genes were shown in red. The metabolic pathway that performs heterologous biosynthesis of resveratrol from d-glucose in E. coli was shown in blue (color figure online)
To date, several studies have already made significant gains for microbial resveratrol production. For example, relevant study demonstrates that supplementation of 15 mM p-coumaric acid and 0.05 mM cerulenin lead to a maximum titer of 2.3 g/L in E. coli [17]. Furthermore, in our research, it is found that 35 mg/L of resveratrol could be obtained from l-tyrosine via modular pathway engineering strategies [28]. As the reliance on supplementation of phenylpropanoids in these strategies is prohibitive during process scale-up, microbial production of resveratrol from cheap and renewable substrate d-glucose has attracted more and more attention.
One relevant study demonstrates that overexpressing acetyl-CoA carboxylase and performing multiple-integration of pathway genes in fed-batch fermentation result in a high resveratrol titer of 415.7 mg/L from d-glucose in Saccharomyces cerevisiae [15]. However, similar achievements are not observed in E. coli, which is often preferred as engineered strains for many metabolites due to its simple genetic background, short cultivate time and low culture condition requirement [2]. Recent study utilizes a site-specific integration strategy to integrate the resveratrol synthetic pathway into the chromosome of E. coli, which results in only 4.6 mg/L of resveratrol from d-glucose [18].
Hence, there is a pressing need to improve de novo resveratrol production titer in E. coli cells. Endogenous central metabolism strongly predominates and competes for carbon and energy sources during the synthesis of malonyl-CoA, which is the major bottleneck of the phenylpropanoid pathway due to both the low basal level and high requirement of this metabolite [24, 27]. The previous efforts to improve malonyl-CoA availability mainly focus on engineering the upstream pathway, such as overexpression of acetyl-CoA carboxylase [32] and the malonate assimilation pathway [14, 28] to increase malonly-CoA supply, or overexpression of acetyl-CoA synthase and deleting acetyl-CoA consumption pathways to increase acetyl-CoA supply [32]. Another relevant efforts have been made in engineering the malonyl-CoA consumption pathway. As inactivating fatty acid biosynthesis through conventional gene knockout strategies is usually lethal to host cells, a specific inhibitor cerulenin repressing fabB and fabF is used to increase intracellular malonyl-CoA [26]. While cerulenin is expensive and cost prohibitive, antisense RNAs are employed to down-regulate fabB and fabF [29] or fabD [34] instead, leading to a 4.5-fold increase in intracellular malonyl-CoA concentration. In our one study, the clustered regularly interspaced short palindromic repeats interference (CRISPRi) system is also used to repress fabB and fabF to improve malonyl-CoA level [27].
All of these studies demonstrate that both manipulation of precursor supply and inactivation of malonyl-CoA consumption pathway would improve the intracellular malonyl-CoA concentration. Based on this, an efficient malonyl-CoA synthesis method involving manipulating both precursor supply and malonyl-CoA consumption is established to increase de novo resveratrol production. To avoid manipulating complex central metabolism when overexpressing acetyl-CoA carboxylase or relational precursor enzymes and deleting enzymes involved in acetyl-CoA depletion [32], a recombinant malonate assimilation pathway from Rhizobium trifolii [14] was introduced to synthesize malonyl-CoA directly from malonate. In the same time, the CRISPRi system, which could simultaneously manipulate multiple genes in E. coli compared to antisense RNAs, was explored to conduct a more systematic study on the effect of down-regulating up to five genes involved in fatty acid biosynthesis pathway.
Moreover, numerous studies demonstrate that the low turnover number of TAL hampers efficient do novo biosynthesis of compounds derived from phenylpropanoid pathway [17, 24]. To overcome this limitation, firstly, the original RgTAL was replaced with another TAL from Trichosporon cutaneum (TcTAL), which exhibited high activities toward l-tyrosine [10]. Secondly, the TcTAL expression was optimized by modifying its mRNA secondary structure of 5′ region. Finally, the combined effect of these genetic modifications increased the final resveratrol titer to 304.5 mg/L. The strategy described here presents developing an alternative source of resveratrol with numerous applications in food, nutraceutical and pharmaceutical industry.
Materials and methods
E. coli strains, plasmids and general techniques
Luria broth (LB) were used for plasmid construction and MOPS minimal medium [22] supplemented with 5 g/L d-glucose and an additional 4 g/L NH4Cl was used for resveratrol production. E. coli JM109 and BL21 (DE3) were used for plasmid construction and resveratrol production, respectively. Various combinations of ampicillin (100 μg/mL), kanamycin (40 μg/mL), chloramphenicol (20 μg/mL), and streptomycin (40 μg/mL) were added when required. The compatible vectors pETDuet-1, pCDFDuet-1, pCOLADuet-1, and pACYCDuet-1 were purchased from Novagen (Darmstadt, Germany). All restriction enzymes and DNA ligase were purchased from Novagen (Darmstadt, Germany). RgTAL from Rhodotorula glutinis [24], TcTAL from Trichosporon cutaneum [10], 4CL from Petroselinum crispum, STS from Vitis vinifera [17], and matB and matC from Rhizobium trifolii [14] were codon-optimized for E. coli expression (http://www.jcat.de/) and synthesized by GenScript (Nanjing, China). DNA sequences for all synthesized genes are shown in the Supplementary Material. UV/vis spectrophotometer (UVmini-1240, Shimadzu, Kyoto, Japan) was used to monitor cell growth by measuring absorbance at 600 nm (OD600). Primers used in the study were shown in Table 1.
Chromosomal integration of tyrA fbr and aroG fbr
The chromosomal integration of tyrA fbr and aroG fbr was performed according to our previous study [31]. Briefly, two mutations of Met-53-Ile in the chorismate mutase domain and Ala-354-Val in the prephenate dehydrogenase domain were performed to obtain tyrA fbr; One mutation of Asp-146-Asn substitution was performed to obtain aroG fbr. Lambda-red recombination-based method [4] was used to integrate tyrA fbr–aroG fbr cassette into lacZ locus of E. coli BL21 (DE3) under T7 promoter. This cassette was further verified by colony PCR and sequencing.
Construction of CRISPRi system to repress single gene
dCas9 [23] was synthesized by GenScript (Nanjing, China) and inserted into NcoI/AvrII site of pACYCDuet-1, resulted in pACYC-dCas9. The T7-dCas9 region including T7 promoter and dCas9 was amplified from pACYC-dCas9 using primers Pf_T7dCas9(KpnI) and Pr_T7dCas9(XhoI) and inserted into KpnI/XhoI site of pACYC-matC-matB [28], which resulted in pACYC-matC-matB-dCas9. The sgRNA chimera consisting of Trc promoter, complementary region for DNA binding, dCas9-binding hairpin and transcription terminator was synthesized (GenScript, Nanjing, China) and inserted into pfoI/BamHI site of pCOLADuet-1. Other sgRNA cassettes with new complementary regions were generated by PCR-based site-directed mutagenesis (TaKaRa MutanBEST Kit, Takara Biotechnology, Dalian, China) with corresponding primers. Primers and the resultant sgRNA expression vectors are shown in Table 2.
Construction of CRISPRi system to repress multiple genes
To repress multiple genes, different sgRNA chimeras were cloned into pCOLADuet-1. Primers Pf_sgRNA(BamHI)/Pr_sgRNA(EcoRI) were used to amplify anti-fabD(medium), anti-fabI(low), anti-fabB(medium) and anti-fabH(low) sgRNA sequences into the BamHI/EcoRI site of pCOLA-fabF(high). This resulted in pCOLA-fabF(high)/fabD(medium), pCOLA-fabF(high)/fabI(low), pCOLA-fabF(high)/fabB(medium) and pCOLA-fabF(high)/fabH(low). Primers Pf_sgRNA(EcoRI)/Pr_sgRNA(HindIII) were used to amplify anti-fabD(medium), anti-fabI(low) and anti-fabH(low) sgRNA sequences into EcoRI/HindIII site of pCOLA-fabF(high)/fabB(medium). This resulted in pCOLA-fabF(high)/fabB(medium)/fabD(medium), pCOLA-fabF(high)/fabB(medium)/fabI(low), pCOLA-fabF(high)/fabB(medium)/fabH(low). Primers Pf_sgRNA(HindIII)/Pr_sgRNA(NdeI) were used to amplify anti-fabD(medium) and anti-fabH(low) sequences into HindIII/NdeI site of pCOLA-fabF(high)/fabB(medium)/fabI(low). This resulted in pCOLA-fabF(high)/fabB(medium)/fabI(low)/fabD(medium) and pCOLA-fabF(high)/fabB(medium)/fabI(low)/fabH(low). Primers Pf_sgRNA(NdeI)/Pr_sgRNA(BglII) were used to amplify anti-fabH(low) sequence into NdeI/BglII site of pCOLA-fabF(high)/fabB(medium)/fabI(low)/fabD(medium). This resulted in pCOLA-fabF(high)/fabB(medium)/fabI(low)/fabD(medium)/fabH(low).
Construction of different TcTAL variants
Primers Pf_Trc(PfoI) and Pr_Trc(FseI) were used to clone Trc promoter from pTrcHis2B and insert into PfoI/FseI site of pCDFDuet-1, resulted in pCDFD-Trc. TAL from Trichosporon cutaneum (TcTAL) [10] was cloned from pUC57-TcTAL (GenScript, Nanjing, China) with primers Pf_TcTAL(NcoI) and Pr_TcTAL(EcoRI) and inserted into pCDFD-Trc, which resulted in pCDFD-Trc-TcTAL. 4CL was cloned from pUC57-4CL (GenScript, Nanjing, China) with primers Pf_4CL (NcoI) and Pr_4CL (HindIII) and inserted into NcoI/HindIII site of pCDFD-Trc, resulted in pCDFD-Trc-4CL. pTrc-4CL region including Trc promoter and 4CL was cloned with primers Pf_Ptrc4CL(EcoRI) and Pr_Ptrc4CL(HindIII) and inserted into EcoRI/HindIII site of pCDFD-Trc-TcTAL. This resulted in pCDFD-Trc-TcTAL-Trc-4CL.
Different TcTAL variants were cloned from pCDFD-Trc-TcTAL-Trc-4CL with primers Pf_RNA(N) and Pr_RNA(XhoI) and inserted into NdeI/XhoI site of pET-22b(+), resulted in pET22b-TcTAL(N) (N = −6.2, −5.8, −5.2, −4.9, −4.6, −4.3, −3.7, −3.0). ‘N’ means free energy value of 5′ region of mRNA. ΔG of original TcTAL was −6.2 kcal/mol. QuickChange II XL Site-Directed Mutagenesis Kit (Agilent Technologies, Santa Clara, CA) was used to construct pCDFD-Trc-TcTAL(N)-Trc-4CL with primers Pf_TAL(N) and Pr_TAL(N) (N = −5.8, −5.2, −4.9, −4.6, −4.3, −3.7, −3.0). Plasmids constructed in the study were shown in Table 3.
Culture conditions
To investigate the silencing effect of various CRISPRi system, cells were first cultured in 25 mL of MOPS medium supplemented with 5 g/L d-glucose and 4 g/L NH4Cl at a starting OD600 of 0.1 (37 °C, 200 rpm). Additional 25 mL of MOPS medium and 0.5 mM IPTG were added after OD600 reached 1.7. Cultures were then transferred at 30 °C. The intracellular concentration of malonyl-CoA and target mRNA level was quantified by harvesting 1 mL of cell culture at mid-log phase of growth. Values of final OD600 were measured after 48 h.
For resveratrol production, cells were first cultured in 25 mL of MOPS medium supplemented with 5 g/L d-glucose and 4 g/L NH4Cl at a starting OD600 of 0.1 (37 °C, 200 rpm). Additional 25 mL of MOPS medium and 1 mM IPTG were added after OD600 reached 1.7. Cultures were then transferred at 30 °C. The resveratrol concentration and final OD600 values were measured after a total fermentation time of 48 h.
RNA preparation and qPCR
Recombinant cells were collected at the mid-log phase of growth by centrifugation and frozen in liquid nitrogen immediately. Total RNA was purified by RNAprep Pure Kit for Cell/Bacteria (TIANGEN, Beijing, China) according to the manufacturer’s instructions and DNase I and RNeasy Mini Kit (Takara, Dalian, China) was employed to remove genomic DNA. RNA quality and quantification were analyzed by NanoDrop ND-1000 Spectrophotometer (NanoDrop Technologies, Wilmington, DE, USA) [20].
cDNAs were synthesized via SuperScriptTM III First-Strand Synthesis System (Invitrogen). Primers Pf_qfabD/Pr_qfabD, Pf_qfabH/Pr_qfabH, Pf_qfabB/Pr_qfabB, Pf_qfabF/Pr_qfabF, Pf_qfabI/Pr_qfabI were used to amplify fabD, fabH, fabB, fabF and fabI with cDNA as templates. Primer 3 software (http://bioinfo.ut.ee/primer3-0.4.0/) was used to design primers, which were further synthesized by GenScript (Nanjing, China). The specificity of designed primers was confirmed by running genomic DNA-amplified products in agarose gel. SYBR green PCR Master Mix (Takara, Dalian, China) and LightCycler 480 II thermal cycler system (Roche, Mannheim, Germany) were used to perform real-time RT-PCR assay [8]. The efficiencies of real-time PCR were verified by five dilutions of cDNA and the purity of samples was confirmed by a negative control without cDNA added. The threshold cycle (CT) values were calculated by LightCycler software based on fluorescent values. Housekeeping rrsD gene was used to normalize qPCR values along with primers Pf_16S and Pr_16S [21]. The comparative \(2^{{ - \Delta \Delta C_{\text{t}} }}\) method was used to calculate relative gene expression [19]. All experiments were conducted in triplicate and mean values were employed for analysis.
Analytical methods
For each experiment, individual triplicate analyses were performed and average values of the triplicates were presented. For quantifying resveratrol concentration, culture supernatant was processed according to the previous report [28] along with Agilent 1100 series high-performance liquid chromatography (HPLC) instrument and a reverse-phase Gemini NX-C18 column (5 × 110 mm). For quantifying malonyl-CoA concentration, cell culture was processed according to the previous report [27] along with liquid chromatography–mass spectrophotometer (Shimadzu, Kyoto, Japan), a reverse-phase Gemini NX-C18 column (5 × 110 mm) and an electrospray ionization (ESI) source.
Results
Construction of de novo resveratrol synthetic pathway
To obtain resveratrol from d-glucose, we first need to construct strains exhibiting enhanced capacity for l-tyrosine synthesis. In E. coli, both the 3-deoxy-d-arabinoheptulosonate 7-phosphate (DAHP) synthase and chorismate mutase/prephenate (CM/PDH) dehydrogenase are bottleneck enzymes [25, 31]. Based on known properties of the aromatic amino acid pathway, the feedback-resistant derivatives of 3-deoxy-d-arabinoheptulosonate-7-phosphate (DAHP) synthase (aroG fbr) [11] and chorismate mutase/prephenate dehydrogenase (tyrA fbr) [12] were overexpressed to increase flux toward l-tyrosine.
Previously, our studies demonstrate that during the synthesis of other natural products such as naringenin [31] and pinocembrin [30] derived from the phenylpropanoid pathway, l-tyrosine or l-phenylalanine is not the limiting precursor. Here, in order to decrease the metabolic burden related to antibiotic cassettes and plasmid-based expression, aroG fbr and tyrA fbr were integrated into lacZ locus of E. coli BL21 (DE3) under T7 promoter.
In previous work [28], production of resveratrol from l-tyrosine in E. coli has been optimized. Based on this, Rhodotorula glutinis TAL (RgTAL) and Petroselinum crispum 4CL under Trc promoter in pCDFDuet-1(pCDF-Trc-RgTAL-Trc-4CL), STS from Vitis vinifera under T7 promoter in pETDuet-1(pET-STS) were overexpressed in an engineered BL21(DE3) strain integrated with aroG fbr and tyrA fbr. It was found that the strain was able to produce resveratrol up to 18.5 mg/L directly from d-glucose.
Engineering malonyl-CoA availability
The supply of malonyl-CoA is often considered as the major bottleneck of the phenylpropanoid pathway. Previous efforts to improve malonyl-CoA availability mainly focus on either the upstream pathway [13, 14, 32] or malonyl-CoA consumption pathway [34]. Here, a combining bioengineering strategy involving manipulating both precursor supply and malonyl-CoA consumption was established to increase intracellular malonyl-CoA level. matB (encoding malonyl-CoA synthetase) and matC (encoding malonate carrier protein) [14, 28] were overexpressed to synthesize malonyl-CoA directly from malonate avoiding manipulating complex central metabolism. The resveratrol titer increased to 31.2 mg/L after introducing this recombinant malonate assimilation pathway.
Fatty acid synthesis is the only process in E. coli which consumes malonyl-CoA [3]. Previously, only fabB and fabF involved in fatty acid biosynthesis were repressed as final target genes by CRISPRi system in combination with other genetic modification in upstream pathway to improve (2S)-naringenin titer [27]. Here, in order to reduce malonyl-CoA consumption to a large extent, fabD, fabH, fabB, fabF and fabI were chosen as target genes as repression of these genes resulted in an increase of intracellular malonyl-CoA level [27]. It was found that repression of fabD, fabH, fabB, fabF and fabI increased resveratrol titer by 80.2, 195.6, 170.3, 216.5 and 123.7%, respectively (Fig. 2a).
The effect of single gene repression toward fatty acid pathway on resveratrol production. a Effect of single genetic perturbation on resveratrol production was investigated. Control mean strains containing resveratrol synthetic pathway without CRISPRi system. The sgRNA-expressing plasmids repressing different genes were then transformed into the control strain. b The changing patterns of intracellular malonyl-CoA concentration and target gene mRNA level from engineered strains were measured. Control mean wild-type strains. The sgRNA-expressing plasmids repressing different genes were then transformed into the control strain. For each experiment, biological triplicate analyses were conducted and the average values were presented. Error bars exhibited SD with 95% CI
To verify the efficiency of this CRISPRi system, the changing pattern of intracellular malonyl-CoA concentration and target gene mRNA level from CRISPRi-regulated strains were measured compared to the control. As seen from Fig. 2b, the transcriptional levels of fabD, fabH, fabB, fabF and fabI decreased by 81.9, 90.5, 91.2, 92.1 and 90.1%, respectively. Repressing fabD, fabH, fabB, fabF and fabI increased the intracellular malonyl-CoA concentration by 130.6, 260.8, 239.6, 290.5 and 198.7%, respectively.
Effect of combinatorial genetic perturbations on resveratrol production
As seen from Fig. 2a, it was found that repressing all the genes except for fabF would lead to a dramatic decrease in the final biomass (by over 50%). This is consistent with our previous study that repressing fabH and fabI even with low efficacy would result in a dramatic decrease in the cell growth. Numerous studies have demonstrated that coordination of cell growth and heterologous molecules production is important for strain performance [27, 29, 34]. Here, in order to achieve lower repressing efficacy, the high or medium repressing efficacy toward each target gene was re-designed to bind at the initial and terminal region of the non-template DNA strand while the low repressing efficacy set at the initial region of template strand (Fig. 3a).
The effect of different gene repressing efficacy on cell growth and malonyl-CoA accumulation. a The binding regions of different sgRNAs were shown. High, medium and low mean high, medium and low repressing efficacy. b Target genes were repressed with high silencing efficacy. c Target genes were repressed with medium silencing efficacy. d Target genes were repressed with low silencing efficacy. Control mean wild-type strain. The sgRNA-expressing plasmids repressing different genes were then transformed into the control strain. For each experiment, biological triplicate analyses were conducted and the average values were presented. Error bars exhibited SD with 95% CI
As seen from Fig. 3, high repressing efficacy toward fabF, medium repressing efficacy toward fabB and fabD and low repressing efficacy toward fabH and fabI increased intracellular malonyl-CoA by 290.5, 141.2, 70.1, 122.4 and 98.2%, respectively, with the final OD600 decreasing by less than 10%. Based on this, the effect of combinatorial genetic perturbations on resveratrol production was investigated (Fig. 4). Firstly, the effect of two genetic perturbations on resveratrol production was explored. Repressing fabF (anti-fabF sgRNA) in combination with fabB, fabD, fabH and fabI produced higher resveratrol titer than single anti-fabF sgRNA and repressing fabF and fabB led to the highest resveratrol titer (138.3 mg/L).
The effect of multiple genetic perturbations on resveratrol production. Control strains contained resveratrol heterologous pathway without CRISPRi system. The sgRNA-expressing plasmids repressing single or multiple genes were further transformed into the control strain to investigate the effect of these systems on resveratrol production. For each experiment, biological triplicate analyses were conducted and the average values were presented. Error bars exhibited SD with 95% CI
Similarly, repressing fabF and fabB (anti-fabF/fabI sgRNA) in combination with fabI, fabD and fabH produced higher resveratrol titer than anti-fabF/fabB sgRNA and repressing fabF, fabI and fabB led to the highest resveratrol titer (169.8 mg/L). Furthermore, it was found that repressing fabF, fabI and fabB (anti-fabF/fabI/fabB sgRNA) in combination with fabH and fabD produced higher resveratrol titer and repressing fabF, fabI, fabB and fabD led to the highest resveratrol titer (187.1 mg/L). However, repressing fabF, fabI, fabB, fabD and fabH produced lower resveratrol titer than the former one.
Rational engineering of tyrosine ammonia lyase by reducing mRNA secondary structure in its 5′ region
Numerous studies have demonstrated that tyrosine ammonia lyase (TAL) suffers from low turnover number [17, 24]. To overcome this limitation, firstly, we resorted to replace RgTAL with another TAL from Trichosporon cutaneum (TcTAL), which exhibited high activities toward l-tyrosine [10]. It was found that this replacement resulted in higher resveratrol production (213.2 mg/L). To further alleviate this bottleneck, the expression of the heterologous gene TcTAL was improved by modifying its 5′ region of mRNA secondary structure, as numerous studies demonstrate that local RNA structure between −4 and +38 of the translation start is highly consistent with expression change [7, 30].
Here, changing synonymous codon usage in the first amino acids of TcTAL was performed to optimize protein expression. For each codon, random mutations in the third base position were introduced with the first and second positions keeping constant. The minimum free energy of folding for 5′ region of mRNA transcript of each construct (ΔG) was calculated by NUPACK software [35] based on the sequence from +1 to +42 (Fig. 5a, b). A total of seven variants spanning a range of free energies from −6.2 to −3.0 kcal/mol were selected. Firstly, different variants were overexpressed by pET-22b(+) to test their activities. It was found that variants with reduced mRNA secondary structure of 5′ region displayed enhanced capacities for p-coumaric acid synthesis (Fig. 5c). These different TAL variants were further used to replace the original TAL in the production strains. As seen from Fig. 5d, the resveratrol production increased with larger value of ΔG and a final titer of 304.5 mg/L was observed. Furthermore, it was found that this engineered strain exhibited a yield of 0.075 g/g (0.075 g resveratrol per g total glucose consumed).
The effect of reducing mRNA secondary structure of TcTAL 5′ region on resveratrol production. a Predicted mRNA secondary structures of 5′ region of different variant TcTAL. Numbers such as −6.2, −5.8 denoted the minimum free energy of folding (ΔG) for the 5′ region of mRNA secondary structure. ΔG of original TcTAL was −6.2 kcal/mol. b The alignment of different TcTAL variant sequences (+1 to +42). c The effect of different TcTAL variants on p-coumaric acid concentrations. Engineered strain cultures were supplemented with 500 mg/L l-tyrosine. Values were reported after 48 h cultivation in MOPS minimal medium. d The effect of different TcTAL variants on resveratrol production. For each experiment, biological triplicate analyses were conducted and the average values were presented. Error bars exhibited SD with 95% CI
Discussion
The conventional processes of extracting resveratrol from plant natural resources are expected to be revolutionized by microbial bioconversion and fermentation [5]. However, broad adoption of microbial resveratrol synthesis is hampered by its costly and inefficient process, typically the requirement of supplementing expensive phenylpropanoic precursors in the media [15, 18]. Here, efficient de novo synthesis of resveratrol by metabolically engineered Escherichia coli was demonstrated. Compared to related studies to produce resveratrol from cheap and renewable substrates, such as d-glucose (4.6 mg/L) [18] or even l-tyrosine (35 mg/L) [28], this fermentation platform described here exhibited a dramatic increase in de novo resveratrol production (304.5 mg/L) (Fig. 5). These improvements would expedite developing an economical and simple process for microbial resveratrol production.
The limiting supply of malonyl-CoA is often considered as the major bottleneck of the phenylpropanoid pathway [16, 34]. Here, an efficient malonyl-CoA synthesis method involving manipulating both precursor supply and malonyl-CoA consumption was established instead of engineering either the upstream pathway or the malonyl-CoA consumption pathway. A recombinant malonate assimilation pathway [14] was developed to synthesize malonyl-CoA directly from malonate to avoid manipulating complex central metabolism to obtain malonyl-CoA from acetyl-CoA. Simultaneously, the CRISPRi system was used to repress five genes involved in fatty acid pathway, which is the only process consuming malonyl-CoA. This significantly improved the intracellular malonyl-CoA concentration and final target product resveratrol titer increased by tenfold compared to the initial strain (Fig. 4). This method demonstrated here could be employed for efficient production of many other malonyl-CoA derived compounds.
In another relevant study, antisense RNA strategy was used to investigate the effect of single fatty acid pathway gene perturbation on malonyl-CoA accumulation. However, it was found that interference of two genes would not further improve the final target product titer [34]. In our one study, the clustered regularly interspaced short palindromic repeats interference (CRISPRi) system was developed to improve intracellular malonyl-CoA level and only fabB and fabF involved in fatty acid biosynthesis were chosen as final target genes [27]. Here, a more systematic study on the effect of repressing fatty acid pathway on malonyl-CoA accumulation was conducted. It was found that repressing fabD, fabH, fabB, fabF, fabI increased resveratrol production (Fig. 2). Furthermore, by coupling genetic modification to cell growth, the combined effect of these genetic perturbations was additive (Fig. 4). This further demonstrates that CRISPRi is a promising genetic tool for use in metabolic engineering.
As demonstrated in numerous studies, the low turnover number of TAL is often considered as another major bottleneck during de novo resveratrol production [17, 28]. Here, in order to overcome this limitation, firstly, the original RgTAL was replaced by TcTAL [10], which led to higher resveratrol production. Furthermore, the expression of the heterologous gene TcTAL was improved by modifying its mRNA secondary structure of 5′ region and variants with reduced mRNA secondary structure displayed enhanced capacities for resveratrol synthesis (Fig. 5). This finally improved resveratrol titer by 61.7%. Although high titer of de novo production of resveratrol has been achieved in E. coli here, this titer is not higher than other reported host such as Saccharomyces cerevisiae [15]. However, it is presumed that the titer of this engineered strain could be enhanced if it is successfully cultured in a fermentor. Overall, this work paves the way to developing a simple and economical process for microbial production of value-added compounds.
References
Baur JA, Sinclair DA (2006) Therapeutic potential of resveratrol: the in vivo evidence. Nat Rev Drug Discov 5:493–506
Becker J, Wittmann C (2016) Systems metabolic engineering of Escherichia coli for the heterologous production of high value molecules—a veteran at new shores. Curr Opin Biotechnol 42:178–188. doi:10.1016/j.copbio.2016.05.004
Cao WJ, Ma WC, Zhang BW, Wang X, Chen KQ, Li Y, Ouyang PK (2016) Improved pinocembrin production in Escherichia coli by engineering fatty acid synthesis. J Ind Microbiol Biotechnol 43:557–566. doi:10.1007/s10295-015-1725-3
Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci 97:6640–6645. doi:10.1073/pnas.120163297
Dziggel C, Schäfer H, Wink M (2017) Tools of pathway reconstruction and production of economically relevant plant secondary metabolites in recombinant microorganisms. Biotechnol J. doi:10.1002/biot.201600145
Ge H, Zhang J, Guo B, He Q, Wang B, He B, Wang C (2007) Resveratrol inhibits macrophage expression of EMMPRIN by activating PPARγ. Vasc Pharmacol 46:114–121
Goodman DB, Church GM, Kosuri S (2013) Causes and effects of N-terminal codon bias in bacterial genes. Science 342:475–479
Guo HW, Madzak C, Du GC, Zhou JW, Chen J (2014) Effects of pyruvate dehydrogenase subunits overexpression on the alpha-ketoglutarate production in Yarrowia lipolytica WSH-Z06. Appl Microbiol Biotechnol 98:7003–7012. doi:10.1007/s00253-014-5745-0
Huang XL, Dai YQ, Cai JX, Zhong NJ, Xiao H, McClements DJ, Hu K (2017) Resveratrol encapsulation in core-shell biopolymer nanoparticles: impact on antioxidant and anticancer activities. Food Hydrocolloids 64:157–165. doi:10.1016/j.foodhyd.2016.10.029
Jendresen CB, Stahlhut SG, Li MJ, Gaspar P, Siedler S, Forster J, Maury J, Borodina I, Nielsen AT (2015) Highly active and specific tyrosine ammonia-lyases from diverse origins enable enhanced production of aromatic compounds in bacteria and Saccharomyces cerevisiae. Appl Environ Microbiol 81:4458–4476. doi:10.1128/aem.00405-15
Kikuchi Y, Tsujimoto K, Kurahashi O (1997) Mutational analysis of the feedback sites of phenylalanine-sensitive 3-deoxy-d-arabino-heptulosonate-7-phosphate synthase of Escherichia coli. Appl Environ Microbiol 63:761–762
Lütke-Eversloh T, Stephanopoulos G (2005) Feedback inhibition of chorismate mutase/prephenate dehydrogenase (TyrA) of Escherichia coli: generation and characterization of tyrosine-insensitive mutants. Appl Environ Microbiol 71:7224–7228. doi:10.1128/AEM.71.11.7224-7228.2005
Leonard E, Lim KH, Saw PN, Koffas MAG (2007) Engineering central metabolic pathways for high-level flavonoid production in Escherichia coli. Appl Environ Microbiol 73:3877–3886. doi:10.1128/AEM.00200-07
Leonard E, Yan Y, Fowler ZL, Li Z, Lim CG, Lim KH, Koffas MAG (2008) Strain improvement of recombinant Escherichia coli for efficient production of plant flavonoids. Mol Pharm 5:257–265. doi:10.1021/mp7001472
Li MJ, Kildegaard KR, Chen Y, Rodriguez A, Borodina I, Nielsen J (2015) De novo production of resveratrol from glucose or ethanol by engineered Saccharomyces cerevisiae. Metab Eng 32:1–11. doi:10.1016/j.ymben.2015.08.007
Liang JL, Guo LQ, Lin JF, He ZQ, Cai FJ, Chen JF (2016) A novel process for obtaining pinosylvin using combinatorial bioengineering in Escherichia coli. World J Microbiol Biotechnol 32:1–10. doi:10.1007/s11274-016-2062-z
Lim CG, Fowler ZL, Hueller T, Schaffer S, Koffas MAG (2011) High-yield resveratrol production in engineered Escherichia coli. Appl Environ Microbiol 77:3451–3460. doi:10.1128/AEM.02186-10
Liu XL, Lin J, Hu HF, Zhou B, Zhu BQ (2016) De novo biosynthesis of resveratrol by site-specific integration of heterologous genes in Escherichia coli. FEMS Microbiol Lett 363:5–18. doi:10.1093/femsle/fnw061
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(T)(−Delta Delta C) method. Methods 25:402–408. doi:10.1006/meth.2001.1262
Mansour S, Bailly J, Delettre J, Bonnarme P (2009) A proteomic and transcriptomic view of amino acids catabolism in the yeast Yarrowia lipolytica. Proteomics 9:4714–4725. doi:10.1002/pmic.200900161
Nakashima N, Tamura T (2009) Conditional gene silencing of multiple genes with antisense RNAs and generation of a mutator strain of Escherichia coli. Nucleic Acids Res 37:e103
Neidhardt FC, Bloch PL, Smith DF (1974) Culture medium for enterobacteria. J Bacteriol 119:736–747
Qi LS, Larson MH, Gilbert LA, Doudna JA, Weissman JS, Arkin AP, Lim WA (2013) Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152:1173–1183
Santos CNS, Koffas MAG, Stephanopoulos G (2011) Optimization of a heterologous pathway for the production of flavonoids from glucose. Metab Eng 13:392–400. doi:10.1016/j.ymben.2011.02.002
Santos CNS, Xiao W, Stephanopoulos G (2012) Rational, combinatorial, and genomic approaches for engineering l-tyrosine production in Escherichia coli. Proc Natl Acad Sci 109:13538–13543. doi:10.1073/pnas.1206346109
van Summeren-Wesenhagen PV, Marienhagen J (2015) Metabolic engineering of Escherichia coli for the synthesis of the plant polyphenol pinosylvin. Appl Environ Microbiol 81:840–849. doi:10.1128/aem.02966-14
Wu JJ, Du GC, Chen J, Zhou JW (2015) Enhancing flavonoid production by systematically tuning the central metabolic pathways based on a CRISPR interference system in Escherichia coli. Sci Rep 5:14. doi:10.1038/srep13477
Wu JJ, Liu PR, Fan YM, Bao H, Du GC, Zhou JW, Chen J (2013) Multivariate modular metabolic engineering of Escherichia coli to produce resveratrol from l-tyrosine. J Biotechnol 167:404–411. doi:10.1016/j.jbiotec.2013.07.030
Wu JJ, Yu O, Du GC, Zhou JW, Chen J (2014) Fine-tuning of the fatty acid pathway by synthetic antisense RNA for enhanced (2S)-naringenin production from L-tyrosine in Escherichia coli. Appl Environ Microbiol 80:7283–7292
Wu JJ, Zhang X, Dong MS, Zhou JW (2016) Stepwise modular pathway engineering of Escherichia coli for efficient one-step production of (2S)-pinocembrin. J Biotechnol 231:183–192. doi:10.1016/j.jbiotec.2016.06.007
Wu JJ, Zhou TT, Du GC, Zhou JW, Chen J (2014) Modular optimization of heterologous pathways for de novo synthesis of (2S)-naringenin in Escherichia coli. PLoS One 9:e101492
Xu P, Ranganathan S, Fowler ZL, Maranas CD, Koffas MAG (2011) Genome-scale metabolic network modeling results in minimal interventions that cooperatively force carbon flux towards malonyl-CoA. Metab Eng 13:578–587. doi:10.1016/j.ymben.2011.06.008
Yang JE, Kim JW, Oh YH, Choi SY, Lee H, Park AR, Shin J, Park SJ, Lee SY (2016) Biosynthesis of poly(2-hydroxyisovalerate-co-lactate) by metabolically engineered Escherichia coli. Biotechnol J 11:1572–1585. doi:10.1002/biot.201600420
Yang YP, Lin YH, Li LY, Linhardt RJ, Yan YJ (2015) Regulating malonyl-CoA metabolism via synthetic antisense RNAs for enhanced biosynthesis of natural products. Metab Eng 29:217–226. doi:10.1016/j.ymben.2015.03.018
Zadeh JN, Steenberg CD, Bois JS, Wolfe BR, Pierce MB, Khan AR, Dirks RM, Pierce NA (2011) NUPACK: analysis and design of nucleic acid systems. J Comput Chem 32:170–173. doi:10.1002/jcc.21596
Acknowledgements
This work was financially supported by the National Natural Science Foundation of China (No. 31601448), Natural Science Foundation of Jiangsu Province (BK20160718), National Natural Science Foundation of China (No. 31601448), the Fundamental Research Funds for the Central Universities (KYZ201654 and KJQN201728) and the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wu, J., Zhou, P., Zhang, X. et al. Efficient de novo synthesis of resveratrol by metabolically engineered Escherichia coli . J Ind Microbiol Biotechnol 44, 1083–1095 (2017). https://doi.org/10.1007/s10295-017-1937-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-017-1937-9