Abstract
Background
Results of previous studies about the prognostic roles of histone H4 lysine 16 acetylation (H4K16ac) and histone H4 lysine 20 trimethylation (H4K20me3) in breast cancer were inconsistent. Cellular experiments revealed the interplays between H4K16ac and H4K20me3, but no population study explored the interaction between them on the prognosis.
Methods
H4K16ac and H4K20me3 levels in tumors were evaluated by immunohistochemistry for 958 breast cancer patients. Hazard ratios for overall survival (OS) and progression-free survival (PFS) were estimated using Cox regression models. Interaction was assessed on multiplicative scale. Concordance index (C-index) was calculated to verify the predictive performance.
Results
The prognostic roles of the low level of H4K16ac or H4K20me3 were significant only in patients with the low level of another marker and their interactions were significant. Moreover, compared with joint high levels of both them, only the combined low levels of both them was associated with a poor prognosis but not the low level of single one. The C-index of the clinicopathological model combined the joint expression of H4K16ac and H4K20me3 [0.739 for OS; 0.672 for PFS] was significantly larger than that of the single clinicopathological model [0.699 for OS, P < 0.001; 0.642 for PFS, P = 0.003] or the model combined with the single H4K16ac [0.712 for OS, P < 0.001; 0.646 for PFS, P < 0.001] or H4K20me3 [0.724 for OS, P = 0.031; 0.662 for PFS, P = 0.006].
Conclusions
There was an interaction between H4K16ac and H4K20me3 on the prognosis of breast cancer and the combination of them was a superior prognostic marker compared to the single one.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer is a highly heterogeneous disease ranging from premalignant hyperproliferation to invasive and metastatic carcinomas [1]. Various types of prognostic factors such as clinical stage, hormone receptor status and human epidermal growth factor receptor 2 (HER2) expression have been used to determine therapeutic approaches [2,3,4]. However, female breast cancer has surpassed lung cancer as the most commonly diagnosed cancer, which further leads to the increase in mortality [5, 6]. Therefore, other promising markers are still needed to predict breast cancer outcomes and optimize the therapies.
Epigenetic alterations such as DNA methylation, histone modifications and non-coding RNAs occur in many cancers [7, 8]. A genome-wide study revealed that the global loss of histone H4 lysine 16 acetylation (H4K16ac) and histone H4 lysine 20 trimethylation (H4K20me3) was a common hallmark of human cancer [9]. H4K16ac was a well-established epigenetic marker of active transcription and its loss was associated with the epigenetic silencing of oncogenes or tumor suppressor genes [10, 11], while H4K20me3 was associated with transcriptional repression and the loss could induce the expression of these genes [12, 13]. When H4K16ac and H4K20me3 co-regulate one gene (either oncogene or tumor suppressor gene), there would be interesting interplays between them [13].
Currently, a population study found that the loss of global H4K16ac was associated with a poor prognosis of breast cancer in only univariate analysis but not multivariate analysis [14]. For the prognostic roles of H4K20me3 in breast cancer, the findings were inconsistent [14, 15]. One study found no significant association [14], while another study found that the loss of global H4K20me3 was associated with a poor breast cancer prognosis [15]. These findings suggested that the prognostic roles of H4K16ac and H4K20me3 in breast cancer may be affected by other factors. Considering about the interplays between H4K16ac and H4K20me3 on breast cancer related genes [13], the prognostic roles of them may be affected by each other. Only one previous study simultaneously considered the prognostic roles of H4K16ac and H4K20me3 [14], but it didn’t explore the modification effects between them.
In the present study, therefore, we aimed to examine the interaction between the global H4K16ac and H4K20me3 in breast cancer tissues on the prognosis and explore the joint associations of them with the prognosis. For clinical use, we further explored that whether the joint expression of H4K16ac and H4K20me3 could improve the prediction power of the prognostic model using known clinical and pathologic factors.
Materials and methods
Study population
A total of 1062 female patients with pathologically diagnosed primary invasive breast cancer and > 1 cm of tumor size in diameter between January 2008 and December 2015 were recruited from the Cancer Center of Sun Yat-sen University in Guangzhou, China. Patients with metastatic tumor and missing information of the expression of H4K16ac and H4K20me3 in tumor tissues (N = 88) were excluded (Fig. S1, Online Resource). Of 974 eligible women, 958 (98.4%) were successfully followed up until Dec 31, 2020. This study was approved by the Ethics Committee of the School of Public Health at Sun Yat-sen University. Informed consent was obtained from each participant.
Baseline data collection
Information on demographic and clinicopathologic characteristics was collected at diagnosis from patients’ medical records, including age, menopausal status, body mass index (BMI), family history of breast cancer, clinical stage, histological grade, estrogen receptor (ER), progesterone receptor (PR) and HER2 status etc. The status of ER, PR, and HER2 was defined according to American Society of Clinical Oncology/College of American Pathologists (ASCO/CAP) guidelines [16].
Tissue microarray and Immunohistochemistry
The expression levels of H4K16ac and H4K20me3 were evaluated with tissue microarrays (TMAs) by immunohistochemistry (IHC). TMAs were constructed as previously described [17]. The TMAs were baked at 60 °C for 2 h and then dewaxed with xylene and ethanol. Then antigen retrieval was accomplished using EDTA (PH 9.0) in super-pressure kettle and endogenous peroxide was blocked using 3% H2O2. Antigen–antibody reactions for H4K16ac and H4K20me3 were performed separately. For H4K16ac, slides were incubated in rabbit monoclonal to H4K16ac [EPR1004] (ab109463, diluted 1:100, Abcam) and then labeled with the EnVision Detection System (Peroxidase/DAB, Rabbit/Mouse) (Dako K5007). For H4K20me3, slides were incubated in rabbit polyclonal to H4K20me3- Chip Grade (ab9053, diluted 1:400, Abcam). Then slides were developed by diaminobenzidine (DAB) and counterstained by hematoxylin. These slides were finally dehydrated and mounted.
IHC stained sections were digitally imaged using Pannoramic Scanner and CaseViewer software. IHC staining was analyzed by an experienced pathologist and scored for staining intensity (0-no staining, 1-weak, 2-moderate and 3-strong) and percentage of tumor cell staining (0–100). IHC scoring was done by H-score which was calculated by multiplying the staining intensity by the percentage of positive cells [18, 19]. Thus, the minimal H-score was 0, whereas the maximum H-score was 300. To avoid the observation variability, the mean value of duplicate scores was adapted for further analysis.
Follow up and Outcomes
Patients were followed up by phone calls or out-patient visits every 3 months in the first year, every 6 months in the second and third year after diagnosis and annually thereafter. Outcomes of interest were overall survival (OS) and progression-free survival (PFS). OS was defined as the time from diagnosis to death and PFS was the time from diagnosis to disease progression including recurrence, metastasis, and death. The deaths were confirmed by calling the first-degree relatives of the patients and searching the Death Registration Reporting Information System of Guangzhou Center for Disease Control and Prevention. Survival status was censored at the latest follow-up date or Dec 31, 2020.
Statistical analysis
Wilcoxon signed rank test was used to compare the expression levels of H4K16ac and H4K20me3 between tumor tissues and adjacent tissues. Kruskal–Wallis test and Mann–Whitney U test were used to test the associations of H4K16ac and H4K20me3 H-score (defined as a continuous variable) with age, menopausal status, BMI, family history, histological grade, tumor size, nodal status, clinical stage and expression of ER, PR and HER2. H4K16ac and H4K20me3 H-score were modeled as continuous variable and fitted in a Cox proportional hazard model using restricted cubic splines with knots at the 5th, 35th, 65th, and 95th percentiles to estimate the hazard ratios (HRs) and 95% confidence bands assuming proportional hazard. Then H4K16ac and H4K20me3 H-score were categorized according to the results of restricted cubic splines and the distribution of H4K16ac and H4K20me3 H-score. The optimal cut-off values of H4K16ac and H4K20me3 were determined by the minimum P value from log-rank chi-square statistics based on PFS using the X-tile 3.6.1 software (Yale University, New Haven, CT, USA) [20]. Kaplan–Meier method was used to estimate the 5-year survival. Cox proportional hazard model was used to estimate HRs and their 95% confidence intervals (CIs) for the associations between various prognostic variables and the survival (OS and PFS). Multiplicative scale was used to estimate the interaction between H4K16ac and H4K20me3 on the prognosis.
A combined model containing the known important breast cancer clinicopathological prognostic factors and the joint levels of H4K16ac and H4K20me3 was constructed for the prediction of the prognosis. The performance of the combined model was measured by concordance index (C-index) [21, 22]. Furthermore, a clinicopathological model containing only the known clinicopathological prognostic factors and the extra models combined only H4K16ac or H4K20me3 were constructed for comparison. Comparisons between the different models were performed with the rcorrp.cens package in Hmisc in R and were evaluated by the C-index [23]. The larger the C-index, the more accurate was the prognostic prediction [24]. All the analyses were conducted using R 3.6.3 and a two-sided P-value below 0.05 was considered as statistical significance.
Results
Low level of H4K16ac and H4K20me3 in breast cancer tissues
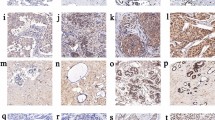
Of 958 women included in the analysis, almost all (99.0%) of them were pathologically diagnosed with invasive ductal carcinoma (IDC). Representative immunohistochemical staining of H4K16ac and H4K20me3 in tumor tissues was shown in Fig. 1. The level of H4K16ac in adjacent normal tissues was available in 552 patients. The H-score of H4K16ac ranged from 0 to 270.0 in both tumor tissues and adjacent tissues, and the median (P25, P75) in tumor tissues [5.0 (0.0, 40.0)] was significantly lower than that in adjacent tissues [60.0 (10.0, 140.0)] (P < 0.001) (Table 1). The level of H4K20me3 in adjacent normal tissues was available in 596 patients and the H-score ranged from 0 to 285.0 in both tumor tissues and adjacent tissues. Similarly, the median (P25, P75) in tumor tissues [90.0 (20.0, 210.0)] was significantly lower than that in adjacent tissues [255.0 (210.0, 255.0)] (P < 0.001).
Demographic and clinicopathological characteristics and the associations with H4K16ac and H4K20me3 in tumor tissues
The median age at diagnosis was 48 years (interquartile range: 41–56) among 958 eligible women. More than half (57.9%) of the women were premenopausal and 55.9% of them had a BMI between 18.5 and 23.9 kg/m2. The majority of the women were diagnosed with low histological grade (grade I/II: 73.4%), early clinical stage (stage I/II: 71.9%), ER-positive (73.3%), PR-positive (72.5%), or HER2-negative (66.8%) (Table 2).
H4K16ac level was lower in tumors with size > 2 cm, or HER2-negative than the level in tumors with size ≤ 2 cm, or HER2-positive (all P < 0.05). No marked differences in H4K16ac level were observed between different age, menopausal status, BMI, family history, histological grade, nodal status, clinical stage, ER status and PR status (Table 2). For H4K20me3, the level was lower in tumors with higher histological grade, size > 2 cm, higher clinical stage, ER-negative or PR-negative (all P < 0.05) than their counterparts. No marked differences in H4K20me3 level were observed between different age, menopausal status, BMI, family history, nodal status, and HER2 status (Table 2). Univariable analysis showed that age, BMI, histological grade, clinical stage and ER status were associated with OS and clinical stage and ER status were associated with PFS (Table S1, Online Resource).
Independent prognostic value of H4K16ac and H4K20me3 in breast cancer tissues
Of the 958 eligible women, 118 died and 199 experienced disease progression with a median follow-up time of 77.7 months (interquartile range: 49.9–108.5). Five-year OS rate and PFS rate were 91.6% and 84.8%, respectively. Nonlinearities of relations of H4K16ac and H4K20me3 in tumor tissues with breast cancer prognosis were shown in Fig. S2 (Online Resource). No significant nonlinearity for the relation between breast cancer prognosis and both H4K16ac (Pnonlinear = 0.296 for OS, Pnonlinear = 0.466 for PFS) and H4K20me3 (Pnonlinear = 0.144 for OS, Pnonlinear = 0.345 for PFS), while there was a significant linear contribution of H4K20me3 to breast cancer prognosis (Ptotal < 0.001 for both OS and PFS). Based on the results of restricted cubic splines, we categorized both H4K16ac and H4K20me3 H-score into two levels using the X-tile software. The optimal cut-off values for H4K16ac and H4K20me3 H-score were 10.0 and 75.0, respectively.
In univariable analysis, significant associations between a poor breast cancer prognosis and the low level of both H4K16ac (HR = 1.51, 95% CI 1.02–2.24 for OS; HR = 1.36, 95% CI 1.01–1.83 for PFS) and H4K20me3 (HR = 2.35, 95% CI 1.60–3.45 for OS; HR = 1.92, 95% CI 1.44–2.55 for PFS) were observed (Table S2, Online Resource). In multivariable analysis, the prognostic effects of the low level of H4K20me3 were still significant for both OS (HR = 1.97, 95% CI 1.29–3.02) and PFS (HR = 1.80, 95% CI 1.31–2.46). Kaplan–Meier survival curves of H4K16ac and H4K20me3 with the prognosis were shown in Fig. S3 (Online Resource).
Joint associations of H4K16ac and H4K20me3 with breast cancer prognosis
Interactions between H4K16ac and H4K20me3 on breast cancer OS (Pinteraction = 0.003) and PFS (Pinteraction = 0.031) were significant (Table 3 and Table 4). Low level of H4K16ac was significantly associated with a poor prognosis (HR = 2.22, 95% CI 1.14–4.35 for OS; HR = 1.71, 95% CI 1.04–2.81 for PFS) only in patients with low level of H4K20me3 (Table 3). Kaplan–Meier survival curves of H4K16ac with the prognosis in patients with different levels of H4K20me3 were shown in Fig. 2. Similarly, Low level of H4K20me3 was significantly associated with a poor prognosis (HR = 3.47, 95% CI 1.86–6.50 for OS; HR = 2.32, 95% CI 1.51–3.57 for PFS) only in patients with low level of H4K16ac (Table 4). Kaplan–Meier survival curves of H4K20me3 with the prognosis in patients with different levels of H4K16ac were shown in Fig. 3. Moreover, compared with the joint high levels of both H4K16ac and H4K20me3, only the combined low levels of both them was associated with the poor OS (HR = 1.81, 95% CI 1.10–2.98), while the single low level of H4K16ac (HR = 0.56, 95% CI 0.28–1.12) or H4K20me3 (HR = 0.80, 95% CI 0.38–1.71) was not associated with the outcomes (Table 5). A similar pattern of association was observed for PFS. Compared with the joint high levels of both H4K16ac and H4K20me3, only the combined low levels of both them was associated with the poor PFS (HR = 1.83, 95% CI 1.24–2.70). Kaplan–Meier survival curves of the combination of H4K16ac and H4K20me3 with the prognosis were shown in Fig. 4.
Prognostic prediction and the comparison
We constructed a clinicopathological model, including age at diagnosis, histological grade, clinical stage, ER and HER2 status, and an extra model combined the joint levels of H4K16ac and H4K20me3. We found that the C-index of the combined model was significantly larger than that of the clinicopathological model (0.739 vs 0.699, P < 0.001 for OS; 0.672 vs 0.642, P = 0.003 for PFS) (Table 6). Moreover, we also constructed two models which were added the single H4K16ac or H4K20me3 based on the clinicopathological model. Notably, the C-index of the combined model was also significantly larger than that of the model added only H4K16ac (0.739 vs 0.712, P < 0.001 for OS; 0.672 vs 0.646, P < 0.001 for PFS) and the model added only H4K20me3 (0.739 vs 0.724, P 0.031 for OS; 0.672 vs 0.662, P = 0.006 for PFS) (Table 6).
Discussion
In this study, we found that the levels of H4K16ac and H4K20me3 in breast cancer tissues were significantly lower than that in adjacent tissues; the low level of H4K16ac was associated with HER2-negative tumors and the low level of H4K20me3 was related to ER-negative or PR-negative tumors. The low level of H4K20me3 was an independent marker of poor prognosis in breast cancer, while the poor prognostic role of low H4K16ac level was observed only in univariate analysis. Importantly, there was an interaction between these two markers on the prognosis. When compared with the joint high levels of both H4K16ac and H4K20me3, only the combined low levels of both them was associated with the poor prognosis, while the single low level of either H4K16ac or H4K20me3 was not associated with the prognosis. Moreover, the joint expression of H4K16ac and H4K20me3 could significantly improve the prediction power of the prognostic model using known clinicopathological factors.
Loss of H4K16ac and H4K20me3 has been reported in multiple types of human tumor cells [9] and the loss of H4K20me3 in breast cancer tissues has also been reported [15]. In this study, we found that there were lower levels of both H4K16ac and H4K20me3 in breast cancer tissues compared with the adjacent tissues. In consistent with our study, previous study has also found that the low level of H4K20me3 correlated with ER negative and PR negative tumors but not with HER2 expression [15]. In addition, we also showed that the low level of H4K16ac was associated with HER2 negative tumors, while another study found no association between them [14]. Underlying mechanisms of the associations of H4K16ac and H4K20me3 levels with these clinicopathological characteristics needs to be further explored.
In consistent with the previous study [14], we also found that the low level of H4K16ac was not associated with the breast cancer prognosis in multivariate analysis; for H4K20me3, we found that its low level was significantly associated with the poor prognosis of breast cancer, which was also consistent with the previous positive findings [15]. However, when stratified by the level of H4K16ac or H4K20me3, the poor prognostic roles of the low level of H4K16ac were observed only among patients with the low level of H4K20me3, and the roles of H4K20me3 were also observed only among patients with the low level of H4K16ac. Considering that there was a negative interplay between H4K16ac and H4K20me3 on breast cancer related genes [13], the prognostic roles of the global H4K16ac or H4K20me3 on breast cancer may also be inhibited by each other. Furthermore, when compared with the joint high levels of both H4K16ac and H4K20me3, only the combined low levels of both them were associated with the poor prognosis, while the single low level of either H4K16ac or H4K20me3 was not associated with the prognosis, which further suggested that the joint effects of H4K16ac and H4K20me3 should be considered in breast cancer prognosis.
TMS1, also known as ASC, was a proapoptotic signaling factor that was subjected to epigenetic silencing in human cancers [25, 26]. It had been found that the silencing of TMS1 was accompanied by loss of the H4K16ac peaks in MDA-MB231 breast cancer cell lines [10]. These findings suggested that the loss of H4K16ac may be important events in the epigenetic silencing of certain tumor suppressor genes in breast cancer. In addition, in cell lines from advanced lung cancer, breast cancer, and melanoma, tensin-3 contributed to cell migration and tumorigenesis [27]. It had been found that depletion of tensin-3 suppressed breast cancer cell invasiveness [12]. Furthermore, silencing of tensin-3 was associated with enrichment of H4K20me3 immediately upstream of the tensin-3 transcription start site, suggesting that the loss of H4K20me3 in tumor cells induced the expression of cancer-promoting genes [12]. Our findings showed that the combined low levels of both H4K16ac and H4K20me3 were associated with the poor prognosis, suggesting that the prognosis of breast cancer patients depended on the overall expression of different genes in cancer cells.
Currently, there are two types of epigenetic drugs approved in the clinics: DNA-methyltransferase inhibitors (DNMTi) and histone deacetylase inhibitors (HDACi) [28]. In breast cancer, HDACis have demonstrated antitumor activity in preclinal studies [29]. However, the combination of exemestane and entinostat (a HDAC inhibitor) did not improve survival in endocrine therapy-resistant advanced hormone receptor-positive, HER2-negative breast cancer in a recent double-blind placebo-controlled phase III trial [30]. Many studies suggested that combination of epigenetic drugs may yield more benefit [31,32,33], which was supported by our finding that the prognostic roles of one histone modification marker could be affected by other markers. Notably, a cellular experiment found that the global levels of H4K16ac and H4K20me3 in breast cancer cell line MCF7 were elevated after garcinol treatment [34]. Garcinol is a natural product obtained from Garcinia indica and has anti-neoplastic properties in many cancers, while use of it is still in pre-clinical stage [35, 36]; our findings further indicated its application prospect.
The first limitation of our analysis is that only patients with tumor > 1 cm were included, which may lead to selective bias. However, the prognosis of patients with tumor ≤ 1 cm was excellent, even with less treatment [37]; thus, it is acceptable to select the patients with tumor > 1 cm as the study population. Second, we didn’t collect the information of treatment which was associated with the outcomes. However, the treatment was determined according to the clinicopathological characteristics and most of the breast cancer patients would comply with the clinical guideline [38]. Therefore, the adjustment of these characteristics in the analysis was able to largely control the confounding effects of the treatment. Third, if intra-tumoral heterogeneity is present, TMAs are not well representative of the whole tumor. However, the whole Hematoxylin and eosin (HE)-stained sections of tissue specimens for every patient were retrieved, followed by re-slicing and re-staining with HE. Representative tumor tissue regions (full of tumor cells) and adjacent normal tissue regions (If available) were marked on the re-stained HE sections. From the marked regions, two tumor tissue cores and one adjacent normal tissue core (If unavailable, it would be replaced with the tumor tissue) with a diameter of 1 mm were selected for the construction of TMA. The results were calculated using the average of 2 or 3 cores. If full faced slides are used, it would also usually be that several typical visions be selected under the microscope to measure the scores.
In conclusion, this study firstly demonstrated that there was an interaction between the global H4K16ac and H4K20me3 on breast cancer prognosis. Moreover, the joint expression of H4K16ac and H4K20me3 could significantly improve the prediction power of the prognostic model using known clinicopathological prognostic factors. It was suggested that the combination of H4K16ac and H4K20me3 was a robust prognostic marker in breast cancer. Since histone modifications were reversible, the use of drugs to alter or restore their levels would help improve patient outcomes.
References
Campbell LL, Polyak K (2007) Breast tumor heterogeneity: cancer stem cells or clonal evolution? Cell Cycle 6:2332–2338. https://doi.org/10.4161/cc.6.19.4914
De Abreu FB, Schwartz GN, Wells WA et al (2014) Personalized therapy for breast cancer. Clin Genet 86:62–67. https://doi.org/10.1111/cge.12381
Kunc M, Peksa R, Cserni G et al (2022) High expression of progesterone receptor may be an adverse prognostic factor in oestrogen receptor-negative/progesterone receptor-positive breast cancer: results of comprehensive re-evaluation of multi-institutional case series. Pathology 54:269–278. https://doi.org/10.1016/j.pathol.2021.10.003
Han Y, Wu Y, Xu H et al (2022) The impact of hormone receptor on the clinical outcomes of HER2-positive breast cancer: a population-based study. Int J Clin Oncol 27:707–716. https://doi.org/10.1007/s10147-022-02115-x
Siegel RL, Miller KD, Fuchs HE et al (2021) Cancer Statistics, 2021. CA Cancer J Clin 71:7–33. https://doi.org/10.3322/caac.21654
Sung H, Ferlay J, Siegel RL et al (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71:209–249. https://doi.org/10.3322/caac.21660
Piekarz RL, Bates SE (2009) Epigenetic modifiers: basic understanding and clinical development. Clin Cancer Res 15:3918–3926. https://doi.org/10.1158/1078-0432.CCR-08-2788
Sharma S, Kelly TK, Jones PA (2010) Epigenetics in cancer. Carcinogenesis 31:27–36. https://doi.org/10.1093/carcin/bgp220
Fraga MF, Ballestar E, Villar-Garea A et al (2005) Loss of acetylation at Lys16 and trimethylation at Lys20 of histone H4 is a common hallmark of human cancer. Nat Genet 37:391–400. https://doi.org/10.1038/ng1531
Kapoor-Vazirani P, Kagey JD, Powell DR et al (2008) Role of hMOF-dependent histone H4 lysine 16 acetylation in the maintenance of TMS1/ASC gene activity. Cancer Res 68:6810–6821. https://doi.org/10.1158/0008-5472.CAN-08-0141
Zhao LJ, Loewenstein PM, Green M (2016) The adenoviral E1A N-terminal domain represses MYC transcription in human cancer cells by targeting both p300 and TRRAP and inhibiting MYC promoter acetylation of H3K18 and H4K16. Genes Cancer 7:98–109. https://doi.org/10.18632/genesandcancer.99
Shinchi Y, Hieda M, Nishioka Y et al (2015) SUV420H2 suppresses breast cancer cell invasion through down regulation of the SH2 domain-containing focal adhesion protein tensin-3. Exp Cell Res 334:90–99. https://doi.org/10.1016/j.yexcr.2015.03.010
Kapoor-Vazirani P, Kagey JD, Vertino PM (2011) SUV420H2-mediated H4K20 trimethylation enforces RNA polymerase II promoter-proximal pausing by blocking hMOF-dependent H4K16 acetylation. Mol Cell Biol 31:1594–1609. https://doi.org/10.1128/MCB.00524-10
Elsheikh SE, Green AR, Rakha EA et al (2009) Global histone modifications in breast cancer correlate with tumor phenotypes, prognostic factors, and patient outcome. Cancer Res 69:3802–3809. https://doi.org/10.1158/0008-5472.CAN-08-3907
Yokoyama Y, Matsumoto A, Hieda M et al (2014) Loss of histone H4K20 trimethylation predicts poor prognosis in breast cancer and is associated with invasive activity. Breast Cancer Res 16:R66. https://doi.org/10.1186/bcr3681
Wolff AC, Hammond ME, Schwartz JN et al (2007) American society of clinical oncology/college of American pathologists guideline recommendations for human epidermal growth factor receptor 2 testing in breast cancer. J Clin Oncol 25:118–145. https://doi.org/10.1200/JCO.2006.09.2775
Chen QX, Yang YZ, Liang ZZ et al (2021) Time-varying effects of FOXA1 on breast cancer prognosis. Breast Cancer Res Treat 187:867–875. https://doi.org/10.1007/s10549-021-06125-7
See SHC, Smith SH, Finkelman BS et al (2023) The role of PRAME and NY-ESO-1 as potential therapeutic and prognostic biomarkers in triple-negative breast carcinomas. Pathol Res Pract 241:154299. https://doi.org/10.1016/j.prp.2022.154299
Li K, Zhao G, Yuan H et al (2022) Upregulated expression of DDX5 predicts recurrence and poor prognosis in breast cancer. Pathol Res Pract 229:153736. https://doi.org/10.1016/j.prp.2021.153736
Camp RL, Dolled-Filhart M, Rimm DL (2004) X-tile: a new bio-informatics tool for biomarker assessment and outcome-based cut-point optimization. Clin Cancer Res 10:7252–7259. https://doi.org/10.1158/1078-0432.CCR-04-0713
Chen D, Liu Z, Liu W et al (2021) Predicting postoperative peritoneal metastasis in gastric cancer with serosal invasion using a collagen nomogram. Nat Commun 12:179. https://doi.org/10.1038/s41467-020-20429-0
Chen L, Zeng H, Du Z et al (2020) Nomogram based on pre-treatment inflammatory biomarkers predicting survival in patients with head and neck soft tissue sarcoma. Cancer Biomark 29:151–161. https://doi.org/10.3233/CBM-201739
Wang Y, Li J, Xia Y et al (2013) Prognostic nomogram for intrahepatic cholangiocarcinoma after partial hepatectomy. J Clin Oncol 31:1188–1195. https://doi.org/10.1200/JCO.2012.41.5984
Huitzil-Melendez FD, Capanu M, O’Reilly EM et al (2010) Advanced hepatocellular carcinoma: which staging systems best predict prognosis? J Clin Oncol 28:2889–2895. https://doi.org/10.1200/JCO.2009.25.9895
Conway KE, McConnell BB, Bowring CE et al (2000) TMS1, a novel proapoptotic caspase recruitment domain protein, is a target of methylation-induced gene silencing in human breast cancers. Cancer Res 60:6236–6242
Parsons MJ, Vertino PM (2006) Dual role of TMS1/ASC in death receptor signaling. Oncogene 25:6948–6958. https://doi.org/10.1038/sj.onc.1209684
Qian X, Li G, Vass WC et al (2009) The Tensin-3 protein, including its SH2 domain, is phosphorylated by Src and contributes to tumorigenesis and metastasis. Cancer Cell 16:246–258. https://doi.org/10.1016/j.ccr.2009.07.031
Pasculli B, Barbano R, Parrella P (2018) Epigenetics of breast cancer: Biology and clinical implication in the era of precision medicine. Semin Cancer Biol 51:22–35. https://doi.org/10.1016/j.semcancer.2018.01.007
Damaskos C, Garmpis N, Valsami S et al (2017) Histone deacetylase inhibitors: an attractive therapeutic strategy against breast cancer. Anticancer Res 37:35–46. https://doi.org/10.21873/anticanres.11286
Connolly RM, Zhao F, Miller KD et al (2021) E2112: randomized phase III trial of endocrine therapy plus entinostat or placebo in hormone receptor-positive advanced breast cancer. A trial of the ECOG-ACRIN cancer research group. J Clin Oncol 39:3171–3181. https://doi.org/10.1200/JCO.21.00944
Arrowsmith CH, Bountra C, Fish PV et al (2012) Epigenetic protein families: a new frontier for drug discovery. Nat Rev Drug Discov 11:384–400. https://doi.org/10.1038/nrd3674
Connolly R, Stearns V (2012) Epigenetics as a therapeutic target in breast cancer. J Mammary Gland Biol Neoplasia 17:191–204. https://doi.org/10.1007/s10911-012-9263-3
Saji S, Kimura-Tsuchiya R (2015) Combination of molecular-targeted drugs with endocrine therapy for hormone-resistant breast cancer. Int J Clin Oncol 20:268–272. https://doi.org/10.1007/s10147-015-0799-2
Collins HM, Abdelghany MK, Messmer M et al (2013) Differential effects of garcinol and curcumin on histone and p53 modifications in tumour cells. BMC Cancer 13:37. https://doi.org/10.1186/1471-2407-13-37
Balasubramanyam K, Altaf M, Varier RA et al (2004) Polyisoprenylated benzophenone, garcinol, a natural histone acetyltransferase inhibitor, represses chromatin transcription and alters global gene expression. J Biol Chem 279:33716–33726. https://doi.org/10.1074/jbc.M402839200
Aggarwal V, Tuli HS, Kaur J et al (2020) Garcinol exhibits anti-neoplastic effects by targeting diverse oncogenic factors in tumor cells. Biomedicines 8:103. https://doi.org/10.3390/biomedicines8050103
Sanchez-Munoz A, Perez-Ruiz E, Jurado JM et al (2011) Outcome of small invasive breast cancer with no axillary lymph node involvement. Breast J 17:32–38. https://doi.org/10.1111/j.1524-4741.2010.01026.x
Xu H, Jin F, Zhang XJ et al (2020) Adherence status to adjuvant endocrine therapy in Chinese women with early breast cancer and its influencing factors: a cross-sectional survey. Cancer Med 9:3703–3713. https://doi.org/10.1002/cam4.3017
Acknowledgements
We sincerely thank the patients who participated in this study, the staff who conducted the baseline and the follow-up data collection, and the medical staff in the breast departments of the Third Affiliated Hospital, and the Cancer Center of Sun Yat-Sen University. We also sincerely thank the funding of National Natural Science Foundation of China (81973115) and Science and Technology Planning Project of Guangdong Province, China (2019B030316002).
Author information
Authors and Affiliations
Contributions
BW, MZ and ZFR designed and directed the study, wrote and/or revised the manuscript. YZY constructed the TMAs and contributed to the IHC. YXR, ZJW, XFZ, JXG, and LYT contributed to digital imaging of IHC-stained sections and the assessment of immunohistochemical expression. BW, MZ and XLG contributed to clinical data collection and curation. BW and MZ participated in the statistical analysis plan and interpretation of results. ZFR provided administrative support and supervision for the study. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interests
The authors declare that they have no conflict of interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary file 1: Figure S1.
Flow chart of the study cohort. Note: there were three people who missed the data of both H4K16ac and H4K20me3 level. H4K16ac, Histone H4 lysine 16 acetylation; H4K20me3, Histone H4 lysine 20 trimethylation. Figure S2. Restricted cubic splines of H4K16ac and H4K20me3 with breast cancer OS (a for H4K16ac, c for H4K20me3) and PFS (b for H4K16ac, d for H4K20me3). H4K16ac, Histone H4 lysine 16 acetylation; H4K20me3, Histone H4 lysine 20 trimethylation; OS, overall survival; PFS, progression-free survival. Figure S3. Kaplan-Meier survival curves of H4K16ac and H4K20me3 with breast cancer OS (a for H4K16ac, c for H4K20me3) and PFS (b for H4K16ac, d for H4K20me3). H4K16ac, Histone H4 lysine 16 acetylation; H4K20me3, Histone H4 lysine 20 trimethylation; OS, overall survival; PFS, progression-free survival. Table S1. Univariate association between the demographic and clinicopathological characteristics and the outcomes. Table S2. Univariate and multivariate association between H4K16ac and H4K20me3 levels in tumor tissues and the outcomes.
About this article
Cite this article
Wang, B., Zhou, M., Gan, Xl. et al. Combined low levels of H4K16ac and H4K20me3 predicts poor prognosis in breast cancer. Int J Clin Oncol 28, 1147–1157 (2023). https://doi.org/10.1007/s10147-023-02378-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10147-023-02378-y