Abstract
Collagen is an abundant structural biopolymer in mammal vertebrates, providing structural support as well as mechanical integrity for connective tissues such as bone, ligament, and tendon. The mechanical behaviours of these tissues are determined by the nanomechanics of their structures at different hierarchies and the role of collagen structures in the extracellular matrix. Some studies revealed that there is significant microstructural difference in the longitudinal direction of the collagen fibril, which challenges the conventional rod-like assumption prevalently adopted in the existing studies. Motivated by this discrepancy, in this study, we investigated the longitudinal heterogeneous nanomechanical properties of type I collagen molecule to probe the origin of the longitudinal heterogeneity of the collagen fibril at the molecular level. A full length type I collagen molecule structure was built based on the experimentally calibrated nanostructure. Then, a suitable strain rate was determined for stretching the three intact ‘gap’ regions and three intact ‘overlap’ regions of the collagen molecule. Further, the nanomechanical properties of the six collagen molecule segments were characterized by performing steered molecular dynamics simulations, using the obtained suitable strain rate in modelling. The results indicate that this computational model can be used to capture the mechanical behaviour of the collagen molecule under physiological stress conditions. Moreover, the ‘gap’ regions show a lower stiffness and undergo a slightly lager strain in the unwinding process, compared to the ‘overlap’ regions of the collagen molecule. This investigation provides insights into the origin of the longitudinal heterogeneity of collagen fibrils at the molecular level and suggests that it is of significant importance to consider the longitudinal heterogeneous mechanical properties of the collagen molecule in the development of coarse-grained models of collagen-related tissues.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Collagen is a ubiquitous structural biopolymer that is critical to various functions in human body, including tissue scaffolding, cell adhesion, cell migration, cancer, angiogenesis, tissue morphogenesis and tissue repair (Kadler et al. 2007). Malfunctions involving this biological polymer due to gene mutations in different types of collagen can result in severe diseases such as premature osteoarthritis (Husar-Memmer et al. 2013), osteogenesis imperfecta (Lindahl et al. 2015), stickler syndrome (Robin et al. 2014) and Ehlers–Danlos syndrome (Leistritz et al. 2011). Understandings of the mechanical properties of collagenous tissues at different hierarchies can improve the diagnosis of related diseases (Kim et al. 2015; Gautieri et al. 2009b, 2012b).
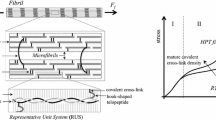
In human, there are at least 29 types of collagens consisting of fibrillar and non-fibrillar collagens. Among those types of fibrillar collagens, type I collagen is the most common in nature (Hulmes 2008) and attracts the most attention among researchers. Type I collagen is prevalent in the extracellular matrix in vertebrates, imparting both structural support and mechanical integrity to connective collagenous tissues. The mechanical properties of these collagenous tissues are mainly attributed to their hierarchical structures, with fibres at the top, then fibrils, then microfibrils, and finally collagen molecules at the bottom level. Collagen molecule, with \(\sim \)1.5 nm in diameter and \(\sim \)300 nm in length, consists of three left-handed polyproline II-type strands intertwining with each other in a right-handed fashion to form a coiled coil triple helix (Jenkins and Raines 2002). Five collagen molecules assemble into a right-handed supermolecule known as microfibril, which further forms fibril with outstanding tensile strength and high flexibility. Multiple fibrils can be organized and cross-linked into fibril networks, which contribute to the tensile resistance of collagenous tissues.
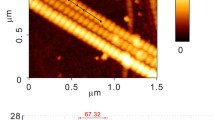
In recent years, extensive experimental and computational efforts have been paid to characterize the elastic properties of collagenous-based tissues at different hierarchies, including tissue (Yang et al. 2015), fibre (Gentleman et al. 2003), fibril and microfibril (Gautieri et al. 2011; Buehler 2006b, 2008; Depalle et al. 2014; Eppell et al. 2006; Shen et al. 2008; Gupta et al. 2004; Rijt et al. 2006; Wenger et al. 2007; Heim et al. 2006), and molecular (Harley et al. 1977; Cusack and Miller 1979; Hofmann et al. 1984; Sasaki and Odajima 1996; Sun et al. 2002; Vesentini et al. 2005; Lorenzo and Caffarena 2005; Buehler 2006a; Stevens 2008; Gautieri et al. 2008, 2009a, 2010, 2012a, 2013; Uzel and Buehler 2009; Pradhan et al. 2011; Zhou et al. 2015). Comparison of elastic modulus of collagen at these hierarchical scales indicates that the stiffness decreases as one marches up the hierarchical level. Detailed information on Young’s modulus of collagen at different hierarchical scales can be found in Table 1 in Sherman et al. (2015). However, only a few of these studies are based on experimentally validated structures of collagen microfibril/molecule (Gautieri et al. 2011; Depalle et al. 2014; Zhou et al. 2015). In previous mechanical modelling studies, collagen fibril is assumed to be a rod-like structure, which is homogeneous along the longitudinal direction. However, a full crystallographic microfibril structure developed in the mid-2000s (Orgel et al. 2006) and other research (Zhou et al. 2015; Robinson and Watson 1952; Weiner and Wagner 1998; Graham et al. 2004; Balooch et al. 2008; Minary-Jolandan and Yu 2009; Wenger and Mesquida 2011; Baldwin et al. 2014; Spitzner et al. 2015; Hodge and Schmitt 1960; Streeter and Leeuw 2011) have challenged the assumption of longitudinal homogeneity. X-ray fibre diffraction experiments revealed that the overlap region of the collagen microfibril is approximately two times less disordered than the gap regions (Orgel et al. 2006). Previous TEM studies implied that minerals prefer to nucleate in the gap region of collagen fibrils in bones (Robinson and Watson 1952; Weiner and Wagner 1998), and a recent atomic force microscopy (AFM) collagen fibril demineralization investigation further confirmed this implication (Balooch et al. 2008). An AFM-based force spectroscopy characterization on individual synthetic collagen fibril showed that the overlap region has a larger cross-sectional area and is less likely to unfold as compared to the gap region (Graham et al. 2004). An AFM indentation study on single type I collagen fibril indicated that the gap region has a 50% smaller Young’s modulus and lower energy dissipations than the overlap region (Minary-Jolandan and Yu 2009). An AFM imaging study on the behaviour of the D-banding structure of collagen fibrils also suggested that there are longitudinal variations in the Poisson’s ratio of collagen fibrils, in which the overlap region has a larger Poisson’s ration than the gap region (Wenger and Mesquida 2011). In contrast, a recent nanomechanical mapping study on hydrated type I collagen fibrils found that the modulus in the overlap region is around 25% larger than in the gap region (Baldwin et al. 2014). A multiset point intermittent contact (MUSIC)-mode AFM investigation on the interaction between the tip and a hydrated type I collagen fibril demonstrated that the gap and overlap regions have different mechanical properties in terms of Young’s modulus, viscoelastic properties and dissipative energy (Spitzner et al. 2015). The overlap/gap density ratio of collagen fibril was proposed to be 5/4 (Hodge and Schmitt 1960). In addition, a recent molecular dynamics (MD) simulation study on the inter-protein interactions in collagen fibrils revealed that the overlap region has a higher average density of water bridges and hydrogen bonds (H-bonds) than the gap region (Streeter and Leeuw 2011). Obviously, all of these studies indicate that there is remarkable difference between the gap and overlap regions in terms of the geometry and mechanical properties. However, the structural basis of this longitudinal mechanical heterogeneity of the collagen fibril is not well understood at the molecular level.
At the molecular level, Bodian et al. (2011) performed MD simulations on the full triple-helical domain of type I collagen molecule by segmenting the protein into 24 overlapping fragments. This study demonstrated that there exists longitudinal structural heterogeneity in the triple-helical domain, which is in good agreement with experimental results (Bodian et al. 2011). More recently, Orgel et al. (2014) investigated the symmetric property of collagen type I and type II triple helix and provided evidence for a symmetric range of 6.0–8.6 nm in native environment. This further confirms the structural heterogeneity of the collagen molecule in the longitudinal direction. In the past few decades, both experimental and computational studies have been conducted to investigate the mechanical properties of single collagen molecules. Detailed information on the adopted techniques, models/samples, strain rates, strain ranges, and the obtained Young’s modulus in those studies are summarized in Table 1. As illustrated in Table 1, the range of the Young’s modulus estimated in previous computational studies is rather broad, varying from 2.4 to 18.82 GPa. This wide range may originate from different strain ranges and strain rates used to determine the elastic properties. In addition, due to the small timescales of MD and the limit of computer power, there is a significant gap between the loading strain rates in computational studies (\(1.25\times 10^{8}\)–\(1.25\times 10^{12}\%\,\hbox {s}^{-1})\) and those employed in experimental characterizations (on order of several or tens of 10–100%\(\,\hbox {s}^{-1})\) (Sun et al. 2004; Bozec and Horton 2005), which tend to be more close to the physiologically relevant deformation rates. Nevertheless, the Young’s modulus of an 8-nm-long collagen molecule decreases with the decrease in the strain rate, but finally converges to approximately 4 GPa for strain rates below \(6.25\times 10^{9}\%\,\hbox {s}^{-1}\) (Gautieri et al. 2009a). This suggests that it is possible to capture the mechanical properties of a single collagen molecule at physiologically relevant deformation rates using atomistic MD simulations, when the strain rate employed is under a critical value (Gautieri et al. 2009a). Therefore, it is of great significance to employ a suitable strain rate, under which the mechanical properties converge, in SMD simulations to obtain the reliable mechanical behaviour and properties of the collagen molecules.
In summary, the longitudinal mechanical properties are heterogeneous in collagen fibrils. To probe the origin of this longitudinal mechanical heterogeneity at the molecular level, Zhou et al. found that 8-nm-long collagen molecule segments in the gap regions of the fibrils have an average Young’s modulus of 4.6 GPa, 45% smaller than that of the overlap regions (Zhou et al. 2015). However, it is not well understood how the longitudinal heterogeneous nanomechanics of the collagen fibril originate from the structural difference among the collagen molecule segments lie in the gap and overlap regions of the collagen fibril. Motivated by this gap, we focus on investigating the mechanical behaviour of the intact gap and overlap regions in type I collagen molecule under tensile loads, aiming to elucidate the structural origin of the longitudinal mechanical heterogeneity of collagen fibrils at the molecular level.
2 Computational model and method
As the building block of collagen fibrils, the super-twisted and right-handed microfibril is found to have a unique repeating arrangement (each repeating arrangement is referred to one D-period), with the overlap region (consisting of 5 molecules) and the gap region (comprising 4 molecules) distributing alternatively along its length direction (Orgel et al. 2006). At the molecular level, the ‘gap’ and ‘overlap’ regions are defined in reference to the microfibril structure, i.e. the ‘overlap’ region of the collagen molecule is referred to the region of the molecule that will be located in the overlap region when arranged into a microfibril, and similarly for the ‘gap’ region.
The signature structure of type I collagen molecule is the unique amino acid sequence within each of the three chains, with Glycine (Gly) being prerequisite at every third residue. Based on the experimentally verified in situ full length type I collagen molecule \(\hbox {C}_{\alpha }\)-atoms model (3HR2), we generated the backbone structure using SABBAC (Maupetit et al. 2006) (http://mobyle.rpbs.univ-paris-diderot.fr/cgi-bin/portal.py#forms::SABBAC), and added the side chains to the backbone structure with Scwrl4 (http://dunbrack.fccc.edu/scwrl4/SCWRL4.php). Then, we mutated the hydroxylated proline into hydroxyproline using Accelrys Discovery Studio Visualization (http://accelrys.com/products/collaborative-science/biovia-discovery-studio/visualization-download.php) to build the full-atomistic collagen molecule structure (Fig. 1). In addition, we built the collagen microfibril structure (Fig. 1) by copying and translating the generated full-atomistic collagen molecule structure using script MakeMultimer.py (http://watcut.uwaterloo.ca/tools/makemultimer/), based on the packing data of collagen molecules in the microfibril determined from X-ray fibre diffraction experiments. Lastly, we extracted the ‘gap’ and ‘overlap’ regions of one collagen molecule located in the second, third and fourth D-period of the developed microfibril. The resulting three ‘gap’ regions (d2gp, d3gp and d4gp) contain 42 Gly-X-Y triplets, while the three ‘overlap’ regions (d2ol, d3ol and d4ol) are composed of 36 Gly-X-Y triplets. To the best of our knowledge, no previous studies investigated the mechanical properties of the collagen molecule intact ‘gap’ and ‘overlap’ regions specifically.
In this study, all MD simulations are carried out using GROMACS 5.0.4 (Berendsen et al. 1995) with GROMOS96 54a7 force field, which has been successfully employed in many previous studies to simulate collagen molecules (Lorenzo and Caffarena 2005; Gautieri et al. 2009a; Zhou et al. 2015).
Each collagen molecule segment is fully solvated using a triclinic SPC (Berendsen et al. 1981) water box, making sure that there is a 1.0 nm water boundary on all sides. For the charged protein models, appropriate number of counter-ions (\(\hbox {Cl}^{-}\) and \(\hbox {Na}^{+}\) ions) are added to neutralize the whole system. Periodic boundary conditions are applied to all the directions. Covalent bond lengths involving hydrogen bonds (H-bonds) are constrained using the SETTLE and LINCS (Hess et al. 1997) algorithms, allowing a time step of 2 fs. Non-bonded interactions are calculated using a cut-off distance for neighbour lists at 1.2 nm, with a switching function between 1.0 and 1.2 nm. The fourth-order Particle-Mesh Ewald sums (PME) method (Essmann et al. 1995) is employed to calculate electrostatic interactions, with a columbic cut-off distance of 1.2 nm. The steepest descent algorithm is applied to minimize the energy of the system. To achieve a good starting point for MD simulations, each model is further equilibrated in the following three steps: firstly, a 30 ns NVT MD simulation is carried out at a temperature of 310 K (\(37\,^{\circ }\hbox {C}\)) using velocity-rescaling algorithm with 1 ps coupling constant; secondly, a 30 ns NPT MD simulation is performed at a pressure of 1 bar and a temperature of 310 K, in which Berendsen barostat with 1 ps coupling constant is employed and the whole protein is held fixed to relax water molecules; finally, another 20 ns NPT MD simulation is used to equilibrate the system with the more accurate Parrinello–Rahman pressure coupling algorithm (Parrinello and Rahman 1981), where only the first and last \(\hbox {C}_{\alpha }\)-atoms of each polypeptide chain are restrained. This results in the equilibrated ‘gap’ and ‘overlap’ regions with a length of approximately 36 and 31 nm, respectively.
To investigate whether the equilibrated ‘gap’ and ‘overlap’ regions carry similar structural disorder degree in comparison with those located within a fibril, we calculated the average helical radius and the average number of residues per turn for both the equilibrated collagen molecule segments and those located in 3HR2.
The radius of the triple helix is calculated as the radius of the circle encompassed by the three \(\hbox {C}_{\alpha }\)-atoms in three polypeptides of the triple helix, and the schematic representation of the definition of radius is illustrated in Gopalakrishnan et al. (2015); the average number of residues per turn was calculated using the algorithm developed in Ravikumar et al. (2007) using MATLAB.
In previous SMD simulation studies on collagen molecules, one end of the collagen molecule is kept fixed and the other end is linked to an elastic spring, moving along the molecular axis with specific velocities. In this study, we tried to use another type of SMD method, where the centre of mass (COM) of the three N-terminal \(\hbox {C}_{\alpha }\)-atoms and the COM of the three C-terminal \(\hbox {C}_{\alpha }\)-atoms are connected by a virtual spring with an elastic constant of \(1000\, \hbox {kJ mol}^{-1}\, \hbox {nm}^{-2}\), which is extended along the molecular axis with a reasonable deformation rate.
All SMD simulations are performed in an NVT ensemble, in which the systems are coupled to a heat bath at 310 K with velocity-rescale algorithm and a pressure coupling constant of 1 ps. In all SMD simulations, the integration time step is 2 fs.
Gautieri et al. reported that the Young’s modulus of an 8-nm-long collagen molecule converges to around 4 GPa for the strain rates under \(6.25 \times 10^{9}\%\,\hbox {s}^{-1}\) Gautieri et al. (2009a). Recently, Pradhan et al. claimed that the length of the collagen molecule has a significant effect on the mechanical properties of the collagen molecule (Pradhan et al. 2011). Therefore, it is essential to determine a suitable strain rate for the three intact ‘gap’ and three intact ‘overlap’ regions with different lengths, under which the mechanical properties of collagen molecule segments under physiological stressed conditions can be achieved. To achieve this, the ‘overlap’ region in the second D-period (d2ol) is stretched under various tensile strain rates by performing SMD simulations. Then, the mechanical response of the ‘gap’ and ‘overlap’ regions under tension is characterized with the determined suitable strain rate, using SMD simulation method. To validate the convergence, we performed three simulations with different initial configurations for each case.
3 Results and discussion
3.1 Validation of equilibrated structure
Table 2 displays the average helical radius and the average number of residues per turn for both the equilibrated collagen molecule segments and those located in 3HR2. As we can see from this Table 2, there is marginal structural difference between the equilibrated ‘gap’ and ‘overlap’ regions and the corresponding segments in 3HR2 in terms of the average helical radius and the average number of residues per turn. This indicates that the equilibrated ‘gap’ and ‘overlap’ regions conserve similar structural disorder degree apart from the amino acid sequence as compared to those within 3HR2. More specifically, the average radius for the equilibrated segments and those located in 3HR2 ranges from 4.55 to 4.90 Å and from 4.67 to 5.08 Å, respectively, while the average number of residues per turn varies between 3.30 and 3.51 and between 3.33 and 3.56, respectively. The ranges for the average helical radius obtained in this study are within the average helical radius range of 2.8–7.3 Å which was reported for glycine residues in the full triple-helical range of human type I collagen molecule in Bodian et al. (2011). In contrast, the average number of residues per turn is smaller than 4.9 which is determined for the full triple-helical range of human type I collagen molecule in Bodian et al. (2011). This discrepancy may come from different initial structures, different simulation boundary conditions and different force fields used in the simulations.
3.2 Critical strain rate in MD models
Figure 2a shows the force–deformation curves for d2ol (31-nm-long) under various tensile strain rates. As indicated in Fig. 2a, the force–strain behaviour of the collagen molecule segment is intimately related to the strain rate, but converges when the strain rate is under \(1.3\times 10^{8}\%\,\hbox {s}^{-1}\). This agrees with the results reported by Gautieri et al. that the deformation behaviour of an 8-nm-long collagen-like molecule under physiological tension can be captured when the strain rate used is smaller than \(6.25\times 10^{9}\%\,\hbox {s}^{-1}\) Gautieri et al. (2009a). Based on the force–strain curves, the tangent Young’s modulus of d2ol was evaluated at the strain of 10.5%, when the initially crimped collagen molecule has been straightened out. Figure 2b displays the Young’s modulus of d2ol under different loading strain rates. As illustrated in Fig. 2b, the Young’s modulus converges to around 3.2 GPa when the strain rate is less than \(1.3\times 10^{8}\%\,\hbox {s}^{-1}\), which further confirms that the mechanical properties of the collagen molecule under physiologically stressed conditions can be accomplished using SMD method with suitable strain rates. This is consistent with the work by Gautieri et al., where the Young’s modulus of an 8-nm-long collagen-like molecule converges to approximately 4 GPa for strain rates smaller than \(6.25\times 10^{9}\%\,\hbox {s}^{-1}\) Gautieri et al. (2009a). We note that the convergent strain rates for the mechanical properties are different between d2ol investigated in this study and the 8-nm-long collagen-like molecule studied in Gautieri et al. (2009a). This difference originates from the fact that d2ol is defined by the amino acid sequence with physiological diversity and is almost four times as long as the collagen-like molecule studied in Gautieri et al. (2009a).
a Force–strain curves for d2ol under different tensile strain rates. The inset figure displays the force–strain curves for d2ol for strains up to 16%. At low strain rate, under which the deformation behaviour converges, the force is nearly negligible until the tensile strain reaches up to around 10.5%, and then it increases nonlinearly and goes up linearly with a higher speed for strains larger than approximately 30%. This indicates that the tensile behaviour of the collagen molecule is highly nonlinear. This may explain the broad range of the Young’s modulus obtained for collagen molecules in previous studies as summarized in Table 1. b Young’s modulus of d2ol with respect to tensile strain rates. Evaluated at the tensile strain of 10.5%, when the initially crimped collagen molecule segment has almost been straightened out. The trend of the curve indicates that the Young’s modulus of the collagen molecule segment decreases as the strain rate goes down, but finally converges to a certain value. Once the Young’s modulus reaches the convergent value (indicated with red dashed line), its fluctuations are within the error of \(\pm 0.1\,\hbox {GPa}\)
3.3 Mechanical response under tensile loading
Figure 3 displays force variations for the three intact ‘gap’ and three intact ‘overlap’ regions of the type I collagen molecule with respect to strains up to 25%. The pulling velocities used for the ‘gap’ and ‘overlap’ regions are approximately 0.0234 and 0.02015 m/s, respectively, based on the length of the molecules. The modelling strategy guarantees a same strain rate (\(6.5\times 10^{7}\%\,\hbox {s}^{-1})\) for both ‘gap’ regions (around 36-nm-long) and ‘overlap’ regions (around 31-nm-long), allowing a reasonable comparison of the tensile behaviours in different regions. The strain rates used in this study are close to half of the smallest strain rate employed in the previous studies as summarized in Table 1, where an 8-nm-long collagen-like molecule was stretched under a strain rate of \(1.25 \times 10^{8}\%\,\hbox {s}^{-1}\). It takes 1.5 months for the ‘gap’ region computational system, which consists of \(\sim 160000\) atoms, to reach the strain of 25% using 64 CPUs. In this study, the length of collagen molecule segment refers to the distance between the COM of the three N-terminal \(\hbox {C}\upalpha \)-atoms and the COM of the three C-terminal \(\hbox {C}\upalpha \)-atoms. The tensile strain is defined as the ratio of elongation to the original length of the collagen molecule segment.
Force–strain relationship for the three intact gap and three intact overlap regions under a tensile strain rate of \(6.5\times 10^{7}\%\,\hbox {s}^{-1}\). The stiffness of the collagen molecule segments increases as the strain increases, showing a highly nonlinear deformation behaviour under tensile loads. When the tensile strains reach up to approximately 12–15%, the force–strain data start to show a zigzag-like appearance. In this process, the stiffness varies among different collagen molecule segments
From Fig. 3, the overall mechanical behaviour of the intact ‘gap’ and ‘overlap’ regions under tensile loads is similar. After unwinding (molecular rotating), each of the six collagen molecule segments undergoes an uncoiling (H-bonds breaking) process under tension. More specifically, the collagen molecule segment is crimped at the beginning. While stretched, the initially crimped collagen segment unwinds until it reaches the strain of around 10–15%. During this period, the tensile force applied on the collagen molecule segment is almost negligible (smaller than 0.35 nN) and the triple-helical structure of the protein is nearly preserved. After having been straightened out, the collagen triple-helical structure starts to uncoil. During this stage, the tensile force on the collagen molecule segment increases gradually as the strain goes up. Here we note that the force–strain data for the six collagen molecule segments fluctuate in this molecule uncoiling stage. This fluctuation has been found in previous studies (Lorenzo and Caffarena 2005; Gautieri et al. 2009a; Zhou et al. 2015) and is inferred to be from the intrinsic property of SMD simulation method. The deformation mechanisms (unwinding followed by uncoiling) of the six collagen molecule fragments found in this study are consistent with the work reported in both experimental study (Fratzl et al. 1998) and molecular modelling (Gautieri et al. 2009a; Zhou et al. 2015; Vesentini et al. 2013). Herein, we did not investigate the third deformation stage (backbone stretching stage) reported in those studies, due to the limited computational resources and the consideration that the cross-links transferring the load between single collagen molecules may rupture before the molecules undergo that much strain.
3.4 Stress–strain response and Young’s modulus
As illustrated in Fig. 3, there is a significant tension stiffening effect in the collagen molecule segments. Hence, it is of great significance to determine the strain range for evaluating the Young’s modulus of those collagen segments. However, it is quite difficult to know the exact strains, at which the collagen molecule segments have been straightened out. In previous studies, two types of methods have been reported to determine the Young’s modulus of collagen molecules. One is linear regression of the stress–strain data (Lorenzo and Caffarena 2005; Pradhan et al. 2011; Zhou et al. 2015), and the other one is the first-order derivative of the fitted stress–strain curve (Buehler 2006a; Gautieri et al. 2009a; Uzel and Buehler 2009). The most crucial and challenging step is how to determine the strain range/point used to derive the Young’s modulus. In this study, the tangent Young’s modulus was calculated as the first-order derivative of the fitted stress–strain curve.
The stress–strain (\(\sigma -\varepsilon )\) data for the three intact ‘gap’ and three intact ‘overlap’ regions are obtained based on the corresponding force–strain data, assuming that those collagen molecule fragments are cylindrical with a diameter of 1.5 nm. Then, a fourth-order polynomial (\(\sigma =\hbox {a}_0 + \hbox {a}_1 \varepsilon + \hbox {a}_2 \varepsilon ^{2} + \hbox {a}_3 \varepsilon ^{3} + \hbox {a}_4 \varepsilon ^{4})\) is fitted to the derived stress–strain data for both strains up to 25% and strains up to 12%, resulting in the fitted stress–strain curves for the six collagen molecule segments displayed in Fig. 4. The fourth-order polynomial fits the stress–strain data well as indicated in Fig 5. Here we only give the ‘gap’ and ‘overlap’ region in the second D-period as an example. As we can see from Fig. 4, when the strains are smaller than 2–6.5%, the stresses within the six collagen molecule segments remain at approximately 0.05–0.06 GPa, which is almost negligible. This is due to the fact that each of the six collagen molecule segments studied has an initially crimped structure. Then, the stresses in the collagen molecule segments increase slowly as the strains go up to \(\sim \)10–15%, when the collagen molecule segments are almost straightened out. During this stage (unwinding stage), the stresses in the ‘overlap’ regions (green, blue and magenta) increase more quickly than those in the ‘gap’ regions (red, cyan and black), showing a higher stiffness as displayed in Fig. 4. This is in good agreement with the structural difference between the ‘gap’ and ‘overlap’ regions as shown in Fig. 1 and reported in the collagen molecule crystal structure (Orgel et al. 2006): the ‘gap’ region is relatively more disordered as compared to the ‘overlap’ region. When having been straightened out, the collagen molecule segments start to uncoil and the tensile stresses increase gradually with a higher loading rate as the strains increase, indicating that the collagen molecule segments are stiffer in this uncoil stage than in the unwinding stage. This higher stiffness is believed to be originated from the effect of inter-chain hydrogen bond breaking process (Gautieri et al. 2009a; Zhou et al. 2015). During this stage, each of the six collagen molecule segments represents different stiffness under tensile loads.
Stress–strain relationship (raw data and fourth-order polynomial fitted curves) for d2ol and d2gp under a tensile strain rate of \(6.5\times 10^{7}\%\,\hbox {s}^{-1}\). The gap and overlap regions show similar deformation behaviour, but the overlap region shows a higher stiffness. The inset figure displays the stress–strain relationship for d2ol and d2gp for strains up to 12%
The tangent Young’s modulus (E) for the six collagen molecule segments with respect to strain is estimated by calculating the first-order derivative (\(E=\hbox { a}_1 + \hbox {2a}_2 \varepsilon + \hbox {3a}_3 \varepsilon ^{2} + \hbox {4a}_4 \varepsilon ^{3})\) of the fitted fourth-order polynomial stress–strain curves as displayed in Fig. 6. As we can see from Fig. 6, the Young’s modulus of the collagen fragments varies with respect to the tensile strain, which indicates that the collagen molecule segments present a highly nonlinear elastic behaviour. This is consistent with the work reported in Gautieri et al. (2009a), Zhou et al. (2015) Gautieri et al. (2013) and Vesentini et al. (2013)). Therefore, it is of great importance to determine the critical strains suitable to derive the Young’s modulus of the collagen segments. To achieve this, we investigated the variation of the inter-chain H-bonds number and the protein potential energy with respect to time during the SMD simulations. We found that both the inter-chain H-bonds number and the protein potential energy remain stable for strains up to \(\sim \)10–15%, where the inter-chain H-bonds number starts to decrease and the protein potential energy starts to increase as the segments undergo further stretching. Then, the tangent Young’s modulus of the collagen molecule segments is calculated at the critical strains, where the segments start to undergo the uncoiling process. Here we take the second D-period as an example. Figure 7 displays the variation of the inter-chain H-bonds number and the protein potential energy with respect to time during the SMD simulations. As indicated in Fig. 7, the number of inter-chain H-bonds and the protein potential energy remain stable for strains up to approximately 10.5%. Therefore, the tangent Young’s modulus of d2ol is calculated as the first-order derivative of the fitted fourth-order polynomial stress–strain curve for d2ol at the strain of 10.5%. The determined critical strain (\(\varepsilon )\) and the evaluated corresponding tangent Young’s modulus (E) of the six collagen molecule segments are summarized in Table 3. The strain and tangent Young’s modulus displayed in Table 3 are averaged from those obtained in three different simulations with different initial structure for each case. As indicated in Table 3 and Fig. 4, the ‘gap’ regions undergo a marginally larger strains than the ‘overlap’ regions in the unwinding stage before having been straightened out. To the best of our knowledge, this has not been reported in the previous literature. This is understandable due to the fact that the three polypeptide chains arrange less in order in the ‘gap’ regions as compared to those in the ‘overlap’ regions. The difference in the unwinding strain ranges between the ‘gap’ and ‘overlap’ regions may originate from their longitudinal structural variation in term of helical radius reported by Bodian et al. (2011). Besides, the estimated tangent Young’s modulus for the six collagen molecule segments ranges between 3.209 and 4.908 GPa. This is in good agreement with the reported experimental values (Harley et al. 1977; Cusack and Miller 1979; Hofmann et al. 1984; Sasaki and Odajima 1996; Sun et al. 2002) as well as those published in MD simulation studies (Vesentini et al. 2005; Lorenzo and Caffarena 2005; Buehler 2006a; Stevens 2008; Gautieri et al. 2008, 2009a; Uzel and Buehler 2009; Gautieri et al. 2010; Pradhan et al. 2011; Zhou et al. 2015; Gautieri et al. 2012a, 2013; Vesentini et al. 2013) as summarized in Table 1. In addition, the Young’s modulus of the six collagen molecule segments is different with each other. This is justified by the fact that the amino acid sequence defining the collagen molecule have a significant effect on its mechanical properties (Vesentini et al. 2005; Uzel and Buehler 2009). This may partly originate from the structural difference including number of residues in each turn and H-bond length and strength, due to different amino acid sequence in the three collagen polypeptides. For detailed information, we refer to work reported in Bodian et al. (2011). This further validates the reliability of the computational model developed in this study.
Number of inter-chain H-bonds and potential energy for d2ol with respect to strain under a tensile strain rate of \(6.5\times 10^{7}\%\,\hbox {s}^{-1}\). In the first unwinding stage, the number of inter-chain H-bonds and potential energy of the collagen segments remain stable until the strain reaches up to around 10.5%. After the collagen molecule segment becomes straight, the inter-chain H-bonds start to break as the strain increases; thus, the number of H-bonds decreases gradually in the uncoiling stage. The profile is consistent with the work reported in Gautieri et al. (2009a). In contrast, the protein potential energy starts to go up, due to the breaking of inter-chain H-bonds and the increase in the Lennard-Jones potential contributions
4 Conclusion
By performing SMD simulations, we investigated the mechanical behaviours of six intact ‘gap’ (approximately 31 nm long) and ‘overlap’ (approximately 36 nm long) regions of a type I collagen molecule using a strain rate of \(6.5\times 10^{7}\%\,\hbox {s}^{-1}\), under which their mechanical behaviours under physiological relevant stress conditions can be achieved. This research focus on characterizing the longitudinal heterogeneous mechanical properties in the ‘gap’ and ‘overlap’ regions in collagen molecules.
This study probed the mechanical properties of collagen molecule segments and investigated the structural origin of the longitudinal heterogeneous mechanical properties of the collagen fibril at the molecular level. The mechanisms of collagen deformation and the quantification of Young’s modulus are consistent with the results reported in previous experimental and numerical studies. Heterogeneous mechanical properties were found along the length direction of the collagen molecule: the ‘gap’ and ‘overlap’ regions have different strain ranges corresponding to the unwinding stage and their tangent Young’s modulus differ from each other. This longitudinal heterogeneity may explain the larger Young’s modulus obtained from the 2D and 3D CG bead-spring models developed previously (Depalle et al. 2014; Buehler 2008), in which the collagen molecule is assumed to be homogeneous along its length direction. The CG bead-spring model for the collagen molecule and fibril with higher accuracy will need to be developed, by considering the entropic effect, the longitudinal heterogeneity of the collagen molecule and using suitable pulling rates, under which the physiological mechanical properties can be achieved.
References
Baldwin SJ, Quigley AS, Clegg C, Kreplak L (2014) Nanomechanical mapping of hydrated rat tail tendon collagen I fibrils. Biophys J 107:1794–1801
Balooch M, Habelitz S, Kinney J, Marshall S, Marshall G (2008) Mechanical properties of mineralized collagen fibrils as influenced by demineralization. J Struct Biol 162:404–410
Berendsen HJ, Postma JP, van Gunsteren WF, Hermans J (1981) Interaction models for water in relation to protein hydration. In: Pullman B (ed) Intermolecular forces. Springer, Berlin, pp 331–342
Berendsen HJ, van der Spoel D, van Drunen R (1995) GROMACS: a message-passing parallel molecular dynamics implementation. Comput Phys Commun 91:43–56
Bodian DL, Radmer RJ, Holbert S, Klein TE (2011) Molecular dynamics simulations of the full triple helical region of collagen type I provide an atomic scale view of the protein’s regional heterogeneity. In: Pacific symposium on biocomputing. pacific symposium on biocomputing, p. 193
Bozec L, Horton M (2005) Topography and mechanical properties of single molecules of type I collagen using atomic force microscopy. Biophys J 88:4223–4231
Buehler MJ (2006a) Atomistic and continuum modeling of mechanical properties of collagen: elasticity, fracture, and self-assembly. J Mater Res 21:1947–1961
Buehler MJ (2006b) Nature designs tough collagen: explaining the nanostructure of collagen fibrils. Proc Natl Acad Sci 103:12285–12290
Buehler MJ (2008) Nanomechanics of collagen fibrils under varying cross-link densities: atomistic and continuum studies. J Mech Behav Biomed Mater 1:59–67
Cusack S, Miller A (1979) Determination of the elastic constants of collagen by Brillouin light scattering. J Mol Biol 135:39–51
Depalle B, Qin Z, Shefelbine SJ, Buehler MJ (2014) Influence of cross-link structure, density and mechanical properties in the mesoscale deformation mechanisms of collagen fibrils. J Mech Behav Biomed Mater 52:1–13
Eppell S, Smith B, Kahn H, Ballarini R (2006) Nano measurements with micro-devices: mechanical properties of hydrated collagen fibrils. J R Soc Interface 3:117–121
Essmann U, Perera L, Berkowitz ML, Darden T, Lee H, Pedersen LG (1995) A smooth particle mesh Ewald method. J Chem Phys 103:8577–8593
Fratzl P, Misof K, Zizak I, Rapp G, Amenitsch H, Bernstorff S (1998) Fibrillar structure and mechanical properties of collagen. J Struct Biol 122:119–122
Gautieri A, Vesentini S, Montevecchi FM, Redaelli A (2008) Mechanical properties of physiological and pathological models of collagen peptides investigated via steered molecular dynamics simulations. J Biomech 41:3073–3077
Gautieri A, Buehler MJ, Redaelli A (2009a) Deformation rate controls elasticity and unfolding pathway of single tropocollagen molecules. J Mech Behav Biomed Mater 2:130–137
Gautieri A, Vesentini S, Redaelli A, Buehler MJ (2009b) Single molecule effects of osteogenesis imperfecta mutations in tropocollagen protein domains. Protein Sci 18:161–168
Gautieri A, Russo A, Vesentini S, Redaelli A, Buehler MJ (2010) Coarse-grained model of collagen molecules using an extended MARTINI force field. J Chem Theory Comput 6:1210–1218
Gautieri A, Vesentini S, Redaelli A, Buehler MJ (2011) Hierarchical structure and nanomechanics of collagen microfibrils from the atomistic scale up. Nano Lett 11:757–766
Gautieri A, Vesentini S, Redaelli A, Buehler MJ (2012a) Viscoelastic properties of model segments of collagen molecules. Matrix Biol 31:141–149
Gautieri A, Vesentini S, Redaelli A, Buehler MJ (2012b) Osteogenesis imperfecta mutations lead to local tropocollagen unfolding and disruption of H-bond network. RSC Adv 2:3890–3896
Gautieri A, Vesentini S, Redaelli A, Ballarini R (2013) Modeling and measuring visco-elastic properties: from collagen molecules to collagen fibrils. Int J Non-Linear Mech 56:25–33
Gentleman E, Lay AN, Dickerson DA, Nauman EA, Livesay GA, Dee KC (2003) Mechanical characterization of collagen fibers and scaffolds for tissue engineering. Biomaterials 24:3805–3813
Gopalakrishnan R, Singam EA, Sundar JV, Subramanian V (2015) Interaction of collagen like peptides with gold nanosurfaces: a molecular dynamics investigation. Phys Chem Chem Phys 17:5172–5186
Graham JS, Vomund AN, Phillips CL, Grandbois M (2004) Structural changes in human type I collagen fibrils investigated by force spectroscopy. Exp Cell Res 299:335–342
Gupta H, Messmer P, Roschger P, Bernstorff S, Klaushofer K, Fratzl P (2004) Synchrotron diffraction study of deformation mechanisms in mineralized tendon. Phys Rev Lett 93:158101
Harley R, James D, Miller A, White J (1977) Phonons and the elastic moduli of collagen and muscle. Nat 267:285–287. doi:10.1038/267285a0
Heim AJ, Matthews WG, Koob TJ (2006) Determination of the elastic modulus of native collagen fibrils via radial indentation. Appl Phys Lett 89:181902
Hess B, Bekker H, Berendsen HJ, Fraaije JG (1997) LINCS: a linear constraint solver for molecular simulations. J Comput Chem 18:1463–1472
Hodge AJ, Schmitt FO (1960) The charge profile of the tropocollagen macromolecule and the packing arrangement in native-type collagen fibrils. Proc Natl Acad Sci USA 46:186
Hofmann H, Voss T, Kühn K, Engel J (1984) Localization of flexible sites in thread-like molecules from electron micrographs: comparison of interstitial, basement membrane and intima collagens. J Mol Biol 172:325–343
Hulmes D (2008) Collagen diversity, synthesis and assembly. In: Fratzl P (ed) Collagen. Springer, Berlin, pp 15–47
Husar-Memmer E, Ekici A, Al Kaissi A, Sticht H, Manger B, Schett G et al (2013) Premature osteoarthritis as presenting sign of type II collagenopathy: a case report and literature review. In: Seminars in arthritis and rheumatism, pp 355–360
Jenkins CL, Raines RT (2002) Insights on the conformational stability of collagen. Nat Prod Rep 19:49–59
Kadler KE, Baldock C, Bella J, Boot-Handford RP (2007) Collagens at a glance. J Cell Sci 120:1955–1958
Kim J-H, Lee G, Won Y, Lee M, Kwak J-S, Chun C-H et al (2015) Matrix cross-linking-mediated mechanotransduction promotes posttraumatic osteoarthritis. Proc Natl Acad Sci 112:9424–9429
Leistritz DF, Pepin MG, Schwarze U, Byers PH (2011) COL3A1 haploinsufficiency results in a variety of Ehlers–Danlos syndrome type IV with delayed onset of complications and longer life expectancy. Genet Med 13:717–722
Lindahl K, Åström E, Rubin C-J, Grigelioniene G, Malmgren B, Ljunggren Ö et al (2015) Genetic epidemiology, prevalence, and genotype-phenotype correlations in the Swedish population with osteogenesis imperfecta. Eur J Hum Genet 23:1042–1050
Lorenzo AC, Caffarena ER (2005) Elastic properties, Young’s modulus determination and structural stability of the tropocollagen molecule: a computational study by steered molecular dynamics. J Biomech 38:1527–1533
Maupetit J, Gautier R, Tufféry P (2006) SABBAC: online Structural Alphabet-based protein BackBone reconstruction from Alpha-Carbon trace. Nucleic Acids Res 34:W147–W151
Minary-Jolandan M, Yu M-F (2009) Nanomechanical heterogeneity in the gap and overlap regions of type I collagen fibrils with implications for bone heterogeneity. Biomacromolecules 10:2565–2570
Orgel JP, Irving TC, Miller A, Wess TJ (2006) Microfibrillar structure of type I collagen in situ. Proc Natl Acad Sci 103:9001–9005
Orgel JP, Persikov AV, Antipova O (2014) Variation in the helical structure of native collagen. PloS ONE 9:e89519
Parrinello M, Rahman A (1981) Polymorphic transitions in single crystals: a new molecular dynamics method. J Appl Phys 52:7182–7190
Pradhan SM, Katti DR, Katti KS (2011) Steered molecular dynamics study of mechanical response of full length and short collagen molecules. J Nanomech Micromech 1:104–110
Ravikumar K, Humphrey J, Hwang W (2007) Spontaneous unwinding of a labile domain in a collagen triple helix. J Mech Mater Struct 2:999–1010
Robin NH, Moran RT, Ala-Kokko L (2014) Stickler syndrome In: Pagon RA, Adam MP, Ardinger HH et al (eds) GeneReviews(r) [Internet], 1993–2016. University of Washington, Seattle. http://www.ncbi.nlm.nih.gov/books/NBK1302/
Robinson R, Watson M (1952) Collagen–crystal relationships in bone as seen in the electron microscope. Anat Rec 114:383–409
Sasaki N, Odajima S (1996) Elongation mechanism of collagen fibrils and force–strain relations of tendon at each level of structural hierarchy. J Biomech 29:1131–1136
Shen ZL, Dodge MR, Kahn H, Ballarini R, Eppell SJ (2008) Stress–strain experiments on individual collagen fibrils. Biophys J 95:3956–3963
Sherman VR, Yang W, Meyers MA (2015) The materials science of collagen. J Mech Behav Biomed Mater 52:22–50
Spitzner E-C, Röper S, Zerson M, Bernstein A, Magerle R (2015) Nanoscale swelling heterogeneities in type I collagen fibrils. ACS Nano 9:5683–5694
Stevens MJ (2008) Simulation of the mechanical strength of a single collagen molecule. Biophys J 95:33–39
Streeter I, de Leeuw NH (2011) A molecular dynamics study of the interprotein interactions in collagen fibrils. Soft Matter 7:3373–3382
Sun Y-L, Luo Z-P, Fertala A, An K-N (2002) Direct quantification of the flexibility of type I collagen monomer. Biochem Biophys Res Commun 295:382–386
Sun Y-L, Luo Z-P, Fertala A, An K-N (2004) Stretching type II collagen with optical tweezers. J Biomech 37:1665–1669
Uzel SG, Buehler MJ (2009) Nanomechanical sequencing of collagen: tropocollagen features heterogeneous elastic properties at the nanoscale. Integr Biol 1:452–459
van der Rijt JA, van der Werf KO, Bennink ML, Dijkstra PJ, Feijen J (2006) Micromechanical testing of individual collagen fibrils. Macromol Biosci 6:697–702
Vesentini S, Fitié CF, Montevecchi FM, Redaelli A (2005) Molecular assessment of the elastic properties of collagen-like homotrimer sequences. Biomech Model Mechanobiol 3:224–234
Vesentini S, Redaelli A, Gautieri A (2013) Nanomechanics of collagen microfibrils. Muscles Ligaments Tendons J 3:23
Weiner S, Wagner HD (1998) The material bone: structure-mechanical function relations. Annu Rev Mater Sci 28:271–298
Wenger MP, Bozec L, Horton MA, Mesquida P (2007) Mechanical properties of collagen fibrils. Biophys J 93:1255–1263
Wenger MP, Mesquida P (2011) Longitudinal variations in the Poisson’s ratio of collagen fibrils. Appl Phys Lett 98:163707
Yang W, Sherman VR, Gludovatz B, Schaible E, Stewart P, Ritchie RO et al (2015) On the tear resistance of skin. Nat Commun 6
Zhou Z, Minary-Jolandan M, Qian D (2015) A simulation study on the significant nanomechanical heterogeneous properties of collagen. Biomech Model Mechanobiol 14(3): 445–457
Acknowledgements
The authors thank Joseph Orgel (Center For Molecular Study Of Condensed Soft Matter, Pritzker Institute of Biomedical Science and Engineering at the Illinois Institute of Technology and Argonne National Lab, USA) for his help on dividing the gap and overlap regions of the collagen molecule. This research was funded by ARC Discovery Project (DP150100828). The High Performance Computer resources provided by Queensland University of Technology (QUT) are gratefully acknowledged. Also the financial support of China Scholarship Council (CSC) scholarship from Chinese government and Top-up scholarship from QUT are greatly appreciated.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Tang, M., Li, T., Gandhi, N.S. et al. Heterogeneous nanomechanical properties of type I collagen in longitudinal direction. Biomech Model Mechanobiol 16, 1023–1033 (2017). https://doi.org/10.1007/s10237-016-0870-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10237-016-0870-6