Abstract
The purpose of this study was to investigate the expression significance, predictive value, immunologic function, and biological role of transmembrane protein 158 (TMEM158) in the development of pan-cancer. To achieve this, we utilized data from multiple databases, including TCGA, GTEx, GEPIA, and TIMER, to collect gene transcriptome, patient prognosis, and tumor immune data. We evaluated the association of TMEM158 with patient prognosis, tumor mutational burden (TMB), and microsatellite instability (MSI) in pan-cancer samples. We performed immune checkpoint gene co-expression analysis and gene set enrichment analysis (GSEA) to better understand the immunologic function of TMEM158. Our findings revealed that TMEM158 was significantly differentially expressed between most types of cancer tissues and their adjacent normal tissues and was associated with prognosis. Moreover, TMEM158 was significantly correlated with TMB, MSI, and tumor immune cell infiltration in multiple cancers. Co-expression analysis of immune checkpoint genes showed that TMEM158 was related to the expression of several common immune checkpoint genes, especially CTLA4 and LAG3. Gene enrichment analysis further revealed that TMEM158 was involved in multiple immune-related biological pathways in pan-cancer. Overall, this systematic pan-cancer analysis suggests that TMEM158 is generally highly expressed in various cancer tissues and is closely related to patient prognosis and survival across multiple cancer types. TMEM158 may serve as a significant predictor of cancer prognosis and modulate immune responses to various types of cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Membrane proteins are a broad class of proteins that can bind or integrate into cell membranes or organelle membranes. These proteins represent approximately 20–30% of the proteome of most organisms and comprise more than 40% of drug targets (Carpenter et al. 2008). Membrane proteins localized on the plasma membrane can be immobilized by ligands and receptors or internalized by small biomolecules. They play critical roles in signal transduction between cells and between cells and the extracellular environment, making them prime targets for drug research and development. Numerous studies have demonstrated that membrane proteins are crucial for the onset and progression of cancer cells (Chitwood and Hegde 2019; Kanai 2022; Banerjee et al. 2022; Fan and Huang 2022). There are three distinct types of membrane proteins based on their binding strength and location: peripheral membrane proteins, integral membrane proteins, and lipid-anchored membrane proteins. Transmembrane proteins (TMEM) are integral membrane proteins that possess at least one transmembrane segment that entirely or partially passes through biological membranes (Marx et al. 2020; Du et al. 2022). Prior research has indicated that TMEM family proteins can impact the activation of glycoprotein ligands on the tumor cell membrane, as well as influence the structure of the bilayer lipid membrane, leading to the activation of the G protein-coupled pathway within the tumor cell membrane. Consequently, the second messenger is activated, resulting in abnormal activation of genes transcribed in the tumor cell nucleus (Schmit and Michiels 2018).
Among the genes in the TMEM family, TMEM158 is a pivotal member also known as BBP, HBBP, RIS1, and P40BBP. Its gene is located on chromosome 3p21.31, which was first identified as an inducible gene of rat sarcoma (Barradas et al. 2002). TMEM158 has been reported to play a role in various tumors by affecting the progression and prognosis of tumor cells through modulating tumor cell invasion, migration, and angiogenesis (Silva et al. 2006). Specifically, Mohammed et al. demonstrated that targeted knockout of TMEM158 reduced the toxic effects of cisplatin on non-small cell lung cancer cells, indicating its relationship with chemotherapy drug sensitivity (Mohammed Ael et al. 2012). In colon cancer development, Iglesias et al. suggested that TMEM158 acts as an oncogene (Iglesias et al. 2006), while Zirn et al. showed that TMEM158 is highly expressed in CTNNB1-mutated Wilms tumors, which may be linked to Ras and Wnt signaling pathways (Zirn et al. 2006). TMEM158 was found to be abnormally upregulated in pancreatic cancer according to Fu et al. Additionally, a further study revealed that TMEM158 activated TGF-1 and PI3K/AKT pathways, promoting pancreatic cancer proliferation, migration, and invasion (Fu et al. 2020). Furthermore, in a study by Tong et al., overexpression of TMEM158 was found to participate in EMT by activating the TGF-β pathway, thereby promoting migration, invasion, and metastasis of breast cancer cells (Tong et al. 2022). A recent study published in August 2022 by Li et al. showed that knockdown of TMEM158 inhibited the proliferation of glioblastoma cells, while overexpression of TMEM158 promoted tumor cell migration by stimulating EMT (Li et al. 2022). Moreover, Cheng et al. observed that TMEM158 was abnormally upregulated in ovarian cancer tissues, and its overexpression played a crucial role in the proliferation, invasion, and adhesion of ovarian cancer cells. Targeted knockout of TMEM158 significantly inhibited the proliferation, invasion, and metastasis of ovarian cancer cells. TMEM158 was found to mediate the malignant biological behavior of ovarian cancer by regulating the cycle and TGF-β pathway, suggesting that it may become a therapeutic target for ovarian cancer (Cheng et al. 2015). Nonetheless, further research is needed to investigate whether TMEM158 affects the occurrence and development of other tumors.
Due to the absence of a comprehensive analysis of TMEM158 in pan-cancer, we gathered expression data from 33 human cancers in the TCGA and GTEx databases to investigate the expression of TMEM158 in different cancer types, as well as its potential correlation with patient prognosis and tumor immunity. We believe this study could offer novel insights and perspectives in understanding the involvement of TMEM158 in tumor progression and immune regulation.
Materials and methods
Data acquisition and processing
In this study, we utilized the RNA-seq data and clinical information of 33 cancer types from the Cancer Genome Atlas (TCGA) database (http://cancergenome.nih.gov). However, due to the absence of normal tissue sequencing data in the TCGA database, and insufficient transcriptome sequencing data for normal tissue in some patients, we sourced RNA-seq data for normal tissue from the Genotype-Tissue Expression (GTEx) database (http://commonfund.nih.gov/GTEx/) (Aguet et al. 2020; Eraslan et al. 2022; Kim-Hellmuth et al. 2020).
mRNA expression analysis of TMEM158 in pan-cancer
In order to enable meaningful comparisons of gene expression levels across different samples, we preprocessed the data by converting the FPKM values to TPM values and normalizing them using Log2 transformation. We subsequently performed an in-depth investigation of the expression patterns of TMEM158 across 33 types of cancer and their corresponding normal tissues. Our analyses revealed significant differences in TMEM158 expression levels across different cancer types, which we visualized using a range of informative and intuitive visualizations.
Prognostic analysis of TMEM158 expression in pan-cancer
We comprehensively analyzed TMEM158 expression in pan-cancer using the Gene Expression Profiling Interactive Analysis (GEPIA) public database. GEPIA is a powerful platform designed to facilitate a comprehensive analysis of gene expression profiles in cancer and normal tissues, and addresses a critical need for dynamic analysis of large-scale cancer genomics data (Tang et al. 2019). By leveraging RNA-sequencing data from TCGA and GTEx, we were able to examine the expression levels and prognostic relevance of TMEM158 across multiple cancer types.
Association of TMEM158 expression with tumor mutational burden and microsatellite instability in pan-cancer
We explored the relationship between TMEM158 expression and two key biomarkers of immunotherapy response, tumor mutational burden (TMB) and microsatellite instability (MSI), in pan-cancer. TMB reflects the relative number of gene mutations within a specific tumor, while MSI results from replication errors that cause changes in the length of microsatellite sequences. These biomarkers have emerged as important predictors of immunotherapy efficacy and are actively being investigated in cancer research. By leveraging TMB data from Thorsson et al. (2018) and MSI data from Bonneville et al. (2017), we investigated the association between TMB/MSI and TMEM158 expression in pan-cancer (Thorsson et al. 2018; Bonneville et al. 2017). Our findings demonstrate a significant correlation between TMEM158 expression and both TMB and MSI, highlighting its potential as a valuable biomarker for guiding immunotherapy and informing cancer treatment decisions.
Biological pathway analysis of TMEM158 in pan-cancer
We conducted a gene set enrichment analysis (GSEA) to investigate the biological pathways associated with TMEM158 in pan-cancer. GSEA is a widely used method for identifying enriched pathways and biological processes based on gene expression data. We selected functional gene sets from the MSigDB database and ranked them based on their correlation with TMEM158 expression levels. Using the GTBA database, we investigated the relationship between TMEM158 and the HALLMARK pathway and visualized the results using heatmaps and bar graphs (http://guotosky.vip:13838/GTBA/). Our analysis provides novel insights into the potential biological mechanisms underlying TMEM158's role in cancer and identifies potential targets for future research.
Immune correlation analysis of TMEM158 in pan-cancer
We conducted an immune correlation analysis to investigate the relationship between TMEM158 expression and immune cell infiltration in pan-cancer. We used the TIMER database, which provides RNA-Seq expression data for immune cells in cancer tissue (http://timer.cistrome.org/) (Li et al. 2020, 2017, 2016). Additionally, we employed four state-of-the-art algorithms from the "immunedeconv" R package, including TIMER, xCell, MCP-counter, and EPIC, to perform a comprehensive analysis of immune cell infiltration. We extracted expression data for eight immune checkpoint genes (SIGLEC15, IDO1, CD274, HAVCR2, PDCD1, CTLA4, LAG3, and PDCD1LG2) and explored their correlation with TMEM158 in pan-cancer. Moreover, we examined the correlation between TMEM158 and cancer-associated fibroblast infiltration, given the important role of fibroblasts in the tumor microenvironment. Our analysis provides novel insights into the potential role of TMEM158 in modulating the immune response in cancer and highlights its possible clinical significance as a therapeutic target.
Statistical analysis
For statistical analysis, we utilized R software version 4.0.3. Furthermore, the software integrated within the online platform was employed to conduct statistical analysis on the online data. Spearman's rank correlation test was used to evaluate the correlation between two sets of data, while the rank sum test was utilized to identify differences between two groups of data. The statistical significance threshold was set at a P value of less than 0.05.
Results
TMEM158 mRNA expression in pan-cancer pathological and normal tissues
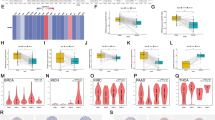
To investigate disparities in TMEM158 mRNA expression across various human cancer types, we utilized RNA-seq data from 33 different cancer types obtained from the TCGA database. Violin plots were used to present the findings. Our results demonstrate that TMEM158 expression is significantly varied between cancerous tissues and adjacent normal tissues in multiple cancer types including BRCA, CESC, CHOL, COAD, ESCA, GBM, HNSC, KICH, KIRC, KIRP, LUAD, LUSC, PCPG, PRAD, READ, STAD, THCA, and UCEC (Fig. 1A-D). To supplement the TCGA data, we included data from the GTEx database for normal tissue samples, which were lacking in the TCGA database. The results from this analysis reveal that TMEM158 expression is significantly different between cancers and adjacent normal tissues in additional cancer types, including DLBC, LGG, LIHC, OV, PAAD, SKCM, TGCT, and UCS (Fig. 1E-H).
TMEM158 mRNA expression in both pathological and normal tissues across a wide range of cancer types. (A-D) According to the TCGA database, the violin plot shows the expression of TMEM158 in 33 types of cancer tissues and normal tissues. (E–H) Utilizing data from the TCGA and GTEx databases, the violin plot illustrates the expression of TMEM158 in cancer tissues and normal tissues. Red indicates tumor tissue and blue indicates normal tissue. Statistical significance was assessed using P values, with *P < 0.05, **P < 0.01, and ***P < 0.001 denoting levels of significance
The mRNA expression of TMEM158 between different stages in multiple cancer types
Clinicians commonly rely on the stage of cancer as a critical clinicopathological indicator to classify cancer and guide treatment decisions. The stage of cancer is based on factors such as tumor size and the extent of cancer spread, and is typically classified using Roman numerals (0, I, II, III, or IV) that have varying clinical implications depending on the cancer type (Amin et al. 2017; Byrd et al. 2021). In this study, the expression of TMEM158 was analyzed across different stages of multiple cancer types, including BLCA, BRCA, KICH, KIRC, KIRP, LIHC, TGCT, and THCA, revealing significant expression differences (Fig. 2A-H). These findings suggest that TMEM158 may serve as a potential biomarker for cancer staging and prognosis.
The overall survival of TMEM158 in pan-cancer studies
Overall survival (OS) has been widely recognized as a critical reference index in clinical diagnosis and treatment (Puisieux et al. 2006). To explore the potential prognostic significance of the TMEM158 gene in pan-cancer patients, we collected RNA-seq data and corresponding patient survival information from the TCGA database for 33 cancer types. Using univariate Cox regression analysis and the "forestplot" package in R, we generated a forest plot to assess the association between TMEM158 expression and pan-cancer survival (Fig. 3A). To confirm our previous findings, we also evaluated the OS of the TMEM158 gene using the GEPIA database (Fig. 3B). Our analysis revealed that low TMEM158 gene expression was significantly associated with better OS in ACC, BLCA, CESC, KIRC, KIRP, LGG, LUAD, MESO, and PAAD patients (Fig. 3C-K). In contrast, low TMEM158 gene expression was positively correlated with poorer OS in SKCM patients (Fig. 3L). These results suggest that TMEM158 may serve as a promising prognostic biomarker for certain cancer types.
The overall survival of TMEM158 in pan-cancer studies. (A) Combining TMEM158 gene expression and patient OS, forest plots show the univariate Cox analysis results of TMEM158 in 33 cancer types. (B) Based on the GEPIA database, the heatmap shows the OS analysis results of TMEM158 in 33 cancer types. Red indicates that TMEM158 acts as a risk factor, while blue indicates that it acts as a protective factor. (C-L) The Kaplan–Meier curves illustrate the OS of TMEM158 in ACC, BLCA, CESC, KIRC, KIRP, LGG, LUAD, MESO, PAAD, and SKCM. Blue represents the low-expression group of TMEM158, while red represents the high-expression group
The disease-free survival of TMEM158 in pan-cancer studies
In recent years, disease-free survival (DFS) has become increasingly significant in clinical practice (Robinson et al. 2014; Ajani et al. 2022). To investigate the DFS of the TMEM158 gene in pan-cancer, we analyzed the TCGA database and conducted a univariate Cox regression analysis for TMEM158 in pan-cancer, which was plotted using the "forestplot" R package (Fig. 4A). To confirm our findings, we further assessed the DFS of TMEM158 in pan-cancer patients using the GEPIA database (Fig. 4B). Our results indicated that lower expression levels of TMEM158 were positively associated with better DFS in BRCA, KIRC, KIRP, and PAAD patients (Fig. 4C-F). However, in SKCM, lower expression of the TMEM158 gene was associated with poorer DFS (Fig. 4G). These findings suggest that TMEM158 may exert its biological effects through different pathways in different cancer types.
The disease-free survival of TMEM158 in pan-cancer studies. (A) Combining TMEM158 gene expression and patients' DFS, forest plots show the univariate Cox analysis results of TMEM158 in 33 cancer types. (B) Based on the GEPIA database, the heatmap shows the results of the DFS analysis of TMEM158 in 33 cancer types. Red indicates TMEM158 acts as a risk factor for this cancer type, and blue indicates TMEM158 acts as a protective factor. (C-G) The Kaplan–Meier curves illustrate the DFS of TMEM158 in BRCA, KIRC, KIRP, PAAD and SKCM. Blue represents the low-expression group of TMEM158, while red represents the high-expression group
Correlation of TMEM158 with immune-related biomarkers and biological signaling pathways in pan-cancer
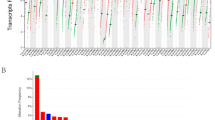
Tumor mutation burden (TMB) and microsatellite instability (MSI) are crucial biomarkers for predicting response to cancer immunotherapy (Schrock et al. 2019; Palmeri et al. 2022; Li et al. 2021; Wang et al. 2021). Using the TCGA database, we calculated TMB and investigated the correlation between TMEM158 and TMB in 33 cancer types. Our analysis revealed that TMEM158 was positively correlated with TMB in four cancer types (THYM, KICH, ACC, and UCS) and negatively correlated with TMB in four cancer types (PRAD, LIHC, CHOL, and ESCA) (Fig. 5A). Subsequently, we examined the correlation between MSI and TMEM158 expression levels. Our analysis demonstrated that MSI was positively correlated with TMEM158 in TGCT, MESO, LUSC, and SARC, but negatively correlated in four cancer types (KICH, ESCA, ACC, and READ) (Fig. 5B). To further investigate the underlying mechanism of TMEM158 expression in pan-cancer, we performed extensive gene set enrichment analysis (GSEA) and visualized the results in a heatmap and a histogram (Fig. 5C-D). Our findings suggest that TMEM158 is associated with abnormal activation of several pathways, including HALLMARK EPITHELIAL MESENCHYMAL TRANSITION, HALLMARK HYPOXIA, HALLMARK IL2 STAT5 SIGNALING, HALLMARK TNFA SIGNALING VIA NFKB, HALLMARK APICAL JUNCTION, and HALLMARK ANGIOGENESIS in pan-cancer. We believe that these results could provide valuable insights for future mechanistic studies.
Correlation of TMEM158 with immune-related biomarkers and biological signaling pathways in pan-cancer. (A) A Spearman correlation analysis of TMEM158 gene expression and TMB was performed. (B) A Spearman correlation analysis of TMEM158 gene expression and MSI. In this chart, the size of the dots represents the correlation coefficient, and the color represents the significance of the p-value. (C-D) The heatmap and bar graph below show the GSEA results of TMEM158 gene in pan-cancer. Purple represents activation, and green represents inhibition. ∗ p < 0.05 and ∗ ∗ p < 0.01
Correlation of TMEM158 with immune cell infiltration and immune checkpoint genes in pan-cancer
Recent advances in cancer research have led to an increased focus on the tumor microenvironment, which plays a critical role in cancer progression and treatment. The tumor microenvironment is highly heterogeneous, particularly in relation to the characteristics of tumor-infiltrating immune cells (Hinshaw and Shevde 2019; Anderson and Simon 2020; Pitt et al. 2016). Therefore, gaining a comprehensive understanding of the tumor microenvironment and its interactions with immune cells is crucial for developing effective cancer therapies. To investigate the correlation between TMEM158 and tumor-infiltrating immune cells, we employed four state-of-the-art algorithms, namely TIMER, xCell, MCP-counter, and EPIC (Fig. 6A-D). Furthermore, most immune checkpoint genes were correlated with TMEM158 expression, especially CTLA4, and LAG3, which are important targets of immune checkpoint inhibitors (Fig. 6E). By elucidating the complex interactions between TMEM158, immune cells, and immune checkpoint genes, these findings contribute to a deeper understanding of the tumor microenvironment and its potential as a therapeutic target in pan-cancer.
Correlation of TMEM158 with immune cell infiltration and immune checkpoint genes in pan-cancer. (A-D) The heatmap showing the correlation between TMEM158 expression and immune cell infiltration in pan-cancer was created using four different algorithms, TIMER, xCell, MCP-counter, and EPIC. (E) The heatmap shows the correlation results between TMEM158 expression in pan-cancer and immune checkpoints, including SIGLEC15, IDO1, CD274, HAVCR2, PDCD1, CTLA4, LAG3, and PDCD1LG2
Correlation of TMEM158 with cancer-associated fibroblast infiltration in pan-cancer
The interaction between tumor cells and stromal cells has gained increasing attention from researchers and is considered a major driver of tumor progression and metastasis. Among the tumor stromal cells, Cancer-Associated Fibroblasts (CAF) are the most abundant and comprise the largest proportion, contributing to the formation of connective tissue proliferation (Biffi and Tuveson 2021; Chen et al. 2021; Wu et al. 2021). In this study, we examined the correlation between TMEM158 and CAFs using the TIMER database (Fig. 7A) and found a significant positive association between TMEM158 and CAFs in over 20 cancer types, including BLCA, BRCA − LumB, DLBC, ESCA, KIRC, OV, PAAD, TGCT, and THCA (Fig. 7B-J). Thus, we believe that our study provides valuable information and credible data for future scientific research.
Correlation of TMEM158 with cancer-associated fibroblast infiltration in pan-cancer. (A) The heatmap displays the correlation between TMEM158 expression and cancer-associated fibroblasts across 33 cancer types. Red denotes a positive correlation, while blue represents a negative correlation. (B-J) The scatter plots illustrate the correlation between TMEM158 expression and cancer-associated fibroblasts in specific cancer types, including BLCA, BRCA − LumB, DLBC, ESCA, KIRC, OV, PAAD, TGCT, and THCA
Discussion
Cancer is an umbrella term for diseases characterized by abnormal cell proliferation, which poses a significant threat to human health worldwide. It is currently the second leading cause of global mortality, with an estimated 10 million deaths in 2020 (Sung et al. 2021). Despite the progress made in basic cancer research and clinical treatments over the past few decades, most patients with metastatic advanced cancer still lack a definitive cure (Zhou and Li 2022; Pich et al. 2022). In recent years, pan-cancer analysis has emerged as a vital approach in cancer research (Zaorsky et al. 2022; Xu et al. 2022). Comparing genomic and cellular changes between various cancer types is a key aspect of this strategy (Combes et al. 2022). Pan-cancer expression analysis is used to explore the association between gene expression and clinical prognosis, and to elucidate the molecular mechanisms underlying tumorigenesis (Weinstein et al. 2013; Omberg et al. 2013). Furthermore, identifying crucial genes that are shared across different cancer types can enhance cancer diagnosis and treatment.
TMEM158 is a member of the transmembrane protein family and has been shown to be involved in various biological functions of tumor cells. However, its role in pan-cancer has not been fully investigated and is therefore a subject worthy of further exploration (Tong et al. 2022; Li et al. 2022). In this study, we analyzed the expression levels of TMEM158 in 33 types of cancer using the TCGA and GTEx databases. Our results demonstrated that TMEM158 was significantly upregulated in more than 20 types of cancer tissues compared to adjacent normal tissues. The dysregulation of gene expression is a hallmark of tumorigenesis, and the identification of differentially expressed genes is crucial for understanding the molecular mechanisms of cancer development. To determine the clinical relevance of TMEM158 expression in pan-cancer, we performed overall and disease-free survival analyses using the TCGA database. We found that high expression of TMEM158 was associated with poor prognosis in patients with nine types of cancer, including ACC, BLCA, CESC, KIRC, KIRP, LGG, LUAD, MESO, and PAAD. Conversely, high expression of TMEM158 in SKCM was negatively correlated with poor patient prognosis. Moreover, based on disease-free survival analyses, high TMEM158 expression was correlated with protective effects in SKCM and risk factors in BRCA, KIRC, KIRP, and PAAD. Taken together, our results suggest that TMEM158 may play distinct biological roles in different types of tumors and may be a potential prognostic biomarker in pan-cancer. Further studies are warranted to elucidate the underlying mechanisms of TMEM158 dysregulation in tumorigenesis and to explore its potential as a therapeutic target in cancer treatment.
TMB, or tumor mutation burden, is defined as the number of somatic mutations in the tumor genome after excluding germline mutations (Chan et al. 2019; Jardim et al. 2021; Sha et al. 2020). It provides a measure of the unique mutations present in tumor cells and is correlated with the production of neoantigens. The higher the TMB value, the greater the likelihood that tumor cells will be recognized by immune cells, leading to a more favorable response to immunotherapy. Thus, this study investigated the relationship between TMEM158 and TMB in pan-cancer. The findings showed a significant positive correlation between TMEM158 and TMB in THYM, KICH, ACC, and UCS. Microsatellite instability (MSI) is another phenomenon that has drawn the attention of oncology researchers (Hause et al. 2016; Chang et al. 2018; Yamamoto et al. 2020). Microsatellites are short DNA sequences that are repeated in tandem throughout the genome. In some cases, microsatellites exhibit instability, resulting in numerous small genetic mutations across the tumor's genome. This phenomenon is referred to as MSI, and tumors can progress through this pathway due to defective mismatch repair (dMMR). In this study, we also investigated the correlation between TMEM158 and MSI. The results demonstrated a significant positive correlation between TMEM158 and MSI in TGCT, MESO, LUSC, and SARC.
Cancer-associated fibroblasts (CAFs) have emerged as critical components of the tumor microenvironment (TME), interacting extensively with cancer cells and influencing other TME constituents (Park et al. 2020). While normal fibroblasts contribute to tissue homeostasis (LeBleu and Neilson 2020), CAFs promote tumor progression by facilitating tumor proliferation, invasion, metastasis, drug resistance, and immunosuppression (Kalluri 2016; D'Arcangelo et al. 2020; Elyada et al. 2019; Hu et al. 2019; Uchihara et al. 2020; Özdemir et al. 2014). Therefore, gaining a deep understanding of the complex nature of CAFs is crucial for prognostic and therapeutic evaluations of cancer patients. In this study, we investigated the correlation between TMEM158 and CAFs in pan-cancer patients. Our results demonstrated a strong association between TMEM158 and CAFs in over twenty types of cancer, suggesting that TMEM158 may be a potential target for CAF depletion or modification of their functions. This approach could have a significant impact on enhancing the therapeutic efficacy of cancer patients and could potentially serve as a novel target for anti-cancer therapy (Hanahan and Weinberg 2011). Furthermore, to explore the underlying molecular mechanisms of TMEM158 in pan-cancer progression, we performed Gene Set Enrichment Analysis (GSEA) using normalized RNA-Seq data from TCGA. Our findings indicated that high TMEM158 gene expression was associated with biological processes such as HALLMARK EPITHELIAL-MESENCHYMAL TRANSITION, HALLMARK HYPOXIA, HALLMARK TNFA SIGNALING VIA NFKB, HALLMARK ANGIOGENESIS, HALLMARK GLYCOLYSIS, and HALLMARK TGF BETA SIGNALING. This discovery further advances our understanding of the role of TMEM158 in pan-cancer tumorigenesis and progression.
In our study, despite obtaining extensive data to demonstrate the prognostic value and immune relevance of TMEM158 in pan-cancer, there are still limitations that need to be addressed. First, all the data analyzed in this study were obtained from public databases, and further validation with large-scale clinical data is required to evaluate the reliability of the constructed survival curves. Second, although TMEM158 was identified as a key gene in this study, additional validation through in vivo and in vitro experiments is necessary to confirm its biological function and mechanism in pan-cancer. Future studies aim to address these limitations and provide a deeper understanding of the role of TMEM158 in pan-cancer progression.
Conclusion
To summarize, this study investigated the expression of TMEM158 in pan-cancer and its prognostic value. The findings indicate that TMEM158 exhibits abnormal expression in pan-cancer and is associated with clinicopathological features and prognosis, particularly in ACC, BLCA, CESC, KIRC, KIRP, LGG, LUAD, MESO, and PAAD. Furthermore, dysregulation of TMEM158 may be related to TMB, MSI, and CAFs in different cancer types, indicating that targeting TMEM158 could be a potential strategy for improving immunotherapy efficacy. However, this study has certain limitations, such as the need for further validation in larger clinical datasets and in vivo and in vitro experiments to explore the biological function and mechanism of TMEM158 in pan-cancer.
Data availability
The data that underpins the findings of this study are obtainable from the corresponding author upon request.
Change history
21 May 2024
This article has been retracted. Please see the Retraction Notice for more detail: https://doi.org/10.1007/s10142-024-01377-0
Abbreviations
- TCGA:
-
The Cancer Genome Atlas
- GTEx:
-
Genotype-Tissue Expression
- GSCA:
-
Gene Set Cancer Analysis
- GSEA:
-
Gene Set Enrichment Analysis
- BRCA:
-
Breast invasive carcinoma
- CESC:
-
Cervical squamous cell carcinoma and endocervical adenocarcinoma
- CHOL:
-
Cholangiocarcinoma
- COAD:
-
Colon adenocarcinoma
- ESCA:
-
Esophageal carcinoma
- GBM:
-
Glioblastoma multiforme
- HNSC:
-
Head and Neck squamous cell carcinoma
- KICH:
-
Kidney Chromophobe
- KIRC:
-
Kidney renal clear cell carcinoma
- KIRP:
-
Kidney renal papillary cell carcinoma
- LUAD:
-
Lung adenocarcinoma
- LUSC:
-
Lung squamous cell carcinoma
- PCPG:
-
Pheochromocytoma and Paraganglioma
- PRAD:
-
Prostate adenocarcinoma
- READ:
-
Rectum adenocarcinoma
- STAD:
-
Stomach adenocarcinoma
- THCA:
-
Thyroid carcinoma
- UCEC:
-
Uterine Corpus Endometrial Carcinoma
- DLBC:
-
Lymphoid Neoplasm Diffuse Large B-cell Lymphoma
- LGG:
-
Brain Lower Grade Glioma
- LIHC:
-
Liver hepatocellular carcinoma
- OV:
-
Ovarian serous cystadenocarcinoma
- PAAD:
-
Pancreatic adenocarcinoma
- SKCM:
-
Skin Cutaneous Melanoma
- TGCT:
-
Testicular Germ Cell Tumors
- UCS:
-
Uterine Carcinosarcoma
- ACC:
-
Adrenocortical carcinoma
- BLCA:
-
Bladder Urothelial Carcinoma
- MESO:
-
Mesothelioma
- TMB:
-
Tumor mutational burden
- MSI:
-
Microsatellite instability
- CAF:
-
Cancer-associated fibroblast
References
Aguet F, Anand S, Ardlie KG, Gabriel S, Getz GA, Graubert A, Hadley K, Handsaker RE, Huang KH, Kashin S, Li X, MacArthur DG, Meier SR, Nedzel JL, Nguyen DT, Segrè AV, Todres E, Balliu B, Barbeira AN, Battle A, Bonazzola R, Brown A, Brown CD, Castel SE, Conrad DF, Cotter DJ, Cox N, Das S, de Goede OM, Dermitzakis ET, Einson J, Engelhardt BE, Eskin E, Eulalio TY, Ferraro NM, Flynn ED, Fresard L, Gamazon ER, Garrido-Martín D, Gay NR, Gloudemans MJ, Guigó R, Hame AR, He Y, Hoffman PJ, Hormozdiari F, Hou L, Im HK, Jo B, Kasela S, Kellis M, Kim-Hellmuth S, Kwong A, Lappalainen T, Li X, Liang Y, Mangul S, Mohammadi P, Montgomery SB, Muñoz-Aguirre M, Nachun DC, Nobel AB, Oliva M, Park Y, Park Y, Parsana P, Rao AS, Reverter F, Rouhana JM, Sabatti C, Saha A, Stephens M, Stranger BE, Strober BJ, Teran NA, Viñuela A, Wang G, Wen X, Wright F, Wucher V, Zou Y, Ferreira PG, Li G, Melé M, Yeger-Lotem E, Barcus ME, Bradbury D, Krubit T, McLean JA, Qi L, Robinson K, Roche NV, Smith AM, Sobin L, Tabor DE, Undale A, Bridge J, Brigham LE, Foster BA, Gillard BM, Hasz R, Hunter M, Johns C, Johnson M, Karasik E, Kopen G, Leinweber WF, McDonald A, Moser MT, Myer K, Ramsey KD, Roe B, Shad S, Thomas JA, Walters G, Washington M, Wheeler J, Jewell SD, Rohrer DC, Valley DR, Davis DA, Mash DC, Branton PA, Barker LK, Gardiner HM, Mosavel M, Siminoff LA, Flicek P, Haeussler M, Juettemann T, Kent WJ, Lee CM, Powell CC, Rosenbloom KR, Ruffier M, Sheppard D, Taylor K, Trevanion SJ, Zerbino DR, Abell NS, Akey J, Chen L, Demanelis K, Doherty JA, Feinberg AP, Hansen KD, Hickey PF, Jasmine F, Jiang L, Kaul R, Kibriya MG, Li JB, Li Q, Lin S, Linder SE, Pierce BL, Rizzardi LF, Skol AD, Smith KS, Snyder M, Stamatoyannopoulos J, Tang H, Wang M, Carithers LJ, Guan P, Koester SE, Little AR, Moore HM, Nierras CR, Rao AK, Vaught JB, Volpi S (2020) The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369(6509):1318–1330. https://doi.org/10.1126/science.aaz1776
Ajani JA, Leung L, Singh P, Kurt M, Kim I, Pourrahmat MM, Kanters S (2022) Disease-free survival as a surrogate endpoint for overall survival in adults with resectable esophageal or gastroesophageal junction cancer: a correlation meta-analysis. Eur J Cancer 170:119–130. https://doi.org/10.1016/j.ejca.2022.04.027
Amin MB, Greene FL, Edge SB, Compton CC, Gershenwald JE, Brookland RK, Meyer L, Gress DM, Byrd DR, Winchester DP (2017) The eighth edition AJCC cancer staging manual: continuing to build a bridge from a population-based to a more "personalized" approach to cancer staging. CA Cancer J Clin 67(2):93–99. https://doi.org/10.3322/caac.21388
Anderson NM, Simon MC (2020) The tumor microenvironment. Curr Biol 30(16):R921-r925. https://doi.org/10.1016/j.cub.2020.06.081
Banerjee S, Lo WC, Majumder P, Roy D, Ghorai M, Shaikh NK, Kant N, Shekhawat MS, Gadekar VS, Ghosh S, Bursal E, Alrumaihi F, Dubey NK, Kumar S, Iqbal D, Alturaiki W, Upadhye VJ, Jha NK, Dey A, Gundamaraju R (2022) Multiple roles for basement membrane proteins in cancer progression and EMT. Eur J Cell Biol 101(2):151220. https://doi.org/10.1016/j.ejcb.2022.151220
Barradas M, Gonos ES, Zebedee Z, Kolettas E, Petropoulou C, Delgado MD, León J, Hara E, Serrano M (2002) Identification of a candidate tumor-suppressor gene specifically activated during Ras-induced senescence. Exp Cell Res 273(2):127–137. https://doi.org/10.1006/excr.2001.5434
Biffi G, Tuveson DA (2021) Diversity and biology of cancer-associated fibroblasts. Physiol Rev 101(1):147–176. https://doi.org/10.1152/physrev.00048.2019
Bonneville R, Krook MA, Kautto EA, Miya J, Wing MR, Chen HZ, Reeser JW, Yu L, Roychowdhury S (2017) Landscape of microsatellite instability across 39 cancer types. JCO Precis Oncol 2017:PO.17.00073. https://doi.org/10.1200/po.17.00073
Byrd DR, Brierley JD, Baker TP, Sullivan DC, Gress DM (2021) Current and future cancer staging after neoadjuvant treatment for solid tumors. CA Cancer J Clin 71(2):140–148. https://doi.org/10.3322/caac.21640
Carpenter EP, Beis K, Cameron AD, Iwata S (2008) Overcoming the challenges of membrane protein crystallography. Curr Opin Struct Biol 18(5):581–586. https://doi.org/10.1016/j.sbi.2008.07.001
Chan TA, Yarchoan M, Jaffee E, Swanton C, Quezada SA, Stenzinger A, Peters S (2019) Development of tumor mutation burden as an immunotherapy biomarker: utility for the oncology clinic. Ann Oncol 30(1):44–56. https://doi.org/10.1093/annonc/mdy495
Chang L, Chang M, Chang HM, Chang F (2018) Microsatellite instability: a predictive biomarker for cancer immunotherapy. Appl Immunohistochem Mol Morphol 26(2):e15–e21. https://doi.org/10.1097/pai.0000000000000575
Chen Y, McAndrews KM, Kalluri R (2021) Clinical and therapeutic relevance of cancer-associated fibroblasts. Nat Rev Clin Oncol 18(12):792–804. https://doi.org/10.1038/s41571-021-00546-5
Cheng Z, Guo J, Chen L, Luo N, Yang W, Qu X (2015) Overexpression of TMEM158 contributes to ovarian carcinogenesis. J Exp Clin Cancer Res 34(1):75. https://doi.org/10.1186/s13046-015-0193-y
Chitwood PJ, Hegde RS (2019) The role of EMC during membrane protein biogenesis. Trends Cell Biol 29(5):371–384. https://doi.org/10.1016/j.tcb.2019.01.007
Combes AJ, Samad B, Tsui J, Chew NW, Yan P, Reeder GC, Kushnoor D, Shen A, Davidson B, Barczak AJ, Adkisson M, Edwards A, Naser M, Barry KC, Courau T, Hammoudi T, Argüello RJ, Rao AA, Olshen AB, Cai C, Zhan J, Davis KC, Kelley RK, Chapman JS, Atreya CE, Patel A, Daud AI, Ha P, Diaz AA, Kratz JR, Collisson EA, Fragiadakis GK, Erle DJ, Boissonnas A, Asthana S, Chan V, Krummel MF (2022) Discovering dominant tumor immune archetypes in a pan-cancer census. Cell 185(1):184-203.e119. https://doi.org/10.1016/j.cell.2021.12.004
D’Arcangelo E, Wu NC, Cadavid JL, McGuigan AP (2020) The life cycle of cancer-associated fibroblasts within the tumour stroma and its importance in disease outcome. Br J Cancer 122(7):931–942. https://doi.org/10.1038/s41416-019-0705-1
Du Y, Zeng X, Yu W, Xie W (2022) A transmembrane protein family gene signature for overall survival prediction in osteosarcoma. Front Genet 13:937300. https://doi.org/10.3389/fgene.2022.937300
Elyada E, Bolisetty M, Laise P, Flynn WF, Courtois ET, Burkhart RA, Teinor JA, Belleau P, Biffi G, Lucito MS, Sivajothi S, Armstrong TD, Engle DD, Yu KH, Hao Y, Wolfgang CL, Park Y, Preall J, Jaffee EM, Califano A, Robson P, Tuveson DA (2019) Cross-species single-cell analysis of pancreatic ductal adenocarcinoma reveals antigen-presenting cancer-associated fibroblasts. Cancer Discov 9(8):1102–1123. https://doi.org/10.1158/2159-8290.Cd-19-0094
Eraslan G, Drokhlyansky E, Anand S, Fiskin E, Subramanian A, Slyper M, Wang J, Van Wittenberghe N, Rouhana JM, Waldman J, Ashenberg O, Lek M, Dionne D, Win TS, Cuoco MS, Kuksenko O, Tsankov AM, Branton PA, Marshall JL, Greka A, Getz G, Segrè AV, Aguet F, Rozenblatt-Rosen O, Ardlie KG, Regev A (2022) Single-nucleus cross-tissue molecular reference maps toward understanding disease gene function. Science 376 (6594):eabl4290. https://doi.org/10.1126/science.abl4290
Fan JJ, Huang X (2022) Ion channels in cancer: orchestrators of electrical signaling and cellular crosstalk. Rev Physiol Biochem Pharmacol 183:103–133. https://doi.org/10.1007/112_2020_48
Fu Y, Yao N, Ding D, Zhang X, Liu H, Ma L, Shi W, Zhu C, Tang L (2020) TMEM158 promotes pancreatic cancer aggressiveness by activation of TGFβ1 and PI3K/AKT signaling pathway. J Cell Physiol 235(3):2761–2775. https://doi.org/10.1002/jcp.29181
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144(5):646–674. https://doi.org/10.1016/j.cell.2011.02.013
Hause RJ, Pritchard CC, Shendure J, Salipante SJ (2016) Classification and characterization of microsatellite instability across 18 cancer types. Nat Med 22(11):1342–1350. https://doi.org/10.1038/nm.4191
Hinshaw DC, Shevde LA (2019) The tumor microenvironment innately modulates cancer progression. Cancer Res 79(18):4557–4566. https://doi.org/10.1158/0008-5472.Can-18-3962
Hu JL, Wang W, Lan XL, Zeng ZC, Liang YS, Yan YR, Song FY, Wang FF, Zhu XH, Liao WJ, Liao WT, Ding YQ, Liang L (2019) CAFs secreted exosomes promote metastasis and chemotherapy resistance by enhancing cell stemness and epithelial-mesenchymal transition in colorectal cancer. Mol Cancer 18(1):91. https://doi.org/10.1186/s12943-019-1019-x
Iglesias D, Fernández-Peralta AM, Nejda N, Daimiel L, Azcoita MM, Oliart S, González-Aguilera JJ (2006) RIS1, a gene with trinucleotide repeats, is a target in the mutator pathway of colorectal carcinogenesis. Cancer Genet Cytogenet 167(2):138–144. https://doi.org/10.1016/j.cancergencyto.2005.12.002
Jardim DL, Goodman A, de Melo GD, Kurzrock R (2021) The challenges of tumor mutational burden as an immunotherapy biomarker. Cancer Cell 39(2):154–173. https://doi.org/10.1016/j.ccell.2020.10.001
Kalluri R (2016) The biology and function of fibroblasts in cancer. Nat Rev Cancer 16(9):582–598. https://doi.org/10.1038/nrc.2016.73
Kanai Y (2022) Amino acid transporter LAT1 (SLC7A5) as a molecular target for cancer diagnosis and therapeutics. Pharmacol Ther 230:107964. https://doi.org/10.1016/j.pharmthera.2021.107964
Kim-Hellmuth S, Aguet F, Oliva M, Muñoz-Aguirre M, Kasela S, Wucher V, Castel SE, Hamel AR, Viñuela A, Roberts AL, Mangul S, Wen X, Wang G, Barbeira AN, Garrido-Martín D, Nadel BB, Zou Y, Bonazzola R, Quan J, Brown A, Martinez-Perez A, Soria JM; GTEx Consortium; Getz G , Dermitzakis ET, Small KS, Stephens M, Xi HS, Im HK, Guigó R, Segrè AV, Stranger BE, Ardlie KG, Lappalainen T (2020) Cell type-specific genetic regulation of gene expression across human tissues. Science 369 (6509 ):eaaz8528. https://doi.org/10.1126/science.aaz8528
LeBleu VS, Neilson EG (2020) Origin and functional heterogeneity of fibroblasts. Faseb j 34(3):3519–3536. https://doi.org/10.1096/fj.201903188R
Li B, Severson E, Pignon JC, Zhao H, Li T, Novak J, Jiang P, Shen H, Aster JC, Rodig S, Signoretti S, Liu JS, Liu XS (2016) Comprehensive analyses of tumor immunity: implications for cancer immunotherapy. Genome Biol 17(1):174. https://doi.org/10.1186/s13059-016-1028-7
Li J, Wang X, Chen L, Zhang J, Zhang Y, Ren X, Sun J, Fan X, Fan J, Li T, Tong L, Yi L, Chen L, Liu J, Shang G, Ren X, Zhang H, Yu S, Ming H, Huang Q, Dong J, Zhang C, Yang X (2022) TMEM158 promotes the proliferation and migration of glioma cells via STAT3 signaling in glioblastomas. Cancer Gene Ther 29(8–9):1117–1129. https://doi.org/10.1038/s41417-021-00414-5
Li T, Fan J, Wang B, Traugh N, Chen Q, Liu JS, Li B, Liu XS (2017) TIMER: a web server for comprehensive analysis of tumor-infiltrating immune cells. Cancer Res 77(21):e108–e110. https://doi.org/10.1158/0008-5472.Can-17-0307
Li T, Fu J, Zeng Z, Cohen D, Li J, Chen Q, Li B, Liu XS (2020) TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res 48 (W1):W509-w514. https://doi.org/10.1093/nar/gkaa407
Li Y, Ma Y, Wu Z, Zeng F, Song B, Zhang Y, Li J, Lui S, Wu M (2021) Tumor mutational burden predicting the efficacy of immune checkpoint inhibitors in colorectal cancer: a systematic review and meta-analysis. Front Immunol 12:751407. https://doi.org/10.3389/fimmu.2021.751407
Marx S, Dal Maso T, Chen JW, Bury M, Wouters J, Michiels C, Le Calvé B (2020) Transmembrane (TMEM) protein family members: poorly characterized even if essential for the metastatic process. Semin Cancer Biol 60:96–106. https://doi.org/10.1016/j.semcancer.2019.08.018
Mohammed Ael S, Eguchi H, Wada S, Koyama N, Shimizu M, Otani K, Ohtaki M, Tanimoto K, Hiyama K, Gaber MS, Nishiyama M (2012) TMEM158 and FBLP1 as novel marker genes of cisplatin sensitivity in non-small cell lung cancer cells. Exp Lung Res 38(9–10):463–474. https://doi.org/10.3109/01902148.2012.731625
Omberg L, Ellrott K, Yuan Y, Kandoth C, Wong C, Kellen MR, Friend SH, Stuart J, Liang H, Margolin AA (2013) Enabling transparent and collaborative computational analysis of 12 tumor types within the cancer genome atlas. Nat Genet 45(10):1121–1126. https://doi.org/10.1038/ng.2761
Özdemir BC, Pentcheva-Hoang T, Carstens JL, Zheng X, Wu CC, Simpson TR, Laklai H, Sugimoto H, Kahlert C, Novitskiy SV, De Jesus-Acosta A, Sharma P, Heidari P, Mahmood U, Chin L, Moses HL, Weaver VM, Maitra A, Allison JP, LeBleu VS, Kalluri R (2014) Depletion of carcinoma-associated fibroblasts and fibrosis induces immunosuppression and accelerates pancreas cancer with reduced survival. Cancer Cell 25(6):719–734. https://doi.org/10.1016/j.ccr.2014.04.005
Palmeri M, Mehnert J, Silk AW, Jabbour SK, Ganesan S, Popli P, Riedlinger G, Stephenson R, de Meritens AB, Leiser A, Mayer T, Chan N, Spencer K, Girda E, Malhotra J, Chan T, Subbiah V, Groisberg R (2022) Real-world application of tumor mutational burden-high (TMB-high) and microsatellite instability (MSI) confirms their utility as immunotherapy biomarkers. ESMO Open 7(1):100336. https://doi.org/10.1016/j.esmoop.2021.100336
Park D, Sahai E, Rullan A (2020) SnapShot: cancer-associated fibroblasts. Cell 181(2):486-486.e481. https://doi.org/10.1016/j.cell.2020.03.013
Pich O, Bailey C, Watkins TBK, Zaccaria S, Jamal-Hanjani M, Swanton C (2022) The translational challenges of precision oncology. Cancer Cell 40(5):458–478. https://doi.org/10.1016/j.ccell.2022.04.002
Pitt JM, Marabelle A, Eggermont A, Soria JC, Kroemer G, Zitvogel L (2016) Targeting the tumor microenvironment: removing obstruction to anticancer immune responses and immunotherapy. Ann Oncol 27(8):1482–1492. https://doi.org/10.1093/annonc/mdw168
Puisieux A, Valsesia-Wittmann S, Ansieau S (2006) A twist for survival and cancer progression. Br J Cancer 94(1):13–17. https://doi.org/10.1038/sj.bjc.6602876
Robinson AG, Booth CM, Eisenhauer EA (2014) Disease-free survival as an end-point in the treatment of solid tumours–perspectives from clinical trials and clinical practice. Eur J Cancer 50(13):2298–2302. https://doi.org/10.1016/j.ejca.2014.05.016
Schmit K, Michiels C (2018) TMEM proteins in cancer: a review. Front Pharmacol 9:1345. https://doi.org/10.3389/fphar.2018.01345
Schrock AB, Ouyang C, Sandhu J, Sokol E, Jin D, Ross JS, Miller VA, Lim D, Amanam I, Chao J, Catenacci D, Cho M, Braiteh F, Klempner SJ, Ali SM, Fakih M (2019) Tumor mutational burden is predictive of response to immune checkpoint inhibitors in MSI-high metastatic colorectal cancer. Ann Oncol 30(7):1096–1103. https://doi.org/10.1093/annonc/mdz134
Sha D, Jin Z, Budczies J, Kluck K, Stenzinger A, Sinicrope FA (2020) Tumor mutational burden as a predictive biomarker in solid tumors. Cancer Discov 10(12):1808–1825. https://doi.org/10.1158/2159-8290.Cd-20-0522
Silva J, Silva JM, Barradas M, García JM, Domínguez G, García V, Peña C, Gallego I, Espinosa R, Serrano M, Bonilla F (2006) Analysis of the candidate tumor suppressor Ris-1 in primary human breast carcinomas. Mutat Res 594(1–2):78–85. https://doi.org/10.1016/j.mrfmmm.2005.07.017
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F (2021) Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71(3):209–249. https://doi.org/10.3322/caac.21660
Tang Z, Kang B, Li C, Chen T, Zhang Z (2019) GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res 47(W1):W556-w560. https://doi.org/10.1093/nar/gkz430
Thorsson V, Gibbs DL, Brown SD, Wolf D, Bortone DS, Ou Yang TH, Porta-Pardo E, Gao GF, Plaisier CL, Eddy JA, Ziv E, Culhane AC, Paull EO, Sivakumar IKA, Gentles AJ, Malhotra R, Farshidfar F, Colaprico A, Parker JS, Mose LE, Vo NS, Liu J, Liu Y, Rader J, Dhankani V, Reynolds SM, Bowlby R, Califano A, Cherniack AD, Anastassiou D, Bedognetti D, Mokrab Y, Newman AM, Rao A, Chen K, Krasnitz A, Hu H, Malta TM, Noushmehr H, Pedamallu CS, Bullman S, Ojesina AI, Lamb A, Zhou W, Shen H, Choueiri TK, Weinstein JN, Guinney J, Saltz J, Holt RA, Rabkin CS, Lazar AJ, Serody JS, Demicco EG, Disis ML, Vincent BG, Shmulevich I (2018) The immune landscape of cancer. Immunity 48(4):812-830.e814. https://doi.org/10.1016/j.immuni.2018.03.023
Tong J, Li H, Hu Y, Zhao Z, Li M (2022) TMEM158 regulates the canonical and non-canonical pathways of TGF-β to mediate EMT in triple-negative breast cancer. J Cancer 13(8):2694–2704. https://doi.org/10.7150/jca.65822
Uchihara T, Miyake K, Yonemura A, Komohara Y, Itoyama R, Koiwa M, Yasuda T, Arima K, Harada K, Eto K, Hayashi H, Iwatsuki M, Iwagami S, Baba Y, Yoshida N, Yashiro M, Masuda M, Ajani JA, Tan P, Baba H, Ishimoto T (2020) Extracellular vesicles from cancer-associated fibroblasts containing annexin A6 Induces FAK-YAP activation by stabilizing β1 Integrin. Enhanc Drug Resist Cancer Res 80(16):3222–3235. https://doi.org/10.1158/0008-5472.Can-19-3803
Wang Y, Tong Z, Zhang W, Zhang W, Buzdin A, Mu X, Yan Q, Zhao X, Chang HH, Duhon M, Zhou X, Zhao G, Chen H, Li X (2021) FDA-approved and emerging next generation predictive biomarkers for immune checkpoint inhibitors in cancer patients. Front Oncol 11:683419. https://doi.org/10.3389/fonc.2021.683419
Weinstein JN, Collisson EA, Mills GB, Shaw KR, Ozenberger BA, Ellrott K, Shmulevich I, Sander C, Stuart JM (2013) The cancer genome atlas pan-cancer analysis project. Nat Genet 45(10):1113–1120. https://doi.org/10.1038/ng.2764
Wu F, Yang J, Liu J, Wang Y, Mu J, Zeng Q, Deng S, Zhou H (2021) Signaling pathways in cancer-associated fibroblasts and targeted therapy for cancer. Signal Transduct Target Ther 6(1):218. https://doi.org/10.1038/s41392-021-00641-0
Xu Y, Shen M, Peng Y, Liu L, Tang L, Yang T, Pu D, Tan W, Zhang W, Liu S (2022) Cell Division Cycle-Associated Protein 3 (CDCA3) Is a potential biomarker for clinical prognosis and immunotherapy in pan-cancer. BioMed Res Int 2022:4632453. https://doi.org/10.1155/2022/4632453
Yamamoto H, Watanabe Y, Maehata T, Imai K, Itoh F (2020) Microsatellite instability in cancer: a novel landscape for diagnostic and therapeutic approach. Arch Toxicol 94(10):3349–3357. https://doi.org/10.1007/s00204-020-02833-z
Zaorsky NG, Wang X, Garrett SM, Lehrer EJ, Lin C, DeGraff DJ, Spratt DE, Trifiletti DM, Kishan AU, Showalter TN, Park HS, Yang JT, Chinchilli VM, Wang M (2022) Pan-cancer analysis of prognostic metastatic phenotypes. Int J Cancer 150(1):132–141. https://doi.org/10.1002/ijc.33744
Zhou Z, Li M (2022) Targeted therapies for cancer. BMC Med 20(1):90. https://doi.org/10.1186/s12916-022-02287-3
Zirn B, Samans B, Wittmann S, Pietsch T, Leuschner I, Graf N, Gessler M (2006) Target genes of the WNT/beta-catenin pathway in Wilms tumors. Genes Chromosome Cancer 45(6):565–574. https://doi.org/10.1002/gcc.20319
Acknowledgements
We express our gratitude to the Cancer Genome Atlas (TCGA) for providing publicly available data.
Funding
This research was funded by the Shandong Provincial Science and Technology Project (No. 2016GSF201054).
Author information
Authors and Affiliations
Contributions
Guibao Li and Jiayi Li designed the research methods and analyzed the data. Haiguang Hou and Jinhao Sun participated in data collection. Zhaoxi Ding and Jiayi Li drafted the manuscript, and Guibao Li and Yingkun Xu revised it. All authors have approved the final version and agree to take responsibility for all aspects of the work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
Not applicable.
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article has been retracted. Please see the retraction notice for more detail: https://doi.org/10.1007/s10142-024-01377-0"
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, J., Hou, H., Sun, J. et al. RETRACTED ARTICLE: Systematic pan-cancer analysis identifies transmembrane protein 158 as a potential therapeutic, prognostic and immunological biomarker. Funct Integr Genomics 23, 105 (2023). https://doi.org/10.1007/s10142-023-01032-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10142-023-01032-0