Abstract
Although magnetic resonance imaging (MRI) has a higher sensitivity of early breast cancer than mammography, the specificity is lower. The purpose of this study was to develop a computer-aided diagnosis (CAD) scheme for distinguishing between benign and malignant breast masses on dynamic contrast material-enhanced MRI (DCE-MRI) by using a deep convolutional neural network (DCNN) with Bayesian optimization. Our database consisted of 56 DCE-MRI examinations for 56 patients, each of which contained five sequential phase images. It included 26 benign and 30 malignant masses. In this study, we first determined a baseline DCNN model from well-known DCNN models in terms of classification performance. The optimum architecture of the DCNN model was determined by changing the hyperparameters of the baseline DCNN model such as the number of layers, the filter size, and the number of filters using Bayesian optimization. As the input of the proposed DCNN model, rectangular regions of interest which include an entire mass were selected from each of DCE-MRI images by an experienced radiologist. Three-fold cross validation method was used for training and testing of the proposed DCNN model. The classification accuracy, the sensitivity, the specificity, the positive predictive value, and the negative predictive value were 92.9% (52/56), 93.3% (28/30), 92.3% (24/26), 93.3% (28/30), and 92.3% (24/26), respectively. These results were substantially greater than those with the conventional method based on handcrafted features and a classifier. The proposed DCNN model achieved high classification performance and would be useful in differential diagnoses of masses in breast DCE-MRI images as a diagnostic aid.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Breast cancer is the most common cancer in women. In the USA, 40,610 women died due to breast cancer in 2017 [1]. Early detection and treatment are very important for breast cancer patients since 95% of patients with early breast cancer can be cured completely [2]. Dynamic contrast enhanced magnetic resonance imaging (DCE-MRI) has become an established clinical imaging modality for diagnosis and staging of breast cancer [3, 4]. The sensitivity of breast DCE-MRI is higher than that of mammography, which is the standard screening modality. Especially in dense breast, the sensitivity has been improved from 33–59% with mammography to 71–94% with DCE-MRI [5,6,7,8,9]. However, the specificity of DCE-MRI, which is typically between 30–70%, is lower than that of mammography [10,11,12,13,14].
A computer-aided diagnosis (CAD) scheme [15] is one of the solutions to improve the specificity of breast MRI. The CAD scheme presents the likelihood of malignancy for a lesion on medical image as a “second opinion” in order to assist radiologists’ diagnosis [15,16,17,18]. In our previous study, we developed the CAD scheme for distinguishing between benign and malignant masses on breast DCE-MRI [19]. The CAD scheme was based on the conventional method with the handcrafted features and a classifier. In the CAD scheme, mass region was segmented from DCE-MRI images. Objective features were then extracted from the segmented mass region to distinguish between benign and malignant masses. Therefore, there was a problem that classification performance was strongly dependent on the segmented mass region.
Recently, deep convolutional neural networks (DCNNs) such as AlexNet, ZFNet, VGG16, and GoogLeNet have been applied to classification tasks [20,21,22,23]. The DCNNs have shown greater classification performances than the conventional methods with the handcrafted features and a classifier in an ImageNet Large Scale Visual Recognition Competition (ILSVRC) [24]. The DCNN can extract complex multi-level objective features from input images due to self-learning ability without the segmentation of target [25,26,27]. Therefore, there is not a problem that the classification performance is influenced from the segmented region. As with many studies [18, 28,29,30,31], the classification accuracy of our previous CAD scheme based on the conventional method might be improved by using the DCNN. The well-known DCNN models such as AlexNet, ZFNet, VGG16, and GoogLeNet were constructed for general images with RGB channels. Therefore, those models can be inadequacy in the classification task for medical images with grayscale channel.
The purpose of this study was to determine the optimum architecture of the DCNN models for distinguishing benign from malignant masses on breast DCE-MRI using Bayesian optimization [32,33,34,35]. In this study, we first determined a baseline DCNN model from well-known DCNN models in term of classification performance. The optimum architecture of the DCNN model was clarified by changing the hyperparameters of the baseline DCNN model such as the number of layers, the filter size, and the number of filters using Bayesian optimization. The usefulness of the optimized DCNN model was evaluated by comparing with conventional DCNN models.
Materials and Methods
The use of the following database was approved by the Institutional Review Board at Ritsumeikan University. The database was stripped of all patient identifiers.
Materials
Our database consisted of 56 DCE-MRI examinations—each of which contained five sequential phase images—that were obtained from 56 patients (mean age: 55.8 years, age range: 20–82 years). These DCE-MRI images were acquired with a 3.0 T MR scanner at Hokuto Hospital (Obihiro, Japan) from October 2009 and July 2015. The patients were excluded if they had undergone breast surgery in the past, size of mass was more than 5 cm. It included 30 malignant and 26 benign breast masses. Table 1 shows the patients’ clinical information. All masses underwent 10 G vacuum-assisted biopsy and/or surgical specimen. After the injection of a contrast agent, four post-contrast series of 3D MRI scans and data acquisitions were sequentially performed after a duration of 0 min, 1 min, 2 min, and 4 min. The one pre-contrast and the four post-contrast series generated images with a spatial resolution of 0.7 × 0.7 × 1.2 mm3, with a data matrix of 512 × 512 pixels. Figure 1 shows an example of pre-contrast and four post-contrast DCE-MRI images. Each of five image scan series consisted of 150 image slices.
Determination of Baseline DCNN Model
To optimize the architecture of DCNN model for distinguishing between benign and malignant masses on DCE-MRI images, we first determined a baseline DCNN model from AlexNet, ZFNet, VGG16, and GoogLeNet in terms of area under the receiver operating characteristic (ROC) curve [36]. Here, we briefly described those DCNN models below.
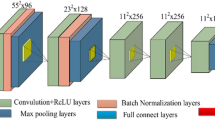
AlexNet consists of five convolutional layers, three max-pooling layers, and three fully connected layers including cross channel normalization layer, rectified linear unit (ReLU) function, and dropout. The convolutional layer and the max-pooling layer play a role of feature extractor, whereas the fully connected layer plays a role of classifier. The first convolutional layer has 96 filters of size 11 × 11 with a stride of four pixels and padding with two pixels. The second convolutional layer has 256 filters of size 5 × 5. The third, fourth, and fifth convolutional layers have 384, 384, and 256 filters with size of 3 × 3, respectively. The number of the units in the first and second fully connected layer is 4,096, whereas that of the units in the third fully connected layer is same as the number of the classes. In this study, the number of units in the third fully connected layer is two (Benign or Malignant).
ZFNet is similar architecture to AlexNet. The difference between ZFNet and AlexNet is only filter size and stride in first convolutional layer. The filter size of the first convolutional layer in ZFNet is 7 × 7, whereas the stride is two pixels. The other parameters of ZFNet are same as AlexNet.
VGG16 consists of 13 convolutional layers with filter size of 3 × 3, and three fully connected layers including ReLU function and dropout. The configurations of fully-connected layers in VGG16 are the same with AlexNet.
GoogLeNet consists of 22 layers with nine inception layers and one fully connected layer. The inception layer has multiple convolution filters [23].
Optimization of DCNN Architecture with Bayesian Optimization
After the determination of baseline DCNN model with the highest area under the ROC curve (AUC), the hyperparameters such as the number of the convolutional layer, the number of filters, and the filter size were optimized in the baseline DCNN model using Bayesian optimization with Gaussian process [32,33,34,35]. The Bayesian optimization is an algorithm for optimizing hyperparameters in a machine learning. Table 2 shows the search range for each hyperparameter in the DCNN model. When the number of the convolutional layer was − 4, it means to remove the final and the fourth from the last convolutional layer in the baseline DCNN model. On the other hand, when the number of the convolutional layer was + 4, it means to add four convolutional layers after the final convolutional layer. With the number of the convolutional layer, the number of the max-pooling layer, and the number of the fully connected layer were 0, the configuration of the DCNN model was same as the baseline DCNN model.
In the Bayesian optimization, four different combinations of hyperparameters were first determined by selecting search value randomly in each hyperparameter. The DCNN model with each combination was then trained independently. With a Gaussian process based on the classification errors for each DCNN model, the combination of hyperparameters was updated to reduce the classification error. By repeating this process 100 times, the optimal combination of hyperparameters was founded efficiently.
Training and Testing of DCNN
The DCNN model was developed and evaluated using MATLAB 2019a on a workstation (CPU: Intel Core i7-7820X processor, RAM: 128 GB, and GPU: NVIDIA GeForce GTX 1080Ti).
A k-fold cross validation method [37] with k = 3 was utilized for the training and testing of the DCNN model. In the method, 56 patients were randomly divided into three groups (A, B, C) so that the number of patients was approximately equal. A group was used for test dataset, whereas the remaining two groups were used for training dataset. This process was repeated three times until every group had been used for test dataset.
The ROIs which include an entire mass were selected from each DCE-MRI image by an experienced radiologist (12 years of experience devoted in breast image diagnosis). For augmenting each training data, each ROI was flipped horizontally and cropped [38] randomly eight times. Thus, the total number of training samples was increased by 16 times. A stochastic gradient descent (SGD) was employed to minimize the loss between the output of the proposed DCNN model and the corresponding teacher signal.
Evaluation of Classification Performance
The classification accuracy, the sensitivity, the specificity, the positive predictive value (PPV), and the negative predictive value (NPV) [39] of the DCNN model were evaluated by using the ensemble average from the testing datasets over the threefold cross validation method. ROC analysis [36] was also used for evaluation of classification performance.
Results
The baseline DCNN model was first determined from AlexNet, ZFNet, VGG16, and GoogLeNet in terms of the AUC. Here, the learning rate, the mini-batch size, and the number of epochs were given by 0.0001, 3, and 15, respectively. The AUC for ZFNet was 0.889, showing to be greater than those for AlexNet (0.867, P = 0.302), VGG16 (0.800, P = 0.050), and GoogLeNet (AUC = 0.827, P = 0.080). Therefore, ZFNet was determined as the baseline DCNN model.
The hyperparameters in the determined baseline DCNN model were optimized using Bayesian optimization. Tables 3 and 4 show the optimal architecture and the parameters determined by Bayesian optimization when each group was used for testing dataset, respectively. The average AUC of the determined DCNN models for three test datasets was 0.945.
Figure 2 compares the ROC curve for the proposed DCNN model with those for AlexNet, ZFNet, VGG16, and GoogLeNet. The AUC for the proposed DCNN model (0.945) was significantly higher than those for the other four DCNN models (AlexNet: 0.867, P = 0.015; ZFNet: 0.889, P = 0.026; VGG16: 0.800, P = 0.002, GoogLeNet: 0.827, P = 0.006). Table 5 shows average classification accuracy among five different DCNN models. All evaluation indices for the proposed DCNN model was the highest among the five different DCNN models.
Figure 3 shows example of correctly classified cases and incorrectly classified cases in DCE-MRI images at 1-min post-contrast. The characteristics of masses incorrectly classified by the proposed DCNN model were as follows: (1) small masses (2 cm or lower in size), (2) malignant masses with regularity in shape, and (3) benign masses with irregularity in shape.
Discussion
To investigate the usefulness of the proposed DCNN model, we compared with the classification performance with our previous method based on the handcrafted features and a classifier [19]. In the previous method, ROI which included an entire mass was selected manually on the DCE-MRI image. The mass region was determined automatically by applying Otsu’s method [40]. Quadratic discriminant analysis (QDA) was employed to distinguish between benign and malignant masses. The four handcrafted features were used for the input of the QDA. With the previous method, the average classification accuracy, the sensitivity, the specificity, the PPV, and the NPV were 75.0% (42/56), 76.7% (23/30), 73.1% (19/26), 76.7% (23/30), and 73.1% (19/26), respectively. Figure 4 compares the ROC curve for the proposed DCNN model with that for the previous method. The AUC for the proposed DCNN model was significantly greater than that for the previous method (0.810, P = 0.01). These results would imply that the features extracted automatically from the proposed DCNN model were more useful for distinguishing between benign and malignant masses when compared with handcrafted features manually obtained in our previous method.
With the proposed DCNN model, the classification accuracy, sensitivity, specificity, PPV, and NPV for 34 masses smaller than 2 cm were 88.2% (30/34), 84.6% (11/13), 90.5% (19/21), 84.6% (11/13), and 90.5% (19/21), whereas those for 22 masses larger than 2 cm were 100% (22/22), 100% (17/17), 100% (5/5), 100% (17/17), and 100% (5/5). We believe that the proposed DCNN model can reduce the number of unnecessary biopsies for masses larger than 2 cm. Patients with masses smaller than 2 cm which are early lesions will be able to undergo follow-up at a short interval. The classification accuracy for those masses might be improved by introducing the differences in growth speed between benign and malignant cases into the proposed method[41,42]. However, further studies are required by use of large data sets to evaluate the computerized method for analysis of changes over time.
In this study, a strategy for optimizing the hyperparameters in the DCNN model was proposed. It is very difficult to construct suitable DCNN model for classification tasks from scratch, because the combination of hyperparameters in the DCNN is infinite. In the proposed strategy, we determined baseline DCNN model from well-known DCNN models, and then optimized architecture and the other parameters of the baseline DCNN model for breast mass classification task using Bayesian optimization. With this strategy, we could easily construct suitable DCNN architecture according to tasks. In the fact, this study has shown that the proposed DCNN model achieved better classification accuracy than that of our previous method and well-known DCNN model. Therefore, we believe that this strategy is effective for optimizing DCNN architecture and its parameters in classification task.
There are some limitations in this study. We used only 56 patient data in this study. Thus, we need to evaluate the strategy for optimizing the hyperparameters and the proposed DCNN model in the further study by using larger dataset. Other limitation is that the proposed DCNN model used two-dimensional (2D) ROIs as the input. In clinical practice, radiologists usually diagnose by considering three-dimensional (3D) information on DCE-MRI. Therefore, 3D-based DCNN model might be more appropriate than 2D. However, 3D-based DCNN model has a lot of parameters to train and requires a large number of training data. In this study with a small dataset, 2D-based DCNN model was employed to distinguish between benign and malignant masses. Finally, the DCNN model determined by the Bayesian optimization might not yield the best classification accuracy because the Bayesian optimization does not evaluate all combinations of hyperparameters. However, we believe that Bayesian optimization is useful for effectively determining appropriate hyperparameters of the DCNN model from infinite combinations.
Conclusion
In this study, we developed the CAD scheme to distinguish benign from malignant masses on breast DCE-MRI images by use of the optimal DCNN model determined with Bayesian optimization. The proposed DCNN model achieved high classification accuracy for masses and would be useful in differential diagnoses of masses as a diagnostic aid.
References
Siegel RL, Miller KD, Jemal A, et al: Cancer statistics, 2017. CA: a cancer journal for clinicians 2017; 67(1): 7–30
Reynolds HE, Jackson VP: Self-referred mammography patients: analysis of patients' characteristics. American Journal of Roentgenology 1991; 157: 481-484
Kuhl CK, Schild HH: Dynamic image interpretation of MRI of the breast. Journal of Magnetic Resonance Imaging 2000; 12(6): 965-974
Gubern-Mérida A, Martí R, Melendez J, Hauth JL, Mann RM, Karssemeijer N, Platel B, et al: Automated localization of breast cancer in DCE-MRI. Medical image analysis 2015; 20(1): 265-274
Leach MO, Boggis CR, Dixon AK, et al: Screening with magnetic resonance imaging and mammography of a UK population at high familial risk of breast cancer: a prospective multicentre cohort study (MARIBS), LANCET 2005; 365: 1769-1778
Peters NH, Borel Rinkes IH, Zuithoff NP, Mali WP, Moons KG, Peeters PH, et al: Meta-analysis of MR imaging in the diagnosis of breast lesions. Radiology 2008; 246(1): 116-124
Sardanelli F, Podo F, D'Agnolo G, et al: Multicenter comparative multimodality surveillance of women at genetic-familial high risk for breast cancer (HIBCRIT study): interim results. Radiology 2007; 242(3): 698-715
Kuhl CK, Schrading S, Leutner CC, et al: Mammography, breast ultrasound, and magnetic resonance imaging for surveillance of women at high familial risk for breast cancer. Journal of Clinical Oncology 2005; 23: 8469–8476
Pisano ED, Hendrick RE, Yaffe MJ, Baum JK, Acharyya S, Cormack JB, D'Orsi CJ, et al: Diagnostic accuracy of digital versus film mammography: exploratory analysis of selected population subgroups in DMIST. Radiology 2008; 246(2): 376-383
Nie K, Chen JH, Yu HJ, Chu Y, Nalcioglu O, Su MY: Quantitative analysis of lesion morphology and texture features for diagnostic prediction in breast MRI. Academic Radiology 2008; 15(12): 1513–1525
Siegmann KC, Kra¨mer B, Claussen C, et al: Current status and new developments in breast MRI. Breast Care (Basel) 2011; 6(2): 87–92
Fusco R, Sansone M, Filice S, Carone G, Amato DM, Sansone C, Petrillo A, et al: Pattern recognition approaches for breast cancer DCE-MRI classification: a systematic review. Journal of medical and biological engineering 2016; 36(4): 449-459
Berg WA, Blume JD, Cormack JB, et al.: Combined screening with ultrasound and mammography vs mammography alone in women at elevated risk of breast cancer. The Journal of the American Medical Association 2008; 299(18): 2151-2163
Chae EY, Kim HH, Cha JH, Shin HJ, Kim H, et al: Evaluation of Screening Whole‐Breast Sonography as a Supplemental Tool in Conjunction with Mammography in Women With Dense Breasts. Journal of Ultrasound in Medicine 2013; 32(9): 1573-1578
Doi K, MacMahon H, Katsuragawa S, Nishikawa RM, Jiang Y, et al: Computer-aided diagnosis in radiology: potential and pitfalls. European journal of Radiology 1999; 31(2): 97–109
Doi K: Computer-aided diagnosis in medical imaging: historical review, current status and future potential. Computerized medical imaging and graphics 2007; 31(4–5): 198–211
Hizukuri A, Nakayama R, Kashikura Y, Takase H, Kawanaka H, Ogawa T, Tsuruoka S, et al: Computerized determination scheme for histological classification of breast mass using objective features corresponding to clinicians’ subjective impressions on ultrasonographic images. Journal of digital imaging 2013; 26(5): 958-970
Hizukuri A, Nakayama R: Computer-Aided Diagnosis Scheme for Determining Histological Classification of Breast Lesions on Ultrasonographic Images Using Convolutional Neural Network. Diagnostics 2018; 8(3): 1-9
Honda E, Nakayama R, Koyama H, Yamashita A, et al: Computer-aided diagnosis scheme for distinguishing between benign and malignant masses in breast DCE-MRI. Journal of digital imaging 2016; 29(3): 388-393
Krizhevsky A, Sutskever I, Hinton GE: Imagenet classification with deep convolutional neural networks. In Advances in Neural Information Processing Systems 2012; 1097–1105
Zeiler MD, Fergus R: Visualizing and understanding convolutional networks. In European conference on computer vision, Springer, Cham, 2014; 818-833
Simonyan K, Zisserman A: Very deep convolutional networks for large-scale image recognition. Int. Conf. on Learning Representations 2015; San Diego, CA
Christian S, et.al.: Going deeper with convolutions, Proceedings of the IEEE conference on computer vision and pattern recognition. 2015
Russakovsky O, Deng J, Su H, Krause J, Satheesh S, Ma S, Huang Z, Karpathy A, Khosla A, Bernstein M, Berg AC, Fei-Fei L, et al: ImageNet Large Scale Visual Recognition Challenge, International Journal of Computer Vision, 2015; 115(3); 211-252
LeCun Y, Bengio Y, Hinton G, et al: Deep learning. nature 2015; 521(7553): 436
Litjens G, Kooi T, Bejnordi BE, Setio AAA, Ciompi F, Ghafoorian M, Sánchez CI, et al: A survey on deep learning in medical image analysis, Medical image analysis 2017; 42: 60-88
Sahiner B, Pezeshk A, Hadjiiski LM, Wang X, Drukker K, Cha KH, Summers RM, Giger ML, et al: Deep learning in medical imaging and radiation therapy, Medical Physics 2019; 46(1)
Bayramoglu N, Juho K, Janne H, et al: Deep learning for magnification independent breast cancer histopathology image classification, 23rd International conference on pattern recognition (ICPR), 2016; 2440–2445
Jadoon MM, Zhang Q, Haq IU, Butt S, Jadoon A, et al: Three-class mammogram classification based on descriptive CNN features, BioMed Research International, 2017, 1–11
Li Q, Cai W, Wang X, Zhou Y, Feng DD, Chen M, et al: Medical image classification with convolutional neural network, 13th International Conference on Control Automation Robotics & Vision (ICARCV), 2014, 844–848
Nejad EM, Affendey LS, Latip RB, Bin Ishak, et al: Classification of histopathology images of breast into benign and malignant using a single-layer convolutional neural network, Proceedings of the International Conference on Imaging, Signal Processing and Communication, 2017, 50–53
Rasmussen CE: Gaussian processes in machine learning. In Summer School on Machine Learning, Springer, Berlin, Heidelberg, 2003, 63-71
Le HT, Phung SL, Bouzerdoum A, Tivive FHC, et al: Human motion classification with micro-doppler radar and bayesian-optimized convolutional neural networks. In 2018 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP) , 2018, 2961–2965
Snoek J, Larochelle H, Adams RP: Practical Bayesian optimization of machine learning algorithms. In Advances in Neural Information Processing Systems, 2012, 2951–2959
Dernoncourt F, Lee JY: Optimizing neural network hyperparameters with Gaussian processes for dialog act classification. In 2016 IEEE Spoken Language Technology Workshop (SLT), 2016, 406–413
Metz CE: ROC methodology in radiologic imaging. Investigative Radiology 1986; 21: 720–733
Peng Y, Jiang Y, Antic T, Giger ML, Eggener SE, Oto A, et al: Validation of quantitative analysis of multiparametric prostate MR images for prostate cancer detection and aggressiveness assessment: a cross-imager study. Radiology 2014; 271(2): 461-471
Cireşan DC, Meier U, Gambardella LM, Schmidhuber J, et al: Deep, big, simple neuralnets for handwritten digit recognition. Neural computation 2010; 22(12): 3207-3220
Langlotz CP: Fundamental measures of diagnostic examination performance: Usefulness for clinicaldecision making and research. Radiology 2003; 228: 3–9
Otsu N: A Threshold Selection Method from Gray-Level Histograms. IEEE Transactions on Systems, Man, and Cybernetics 1979; 9(1): 62–66
Nakayama R, Uchiyama Y, Watanabe R, Katsuragawa S, Namba K, Doi K, et al: Computer‐aided diagnosis scheme for histological classification of clustered microcalcifications on magnification mammograms. Medical physics 2004; 31(4): 789–799
Nakayama R, Watanabe R, Namba K, Takeda K, Yamamoto K, Katsuragawa S, Doi K, et al: Computer-aided diagnosis scheme for identifying histological classification of clustered microcalcifications by use of follow-up magnification mammograms. Academic radiology 2006; 13(10): 1219–1228
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Hizukuri, A., Nakayama, R., Nara, M. et al. Computer-Aided Diagnosis Scheme for Distinguishing Between Benign and Malignant Masses on Breast DCE-MRI Images Using Deep Convolutional Neural Network with Bayesian Optimization. J Digit Imaging 34, 116–123 (2021). https://doi.org/10.1007/s10278-020-00394-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10278-020-00394-2