Abstract
Pinosylvin as a bioactive stilbene is of great interest for food supplements and pharmaceuticals development. In comparison to conventional extraction of pinosylvin from plant sources, biosynthesis engineering of microbial cell factories is a sustainable and flexible alternative method. Current synthetic strategies often require expensive phenylpropanoic precursor and inducer, which are not available for large-scale fermentation process. In this study, three bioengineering strategies were described to the development of a simple and economical process for pinosylvin biosynthesis in Escherichia coli. Firstly, we evaluated different construct environments to give a highly efficient constitutive system for enzymes of pinosylvin pathway expression: 4-coumarate: coenzyme A ligase (4CL) and stilbene synthase (STS). Secondly, malonyl coenzyme A (malonyl-CoA) is a key precursor of pinosylvin bioproduction and at low level in E. coli cell. Thus clustered regularly interspaced short palindromic repeats interference (CRISPRi) was explored to inactivate malonyl-CoA consumption pathway to increase its availability. The resulting pinosylvin content in engineered E. coli was obtained a 1.9-fold increase depending on the repression of fabD (encoding malonyl-CoA-ACP transacylase) gene. Eventually, a phenylalanine over-producing E. coli consisting phenylalanine ammonia lyase was introduced to produce the precursor of pinosylvin, trans-cinnamic acid, the crude extraction of cultural medium was used as supplementation for pinosylvin bioproduction. Using these combinatorial processes, 47.49 mg/L pinosylvin was produced from glycerol.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
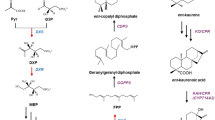
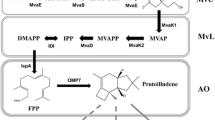
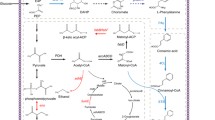
Pinosylvin is a natural secondary metabolite that is biosynthesized through a general phenylalanine metabolism pathway in genus Pinus (Rivière et al. 2012). Recent studies have exhibited that pinosylvin possessed benefits for human health, including anti-antioxidant, anti-cancer, cardioprotection, anti-inflammation, and other activities (Jancinova et al. 2012; Park et al. 2012; Park et al. 2013; Yeo et al. 2013; Koskela et al. 2014; Laavola et al. 2015). However, pinosylvin is hard to vast purification via phytoextraction because of low concentrations in plants and various isomers or similar compounds were need to separation (Conde et al. 2014). Biosynthesis provides an alternative approach to pinosylvin production. The engineering strategy for the biosynthetic pathway of pinosylvin in E. coli cell is shown in Fig. 1a. This strategy involves embedding phenylalanine ammonia lyase (PAL) to convert endogenous/extrinsic l-phenylalanine into trans-cinnamic acid. The trans-cinnamic acid is then converted into trans-cinnamoyl-CoA, through 4-coumarate: CoA ligase (4CL). Stilbene synthase (STS) catalyzes the stepwise condensation of three molecules of intracellular malony-CoA with one molecule of trans-cinnamoyl-CoA.
Engineered pathways for pinosylvin synthesis by recombinant Escherichia coli. a Strategy in previous studys. PAL phenylalanine ammonia lyase; 4CL 4-coumarate: CoA ligase, STS stilbene synthase. b Strategy in this study. FABB β-ketoacyl-acyl carrier protein synthases I, FABD malonyl-CoA-ACP transacylase, FABF β-ketoacyl-acyl carrier protein synthases II
To data, microbial cell factories for pinosylvin bioproduction have been developed but efficient conversion of aromatic amino acids to pinosylvin is the most important limiting factor (Katsuyama et al. 2007; Wang et al. 2015). All of the study required inducer of gene expression and expensive chemicals as precursors, typically l-phenylalanine or trans-cinnamic acid in comparison to glycerol. Recently, a feasible strategy for increasing the production of pinosylvin was improving the content of intracellular malonyl-CoA by the addition of the fatty acid production inhibitor cerulenin, the titer of pinosylvin has 20-fold enhancement (van Summeren-Wesenhagen and Marienhagen 2015). However, cerulenin is an expensive inducer and cost prohibitive for a production scale fermentation process in terms of food safety. Therefore, another metabolic regulation strategy was investigated to engineered strains with improved levels of malonyl-CoA. A study was introducing matB (encoding malonyl-CoA synthetase) and matC (encoding dicarboxylate carrier protein) into E. coli to uptake the malonate and to convert it into malonyl-CoA (Leonard et al. 2008; Wu et al. 2013). In addition, the stilbene resveratrol production in E. coli showed a 1.6-fold increase through overexpression of pdh (encoding pyruvate dehydrogenase multi-enzyme complex), pgk (encoding phosphoglycerate kinase), and gapA (encoding glyceraldehyde-3-phosphate dehydrogenase) and deletion of fumC (encoding fumarase) (Bhan et al. 2013).Whereas the strain cannot bear the burden of multi-gene efficient expression in one or two plasmids. Sometimes, strategy of gene deletion is limited because genes associated with the main metabolic pathway cannot be knocked out. Regulating malonyl-CoA metabolism through synthetic antisense RNAs was reported, a 2.5-fold increase in stilbene resveratrol concentration when fabD gene expression is inhibited (Yang et al. 2015). The advantage of this strategy is that metabolic regulation would not influence the growth of strains.
A similar method with antisense RNAs system was Clustered regularly interspaced short palindromic repeats interference (CRISPRi), an emerging gene repression system, was applied on prokaryotic metabolic engineering in recent studies (Ji et al. 2014; Wu et al. 2014; Lv et al. 2015). Two functional modules have been introduced into E. coli for genome regulation without genome editing through flexible CRISPR systems (Qi et al. 2013). A catalytically dead Cas9 mutant (dCas9) acts as a simple RNA-guided DNA-binding complex. A small guide RNA (sgRNA) was coexpressed with target-specific complementary region, Cas9-binding hairpin, and transcription terminator. The silencing effects of dCas9-sgRNA can be induced and reversed as required by using CRISPRi-based knockdown of gene expression. And application of CRISPRi for improving stilbenes production has never been reported.
In this study, a combinatorial bioengineering approach was employed for the production of pinosylvin in E. coli directly from glycerol without the addition of expensive inducers and precursors (Fig. 1b). BioBrick assembly and strains optimization were used to increase production ability of pinosylvin in E. coli. Application of the CRISPRi system to regulation of gene expression related to prokaryotic fatty acid metabolism, the engineered E. coli strain with fabD interfered shows a 1.9-fold increase in pinosylvin concentration. For production of trans-cinnamic acid supplemented to the medium, an l-phenylalanine over-producing E. coli carrying the pal gene with constitutive system was employed. Combination of the optimized E. coli cells and trans-cinnamic acid extract at 37 °C for 12 in shake-flask culture yielded 47.49 mg/L pinosylvin.
Materials and methods
Materials and reagents
E. coli DH5α and other strains (listed in Table 1) were used for plasmid cloning and recombinant molecule production. PrimeSTAR® HS DNA Polymerase, ExTaq Polymerase, pMD18-T, restriction enzymes, T4 DNA ligase and Anhydrotetracycline were purchased from Takara Biochemicals Inc (Japan). Plasmid pQE-30 was purchased from Qiagen Inc (Germany). Plasmid pdCas9_bacteria and pgRNA_bacteria was purchased from Addgene Inc (USA). Plasmid DNA was prepared from stock strains by using a Tiangen plasmid miniprep kit (China), while fragment DNA was isolated through gel extraction by a Tiangen gel DNA recovery kit (China). Total RNA was isolated by using the Tiangen TRNzol Universal Reagent Kit (China). The Tiangen Fastquant RT Kit (China) was used to synthesize the cDNA for mRNA analysis. Real-time PCR (RT-PCR) was carried out for mRNA analysis with Tiangen SuperReal PreMix Plus (SYBR Green) (China). The trans-cinnamic acid and pinosylvin standards were purchased from Sigma-Aldrich (USA).
Genetic constructs
The constitutive gap promoter (the endogenic promoter of glyceraldehyde-3-phosphate dehydrogenase gene in E. coli) was cloned from pMD18-pGAP. The rrnB T1 terminator was cloned from pQE-30. The At4cl fragment (Accession code: NM_104046.2) was cloned from pMD18- At4cl. The Vvsts fragment (Accession code: KC417319.1) was cloned from pMD18-Vvsts. The total expression fragment (fusion of three or four DNA fragments) was amplified though overlap extension PCR. Different sequences of ribosome binding site (RBS) or additional nucleotide sequence were added by specific primers. The total sgRNA expression fragment (two DNA fragments fusion) was amplified by overlap extension PCR.
All constructed plasmids were verified by both colony PCR and sequencing. All primers, RBS sequences (Ribosome Binding Site), PAM sequences (Protospacer Adjacent Motif, sequence: NGG) for sgRNA design and additional sequences were listed in Supplementary data Table 1. All plasmids used in this study are listed in Table 1.
Preparation of trans-cinnamic acid extraction
The cells of E. coli ATCC31884 harboring pQE30 with Rgpal (Accession code: KF770992) was pre-cultured overnight at 37 °C in 10 mL of LB medium along with 100 μg/mL ampicillin antibiotics. Pre-inoculums were diluted into 100 mL of fresh YM9 medium (1xM9 salts, 10 g/L yeast extract, 5 % glycerol, and 42 g/L Morpholinepropanesulfonic acid, pH 7.0) with 100 μg/mL ampicillin antibiotics according to the 1 % volume fraction of bacteria liquid. Shake-flask culture was performed at 200 rpm and 37 °C. Sample was centrifuged at 12,000 rpm (micro liter rotor size: 24 × 2 mL) for 5 min, and the supernatants were extracted with twice volume of ethyl acetate after 12 h of culture. The organic phase was volatilized by rotary evaporator at 40 °C. The sediments were dissolved in 2 mL Tris–HCl (1 mM, pH 9.5) for preparation of pinosylvin synthesis.
Pinosylvin bioproduction through engineered E. coli
The culture medium consisted of YM9 medium along with relevant antibiotics. Recombinant E. coli cells containing different expression vector were pre-cultured overnight at 37 °C in 100 mL LB medium. Pre-inoculums were collected by centrifugation and resuspended into 100 mL of fresh YM9 medium. For dCAS9 expression, 1 μM of aTc (Anhydrotetracycline) was supplemented at this time. Shake flask fermentation were performed at 200 rpm, 37 °C additionally supplemented with trans-cinnamic acid extraction. Samples were taken every few hours and used for the analysis of the cell growth, product accumulation and gene expression.
Expression analysis by real-time PCR
The gene expression levels of fabB, fabD, fabF were analyzed using a Bio-Rad CFX96 real-time PCR detection system (USA). 16S rRNA (Accession code: J01859.1) as the inner standard. Primers were designed from the full-length cDNA sequences (listed in Supplementary data Table 1). Real-time PCR was performed in a 25 μL reaction volume with 0.3 μM of each primer. PCR protocols were as follows: 1 cycle of 15 min at 95 °C; 40 cycles with a denaturing time of 10 s at 94 °C, an annealing time of 20 s at 60 °C, and an elongation time of 20 s at 72 °C. All samples were prepared with three parallel groups to obtain results of ΔCt values from the outputs of RT-PCR.
HPLC analysis of pinosylvin and trans-cinnamic acid
All samples were extracted with twice volume of ethyl acetate. The mixture was centrifuged at 12,000 rpm (micro liter rotor size: 24 × 2 mL) for 5 min and the supernatants were analyzed on an HPLC system with a phenomenex ODS reverse phase C-18 column (5 μm, 250 × 4.6 mm) and LC2030 UV Detector. The solvent profile was acetonitrile (buffer A) and water (buffer B) as the mobile phases with a flow rate of 1.0 mL per minute. The injection volume was 20 μL. For pinosylvin detection, samples were separated with the mobile phase composition profile fixed at 40 % buffer A and 60 % buffer B. The characteristic UV absorption spectrum was 294 nm. The metabolites were confirmed by the retention time compared to those of authentic standards which retention time was 12.32 min. trans-cinnamic acid was detected under the mobile phase composition profile fixed at 40 % buffer A and 60 % buffer B (0.1 % acetic acid). The characteristic UV absorption spectrum was 272 nm and the retention time was 5.56 min.
Results
Identifying the best constitutive BioBrick combination for pinosylvin bioproduction
RBS is an effective control element for translation and thereby protein expression in E. coli (Qi et al. 2012). The replication region determines the copy number of vector as well as the expression level of heterogeneous genes. The high-copy plasmid does not always have maximum gene expression because of metabolic burden effects (Jones et al. 2000). The BioBrick assembly contains a high-copy number plasmid background with a constitutive gap promoter operon, and was suitable for stilbene resveratrol pathway assembly in E. coli BW27784 (Lim et al. 2011). In this study, we used pGAP, Vvsts, and At4cl to reconstruct different constitutive plasmids to further study BioBricks combinatorial optimization.
To determine the best RBS and replication region assembly, four plasmids with different RBS and replication regions (Fig. 2a) were transferred into E. coli BW27784 for shake flask cultivation supplemented with 0.5 mM trans-cinnamic acid. When analyzing the supernatant of fermentation broth by HPLC, a peak was detected at the same retention time of pinosylvin and trans-cinnamic acid standard (Supporting Information Fig. 1). With the same RBS, the production with moderate copy number plasmid was approximately sixfold greater than that with the high copy number plasmid (comparison between strains P1 and P2) because the high copy number of vector was not suitable for protein functional folding and plasmid stability (Jones et al. 2000). The production with RBS3 plasmid was approximately 1.16-fold greater than that with the RBS1 plasmid (comparison between strain P4 and P2 at the same replication region) because the strong RBS of pET system took advantage of ribosome binding and transcriptional initiation (Holleley and Geerts 2009). However, no product was detected in strain P3 with RBS2 sequence from reference construct RL027A (Lentini et al. 2013). Therefore, different BioBrick compositions directly influenced the titers of metabolic production. RBS3 with moderate-copy number constructs was used for subsequent experiments.
Pinosylvin bioproduction in E. coli containing different constitutive biobrick combination. a The effect of configuration of four plasmids with different RBS and replication region. Data are plotted relative to strain P1. b The effect of configuration of four plasmids with different polycistron transcriptional assembly. Data are plotted relative to strain P4
We then observed pinosylvin production from different configurations of polycistronic transcription assembly. The influence of upstream sequence composition of RBS on production levels is shown in Fig. 2b. The constructs with no spacer (strain P4) and with a NcoI (8 bp) spacer (strain P5) extra restriction enzyme cutting site both resulted in higher relative production titers when compared with the constructs with a 31 bp spacer (strain P6), which was the optimal sequence between two continuous heterogeneous gene expressions from the reference construct RL027A (Lentini et al. 2013). We aimed to answer the following question: which combination of two gene fragments was better for bioproduction. Through the comparison of the strains P4, P7, and P8, the highest pinosylvin production was obtained in E. coli BW27784 (strain P7) with the double-promoter construct in opposite expression direction. These results were similar to the exploration of optimal plasmid configuration (Huang et al. 2013).
Determining the best strain for pinosylvin biosynthesis
We examined the plasmid pA7 in seven general lab strains (Table 1) and strain-to-strain variations in pinosylvin titer (Fig. 3) to determine the ideal strain for pinosylvin production. The maximum pinosylvin production was approximately threefold higher than the minimum. E. coli Dh5α with pA7 produced the highest yield of pinosylvin among the strains. The pinosylvin biosynthetic capacity differed among strains. E. coli Dh5α cells displayed preferable metabolic capability and were used for subsequent experiments.
Effect of gene repression with CRISPRi on pinosylvin production
In the fatty acid biosynthesis, the malonyl-CoA: ACP transacylase encoded by fabD was a key gene in malonyl-CoA consumption, and the strain growth was limited without it (Janßen and Steinbüchel 2014). The fabB and fabF (encoding β-ketoacyl-acyl carrier protein synthases [KAS] I and II) gene products were the suppressive target of cerulenin (Leonard et al. 2008; Lim et al. 2011). Therefore we selected these E. coli chromosomal genes (fabB, fabD and fabF) as target genes to demonstrate the effectiveness of CRISPRi on the repression of fatty acid biosynthesis and enrichment of pinosylvin production. An engineering strain co-expression of the mutated dCas9 protein and different sgRNA Biobrick was constructed (Fig. 4a). Whole sgRNA Biobrick contains four parts: a 36 bp constitutive promoter J_23119, a 43 bp hairpin region for dCas9 protein binding, a 38 bp tracrRNA terminator and a 20 bp DNA based-pairing region, different gene has different pairing region. In addition, the dCAS9 protein encoded by another plasmid pdCas9_bacteria was conveniently induced by 1 μM aTc.
Metabolic regulation of E. coli for pinosylvin bioproduction using CRIPSPi strategy. a The plasmids used for dCAS9 expression and sgRNA-based repression. Process of the CRISPRi system to block transcription initiation. b Gene expression in recombinant E. coli. C0, Control; C1, anti-fabB; C2, anti-fabD; C3, anti-fabF. c The effect of configuration of three plasmids with different targeted-sgRNA assembly. Data are plotted relative to strain C0
Real-time PCR results indicated that the CRISPRi system functioned at the mRNA level to regulate reduction of fabB, fabD and fabF gene expression (Fig. 4b). The sgRNA anti-fabB, anti-fabD and anti-fabF were able to bind with their respective targets with different efficiencies ranging from 79.9 to 99.5 % repression. Thus, the CRISPRi system was successfully established as a technique for bacterial fatty acid biosynthesis regulation. Compared with the control group strain C0 with non-target sgRNA construct, repressing fabB resulted in a 1.5-fold increase in pinosylvin production (Fig. 4c). The highest increase of pinosylvin yield in strain C2 with the anti-fabD construct was 1.9-fold that of the control group strain C0. However, for unknown reason, anti-fabF in the strain C3 was no obvious effect on production improvement.
Next we have observed the cell dynamics during pinosylvin bioproduction compared with strain C0 (without sgRNA) and strain C2. Obviously, the biomass of the strain C2 was low after induction of 1 μM aTc because of the inhibition of endogenous fatty acid synthesis. After fermentation for 12 h, the inducer was washed from the growth media, and cells started to grow and propagate (Fig. 5a). These results demonstrate that the silencing effects of dCas9-sgRNA for fabD can be induced and reversed. Furthermore, this method can be developed into a biological switch for pinosylvin or other stilbene chemical biosynthesis. Finally, we tested the biosynthetic capacity of pinosylvin in engineered strain C2. Different concentrations of trans-cinnamic acid, ranging from 0.25 to 4 mM, were fed into the medium. As shown in Fig. 5b, after 12 h of culture, the pinosylvin production was measured by HPLC analysis. The result suggested that 0.5 mM trans-cinnamic acid can be converted to ~0.22 mM (47.49 ± 2.57 mg/L) pinosylvin. Addition of substrates failed to increase the production.
Pinosylvin bioproduction from trans-cinnamic acid extraction
In previous study, engineered E. coli cell for pinosylvin bioproduction also using the isopropyl-β-D-1-thiogalactopyranoside, IPTG for gene expression and low efficient conversion of aromatic amino acids to pinosylvin, up to 13 mg/L production was detected in medium (Katsuyama et al. 2007; van Summeren-Wesenhagen and Marienhagen 2015; Wang et al. 2015). When the engineered strain P7 harboring Vvsts and At4cl in double-promoter construct with opposite expression direction, higher efficient conversion of trans-cinnamic acid to pinosylvin (production was 25.05 ± 2.55 mg/L) was explored. However, the addition of trans-cinnamic acid was not suitable for larger fermentation in terms of food safety, because of so many trans-cinnamic acid is obtained by chemical synthesis routes by benzaldehyde from petroleum industry. Thus we first examined stepwise for pinosylvin bioproduction. To reduce the cost of substrate, we selected a phenylalanine overproduction E. coli ATCC31884, and engineered the strain with PAL expression to trans-cinnamic acid production from its phenylalanine (strain D1). After shake-flask culture for 12 h, 165.57 ± 4.85 mg/L trans-cinnamic acid was detected in medium. According to the transformational capacity of trans-cinnamic acid (Fig. 5b) in engineered strain C2, the crude extract contained 0.5 mM (74 mg/L) trans-cinnamic acid was added into the culture medium of engineered strain C2. Finally, the yield of pinosylvin was approximate 47 mg/L by HPLC detection. These results suggested that we successfully build up a process for pinosylvin biosynthesis from trans-cinnamic acid extraction in Escherichia coli.
Discussion
Phenylpropanoids, which include stilbenoids, flavonoids, curcuminoids, and cinnamyl anthranilate, are a diverse group of secondary metabolites synthesized from l-phenylalanine or l-tyrosine; these compounds exhibit various bioactivities. Recently, much more study for natural product synthesis by different artificial biological systems (Zhou et al. 2014). For solution to the existing problem about high cost and low conversion efficiency, these work systematically demonstrated that a combinatorial strategy for pin pinosylvin bioproduction.
BioBrick assembly and metabolic regulation are essential to facilitate the construction of microbial cell factories for phenylpropanoid bioproduction. We have successfully improved the biosynthetic capacity of pinosylvin in E. coli by BioBrick combinatorial optimization. The engineered E. coli Dh5α carrying Vvsts and At4cl in double GAP promoter construct with opposite expression direction, which displays higher efficient conversion of trans-cinnamic acid to pinosylvin.
The CRISPRi system offers the possibility of repressing expression of genes related with fatty acid metabolism, thus avoiding labor-intensive multiple gene expression and deletion. In addition, CRISPRi avoids the disadvantage of the traditional gene knockout method, especially the fabD gene was repressed and not lead to strain death. Just like a function of switch, CRISPRi can be used to repress the target gene any time, and its effects are reversible. Although this improvement of production was less than that obtained when cerulenin was used (van Summeren-Wesenhagen and Marienhagen 2015), this method could be an alternative to the fabB and fabF gene repression in E. coli. The CRISPRi system works to regulate expression of fabD gene, a maximum increasement (1.9-fold) of pinosylvin was investigated. To our knowledge, this is the first report to explore the possibility of using CRISPRi system for improving stilbene production in E. coli. In comparison to another similar function method, synthetic antisense RNAs, the stages of gene regulatory level in the genome is different. The synthetic antisense RNAs system controls the expression of target genes in trans at the post-transcriptional level (Na et al. 2013), while the CRISPRi system influences the gene coding sequence to block transcription initiation or elongation (Qi et al. 2013). So we also want to have experiment to find out a more flexible and economical method further.
Finally, we also demonstrated the feasibility of rational design of stepwise pinosylvin biosynthesis from glycerol. However, the yield was not satisfactory for industrial-scale production. A larger scale fermentation for improving pinosylvin bioproduction will be experiment in the future. The trans-cinnamic acid was usually used as the starting precursor to produce phenylpropanoids, such as naringenin, pinocembrin, chrysin and so on (Kong 2015). Thus we believe that these combinatorial processed also could be a sustainable and flexible alternative method for microbial production of phenylpropanoids.
References
Bhan N, Xu P, Khalidi O, Koffas MAG (2013) Redirecting carbon flux into malonyl-CoA to improve resveratrol titers: proof of concept for genetic interventions predicted by OptForce computational framework. Chem Eng Sci 103:109–114. doi:10.1016/j.ces.2012.10.009
Conde E, Fang W, Hemming J et al (2014) Recovery of bioactive compounds from Pinus pinaster wood by consecutive extraction stages. Wood Sci Technol 48:311–323. doi:10.1007/s00226-013-0604-1
Holleley CE, Geerts PG (2009) Multiplex manager 1.0: a cross-platform computer program that plans and optimizes multiplex PCR. Biotechniques 46:511–517. doi:10.2144/000113156
Huang Q, Lin Y, Yan Y (2013) Caffeic acid production enhancement by engineering a phenylalanine over-producing Escherichia coli strain. Biotechnol Bioeng 110:3188–3196. doi:10.1002/bit.24988
Jancinova V, Perecko T, Nosal R et al (2012) The natural stilbenoid pinosylvin and activated neutrophils: effects on oxidative burst, protein kinase C, apoptosis and efficiency in adjuvant arthritis. Acta Pharmacol Sin 33:1285–1292. doi:10.1038/aps.2012.77
Janßen HJ, Steinbüchel A (2014) Fatty acid synthesis in Escherichia coli and its applications towards the production of fatty acid based biofuels. Biotechnol Biofuels 7:7. doi:10.1186/1754-6834-7-7
Ji W, Lee D, Wong E et al (2014) Specific gene repression by CRISPRi system transferred through bacterial conjugation. ACS Synth Biol 3:929–931. doi:10.1021/sb500036q
Jones KL, Kim SW, Keasling JD (2000) Low-copy plasmids can perform as well as or better than high-copy plasmids for metabolic engineering of bacteria. Metab Eng 2:328–338. doi:10.1006/mben.2000.0161
Katsuyama Y, Funa N, Horinouchi S (2007) Precursor-directed biosynthesis of stilbene methyl ethers in Escherichia coli. Biotechnol J 2:1286–1293. doi:10.1002/biot.200700098
Kong J (2015) Phenylalanine ammonia-lyase, a key component used for phenylpropanoids production by metabolic engineering. RSC Adv 5:62587–62603. doi:10.1039/C5RA08196C
Koskela A, Reinisalo M, Hyttinen JMT et al (2014) Pinosylvin-mediated protection against oxidative stress in human retinal pigment epithelial cells. Mol Vis 20:760–769
Laavola M, Nieminen R, Leppänen T et al (2015) Pinosylvin and monomethylpinosylvin, constituents of an extract from the knot of Pinus sylvestris, reduce inflammatory gene expression and inflammatory responses in vivo. J Agric Food Chem 63:3445–3453. doi:10.1021/jf504606m
Lentini R, Forlin M, Martini L et al (2013) Fluorescent proteins and in vitro genetic organization for cell-free synthetic biology. ACS Synth Biol 2:482–489. doi:10.1021/sb400003y
Leonard E, Yan Y, Fowler ZL et al (2008) Strain improvement of recombinant Escherichia coli for efficient production of plant flavonoids. Mol Pharm 5:257–265. doi:10.1021/mp7001472
Lim CG, Fowler ZL, Hueller T et al (2011) High-yield resveratrol production in engineered Escherichia coli. Appl Environ Microbiol 77:3451–3460. doi:10.1128/AEM.02186-10
Lv L, Ren Y, Chen J et al (2015) Application of CRISPRi for prokaryotic metabolic engineering involving multiple genes, a case study: controllable P(3HB-co-4HB) biosynthesis. Metab Eng 29:1–9. doi:10.1016/j.ymben.2015.03.013
Na D, Yoo SM, Chung H et al (2013) Metabolic engineering of Escherichia coli using synthetic small regulatory RNAs. Nat Biotechnol 31:170–174. doi:10.1038/nbt.2461
Park E-J, Park HJ, Chung H-J et al (2012) Antimetastatic activity of pinosylvin, a natural stilbenoid, is associated with the suppression of matrix metalloproteinases. J Nutr Biochem 23:946–952. doi:10.1016/j.jnutbio.2011.04.021
Park EJ, Chung HJ, Park HJ et al (2013) Suppression of Src/ERK and GSK-3/beta-catenin signaling by pinosylvin inhibits the growth of human colorectal cancer cells. Food Chem Toxicol 55:424–433. doi:10.1016/j.fct.2013.01.007
Qi L, Haurwitz RE, Shao W et al (2012) RNA processing enables predictable programming of gene expression. Nat Biotechnol 30:1002–1006. doi:10.1038/nbt.2355
Qi LS, Larson MH, Gilbert LA et al (2013) Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152:1173–1183. doi:10.1016/j.cell.2013.02.022
Rivière C, Pawlus AD, Mérillon JM (2012) Natural stilbenoids: distribution in the plant kingdom and chemotaxonomic interest in Vitaceae. Nat Prod Rep 29:1317–1333. doi:10.1039/c2np20049j
van Summeren-Wesenhagen PV, Marienhagen J (2015) Metabolic engineering of Escherichia coli for the synthesis of the plant polyphenol pinosylvin. Appl Environ Microbiol 81:840–849. doi:10.1128/AEM.02966-14
Wang S, Zhang S, Xiao A et al (2015) Metabolic engineering of Escherichia coli for the biosynthesis of various phenylpropanoid derivatives. Metab Eng 29:153–159. doi:10.1016/j.ymben.2015.03.011
Wu J, Liu P, Fan Y et al (2013) Multivariate modular metabolic engineering of Escherichia coli to produce resveratrol from l-tyrosine. J Biotechnol 167:404–411. doi:10.1016/j.jbiotec.2013.07.030
Wu H, Wang Y, Wang Y et al (2014) Quantitatively relating gene expression to light intensity via the serial connection of blue light sensor and CRISPRi. ACS Synth Biol 3:979–982. doi:10.1021/sb500059x
Yang Y, Lin Y, Li L et al (2015) Regulating malonyl-CoA metabolism via synthetic antisense RNAs for enhanced biosynthesis of natural products. Metab Eng 29:217–226. doi:10.1016/j.ymben.2015.03.018
Yeo SCM, Luo W, Wu J et al (2013) Quantification of pinosylvin in rat plasma by liquid chromatography-tandem mass spectrometry: application to a pre-clinical pharmacokinetic study. J Chromatogr B Analyt Technol Biomed Life Sci 931:68–74. doi:10.1016/j.jchromb.2013.05.023
Zhou J, Du G, Chen J (2014) Novel fermentation processes for manufacturing plant natural products. Curr Opin Biotechnol 25:17–23. doi:10.1016/j.copbio.2013.08.009
Acknowledgments
This work was supported by the National Natural Science Foundation of China (Grant Nos. 31071837, 31272217, and 31372116); the Projects of Science and Technology of Guangdong Province (Grant No. 2013B010404041 and 2014B050505018).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liang, Jl., Guo, Lq., Lin, Jf. et al. A novel process for obtaining pinosylvin using combinatorial bioengineering in Escherichia coli . World J Microbiol Biotechnol 32, 102 (2016). https://doi.org/10.1007/s11274-016-2062-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-016-2062-z