Abstract
Schisandra chinensis is a monoecious plant with unisex flowers. The fruit of S. chinensis is of high medical with economic value. The yield of S. chinensis fruit is related to the ratio of its female and male flowers. However, there is little research on its floral development and sex differentiation. To elucidate the possible mechanism for the sex differentiation of S. chinensis, we collected 18 samples of female and male flowers from three developmental stages and performed a comparative RNA-seq analysis aimed at identifying differentially expressed genes (DEGs) that may be related to sex differentiation. The results showed 936, 7179, and 6890 differentially expressed genes between female and male flowers at three developmental stages, respectively, and 466 candidate genes may play roles in sex differentiation. KEGG analysis showed genes involved in the flavonoid biosynthesis pathway and DNA replication pathway were essential for the development of female flowers. 51 MADS-box genes and 10 YABBY genes were identified in S. chinensis. The DEGs analysis indicated that MADS-box and YABBY genes were strongly related to the sex determination of S. chinensis. RT-qPCR confirmed the RNA-seq results of 20 differentially expressed genes, including three male-biased genes and 17 female-biased genes. A possible regulatory model of sex differentiation in S. chinensis was proposed according to our results. This study helps reveal the sex-differentiation mechanism of S. chinensis and lays the foundation for regulating the male–female ratio of S. chinensis in the future.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The sex-determination mechanism of plants is complex and regulated by multiple factors such as sex-determination genes, sex chromosomes, phytohormones, environmental factors, and epigenetic regulation. Most theorizing about sex determination in plants has focused on dioecious species, but monoecious species also matter, the sex of monecious or hermaphroditic species is determined at the level of modules, tissues, or cells. The study of flower sex differentiation in monoecious plants is the link between monoecy and dioecy to explore plant evolution (Pannell 2017). Schisandra chinensis which belongs to Schisandraceae, is a monoecious plant with male and female unisexual flowers (Cao et al. 2015). The fruit of S. chinensis is the traditional Chinese herb named “Wuweizi” or “Beiwuweizi” used as an astringent tonic to astringe the lungs and the kidneys, replenish energy, promote the production of body fluids, tonify the kidney, and induce sedation (Yang et al. 2022). The fruit of S. chinensis is also used as fruit or health food of high medical and economic value. The fruit yield of S. chinensis was affected by the ratio of female and male flowers, so the study on the development of male and female flowers and sex-related genes is of great significance for the subsequent regulation of this ratio. In addition, there are both monoecious plants and dioecious plants in Schisandraceae, so it is of great significance to study the flower differentiation of S. chinensis for the evolution of Schisandraceae.
With the rapid development of next-generation sequencing technology, various sequencing methods are available for the study of plant sex determination. Whole-genome sequencing (WGS) of non-model organisms is now widely accessible and has allowed a range of questions, including plant sex determination, in the field of molecular ecology to be investigated with greater power (Vekemans et al. 2021). Zhou et al. applied multiomics analysis including WGS, RNA-seq, and assay for Transposase Accessible Chromatin with high-throughput sequencing (ATAC-seq) to investigate the sex determination of areca palm (Areca catechu) and revealed a region on pseudochromosome 15 harbored sex-related genes, including CYP703, LOG, GPAT, AMS, and BiP (Zhou et al. 2022). However, the method based on WGS is costly and not suitable for every species, especially those with large genomes, such as plants from the Schisandraceae, which have a genome size of about 8G (Zhang et al. 2013). In contrast, RNA-Seq can instead be utilized as a powerful and cost-effective approach for the study of sex determination and has widely been used in the identification of DEGs associated with sex differentiation in plants (Harkess and Leebens-Mack 2016). RNA-seq can provide a molecular basis for revealing differences between male and female flowers at the transcriptional levels and facilitate genetic research on sex determination in non-model plants, such as the monoecious plants Jatropha curcas, Vernicia fordii, and Castanea henryi (Zhao et al. 2020; Alsubaie et al. 2023). In Jatropha curcas, DEGs such as KNAT, MYC2, SRS5, SVP, TFL1, and TS2 may participate in the sex determination (Chen et al. 2016). In Vernicia fordii, 310 and 298 DEGs showed high expression levels in male and female flowers (Mao et al. 2017). In Castanea henryi, WRKY47, ERF021, MYB4, AGL11/15, DEF, and SEP1 were critical regulators of sex determination (Wu et al. 2023). Therefore, we suppose that RNA-seq is a feasible tool to explore the sex determination mechanism.
Liu et al. used RNA-seq to identify the sex-determining genes of S. chinensis using flower buds about to bloom, and the results showed that phytohormones including auxin and jasmonate (JA), sucrose may contribute to the development of S. chinensis flowers. AG may be critical for the sex determination of S. chinensis (Liu et al. 2022). The members of MADS-box and YABBY gene families have been reported to be strongly associated with sex differentiation and determination of plants. MADS-box genes such as AP3, PI, and STK are candidate genes in the sex determination of Areca catechu, Populus tremula, and Zanthoxylum armatum (Zhou et al. 2022; Leite Montalvão et al. 2022; Hui et al. 2022). The YABBY family genes such as CRABS CLAW (CRC) and INNER NO OUTER (INO) are important for female organ development. CRC controls the carpel determinacy in cucurbits (Zhang et al. 2022a). INO could regulate the development of integument and be essential for female fertility in pomegranate (Chen et al. 2017). MADS-box and YABBY transcription factors play essential roles in the sex determination of plants as the sex switch (Zhang et al. 2022a, 2022b). Besides the MADS-box and YABBY genes, pathways including flavonoid biosynthesis, DNA replication, and sugar metabolism have been revealed to participate in male or female organ development (Feng et al. 2020; Pawełkowicz et al. 2019; Wang et al. 2019b). To further identify the possible genes that participate in the flower development and sex determination of S. chinensis, we performed RNA-seq and the DEGs analysis between female and male flowers at three developmental stages. Extensive research has shown that MADS-box and YABBY transcription factors play essential roles in the sex determination of plants (Arora et al. 2007; Romanova et al. 2021). Based on transcriptome data, two gene family analyses were conducted to identify possible MADS-box and YABBY genes. The results of this study can lay a foundation for further exploring the sexual differentiation mechanism of S. chinensis and provide valuable information for the study of evolutionary patterns of flowers of Schisandraceae and plant sexual differentiation.
Materials and methods
Plant materials and sample collection
Three monoecious S. chinensis were cultivated in the Beijing Medical Botanical Garden (116.27°E, 40.03°N). Male and female S. chinensis flowers were collected on April 14, 18, and 24, 2022. Three developmental stages of S. chinensis flowers expressed different morphological patterns (Fig. 1). At the first stage (April 14), the male and female flowers of S. chinensis cannot be distinguished by direct observation. The immature stamens and pistils were observed in male and female flower buds under the anatomical microscope and scanning electron microscope (Fig. 1g ~ n). At the second stage, S. chinensis flowers were fully developed, but not fully flowering (Fig. 1b, e), while at the third stage were fully open (Fig. 1c, f). Samples were grouped into F1 vs. M1, F2 vs. M2, and F3 vs. M3, which represented DEGs between female and male flower buds at the first, second, and third developmental stages, respectively. Each group contained three biological replicates and each biological replicate contained at least five flower buds. The samples for RNA-seq were immediately frozen in liquid nitrogen after the collection and stored at a − 80 °C freezer. The samples for observation were stored in FAA fixative solution (70% ethanol: glacial acetic acid: 38% formaldehyde = 18:1:1).
Male and female flowers of S. chinensis at the different developmental stages. a–c Male buds at stages 1–3. d–f Female buds at stages 1–3. g–h The male buds at stage 1 under the microscope. i The male bud at stage 1 under the scanning electron microscope. j The tissue section of a male bud of stage 1. k–l The female buds at stage 1 under the microscope. m The female bud at stage 1 under the scanning electron microscope. n The Tissue section of a female bud of stage 1. Scale. g, h, j, k, l, n 50 mm, i, m 200 μm
RNA extraction, library construction, and Illumina sequencing
The total RNA of samples was extracted using TRIzol reagent (Invitrogen, Carlsbad, CA, USA). The integrity and quality of the RNA were determined by the Bionanlyzer 2100 system (Agilent Technologies, Santa Clara, CA, USA). One microgram of total RNA was used for the transcriptome library construction. We used magnetic beads containing Oligo (dT) by A-T base pairing with polyA to separate mRNA. Then, we used a fragmentation buffer to fragmentize mRNA. cDNA was synthesized by using the SuperScript double-stranded cDNA Synthesis Kit (Invitrogen, Carlsbad, CA, USA) and random hexamer primers (Illumina, San Diego, CA, USA). The synthesized cDNA was then end-repaired, phosphorylated, and the “A” base added according to Illumina’s library construction protocol. Libraries were size-selected for cDNA target fragments of 300 bp on 2% low range ultra agarose followed by PCR amplified using Phusion DNA polymerase (NEB) for 15 PCR cycles. After quality inspection, the paired-end RNA-seq sequencing library was sequenced with the Illumina NovaSeq 6000 platform (Illumina, San Diego, CA, USA). Shanghai Majorbio Bio-pharm Biotechnology Co., Ltd. (Shanghai, China) was assigned to perform high-throughput sequencing.
Quality control, de novo transcriptome assembly, function annotation
Fastp v0.19.5 with default parameters was used for quality control (Chen et al. 2018). The de novo assembly of transcripts was finished by Trinity v2.8.5 (Grabherr et al. 2011). CD-HIT v4.5.7 was used for the optimization of assembled transcripts (Fu et al. 2012). The quality of transcripts was assessed by BUSCO v3.0.2 and TransRate v1.0.3 (Simão et al. 2015; Smith-Unna et al. 2016). All assembled transcripts were compared with the public databases including NCBI protein non-redundant (NR), Swiss-Prot, Pfam, Clusters of Orthologous Groups of proteins (COG), Gene Ontology (GO), and Kyoto Encyclopedia of Genes and Genomes (KEGG) using blast + v.2.9.0 and diamond v0.9.24, with a threshold e value < 1e−5 (Camacho et al. 2009; Buchfink et al. 2015).
Differential expression analysis
Gene expression levels were calculated and normalized using the method of transcript per kilobase per million mapped reads (TPM) by using RSEM v1.3.1 (Li and Dewey 2011). DESeq2 R package v1.24.0 was used to identify differentially expressed genes between male and female flowers in three different stages (Love et al. 2014). We compared three groups, F1 vs. M1, F2 vs. M2, and F3 vs. M3. Genes with | Log2 (fold change) |> 1 and a false discovery rate (FDR) of < 0.05 in comparison were considered as significant DEGs. DEGs were then subjected to enrichment analysis of GO functions and KEGG pathways.
Identification and expression pattern analysis of MADS-box and YABBY gene families members
Blast (https://blast.ncbi.nlm.nih.gov/Blast.cgi) and HMMER v3.0 (Potter et al. 2018) were used to identify possible members of MADS-box and YABBY gene families based on RNA-seq data. MADS-box sequences of Arabidopsis were downloaded from The Arabidopsis Information Resource 10 database (TAIR10) (www.arabidopsis.org/) (Berardini et al. 2015). Arabidopsis and rice YABBY sequences were downloaded from the Plant Transcription Factor database (http://planttfdb.gao-lab.org/). MADS-box HMMER models, SRF-TF (PF00319) and K-box (PF01486), and YABBY HMMER model (PF04690) were obtained from Pfam protein families database (http://pfam.xfam.org/) (Mistry et al. 2021). The filter threshold for both blast and HMMER was set to 1e−5. The intersection of blast and HMMER results were considered as candidate S. chinensis MADS-box and YABBY genes. All predicted MADS-box and YABBY sequences were verified using SMART (http://smart.embl-heidelberg.de) and CDD (https://www.ncbi.nlm.nih.gov/cdd) (Letunic et al. 2021; Lu et al. 2020).
For the MADS-box gene family, besides Arabidopsis thaliana and Oryza sativa MADS-box genes, we also download MADS-box sequences homologous to Schisandra chinensis to construct the phylogenetic tree, including Asarum europaeum, Amborella trichopoda, Schisandra sphenanthera, Kadsura japonica, Illicium anisatum, Illicium floridanum, Illicium henryi, Illicium parviflorum, and Magnolia stellata. . MADS-box genes in grapevine (VvMADS) have been well studied (Díaz-Riquelme et al. 2009). Therefore, we also used VvMADS genes to perform the phylogenetic analysis. For the YABBY gene family, reported YABBY sequences of Arabidopsis thaliana and Oryza sativa were used to build the phylogenetic tree. All protein sequences were aligned using MUSCLE, and trimAI was used to trim the results of alignment (Edgar 2004; Capella-Gutiérrez et al. 2009). IQtree was used to construct a phylogenetic tree with 5000 bootstrap replications (Nguyen et al. 2015). The software ModelFinder helped search for the optimal tree models for phylogenetic analysis (Kalyaanamoorthy et al. 2017). The optimal tree models of MADS-box and YABBY genes were set to LG + G4 and VT + G4, respectively. LG and VT were two amino acid replacement matrixes (Le and Gascuel 2008; Vingron 2000). G was the Gamma model, one rare heterogeneity across-sites model proposed by Yang (1994). LG + G4 indicated that the LG matrix and discrete Gamma model with four rare categories. VT + G4 indicated that the LG matrix and discrete Gamma model with four rare categories. The phylogenetic trees of MADS-box genes and YABBY genes were constructed on the toolkit TBtools and annotated by the iTOL (Chen et al. 2020; Letunic and Bork 2021). The source of all reference sequences is listed in Table S1.
Quantitative real-time PCR analysis
qPCR primers for the validation genes were designed by the Prime Primer 5 software (Singh et al. 1998). We selected the TUBA gene as the reference gene according to the previous study (Liu et al. 2022). Twenty genes were selected to validate their expression patterns according to their functions. AFO, YABBY2, INO, YABBY5-1, and YABBY5-2 belonged to the YABBY gene family, AeAP3-2, STK belonged to the MADS-box gene family, and AIL-1, ANT belonged to AP2 gene family. TT2, TT3, TT4, TT6-1, TT6-2, TT12, and MYB1 participated in the flavonoid biosynthesis. GATA18 was involved in the development of female flowers and GAMYB, CEP1, and MST4 participated in pollen development. The detailed information on forward and reverse primers is listed in Table S2. All qPCR experiments were completed using three biological replicates with three technical replicates each. We used 1μg sample RNA as the template for reverse transcription experiments. The reverse transcription experiments were performed using FastKing RT Kit with gDNase (Tiangen, Beijing, China) according to the instructions. Then, we completed the real-time qPCR experiments with 1 μg cDNA, Talent qPCR Premix Kit (SYBR Green) (Tiangen, Beijing, China) and analyzed on a BIORAD CFX96 Real-Time System. The procedure was initiated at 95 °C for 3 min, followed by 40 cycles of 95 °C for 5 s, 55 °C for 10 s, and 72 °C for 15 s. The relative expression levels were calculated using the 2−ΔΔCt method (Livak and Schmittgen 2001).
Protein–protein interaction prediction
To elucidate the candidate gene co-expression network related to the flower development and sex differentiation of S. chinensis, a protein–protein interaction (PPI) prediction was performed. The string database (https://cn.string-db.org/) was used for the PPI prediction (Szklarczyk et al. 2023). The coding sequences of 466 DEGs were translated to protein sequences and then uploaded to this database. Arabidopsis thaliana was selected as the reference species. The confidence value was set as 0.4 (medium). The prediction result was exported to cytoscape v3.9.1 to draw the PPI network (Shannon et al. 2003).
Results
Results of transcriptome assembly and annotation
Eighteen cDNA libraries from male and female flowers of S. chinensis at three developmental stages were sequenced. After filtering the adaptors and low-quality sequences, 125.51 Gb clean data were obtained from 18 samples. Each sample contained more than 6.03 Gb clean data. The Q20 ratio of each sample was above 97.21%, and the Q30 ratio was above 92.46%. The GC content of 18 samples ranged from 46.37 to 47.16% (Table S3). A total of 149,495 transcripts and 96,694 unigenes were obtained from the de-novo assembly (Table S4). The N50 of transcripts and unigenes were 1,473 bp and 1,352 bp. BUSCO scores of transcripts and unigenes were 93.0% and 86.0%. TransRate scores of transcripts and unigenes were 0.29% and 0.23%. The results above indicated accurate splicing.
Clustering and heatmap analysis, combined with PCA analysis showed that female and male samples could be well separated (Fig. 2a, b). Samples at the first stage were separated from those at the second and third stages, indicating that samples at the second and third stages showed similar expression patterns. All expressed unigenes were compared with the major six databases (GO, KEGG, eggNOG, NR, Swiss-Prot, and Pfam (Fig. 2c). Most annotated unigenes were obtained from the NCBI NR database (37782 unigenes, 39.07%), and the minimum annotated unigenes were obtained from the KEGG database (12536 unigenes, 12.96%). Based on the species distribution analysis of the NR database, the top hit species was Tetracentron sinense (10.24%), belonging to the Trochodendrales.
Results of transcriptome assembly and annotation. a Clustering and heatmap analysis. The sample distances are represented by variations from dark blue (low similarity) to dark red ( similarity). b PCA for all 18 samples based on the expression values of transcriptome-wide expression profiles. c The profile of annotation results based on the major six databases (GO, KEGG, eggNOG, NR, Swiss-Prot, and Pfam)
DEGs analysis, GO, and KEGG enrichment analysis
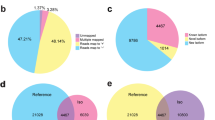
To identify candidate genes that may engage in the flower development and sex differentiation of S. chinensis, we compared the transcriptome profiles between F1 and M1, F2 and M2, and F3 and M3. The total numbers of DEGs in F1 vs. M1, F2 vs. M2, and F3 vs. M3 were 936, 7179, and 6890, respectively (Fig. 3a, b). A total of 466 DEGs were commonly expressed in S. chinensis flowers at three developmental stages.
Results of DEGs, GO, and KEGG enrichment analyses. a DEGs in S. chinensis flowers at three developmental stages. b Venn diagram showing overlaps between the sets of sex-biased genes (|log2(foldchange) |> 1, p.adj < 0.05). c–e GO enrichment analysis of DEGs (1) F1 vs. M1 (2) F2 vs. M2 (3) F3 vs. M3. f–h KEGG enrichment analysis of DEGs (1) F1 vs. M1 (2) F2 vs. M2 (3) F3 vs. M3
GO enrichment analysis revealed 118, 198, and 190 significantly enriched GO terms (FDR < 0.05) at stages 1 to 3, respectively (Fig. 3c, d, e). At the first stage, cell fate commitment (GO:0045165) was the most significantly enriched term. At the second stage, nucleosome assembly (GO:0006334) was the most significantly enriched term. At the third stage, mitotic cell cycle phase transition (GO:0044772) was the most significantly enriched term.
KEGG enrichment analysis revealed that 4, 6, and 7 KEGG pathways were significantly enriched (FDR < 0.05) at stages 1 to 3, respectively (Fig. 3f, g, h). At the first stage, flavonoid biosynthesis (ko00941), fatty acid elongation (ko00062), DNA replication (ko03030), and phenylpropanoid biosynthesis (ko00940) were significantly enriched. At the second stage, ribosome (ko03010), DNA replication (ko03030), starch and sucrose metabolism (ko00500), lysine biosynthesis (ko00300), glycerolipid metabolism (ko00561), and flavonoid biosynthesis (ko00941) were significantly enriched. At the third stage, ribosome (ko03010), lysine biosynthesis (ko00300), glycerolipid metabolism (ko00561), phenylpropanoid biosynthesis (ko00940), DNA replication (ko03030), base excision repair (ko03410), and isoquinoline alkaloid biosynthesis (ko00950) were significantly enriched.
Possible MADS-box gene family members in S. chinensis flowers
Through the above method, we finally obtained 51 MADS-box genes from S. chinensis and divided these genes into several clades according to previous studies (Fig. 4a, Table 1, S5) (Duan et al. 2022; Schilling et al. 2020; Ye et al. 2022). Most common MADS clades can be found in S. chinensis, such as AP1 (class A), AP3/PI (class B), AG (class C), STK (class D), and SEP (class E), which act as flowers organ identity genes (Ali et al. 2019). However, clades like FLC, FUL, ANR, and SVP were not identified in this study. On the contrary, the TM8 gene and OsMADS32 gene, which were not found in Arabidopsis, could be found in S. chinensis (Wang et al. 2020a).
Results of two gene families analyses. MADS-box gene family analysis: a Maximum likelihood tree of MIKC MADS-box genes. 42 MIKC genes were divided into 13 clades. b Heatmap of MIKC genes in S. chinensis female and male flowers at three developmental stages. YABBY gene family analysis: c Maximum likelihood tree of YABBY genes identified in S. chinensis flowers. d Heatmap of YABBY genes in S. chinensis female and male flowers at three developmental stages
Then, we analyzed differentially expressed MADS-box genes in male and female flowers (Fig. 4b). All B-class genes (AP3/PI) had higher expression in male flowers, similar to the results of previous studies (Ren et al. 2021; Wang et al. 2019c). One AG (class C) gene, two STK (class D) genes, and two Bsister genes were upregulated in female flowers. Most SEP (class E) genes showed an extremely low expression level, while we found one SEP gene and one AGL6 gene showed a higher expression level in the female flowers. MIKC* is another important clade of the MADS-box gene family (Liu et al. 2013). Our results suggested that MIKC* genes were mainly upregulated in male flowers.
Possible YABBY gene family members in S. chinensis flowers
We successfully identified 10 YABBY genes from the transcriptome data (Table S6). Based on the classification of YABBY genes in Arabidopsis and rice, we found that all YABBY clades can be found in S. chinensis (Fig. 4c, Table 2). However, YABBY3 genes could not be found in S. chinensis.
Expression analysis showed that most YABBY genes were upregulated in female flowers (Fig. 4d). DL and FIL showed high expression at the early flowering stage. With the development of flowering, the expression level reduced gradually. YABBY2 genes had high expression in male and female flowers. INO gene showed a female-specific expression pattern. Like the expression of DL and FIL genes, two YABBY5 genes had high expression at the first stage, while one YABBY5 gene showed high expression at all stages. Five YABBY genes were regulated in female flowers at all stages.
Sex-biased genes of S. chinensis
The identification of sex-biased genes, which exhibit significantly higher expression in flowers of one sex than in the other sex, can help explore the sex determination mechanism (Harkess et al. 2015; Li et al. 2020). Four hundred and sixty-six DEGs were identified between female and male flowers at three developmental stages, which included 390 female-biased genes, 66 male-biased genes, and 10 no-biased genes (Fig. 5a). In these genes, 52 transcription factors (TFs) were identified, including 42 female-biased TFs, eight male-biased TFs, and two no-biased TFs. The gene family including more than three members were MYB (7), bHLH (7), NAC (6), YABBY (5), LBD (4), GRF (4), AP2/ERF (4), and MADS (3). Notably, most members of the NAC gene family showed a male-biased expression pattern (Fig. 5b). The top five enriched GO terms were centromere complex assembly (GO:0034508), meiotic chromosome separation (GO:0051307), chromosome, centromeric region (GO:0000775), protein-DNA complex (GO:0097522), and nucleosome (GO:0000786) (Fig. 5c). These genes were significantly enriched in two pathways, DNA replication (ko03030), and flavonoid biosynthesis (ko00941) (Fig. 5d).
Validation of DEGs using qRT-PCR analysis
To validate the gene expression revealed by the RNA-seq analysis, 20 DEGs were selected for validation experiments. These genes, including ScAFO, ScYABBY2, ScINO, ScYABBY5-1, ScYABBY5-2, ScAeAP3-2, ScSTK, ScAIL-1, ScANT, ScGATA18, ScTT2, ScTT3, ScTT4, ScTT6-1, ScTT6-2, ScTT12, ScMYB1, ScGAMYB, ScCEP1, and ScMST4. These genes were reported to participate in the development of male and female flowers. The results showed that the expression patterns of these genes in male and female flowers were similar between RNA-seq and qPCR methods, and the correlation ratio (R2) was 0.807 (Fig. S1), suggesting that the RNA-seq analysis results were reliable. The expression of ScGAMYB, ScCEP1, and ScMST4 were downregulated in female flowers, while the expression of ScAFO, ScYABBY2, ScINO, ScYABBY5-1, ScYABBY5-2, ScAeAP3-2, ScSTK, ScAIL-1, ScANT, ScTT2, ScTT3, ScTT4, ScTT6-1, ScTT6-2, ScTT12, ScMYB1, and ScGATA18 were upregulated in female flowers (Fig. 6). ScYABBY2, ScINO, ScSTK, ScAeAP3-2, ScANT, ScMYB1, and ScGATA18 showed female-specific expression pattern, while ScCEP1 and ScMST4 were only expressed in male flowers.
qRT-PCR validation of DEGs. Male-biased genes included GAMYB, CEP1, and MST4. Female-biased genes included AFO, YABBY2, INO, YABBY5-1, YABBY5-2, AeAP3-2, STK, AIL-1, ANT, GATA18, TT2, TT3, TT4, TT6-1, TT6-2, TT12, MYB1. The y-axis indicated the expression level (2−△△.Ct). The error bar indicated the standard error. *: p < 0.05; **: p < 0.01; **** p < 0.001
Protein–protein interaction prediction
A total of 76 DEGs with medium confidence (0.400) were filtered to construct the PPI network. These genes could be divided into four groups according to their functions and clusters, including genes related to the biosynthesis of flavonoids (TT2, TT4, TT5, TT12, etc.), genes related to stomata development (FAMA, STOMAGEN, TMM, SPCH, etc.), genes related to flower development (SUP, NTT, INO, BEL1, ANT, LFY, HEC3, etc.), genes related to DNA replication and cell cycle (MCM3, MCM5, HMGB6, etc.) (Fig. 7). Genes with darker colors showed higher degree scores, suggesting that these genes were more likely to participate in the sex differentiation of S. chinensis. YABBY genes including INO and AFO, MADS-box genes STK and TT16, flower development-related genes LFY, ANT, BEL1, SUP, and DNA replication-related genes including MCM3, MCM5, PCNA2, PRL, HTA6, and HMGB6 may play a key role in the flower development and sex differentiation of S. chinensis.
Discussion
MADS-box genes regulate the flower development of S. chinensis
MADS-box genes are strongly related to the formation of flowers. The C-class gene AG was reported to be a candidate gene involved in the sex differentiation of S. chinensis (Liu et al. 2022). To further explore MADS-box genes potentially involved in the flower development and sex differentiation of S. chinensis, we performed a gene family analysis based on the RNA-seq data.
In this study, we found that the expression pattern of B, C, and D genes in S. chinensis was similar in Arabidopsis. B-class genes including AP3 and PI control the identity of petals and stamens (Whipple et al. 2004). Five B-class genes in S. chinensis (ScMADS6, 8, 9, 10, 43), were upregulated in male flowers, indicating that B-class genes may regulate male organ development. The C-class gene AG could regulate the identity of stamens and carpels (Yamaguchi et al. 2006). In S. chinensis, the AG gene (ScMADS30) had expression in both male and female flowers. Consistent with Liu’s findings, the expression level in female flowers was higher in male flowers (Liu et al. 2022). Therefore, we speculated that the AG gene ScMADS30 could be a candidate gene in the sex differentiation of S. chinensis.
D-class genes including STK and SHP1/2 are responsible for the ovule identity (Rodríguez-Cazorla et al. 2018). Two STK genes ScMADS16 and ScMADS17 had high expression in female flowers. Besides D-class genes, Bsister MADS-box genes also act as the key regulator in female organ development (Mizzotti et al. 2012). We identified two Bsister genes ScMADS38 and ScMADS39 in S. chinensis. They were homologs to AeAP3-2, the Bsister genes identified in Asarum europaeum (Kramer and Irish 2000). It was reported that AeAP3-2 could be involved in carpel and ovule development. We found that ScMADS38 and ScMADS39 were only expressed in female flowers. This result indicated that ScMADS38 and ScMADS39 were essential for female flower development and may participate in the sex differentiation of S. chinensis. AG, STK, and AeAP3-2 mutually regulated the development of female organs (Pinyopich et al. 2003). AG and STK genes are responsible for the specification of ovule identity (Pelayo et al. 2021). STK and AeAP3-2 genes are essential for female gametophyte development (Mizzotti et al. 2012). ScMADS16, ScMADS17, and ScMADS38 were significantly upregulated in female flowers. Overall, we inferred that C, D-class genes, and Bsister MADS-box genes may be crucial for the female flower development of S. chinensis.
E-class genes including SEP1, SEP2, SEP3, SEP4, and AGL6 were involved in the development of sepals, petals, stamens, carpels, and ovules (Morel et al. 2019). The expression analysis showed that most E-class genes identified had a very low expression in flowers, and none of these genes were significantly upregulated. Four E-class genes, ScMADS20 (AGL6), ScMADS37 (SEP1), ScMADS45 (SEP2), and ScMADS46 (SEP3) had high expression in flowers during the flowering process. In Arabidopsis thaliana, SEP genes can control the specification of stamen identity with B-class genes (AP3/PI) and C-class gene (AG). ScMADS37 showed no obvious biased expression in flowers. ScMADS45 and ScMADS46 were downregulated in female flowers. Considering the B-class genes were upregulated in male flowers and C-class were upregulated in female flowers, we speculated that SEP2 and SEP3 may regulate the formation of stamens with B-class genes. Besides SEP genes, we also identified an AGL6 gene, ScMADS20, which exhibited higher expression in female flowers. The AGL6 gene had higher expression in carpels than in stamens and may participate in the ovule development of petunia (Petunia hybrida). We suggested that the AGL6 gene may participate in pistil development (Rijpkema et al. 2009).
Besides ABCDE-class genes, we also identified five genes belonging to MIKC*-type (ScMADS12, 13, 18, 19, 35). MIKC* genes are the other important clades of MIKC genes. In Arabidopsis thaliana, MIKC* genes include AGL30, AGL65, AGL66, AGL103, and AGL104. MIKC* genes are enriched in mature pollens. The mutation of MIKC* genes could lead to reduced male fertility. In rice, MIKC* genes were specifically expressed in late pollen development (Adamczyk et al. 2007). Four MIKC* genes identified in S. chinensis except ScMADS12 had high expression in the male flowers and almost had no expression in the early flower development, suggesting that MIKC* genes may influence the development and maturation of pollens. ScMADS12 had higher expression than other MIKC* genes in flowers and was significantly upregulated in female flowers at the late flowering stage. This confusing result hinted that the role of ScMADS12 needed further investigation.
Based on our results, we tend to agree that the B, C, and D-class genes may play similar roles in the flower development of S. chinensis. B-class genes participated in the formation and development of stamens and C-class and D-class genes are responsible for the development of pistils. Besides, the Bsister genes may play essential roles in the regulation of ovules. E-class genes including SEP-like genes and AGL6-like genes may regulate the flower development together with B, C, and D-class genes. MIKC* genes are most likely to participate in the development of pollen. Combined with the expression pattern, we speculated that ScMADS16, ScMADS17, and ScMADS38 may participate in the sex differentiation of S. chinensis.
YABBY genes are essential for the female flower development in S. chinensis
The YABBY family is plant-specific and characterized by a C2-C2 zinc finger domain at the N-terminus and a YABBY domain at the C-terminus (Bowman 2000). YABBY family genes can regulate the development of lateral organs and abaxial-adaxial polarity (Siegfried et al. 1999). The YABBY gene family could be divided into five clades including CRC/DL, YABBY1/YABBY3, YABBY2, INO, and YABBY5 (Buttar et al. 2020). Accumulated evidence indicates that YABBY genes may be associated with plant sex determination. YABBY1(FIL/AFO), YABBY2, and YABBY5 may play an important role in the sex determination of spinach (Li et al. 2020). INO may be related to the sex differentiation and determination of female organs in Tapiscia sinensis (Xin et al. 2019). Considering the vital role YABBY genes play in the sex determination of flowers; we also performed the gene family analysis to identify possible YABBY genes in S. chinensis based on the transcriptome data.
In our study, we identified ten YABBY genes. Compared to YABBY genes in Arabidopsis, The CRC gene and the YABBY3 gene could not be found. However, we identified a DL gene, which is a homolog to CRC in rice. By the analysis of expression pattern, we found that most YABBY genes in S. chinensis had higher expression levels in female flowers, and five YABBY genes like ScFIL1-2, ScYABBY2, ScINO, ScYABBY5-1, and ScYABBY5-2 were significantly upregulated in female flowers during three developmental stages. DL is essential for the specification of carpels in rice (Yamaguchi et al. 2004). The mutation of DL could lead to the formation of ectopic stamen (Nagasawa et al. 2003). ScDL had higher expression in female flowers than in male flowers, indicating that ScDL may be associated with carpel development in S. chinensis.
The YABBY1 gene, also called AFO or FIL, controls the determination of floral meristem identity and development of the floral meristem (Sawa et al. 1999). The homolog to YABBY1 in cucumber had a high expression level at all ovary developmental stages (Yin et al. 2022). YABBY1 may be a candidate gene for the gynoecium development in Silene latifolia (Bačovský et al. 2022). Three YABBY1 genes, ScFIL-1, ScFIL-2, and ScFIL-3, were identified in this study. ScFIL-1 and ScFIL-2 only had high expression levels in female flowers at the early flowering stage, implying that these genes may be involved in the early female flower development. ScFIL-3 had a high expression level in male flowers at the late flowering stage. In general, we suggested that YABBY1 genes showed female bias. The YABBY2 gene is expressed in the abaxial tissue of the carpel in Arabidopsis and Amborella trichopoda (Siegfried et al. 1999; Yamada et al. 2004). Two YABBY2 genes, ScYABBY2-1, and ScYABBY2-2, had higher expression in male and female flowers, respectively, indicating that YABBY2 genes could be required for flower development of S. chinensis.
INO demonstrated a female-limited expression pattern in S. chinensis flowers. In Arabidopsis thaliana, INO is crucial for the formation of the outer integument of ovules (Skinner et al. 2023). In pomegranates (Punica granatum), the expression level of INO is higher in bisexual flowers than in male flowers, suggesting that INO may potentially regulate the differentiation and development of ovules (Chen et al. 2017). In Tapiscia sinensis, INO acts as an essential regulatory factor in pistil development (Xin et al. 2019). The INO gene is crucial to female flower development, and we infer that ScINO may regulate the gynoecium development of S. chinensis.
YABBY5 is involved in the formation of spikelet in rice (Dubos et al. 2010). In spinach (Spinacia oleracea), YABBY5 acted as a pivotal gene in the development of female flowers (Li et al. 2020). Three ScYABBY5 genes showed higher expression in female flowers than in male flowers and had no expression in male flowers at the last two stages. It implied that ScYABBY5 genes were essential to the development of male and female flowers at the early stage and more associated with the female flower development in S. chinensis. Combined the result of RNA-seq analysis and RT-qPCR, we speculated that ScFIL-2, ScYABBY2-2, ScINO, ScYABBY5-1, and ScYABBY5-2 could potentially serve as key regulators in the gynoecium development and candidate genes in the sex differentiation of S. chinensis.
Multiple transcription factors contribute to the flower development and may be related to the sex differentiation of S. chinensis
In addition to MADS-box and YABBY transcription factors, we also identified genes belonging to other gene families that may be related to the sex differentiation of S. chinensis. In 466 DEGs significantly expressed during the flowering stages, transcription factors accounted for ~ 11%. In these transcription factors, the MYB gene family had the most members. As the gene family, which is large, functionally diverse, and represented in all eukaryotes, the role MYB transcription factors play in the flower development of S. chinensis cannot be ignored (Dubos et al. 2010). MYB transcription factors seemed to exhibit high expression in male and female flowers. GAMYB and DIV had higher expression in male flowers, while RAD had higher expression in female flowers. GAMYB, a part of the GA signal pathway, contributed to the anther development. GAMYB regulates early anther development and is essential for the formation of exine and Ubisch bodies in anthers (Alonso-Peral et al. 2010; Aya et al. 2009). GAMYB can interact with the DNA-specific motifs of GA-upregulated genes of anthers and regulate their expression. The DIV and RAD genes are responsible for the dorsoventral asymmetry of flowers in Antirrhinum majus (Galego and Almeida 2002; Sengupta and Hileman 2022). In snapdragons, RAD and DIV are expressed in ovaries and developing fruit. However, DIV was downregulated in female flowers, indicating that DIV might influence the development of stamens in S. chinensis. In hexaploid persimmon, ectopic overexpression of RAD could result in gynoecium hypergrowth. In Plukenetia volubilis, two RAD-like genes RAD1 and RAD2 had higher expression levels in female flowers than in male flowers (Fu et al. 2018; Masuda et al. 2022). Similarly, one RAD-like gene, ScRAD1, appears to express exclusively in female flowers. These results suggest that ScRAD1 could be involved in the gynoecium development and might serve as a candidate gene in the sex differentiation of S. chinensis. Basic helix-loop-helix (bHLH) proteins, another large family of transcription factors, also play a role in the flower development of plants. Liu et al. found that bHLH transcription factors including SPT, HEC, DYT1, and AMS could be involved in the sex differentiation and determination of S. chinensis (Liu et al. 2022). This study identified a HEC-like gene, ScHEC3, upregulated in female flowers. In Arabidopsis, the HEC3 gene contributes to the gynoecium development. In Plukenetia volubilis, HEC-like genes HEC1 and HEC2 have higher expression in female flowers (Gremski et al. 2007; Peréz-Mesa et al. 2020). ScHEC3 was only expressed in female flowers, suggesting that ScHEC3 may be crucial to gynoecium development and possibly be associated with the sex differentiation of S. chinensis.
Besides HEC3, we also identified many transcription factors related to gynoecium development. The SUP and NTT genes, both from the C2H2 zinc finger gene family, are essential for the development of female flowers. In Arabidopsis thaliana, SUP controls the determinacy of the floral meristem and is required for normal ovule development (Sakai et al. 2000, 1995). SUP could regulate the expression of B-class genes including AP3 and PI, and a mutation of SUP might lead to the development of extra stamens (Sakai et al. 2000). Accumulated research showed that SUP may be involved in plant sex differentiation (Bačovský et al. 2022; Fu et al. 2018; Khadka et al. 2019). NTT, specifically expressed in the transmitting tract, is required for normal differentiation of the ovary transmitting tract cells and pollen tube growth (Crawford et al. 2007). ANT, an AP2 family gene, is essential to the gynoecium development (Klucher et al. 1996). ANT could regulate the autoinduction and expression of INO (Baker et al. 1997). GRF4 and GRF6, members of the growth-regulation factors, act as regulators involved in plant hormone signaling and metabolism (Zhang et al. 2022c). In rice, GRF genes were initially identified as gibberellin-induced (Lu et al. 2021). GRF genes are involved in gynoecium development in persimmon (Yang et al. 2019). WOX1 and WOX9, from lateral organ boundaries domain protein, are crucial for ovule development. WOX1 may regulate the outgrowth of ovules and WOX9 is required for the correct patterning and curvature of the ovule in Arabidopsis. WOX9 could regulate the expression of INO and is involved in the transcriptional activation of a subset of cytokinin response factors (Niu et al. 2018; Petrella et al. 2022). CUC3, a NAC family gene, participates in ovule development in Arabidopsis (Gonçalves et al. 2015). GATA18 is essential to the gynoecium development of Silene latifolia and can modulate cytokinin homeostasis in organ boundaries by regulating CKX3 expression (Bačovský et al. 2022). These transcription factors showed female-specific expression patterns, indicating that they might play a key role in the gynoecium development of S. chinensis.
Auxin and cytokinin participate in the female flower development of S. chinensis
Accumulated evidence indicates that many genes involved in phytohormones biosynthesis and signal pathway act as sex determination factors in many plants. Even though the plant hormones pathway (ko04075) was not the significantly enriched pathway, there were still some certain genes related to the phytohormone identified in 466 DEGs, including eight auxin-related genes and two cytokinin-related genes.
Auxin, essential for the initiation and development of flowers (Yamaguchi et al. 2013), was observed to play a vital role in the sex determination of many plants, such as Carica papaya, Jatropha curcas, and Spinacia oleracea (Li et al. 2020; Xu et al. 2016; Zhou et al. 2019). Auxin-related genes, including auxin efflux carrier component 1,6 (PIN1,6), auxin-induced protein 6B (AX6B), auxin transporter-like protein 2 (LAX2), YUCCA2, auxin-responsive protein SAUR32, auxin-responsive protein IAA7 (IAA7) VAN3-binding protein (VAB), had higher expression in female flowers. Liu’s study pointed out that auxin may contribute to the development of male flowers, while our study suggested auxin also participates in the female flower development of S. chinensis (Liu et al. 2022).
Cytokinin is another common phytohormone. Exogenous application of cytokinin and its homologs can promote the formation of female flowers in many species, such as Castanea henryi, Sapium sebiferum, and Plukenetia volubilis (Luo et al. 2020; Wu et al. 2022; Ni et al. 2018). Cytokinin-degrading cytokinin oxidase/dehydrogenase (CKX) is essential for the female cone development of Dacrydium pectinatum (Wang et al. 2022). CKX3 can regulate the activity of the reproductive meristems of Arabidopsis thaliana (Bartrina et al. 2011). CYP735A2 is a cytochrome P450 monooxygenase (P450s) that catalyzes the biosynthesis of trans-Zeatin. CKX3 and CYP735A2 were upregulated in female flowers. Combined with function and expression pattern, we inferred that auxin and cytokinin may contribute to the female flower development in S. chinensis.
MCM proteins may participate in the ovule development of S. chinensis
DNA unwinding is the process in which the double-stranded DNA (dsDNA) is untwisted into two single-stranded DNAs (ssDNA) and catalyzed by DNA helicase. In eukaryotes, the DNA helicase consists of six minichromosome maintenance proteins (MCM), MCM2, MCM3, MCM4, MCM5, MCM6, and MCM7, which are highly conserved and form a hexamer (Zhang et al. 2020). Accumulated evidence indicated that MCM proteins may participate in the development and sex determination of flowers. Heterozygous MCM7 mutants caused ovule abortion in Arabidopsis thaliana (Herridge et al. 2014). It was reported that MCM protein genes MCM2 and MCM6 may play a significant role in the sex determination in Cucumis sativus (Wang et al. 2019b). MCM2 and MCM6 also participated in the female flower development of Phellodendron amurense (He et al. 2023). Four DNA helicase genes, MCM3, MCM5, MCM6, and MCM7, showed higher expression in female flowers and had only expression in male flowers at the early flowering stage. We suggest that MCM protein genes might engage in the development of ovules in S. chinensis.
Flavonoid biosynthesis pathway is essential for the male and female gametophytes of S. chinensis
KEGG analyses of 466 DEGs showed that eight DEGs were significantly enriched in the flavonoid biosynthesis pathway (ko00941), suggesting that the flavonoid metabolism might be a key difference between female and male flowers in S. chinensis. Besides these enriched genes, we also identified three genes related to flavonoid biosynthesis, MYB1, MYB123 (TT2), and DTX41 (TT12). Among these genes, DFRA (TT3), CHS (TT4), FL3H (TT6), and TT12 had high expression in both male and female flowers (TPM > 10). Accumulated evidence showed that flavonoids play a part in pollen development and are crucial for male fertility. In Arabidopsis, the DFRA-like gene DRL1 is essential for male fertility. The mutation of DRL1 could lead to impaired pollen formation (Tang et al. 2009). TT4, a key enzyme in the flavonoid biosynthesis pathway, plays a key role in the male fertility of rice (Wang et al. 2020b). The expression of TT4 was strongly inhibited in cytoplasmic male sterility (CMS) Raphanus sativus (Yang et al. 2008).
In addition to their potential roles in male fertility, flavonoids are also related to the development of female flowers. In Prunus mume, genes involved in flavonoid biosynthesis exhibited lower expression in abnormal pistils (Shi et al. 2012). The content of flavonoids was higher in Broussonetia papyrifera female flowers (Jiao et al. 2022). In our study, genes related to flavonoid biosynthesis were significantly upregulated in female flowers, suggesting that flavonoids may be more associated with the development of female flowers (Xia et al. 2021). Flavonoids are essential to the development of seeds and the control of seed size. Seed formation, a key process in plant reproduction, starts with megagametophyte development in the ovule (Ehlers et al. 2016). Endothelium differentiates from the inner layer of integument in the ovule. The formation of endothelium is one of the basic features of ovule development in tomatoes (Chaban et al. 2019). Proanthocyanins (PAs), a type of flavonoid in Arabidopsis, are accumulated in the endothelium (Xu et al. 2014). PAs are essential to ovule and seed development and important for female fertility. The TT12 gene acts as a flavonoid/H+-antiporter that controls the vacuolar sequestration of flavonoids in the seed coat endothelium (Debeaujon et al. 2001). The TT16 gene, encoding ARABIDOPSIS BSISTER (ABS) MADS-box domain protein. TT16, together with STK, is required for the endothelium formation. TT16 is expressed mainly in ovules and is necessary for proanthocyanin accumulation in the endothelium of the seed coat (Nesi et al. 2002). MYB1 acts as a transcription activator involved in the regulation of anthocyanin biosynthesis, activating genes such as TT3 (Liu et al. 2017). TT2 might activate the promoters of genes encoding the dedicated enzymes for anthocyanin biosynthesis (Wang et al. 2019a). Considering the function and expression pattern of genes related to flavonoid biosynthesis, we speculate that flavonoids might be related to the development of female gametophytes in S. chinensis. However, due to the limitations of our study, which specific type of flavonoids plays a crucial role in the S. chinensis female gametophytes needs further exploration.
Protein interaction network reveals potential genes involved in the sex differentiation of S. chinensis
In this study, 76 DEGs with medium confidence were selected for the construction of the PPI network. These genes are associated with flower development, DNA replication, flavonoid biosynthesis, and stomata development and may potentially participate in the sex differentiation of S. chinensis. Genes related to flower development including LFY, PIN1, and CKX3 are crucial for the initiation of flowers and the early development of gynoecium (Zúñiga-Mayo et al. 2019). Two MADS-box genes STK and TT16 are essential for the ovule and endothelium formation in Arabidopsis thaliana (Ehlers et al. 2016). Three YABBY genes AFO, INO, and YABBY5 are essential to flower development. INO may interact with BEL1, SUP, HEC3, LFY, ANT, STIP, and CUC3 to participate in the development of female organs (Villanueva et al. 1999). Flavonoids are essential to the normal male and female gametophyte. The flavonoid biosynthesis-related genes including DFR, F3H, MYB1, TT2, TT4, TT5, TT12, and TT16 might participate in the normal gametophyte development of S. chinensis. Free nuclear mitosis of megagametophytes is vital for normal ovule development. The abnormal expression of genes related to DNA replication such as MCM2~7 could lead to the arrest of the cell cycle and ovule abortion (Zhang et al. 2020). DNA replication–related genes including MCM3, MCM5, PCNA2, PRL, HTA6, and HMGB6 may play a potential role in the ovule development of S. chinensis. The stomata are essential for the photosynthesis process in Arabidopsis. Besides leaves, floral organs are also photosynthetically active (Brazel et al. 2023). The fruit stomata are important to seed development (Lugassi et al. 2020). Nevertheless, there is limited research on the relationship between stomata and flower sex differentiation, indicating that the role stomata may play in the sex differentiation of S. chinensis needs further investigation.
Conclusion
Our study successfully identified 466 DEGs that may be involved in the flower development and sex differentiation of S. chinensis. The KEGG enrichment analysis showed that flavonoid biosynthesis and DNA replication pathways were strongly related to the development of S. chinensis female flowers. MADS-box and YABBY gene family analyses provided insight into potential genes that influence flower development. The development of the stamen could be regulated by B, C, and E-class genes while C, D, and E-class genes along with Bsister genes could be crucial to the development of the gynoecium. The majority of YABBY genes exhibited increased expression in female flowers, suggesting that YABBY genes may be vital to the development of female flowers. In addition, auxin and cytokinin were thought to be required for the information of ovule primordia. Combined with quantitative real-time PCR validation experiment results, AFO, INO, YABBY2, YABBY5-1, YABBY5-2, AeAP3-2, STK, AIL-1, ANT, GATA18, TT2, TT3, TT4, TT6-1, TT6-2, TT12, MYB1, GAMYB, CEP1, and MST4 were preliminarily considered to be the key genes for the flower development and candidates for sex differentiation of S. chinensis. In summary, we have proposed a potential model for sex differentiation in S. chinensis (Fig. 8), which could contribute to the comprehension of the sex differentiation mechanism.
A proposed model for the elucidation of sex differentation in S. chinensis. Genes with blue colors represented male-specific genes. Genes with green colors represented female-specific genes. Genes with orange colors had high expression in both male and female flowers and were upregulated in female flowers. Genes with grey colors represented upregulated partially in male and female flowers
Data availability
All fastq raw sequence read data have been uploaded to the NCBI Sequence Read Archive (SRA) under accession number PRJNA1027980.
References
Adamczyk BJ, Lehti-Shiu MD, Fernandez DE (2007) The MADS domain factors AGL15 and AGL18 act redundantly as repressors of the floral transition in Arabidopsis. Plant J 50:1007–1019. https://doi.org/10.1111/j.1365-313X.2007.03105.x
Ali Z, Raza Q, Atif RM, Aslam U, Ajmal M, Chung G (2019) Genetic and molecular control of floral organ identity in cereals. Int J Mol Sci 20:2743. https://doi.org/10.3390/ijms20112743
Alonso-Peral MM, Li J, Li Y, Allen RS, Schnippenkoetter W, Ohms S, White RG, Millar AA (2010) The microRNA159-regulated GAMYB-like genes inhibit growth and promote programmed cell death in Arabidopsis. Plant Physiol 154:757–771. https://doi.org/10.1104/pp.110.160630
Alsubaie B, Kharabian-Masouleh A, Furtado A, Al-Dossary O, Al-Mssallem I, Henry RJ (2023) Highly sex specific gene expression in jojoba. BMC Plant Biol 23:440. https://doi.org/10.1186/s12870-023-04444-z
Arora R, Agarwal P, Ray S, Singh AK, Singh VP, Tyagi AK, Kapoor S (2007) MADS-box gene family in rice: genome-wide identification, organization and expression profiling during reproductive development and stress. BMC Genomics 8:242. https://doi.org/10.1186/1471-2164-8-242
Aya K, Ueguchi-Tanaka M, Kondo M, Hamada K, Yano K, Nishimura M, Matsuoka M (2009) Gibberellin modulates anther development in rice via the transcriptional regulation of GAMYB. Plant Cell 21:1453–1472. https://doi.org/10.1105/tpc.108.062935
Bačovský V, Čegan R, Tihlaříková E, Neděla V, Hudzieczek V, Smrža L, Janíček T, Beneš V, Hobza R (2022) Chemical genetics in Silene latifolia elucidate regulatory pathways involved in gynoecium development. J Exp Bot 73:2354–2368. https://doi.org/10.1093/jxb/erab538
Baker SC, Robinson-Beers K, Villanueva JM, Gaiser JC, Gasser CS (1997) Interactions among genes regulating ovule development in Arabidopsis thaliana. Genetics 145:1109–1124. https://doi.org/10.1093/genetics/145.4.1109
Bartrina I, Otto E, Strnad M, Werner T, Schmülling T (2011) Cytokinin regulates the activity of reproductive meristems, flower organ size, ovule formation, and thus seed yield in Arabidopsis thaliana. Plant Cell 23:69–80. https://doi.org/10.1105/tpc.110.079079
Berardini TZ, Reiser L, Li D, Mezheritsky Y, Muller R, Strait E, Huala E (2015) The Arabidopsis information resource: making and mining the “gold standard” annotated reference plant genome. Genesis 53:474–485. https://doi.org/10.1002/dvg.22877
Bowman JL (2000) The YABBY gene family and abaxial cell fate. Curr Opin Plant Biol 3:17–22. https://doi.org/10.1016/s1369-5266(99)00035-7
Brazel AJ, Fattorini R, McCarthy J, Franzen R, Rümpler F, Coupland G, Ó’Maoiléidigh DS (2023) AGAMOUS mediates timing of guard cell formation during gynoecium development. PLoS Genet 19:e1011000. https://doi.org/10.1371/journal.pgen.1011000
Buchfink B, Xie C, Huson DH (2015) Fast and sensitive protein alignment using DIAMOND. Nat Methods 12(1):59–60. https://doi.org/10.1038/nmeth.3176
Buttar ZA, Yang Y, Sharif R, Wu SN, Xie Y, Wang CJA (2020) Genome Wide Identification, characterization, and expression analysis of YABBY-gene family in wheat (Triticum aestivum L.). Agronomy 10:1189. https://doi.org/10.3390/agronomy10081189
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST+: architecture and applications. BMC Bioinformatics 10:421. https://doi.org/10.1186/1471-2105-10-421
Cao WL, Song Q, Meng XC (2015) Regulation on female-male flower ratio of monoecious plant Schisandra chinensis based on reactive oxygen species (ROS). Zhong Yao Cai 38:211–214. https://doi.org/10.13863/j.issn1001-4454.2015.02.001
Capella-Gutiérrez S, Silla-Martínez JM, Gabaldón T (2009) trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25:1972–1973. https://doi.org/10.1093/bioinformatics/btp348
Chaban I, Baranova E, Kononenko N, Khaliluev M, Smirnova E (2019) Distinct differentiation characteristics of endothelium determine its ability to form pseudo-embryos in tomato ovules. Int J Mol Sci 21:12. https://doi.org/10.3390/ijms21010012
Chen MS, Pan BZ, Fu Q, Tao YB, Martínez-Herrera J, Niu L, Ni J, Dong Y, Zhao ML, Xu ZF (2016) Comparative transcriptome analysis between gynoecious and monoecious plants identifies regulatory networks controlling sex determination in Jatropha curcas. Front Plant Sci 7:1953. https://doi.org/10.3389/fpls.2016.01953
Chen L, Zhang J, Li H, Niu J, Xue H, Liu B, Wang Q, Luo X, Zhang F, Zhao D, Cao S (2017) Transcriptomic analysis reveals candidate genes for female sterility in pomegranate flowers. Front Plant Sci 8:1430. https://doi.org/10.3389/fpls.2017.01430
Chen S, Zhou Y, Chen Y, Gu J (2018) fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34:i884–i890. https://doi.org/10.1093/bioinformatics/bty560
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, Xia R (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13:1194–1202. https://doi.org/10.1016/j.molp.2020.06.009
Crawford BC, Ditta G, Yanofsky MF (2007) The NTT gene is required for transmitting-tract development in carpels of Arabidopsis thaliana. Curr Biol 17:1101–1108. https://doi.org/10.1016/j.cub.2007.05.079
Debeaujon I, Peeters AJ, Léon-Kloosterziel KM, Koornneef M (2001) The TRANSPARENT TESTA12 gene of Arabidopsis encodes a multidrug secondary transporter-like protein required for flavonoid sequestration in vacuoles of the seed coat endothelium. Plant Cell 13:853–871. https://doi.org/10.1105/tpc.13.4.853
Díaz-Riquelme J, Lijavetzky D, Martínez-Zapater JM, Carmona MJ (2009) Genome-wide analysis of MIKCC-type MADS box genes in grapevine. Plant Physiol 149:354–369. https://doi.org/10.1104/pp.108.131052
Duan K, Fu H, Fang D, Wang K, Zhang W, Liu H, Sahu SK, Chen X (2022) Genome-wide analysis of the MADS-Box gene family in holoparasitic plants (Balanophora subcupularis and Balanophora fungosa var. globosa). Front Plant Sci 13:846697. https://doi.org/10.3389/fpls.2022.846697
Dubos C, Stracke R, Grotewold E, Weisshaar B, Martin C, Lepiniec L (2010) MYB transcription factors in Arabidopsis. Trends Plant Sci 15:573–581. https://doi.org/10.1016/j.tplants.2010.06.005
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797. https://doi.org/10.1093/nar/gkh340
Ehlers K, Bhide AS, Tekleyohans DG, Wittkop B, Snowdon RJ, Becker A (2016) The MADS Box genes ABS, SHP1, and SHP2 are essential for the coordination of cell divisions in ovule and seed coat development and for endosperm formation in Arabidopsis thaliana. PLoS One 11:e0165075. https://doi.org/10.1371/journal.pone.0165075
Feng X, Yang Z, Xiu-Rong W, Ying W (2020) Transcriptomic differences between male and female Trachycarpus fortunei. Sci Rep-UK 10:12338. https://doi.org/10.1038/s41598-020-69107-7
Fu L, Niu B, Zhu Z, Wu S, Li W (2012) CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28:3150–3152. https://doi.org/10.1093/bioinformatics/bts565
Fu Q, Niu L, Chen MS, Tao YB, Wang X, He H, Pan BZ, Xu ZF (2018) De novo transcriptome assembly and comparative analysis between male and benzyladenine-induced female inflorescence buds of Plukenetia volubilis. J Plant Physiol 221:107–118. https://doi.org/10.1016/j.jplph.2017.12.006
Galego L, Almeida J (2002) Role of DIVARICATA in the control of dorsoventral asymmetry in Antirrhinum flowers. Gene Dev 16:880–891. https://doi.org/10.1101/gad.221002
Gonçalves B, Hasson A, Belcram K, Cortizo M, Morin H, Nikovics K, Vialette-Guiraud A, Takeda S, Aida M, Laufs P, Arnaud N (2015) A conserved role for CUP-SHAPED COTYLEDON genes during ovule development. Plant J 83:732–742. https://doi.org/10.1111/tpj.12923
Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I, Adiconis X, Fan L, Raychowdhury R, Zeng Q, Chen Z, Mauceli E, Hacohen N, Gnirke A, Rhind N, di Palma F, Birren BW, Nusbaum C, Lindblad-Toh K, Friedman N, Regev A (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29:644–652. https://doi.org/10.1038/nbt.1883
Gremski K, Ditta G, Yanofsky MF (2007) The HECATE genes regulate female reproductive tract development in Arabidopsis thaliana. Development 134:3593–3601. https://doi.org/10.1242/dev.011510
Harkess A, Leebens-Mack J (2016) A century of sex determination in flowering plants. J Hered 108:69–77. https://doi.org/10.1093/jhered/esw060
Harkess A, Mercati F, Shan HY, Sunseri F, Falavigna A, Leebens-Mack J (2015) Sex-biased gene expression in dioecious garden asparagus (Asparagus officinalis). New Phytol 207:883–892. https://doi.org/10.1111/nph.13389
He L, Fan Y, Zhang Z, Wei X, Yu J (2023) Identifying genes associated with female flower development of Phellodendron amurense Rupr. using a transcriptomics approach. Genes-Basal 14:661. https://doi.org/10.3390/genes14030661
Herridge RP, Day RC, Macknight RC (2014) The role of the MCM2–7 helicase complex during Arabidopsis seed development. Plant Mol Biol 86:69–84. https://doi.org/10.1007/s11103-014-0213-x
Hui W, Fan J, Liu X, Zhao F, Saba T, Wang J, Wu A, Zhang X, Zhang J, Zhong Y, Chen G, Gong W (2022) Integrated transcriptome and plant growth substance profiles to identify the regulatory factors involved in floral sex differentiation in Zanthoxylum armatum DC. Front Plant Sci 13:976338. https://doi.org/10.3389/fpls.2022.976338
Jiao P, Chaoyang L, Wenhan Z, Jingyi D, Yunlin Z, Zhenggang X (2022) Integrative metabolome and transcriptome analysis of flavonoid biosynthesis genes in Broussonetia papyrifera leaves from the perspective of sex differentiation. Front Plant Sci 13:900030. https://doi.org/10.3389/fpls.2022.900030
Kalyaanamoorthy S, Minh BQ, Wong TKF, von Haeseler A, Jermiin LS (2017) ModelFinder: fast model selection for accurate phylogenetic estimates. Nat Methods 14:587–589. https://doi.org/10.1038/nmeth.4285
Khadka J, Yadav NS, Guy M, Grafi G, Golan-Goldhirsh A (2019) Epigenetic aspects of floral homeotic genes in relation to sexual dimorphism in the dioecious plant Mercurialis annua. J Exp Bot 70:6245–6259. https://doi.org/10.1093/jxb/erz379
Klucher KM, Chow H, Reiser L, Fischer RL (1996) The AINTEGUMENTA gene of Arabidopsis required for ovule and female gametophyte development is related to the floral homeotic gene APETALA2. Plant Cell 8:137–153. https://doi.org/10.1105/tpc.8.2.137
Kramer EM, Irish VF (2000) Evolution of the petal and stamen developmental programs: evidence from comparative studies of the lower eudicots and basal angiosperms. Int J Plant Sci 161:S29–S40. https://doi.org/10.1086/317576
Le SQ, Gascuel O (2008) An improved general amino acid replacement matrix. Mol Biol Evol 25:1307–1320. https://doi.org/10.1093/molbev/msn067
Leite Montalvão AP, Kersten B, Kim G, Fladung M, Müller NA (2022) ARR17 controls dioecy in Populus by repressing B-class MADS-box gene expression. Philos Trans R Soc Lond B Biol Sci 377:20210217. https://doi.org/10.1098/rstb.2021.0217
Letunic I, Bork P (2021) Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res 49:W293–W296. https://doi.org/10.1093/nar/gkab301
Letunic I, Khedkar S, Bork P (2021) SMART: recent updates, new developments and status in 2020. Nucleic Acids Res 49:D458-d460. https://doi.org/10.1093/nar/gkaa937
Li B, Dewey CN (2011) RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12:323. https://doi.org/10.1186/1471-2105-12-323
Li N, Meng Z, Tao M, Wang Y, Zhang Y, Li S, Gao W, Deng C (2020) Comparative transcriptome analysis of male and female flowers in Spinacia oleracea L. BMC Genomics 21:850. https://doi.org/10.1186/s12864-020-07277-4
Liu Y, Cui S, Wu F, Yan S, Lin X, Du X, Chong K, Schilling S, Theißen G, Meng Z (2013) Functional conservation of MIKC*-Type MADS box genes in Arabidopsis and rice pollen maturation. Plant Cell 25:1288–1303. https://doi.org/10.1105/tpc.113.110049
Liu Y, Zhou B, Qi Y, Chen X, Liu C, Liu Z, Ren X (2017) Expression differences of pigment structural genes and transcription factors explain flesh coloration in three contrasting kiwifruit cultivars. Front Plant Sci 8:1507. https://doi.org/10.3389/fpls.2017.01507
Liu X, Zhang L, Yang S (2022) Analysis of floral organ development and sex determination in Schisandra chinensis by scanning electron microscopy and rna-sequencing. Life-Basel 12:1260. https://doi.org/10.3390/life12081260
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550. https://doi.org/10.1186/s13059-014-0550-8
Lu S, Wang J, Chitsaz F, Derbyshire MK, Geer RC, Gonzales NR, Gwadz M, Hurwitz DI, Marchler GH, Song JS, Thanki N, Yamashita RA, Yang M, Zhang D, Zheng C, Lanczycki CJ, Marchler-Bauer A (2020) CDD/SPARCLE: the conserved domain database in 2020. Nucleic Acids Res 48:D265-d268. https://doi.org/10.1093/nar/gkz991
Lu Y, Zeng J, Liu Q (2021) The rice miR396-GRF-GIF-SWI/SNF module: a player in GA signaling. Front Plant Sci 12:786641. https://doi.org/10.3389/fpls.2021.786641
Lugassi N, Kelly G, Arad T, Farkash C, Yaniv Y, Yeselson Y, Schaffer AA, Raveh E, Granot D, Carmi N (2020) Expression of hexokinase in stomata of citrus fruit reduces fruit transpiration and affects seed development. Front Plant Sci 11:255. https://doi.org/10.3389/fpls.2020.00255
Luo Y, Pan BZ, Li L, Yang CX, Xu ZF (2020) Developmental basis for flower sex determination and effects of cytokinin on sex determination in Plukenetia volubilis (Euphorbiaceae). Plant Reprod 33:21–34. https://doi.org/10.1007/s00497-019-00382-9
Mao Y, Liu W, Chen X, Xu Y, Lu W, Hou J, Ni J, Wang Y, Wu L (2017) Flower development and sex determination between male and female flowers in Vernicia fordii. Front Plant Sci 8:1291. https://doi.org/10.3389/fpls.2017.01291
Masuda K, Ikeda Y, Matsuura T, Kawakatsu T, Tao R, Kubo Y, Ushijima K, Henry IM, Akagi T (2022) Reinvention of hermaphroditism via activation of a RADIALIS-like gene in hexaploid persimmon. Nat Plants 8:217–224. https://doi.org/10.1038/s41477-022-01107-z
Mistry J, Chuguransky S, Williams L, Qureshi M, Salazar GA, Sonnhammer ELL, Tosatto SCE, Paladin L, Raj S, Richardson LJ, Finn RD, Bateman A (2021) Pfam: The protein families database in 2021. Nucleic Acids Res 49:D412-d419. https://doi.org/10.1093/nar/gkaa913
Mizzotti C, Mendes MA, Caporali E, Schnittger A, Kater MM, Battaglia R, Colombo L (2012) The MADS box genes SEEDSTICK and ARABIDOPSIS Bsister play a maternal role in fertilization and seed development. Plant J 70:409–420. https://doi.org/10.1111/j.1365-313X.2011.04878.x
Morel P, Chambrier P, Boltz V, Chamot S, Rozier F, Rodrigues Bento S, Trehin C, Monniaux M, Zethof J, Vandenbussche M (2019) Divergent functional diversification patterns in the SEP/AGL6/AP1 MADS-box transcription factor superclade. Plant Cell 31:3033–3056. https://doi.org/10.1105/tpc.19.00162
Nagasawa N, Miyoshi M, Sano Y, Satoh H, Hirano H, Sakai H, Nagato Y (2003) SUPERWOMAN1 and DROOPING LEAF genes control floral organ identity in rice. Development 130:705–718. https://doi.org/10.1242/dev.00294
Nesi N, Debeaujon I, Jond C, Stewart AJ, Jenkins GI, Caboche M, Lepiniec L (2002) The TRANSPARENT TESTA16 locus encodes the ARABIDOPSIS BSISTER MADS domain protein and is required for proper development and pigmentation of the seed coat. Plant Cell 14:2463–2479. https://doi.org/10.1105/tpc.004127
Nguyen LT, Schmidt HA, von Haeseler A, Minh BQ (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32:268–274. https://doi.org/10.1093/molbev/msu300
Ni J, Shah FA, Liu W, Wang Q, Wang D, Zhao W, Lu W, Huang S, Fu S, Wu L (2018) Comparative transcriptome analysis reveals the regulatory networks of cytokinin in promoting the floral feminization in the oil plant Sapium sebiferum. BMC Plant Biol 18(1):96. https://doi.org/10.1186/s12870-018-1314-5
Niu H, Liu X, Tong C, Wang H, Li S, Lu L, Pan Y, Zhang X, Weng Y, Li Z (2018) The WUSCHEL-related homeobox1 gene of cucumber regulates reproductive organ development. J Exp Bot 69:5373–5387. https://doi.org/10.1093/jxb/ery329
Pannell JR (2017) Plant sex determination. Curr Bio 27:R191–R197. https://doi.org/10.1016/j.cub.2017.01.052
Pawełkowicz M, Pryszcz L, Skarzyńska A, Wóycicki RK, Posyniak K, Rymuszka J, Przybecki Z, Pląder W (2019) Comparative transcriptome analysis reveals new molecular pathways for cucumber genes related to sex determination. Plant Reprod 32:193–216. https://doi.org/10.1007/s00497-019-00362-z
Pelayo MA, Yamaguchi N, Ito T (2021) One factor, many systems: the floral homeotic protein AGAMOUS and its epigenetic regulatory mechanisms. Cur Opin Plant Biology 61:102009. https://doi.org/10.1016/j.pbi.2021.102009
Peréz-Mesa P, Ortíz-Ramírez CI, González F, Ferrándiz C, Pabón-Mora N (2020) Expression of gynoecium patterning transcription factors in Aristolochia fimbriata (Aristolochiaceae) and their contribution to gynostemium development. EvoDevo 11:4. https://doi.org/10.1186/s13227-020-00149-8
Petrella R, Gabrieli F, Cavalleri A, Schneitz K, Colombo L, Cucinotta M (2022) Pivotal role of STIP in ovule pattern formation and female germline development in Arabidopsis thaliana. Development 149:dev201184. https://doi.org/10.1242/dev.201184
Pinyopich A, Ditta GS, Savidge B, Liljegren SJ, Baumann E, Wisman E, Yanofsky MF (2003) Assessing the redundancy of MADS-box genes during carpel and ovule development. Nature 424:85–88. https://doi.org/10.1038/nature01741
Potter SC, Luciani A, Eddy SR, Park Y, Lopez R, Finn RD (2018) HMMER web server: 2018 update. Nucleic Acids ResS 46:W200-w204. https://doi.org/10.1093/nar/gky448
Ren L, Sun H, Dai S, Feng S, Qiao K, Wang J, Gong S, Zhou A (2021) Identification and characterization of MIKC(c)-Type MADS-box genes in the flower organs of Adonis amurensis. Int J Mol Sci 22:9362. https://doi.org/10.3390/ijms22179362
Rijpkema AS, Zethof J, Gerats T, Vandenbussche M (2009) The petunia AGL6 gene has a SEPALLATA-like function in floral patterning. Plant J 60:1–9. https://doi.org/10.1111/j.1365-313X.2009.03917.x
Rodríguez-Cazorla E, Ortuño-Miquel S, Candela H, Bailey-Steinitz LJ, Yanofsky MF, Martínez-Laborda A, Ripoll JJ, Vera A (2018) Ovule identity mediated by pre-mRNA processing in Arabidopsis. PLoS Genet 14:e1007182. https://doi.org/10.1371/journal.pgen.1007182
Romanova MA, Maksimova AI, Pawlowski K, Voitsekhovskaja OV (2021) YABBY genes in the development and evolution of land plants. Int J Mol Sci 22:4139. https://doi.org/10.3390/ijms22084139
Sakai H, Medrano LJ, Meyerowitz EM (1995) Role of SUPERMAN in maintaining Arabidopsis floral whorl boundaries. Nature 378:199–203. https://doi.org/10.1038/378199a0
Sakai H, Krizek BA, Jacobsen SE, Meyerowitz EM (2000) Regulation of SUP expression identifies multiple regulators involved in arabidopsis floral meristem development. Plant Cell 12:1607–1618. https://doi.org/10.1105/tpc.12.9.1607
Sawa S, Watanabe K, Goto K, Liu YG, Shibata D, Kanaya E, Morita EH, Okada K (1999) FILAMENTOUS FLOWER, a meristem and organ identity gene of Arabidopsis, encodes a protein with a zinc finger and HMG-related domains. Gene Dev 13:1079–1088. https://doi.org/10.1101/gad.13.9.1079
Schilling S, Kennedy A, Pan S, Jermiin LS, Melzer R (2020) Genome-wide analysis of MIKC-type MADS-box genes in wheat: pervasive duplications, functional conservation and putative neofunctionalization. New Phytol 225:511–529. https://doi.org/10.1111/nph.16122
Sengupta A, Hileman LC (2022) A CYC-RAD-DIV-DRIF interaction likely pre-dates the origin of floral monosymmetry in Lamiales. EvoDevo 13:3. https://doi.org/10.1186/s13227-021-00187-w
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504. https://doi.org/10.1101/gr.1239303
Shi T, Zhuang W, Zhang Z, Sun H, Wang L, Gao Z (2012) Comparative proteomic analysis of pistil abortion in Japanese apricot (Prunus mume Sieb. et Zucc). J Plant Physiol 169:1301–1310. https://doi.org/10.1016/j.jplph.2012.05.009
Siegfried KR, Eshed Y, Baum SF, Otsuga D, Drews GN, Bowman JL (1999) Members of the YABBY gene family specify abaxial cell fate in Arabidopsis. Development 126:4117–4128. https://doi.org/10.1242/dev.126.18.4117
Simão FA, Waterhouse RM, Ioannidis P, Kriventseva EV, Zdobnov EM (2015) BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31:3210–3212. https://doi.org/10.1093/bioinformatics/btv351
Singh VK, Mangalam AK, Dwivedi S, Naik S (1998) Primer premier: program for design of degenerate primers from a protein sequence. BioTechniques 24:318–319. https://doi.org/10.2144/98242pf02
Skinner DJ, Dang T, Gasser CS (2023) The Arabidopsis INNER NO OUTER (INO) gene acts exclusively and quantitatively in regulation of ovule outer integument development. Plant direct 7(2):e485. https://doi.org/10.1002/pld3.485
Smith-Unna R, Boursnell C, Patro R, Hibberd JM, Kelly S (2016) TransRate: reference-free quality assessment of de novo transcriptome assemblies. Genome Res 26:1134–1144. https://doi.org/10.1101/gr.196469.115
Szklarczyk D, Kirsch R, Koutrouli M, Nastou K, Mehryary F, Hachilif R, Gable AL, Fang T, Doncheva NT, Pyysalo S, Bork P, Jensen LJ, von Mering C (2023) The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res 51:D638-d646. https://doi.org/10.1093/nar/gkac1000
Tang LK, Chu H, Yip WK, Yeung EC, Lo C (2009) An anther-specific dihydroflavonol 4-reductase-like gene (DRL1) is essential for male fertility in Arabidopsis. New Phytol 181:576–587. https://doi.org/10.1111/j.1469-8137.2008.02692.x
Vekemans X, Castric V, Hipperson H, Müller NA, Westerdahl H, Cronk Q (2021) Whole-genome sequencing and genome regions of special interest: Lessons from major histocompatibility complex, sex determination, and plant self-incompatibility. Mol Eco 30:6072–6086. https://doi.org/10.1111/mec.16020
Villanueva JM, Broadhvest J, Hauser BA, Meister RJ, Schneitz K, Gasser CS (1999) INNER NO OUTER regulates abaxial- adaxial patterning in Arabidopsis ovules. Gene Dev 13:3160–3169. https://doi.org/10.1101/gad.13.23.3160
Vingron TMAM (2000) Modeling amino acid replacement. J Comput Biol 7:761–776. https://doi.org/10.1089/10665270050514918
Wang L, Tang W, Hu Y, Zhang Y, Sun J, Guo X, Lu H, Yang Y, Fang C, Niu X, Yue J, Fei Z, Liu Y (2019) A MYB/bHLH complex regulates tissue-specific anthocyanin biosynthesis in the inner pericarp of red-centered kiwifruit Actinidia chinensis cv. Hongyang. Plant J 99:359–378. https://doi.org/10.1111/tpj.14330
Wang R, Lin Y, Jin Q, Yao C, Zhong Y, Wu T (2019) RNA-Seq analysis of gynoecious and weak female cucumber revealing the cell cycle pathway may regulate sex determination in cucumber. Gene 687:289–297. https://doi.org/10.1016/j.gene.2018.11.071
Wang Y, Zhang J, Hu Z, Guo X, Tian S, Chen G (2019) Genome-wide analysis of the MADS-Box transcription factor family in Solanum lycopersicum. Int J Mol Sci 20:2961. https://doi.org/10.3390/ijms20122961
Wang D, Hao Z, Long X, Wang Z, Zheng X, Ye D, Peng Y, Wu W, Hu X, Wang G, Zheng R, Shi J, Chen J (2020) The transcriptome of Cunninghamia lanceolata male/female cone reveal the association between MIKC MADS-box genes and reproductive organs development. BMC Plant Biol 20:508. https://doi.org/10.1186/s12870-020-02634-7
Wang L, Lam PY, Lui ACW, Zhu FY, Chen MX, Liu H, Zhang J, Lo C (2020) Flavonoids are indispensable for complete male fertility in rice. J Exp Bot 71:4715–4728. https://doi.org/10.1093/jxb/eraa204
Wang E, Lu W, Liang H, Zhang X, Huo S, Song X, Wang J, Zhao Y (2022) Morpho-histology, endogenous hormone dynamics, and transcriptome profiling in Dacrydium pectinatum during female cone development. Front Plant Sci 13:954788. https://doi.org/10.3389/fpls.2022.954788
Whipple CJ, Ciceri P, Padilla CM, Ambrose BA, Bandong SL, Schmidt RJ (2004) Conservation of B-class floral homeotic gene function between maize and Arabidopsis. Development 131:6083–6091. https://doi.org/10.1242/dev.01523
Wu GL, Zhu ZJ, Qiu Q, Fan XM, Yuan DY (2022) Transcriptome analysis reveals the regulatory networks of cytokinin in promoting floral feminization in Castanea henryi. Int J Mol Sci 23:6389. https://doi.org/10.3390/ijms23126389
Wu G, Tian X, Qiu Q, Zhang Y, Fan X, Yuan D (2023) Dynamic cytological and transcriptomic analyses provide novel insights into the mechanisms of sex determination in Castanea henryi. Front Plant Sci 14:1257541. https://doi.org/10.3389/fpls.2023.1257541
Xia J, Wang D, Peng Y, Wang W, Wang Q, Xu Y, Li T, Zhang K, Li J, Xu X (2021) Genome-wide analysis of the YABBY transcription factor family in rapeseed (Brassica napus L.). Genes-Basel 12:981. https://doi.org/10.3390/genes12070981
Xin GL, Liu JQ, Liu J, Ren XL, Du XM, Liu WZ (2019) Anatomy and RNA-Seq reveal important gene pathways regulating sex differentiation in a functionally Androdioecious tree. Tapiscia sinensis. BMC Plant Biol 19:554. https://doi.org/10.1186/s12870-019-2081-7
Xu W, Grain D, Bobet S, Le Gourrierec J, Thévenin J, Kelemen Z, Lepiniec L, Dubos C (2014) Complexity and robustness of the flavonoid transcriptional regulatory network revealed by comprehensive analyses of MYB-bHLH-WDR complexes and their targets in Arabidopsis seed. New Phytol 202:132–144. https://doi.org/10.1111/nph.12620
Xu G, Huang J, Yang Y, Yao YA (2016) Transcriptome analysis of flower sex differentiation in Jatropha curcas L. using RNA sequencing. PLoS One 11:e0145613. https://doi.org/10.1371/journal.pone.0145613
Yamada T, Ito M, Kato M (2004) YABBY2-homologue expression in lateral organs of Amborella trichopoda (Amborellaceae). Int J Plant Sci 165:917–924. https://doi.org/10.1086/423793
Yamaguchi T, Nagasawa N, Kawasaki S, Matsuoka M, Nagato Y, Hirano HY (2004) The YABBY gene DROOPING LEAF regulates carpel specification and midrib development in Oryza sativa. Plant Cell 16:500–509. https://doi.org/10.1105/tpc.018044
Yamaguchi T, Lee DY, Miyao A, Hirochika H, An G, Hirano HY (2006) Functional diversification of the two C-class MADS box genes OSMADS3 and OSMADS58 in Oryza sativa. Plant Cell 18:15–28. https://doi.org/10.1105/tpc.105.037200
Yamaguchi N, Wu MF, Winter CM, Berns MC, Nole-Wilson S, Yamaguchi A, Coupland G, Krizek BA, Wagner D (2013) A molecular framework for auxin-mediated initiation of flower primordia. Dev Cell 24:271–282. https://doi.org/10.1016/j.devcel.2012.12.017
Yang Z(1994)Maximum likelihood phylogenetic estimation from DNA sequences with variable rates over sites: Approximate methods. J Mol Evol 39, 306–314. https://doi.org/10.1007/BF00160154
Yang S, Terachi T, Yamagishi H (2008) Inhibition of chalcone synthase expression in anthers of Raphanus sativus with Ogura male sterile cytoplasm. Ann Bot 102:483–489. https://doi.org/10.1093/aob/mcn116
Yang HW, Akagi T, Kawakatsu T, Tao R (2019) Gene networks orchestrated by MeGI: a single-factor mechanism underlying sex determination in persimmon. Plant J 98:97–111. https://doi.org/10.1111/tpj.14202
Yang K, Qiu J, Huang Z, Yu Z, Wang W, Hu H, You Y (2022) A comprehensive review of ethnopharmacology, phytochemistry, pharmacology, and pharmacokinetics of Schisandra chinensis (Turcz.) Baill. and Schisandra sphenanthera Rehd. et Wils. J Ethnopharmacol 284:114759. https://doi.org/10.1016/j.jep.2021.114759
Ye LX, Luo MM, Wang Z, Bai FX, Luo X, Gao L, Peng J, Chen QH, Zhang L (2022) Genome-wide analysis of MADS-box gene family in kiwifruit (Actinidia chinensis var. chinensis) and their potential role in floral sex differentiation. Front Genet 13:1043178. https://doi.org/10.3389/fgene.2022.1043178
Yin S, Li S, Gao Y, Bartholomew ES, Wang R, Yang H, Liu C, Chen X, Wang Y, Liu X, Ren H (2022) Genome-wide identification of YABBY gene family in cucurbitaceae and expression analysis in cucumber (Cucumis sativus L.). Genes-Basel 13:467. https://doi.org/10.3390/genes13030467
Zhang L, Cao B, Bai C (2013) New reports of nuclear DNA content for 66 traditional Chinese medicinal plant taxa in China. Caryologia-Basel 66:375–383. https://doi.org/10.1080/00087114.2013.859443
Zhang M, Li W, Feng J, Gong Z, Yao Y, Zheng C (2020) Integrative transcriptomics and proteomics analysis constructs a new molecular model for ovule abortion in the female-sterile line of Pinus tabuliformis Carr. Plant Sci 294:110462. https://doi.org/10.1016/j.plantsci.2020.110462
Zhang S, Tan F-Q, Chung C-H, Slavkovic F, Devani RS, Troadec C, Marcel F, Morin H, Camps C, Gomez Roldan MV, Benhamed M, Dogimont C, Boualem A, Bendahmane A (2022) The control of carpel determinacy pathway leads to sex determination in cucurbits. Science 378:543–549. https://doi.org/10.1126/science.add4250
Zhang X, Pan L, Guo W, Li Y, Wang W (2022) A convergent mechanism of sex determination in dioecious plants: distinct sex-determining genes display converged regulation on floral B-class genes. Front Plant Sci 13:953445. https://doi.org/10.3389/fpls.2022.953445
Zhang Z, Quan S, Niu J, Guo C, Kang C, Liu J, Yuan X (2022) Comprehensive identification and analyses of the GRF gene family in the whole-genome of four Juglandaceae species. Int J Mol Sci 23:12663. https://doi.org/10.3390/ijms232012663
Zhao ML, Chen MS, Ni J, Xu CJ, Yang Q, Xu ZF (2020) Comparative transcriptome analysis of gynoecious and monoecious inflorescences reveals regulators involved in male flower development in the woody perennial plant Jatropha curcas. Plant Reprod 33:191–204. https://doi.org/10.1007/s00497-020-00396-8
Zhou P, Fatima M, Ma X, Liu J, Ming R (2019) Auxin regulation involved in gynoecium morphogenesis of papaya flowers. Hortic Res 6:119. https://doi.org/10.1038/s41438-019-0205-8
Zhou G, Yin H, Chen F, Wang Y, Gao Q, Yang F, He C, Zhang L, Wan Y (2022) The genome of Areca catechu provides insights into sex determination of monoecious plants. New Phytol 236:2327–2343. https://doi.org/10.1111/nph.18471
Zúñiga-Mayo VM, Gómez-Felipe A, Herrera-Ubaldo H, de Folter S (2019) Gynoecium development: networks in Arabidopsis and beyond. J Exp Bot 70:1447–1460. https://doi.org/10.1093/jxb/erz026
Acknowledgements
The authors appreciate Prof. Zhao Zhang’s help in sample collection, and Li-Hong He, Rui Xu, and Rui-Lin Xiong’s help in the experiment.
Funding
This work was financially supported by the National Natural Science Foundation of China (Grant No. 82274050) and the Chinese Academy of Medical Sciences Innovation Fund for Medical Sciences (Grant No. 2021-I2M-1–032).
Author information
Authors and Affiliations
Contributions
JC and XW collected samples. JC performed the experiments, analyzed the results, and wrote the manuscript. YC assisted in the experiment operation. XW was involved in the design of these experiments and data interpretation. YQ, BZ, and HL conceived and directed the experiments and writing. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.

Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cheng, JL., Wei, XP., Chen, Y. et al. Comparative transcriptome analysis reveals candidate genes related to the sex differentiation of Schisandra chinensis. Funct Integr Genomics 23, 344 (2023). https://doi.org/10.1007/s10142-023-01264-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10142-023-01264-0