Abstract
Carbohydrate-active enzymes (CAZymes) have significant biotechnological potential as agents for degradation or modification of polysaccharides/glycans. As marine macroalgae are known to be rich in various types of polysaccharides, seaweed-associated bacteria are likely to be a good source of these CAZymes. A genomics approach can be used to explore CAZyme abundance and diversity, but it can also provide deep insights into the biology of CAZyme producers and, in particular, into molecular mechanisms that mediate their interaction with their hosts. In this study, a Gram-negative, aerobic, rod-shaped, carrageenolytic, and culturable marine bacterium designated as AOL6 was isolated from a diseased thallus of a carrageenan-producing farmed rhodophyte, Kappaphycus alvarezii (Gigartinales, Rhodophyta). The whole genome of this bacterium was sequenced and characterized. Sequence reads were assembled producing a high-quality genome assembly. The estimated genome size of the bacterium is 4.4 Mb and a G+C content of 52%. Molecular phylogenetic analysis based on a complete sequence of 16S rRNA, rpoB, and a set of 38 single-copy genes suggests that the bacterium is an unknown species and represents a novel genus in the family Cellvibrionaceae that is most closely related to the genera Teredinibacter and Saccharophagus. Genome comparison with T. turnerae T7901 and S. degradans 2-40 reveals several features shared by the three species, including a large number of CAZymes that comprised > 5% of the total number of protein-coding genes. The high proportion of CAZymes found in the AOL6 genome exceeds that of other known carbohydrate degraders, suggesting a significant capacity to degrade a range of polysaccharides including κ-carrageenan; 34% of these CAZymes have signal peptide sequences for secretion. Three putative κ-carrageenase-encoding genes were identified from the genome of the bacterium via in silico analysis, consistent with the results of the zymography assay (with κ-carrageenan as substrate). Genome analysis also indicated that AOL6 relies exclusively on type 2 secretion system (T2SS) for secreting proteins (possibly including glycoside hydrolases). In relation to T2SS, the product of the pilZ gene was predicted to be highly expressed, suggesting specialization for cell adhesion and secretion of virulence factors. The assignment of proteins to clusters of orthologous groups (COGs) revealed a pattern characteristic of r-strategists. Majority of two-component system proteins identified in the AOL6 genome were also predicted to be involved in chemotaxis and surface colonization. These genomic features suggest that AOL6 is an opportunistic pathogen, adapted to colonizing polysaccharide-rich hosts, including carrageenophytes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Marine bacteria, particularly many species in Gammaproteobacteria, Bacteroidetes, and Planctomycetes, are key players in the global ocean carbon cycle (Michel et al. 2006). These marine bacteria produce diverse carbohydrate-active enzymes (CAZymes) such as glycoside hydrolases, carbohydrate esterases, and polysaccharide lyases, many of which are secreted out of the cell. The specific biochemical function of many of these enzymes remains to be elucidated but their characterization holds the potential for understanding complex polysaccharide bioconversion processes (Hehemann et al. 2014). The role of CAZyme-producing bacteria in the marine environment is particularly important given the diversity and quantity of polysaccharides produced by marine macroalgae (especially phaeophytes and rhodophytes) coupled with their large biomass and subsequent ease of harvestability—providing an important carbon source for several marine bacteria (Rhein-Knudsen et al. 2015). CAZyme-producing bacteria can therefore be opportunistic pathogens or saprophytes rather than commensal or mutualistic macroalgal symbionts (Egan et al. 2013).
Agar and carrageenan are among the major forms of polysaccharides in the marine environment, being the main structural material that make up the cell wall and intercellular matrix in many species of red algae (De Ruiter and Rudolph 1997; Hehemann et al. 2014). Various subtypes of these polysaccharides exist, and it is likely that various types of agarases and carrageenases that are specific to these subtypes have already evolved. In the marine environment, bacteria use these enzymes to degrade the polysaccharides that make up the cell wall of their seaweed hosts; hydrolysis of polysaccharides can either provide the bacteria with carbon sources or serve as an important mechanism for the bacteria to infect their hosts. A number of these enzymes have already been isolated and at least partially characterized (Michel et al. 2001; Chauhan and Saxena 2016). However, many carrageenases have not been purified to homogeneity and are not very well characterized (De Ruiter and Rudolph 1997), a situation that remains true today. Proteobacteria and Bacteroidetes are the two bacterial phyla under which most carrageenan-degrading isolates are classified (Chauhan and Saxena 2016).
The growing biotechnological interest in oligosaccharides as potential therapeutic products has also provided impetus to search for CAZyme-producing marine bacteria. Agarases and carrageenases are logical targets considering the amount of the agar and carrageenan that are produced globally via industrial-scale extraction of these polysaccharides from seaweeds (farmed or harvested from the wild) (Rhein-Knudsen et al. 2015). Oligocarrageenans, for example, have been reported to exhibit diverse bioactivities (Yuan et al. 2005; Hu et al. 2006; Yuan et al. 2006a, b; Kalitnik et al. 2012; Wang et al. 2012). Isozymes with slightly different enzymatic specificities may hold the potential for converting carrageenans into various types of oligosaccharides with different bioactivities or applications. For instance, κ-carrageenases exhibit diverse physicochemical properties (Potin et al. 1991; Khambhaty et al. 2007; Zhou et al. 2008; Sun et al. 2010; Liu et al. 2011; Li et al. 2013; Chauhan and Saxena 2016; Liu et al. 2016; Zhu and Ning 2016). The potential structural and functional diversity of the oligosaccharides resulting from enzymatic hydrolysis by these carrageenases is therefore potentially high. Equally important are further applications of CAZymes in cellulosic biomass conversion for bioethanol production and protoplast isolation for production of improved algal strains (Hehemann et al. 2014; Chauhan and Saxena 2016).
Traditionally, bioprospecting for carrageenases involves isolation and purification of enzymes from bacterial culture. In this study, whole genome sequencing of a carrageenase-positive rhodophyte-associated taxonomically novel isolate was used to discover CAZymes, particularly carrageenases. The genomic approach not only allows the discovery of the entire complement of putative CAZymes that a species can potentially produce (whether the genes are expressed or not in culture), but also provides important insights into the biology and ecology of the species, and hence the contexts in which the CAZymes play their roles, e.g., host-bacterial interactions. To our knowledge, this report is among the first to characterize the genome of a carrageenolytic marine bacterium.
Materials and Methods
Provenance of the Bacterium
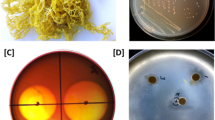
The Gram-negative, aerobic, mesophilic, pigmented, rod-shaped, motile, and carrageenolytic marine bacterium designated as isolate AOL6 was isolated from a thallus of a diseased seaweed Kappaphycus alvarezii. The seaweed was obtained from a seaweed farm in Calatagan, Batangas, Philippines (13° 50′ 0.5′′ N, 120° 37′ 6.8″ E). Carrageenolytic marine bacteria were selectively grown in marine broth (40 g/L) with 1.5% κ-carrageenan. The semi-refined κ-carrageenan powder was obtained from Shemberg in Mandaue City, Cebu, 6014, Philippines. Clearing zones and liquefaction of the substrate κ-carrageenan indicated enzymatic activity. Preliminary molecular identification based on 27F and 1492R universal bacterial primers (Lane 1991) for 16S/SSU rRNA was performed to confirm the identity and purity of the isolate. The morphological characteristics were observed using Hitachi S-510 scanning electron microscope. Meanwhile, Gram-staining, capsule staining, and motility test were likewise performed. For biochemical characterization, API 20E and BIOLOG Gen III were used. The isolate was deposited at the University of the Philippines Los Baños Philippine National Collection of Microorganisms with Accession/Collection No. BIOTECH 10559 and at the University of Santo Tomas Collection of Microbial Strains in Manila, Philippines, with Accession/Collection No. UST CMS 5002.
DNA Extraction, Genome Sequencing and Assembly
Genomic DNA was extracted from an isolate AOL6 culture (mid-exponential phase) using Vivantis GF-1 Bacterial DNA Extraction Kit following manufacturer’s instructions. The quality of the extracted DNA was assessed by agarose gel electrophoresis and UV-Vis spectrophotometry. High-quality DNA was subjected to pyrosequencing using Roche 454 GS Junior Sequencer. An appropriate amount (800 ng) of DNA was used to construct the library following manufacturer’s protocol for fragmentation, purification, blunt-ending, adapter ligation, and size selection. The quality and quantity of the library were assessed by 1% agarose gel electrophoresis stained using Invitrogen’s SYBR Gold DNA stain and by fluorescence spectroscopy (molecules/μL) using BIOTEK microplate reader, respectively. Then, 10−7 dilution (TE buffer) of DNA library was prepared. Clonal amplification of DNA library was performed by emulsion PCR with copy to bead ratio of 1.5 cpb (copy per bead). Clonally amplified DNA beads were recovered, enriched, loaded to the PicoTiter Plate, and subjected to 200 cycles of pyrosequencing. De novo genome assembly including genome size estimation was performed using Roche GS De Novo assembler (Newbler 2.7). Assembly metrics was assessed using PRINSEQ (Schmieder and Edwards 2011). The k-mer depth (k = 11, D’ = 18) was estimated by k-mer counting methods such as Jellyfish (Marçais and Kingsford 2011) and KMC 2 (Deorowicz et al. 2015). Subsequently, the read depth (D) was calculated based on k-mer depth (D = D’l / (l – k + 1) where l is the average read length). Together, the k-mer and read depths were used to calculate the estimated genome size by diving the total number of k-mers by the k-mer depth (G = Nbase / D = Nread (l – k + 1) / D’ where G is the estimated genome size, Nbase is the number of bases, Nread is the number of reads, D’ is the k-mer depth at the peak of the k-mer histogram, and Nk-mer = (l – k + 1) is the number of k-mers in a read) (Sohn and Nam 2018). This Whole Genome Shotgun project has been deposited at DDBJ/ENA/GenBank under the accession NOXC00000000. The version described in this paper is version NOXC01000000. The SSU (16S) rRNA and LSU (23S) rRNA sequences were deposited in GenBank with accession numbers MF928089 and MF928709, respectively. Likewise, the rpoB (DNA-dependent RNA polymerase subunit beta) sequence was deposited in GenBank with accession number MG934211. Unless stated otherwise, data handling and analysis were performed using custom-made Python and R scripts.
Genome Annotation
Genome annotation was carried out using automated and manual workflows. Automated genome annotation was performed using Prokka 1.12 (Seemann 2014). Structural and functional annotations were manually performed following the prediction of coding sequences (CDSs). The list of CDS was predicted from the genome assembly using Glimmer 3.02 (Salzberg et al. 1998; Delcher et al. 1999). The predicted CDS were indexed and retrieved from the assembly. CDSs were translated using EMBOSS TRANSEQ (Rice et al. 2000) at Frame 1 using NCBI Codon Table 11 (Bacterial, Archaeal and Plastid Genetic Code). Translated CDS were annotated by BLASTp (Altschul et al. 1990) search against GenBank NR, trEMBL, and SwissProt protein databases with an e-value threshold of 1E-10. Enzyme Commission (EC) numbers were assigned to CDS using Protein Sequence Analysis Toolkit (PSAT) (Leung et al. 2016). Identification of clusters of orthologous groups (COGs) and gene ontology term enrichment (GO enrichment) were performed using EggNOG4.5 database (Huerta-Cepas et al. 2016). KEGG pathway maps (map03070 and map02040 with Copyright Permission Ref. No. 180121) were reconstructed using BLASTKOALA (Kanehisa et al. 2016). Structural features such as rRNA and tRNA were predicted using RNAmmer (Lagesen et al. 2007) and ARAGORN (Laslett and Canback 2004), respectively. CAZymes were identified and annotated using dbCAN (Yin et al. 2012) and SignalP 4.1 (Petersen et al. 2011). BLASTp-NR result was analyzed using MEGAN (MEtaGenome ANalyzer) (Huson et al. 2007) highlighting notable features such as microbial attributes. Prokaryotic two-component systems (TCS) were identified using P2RP (Barakat et al. 2013) or retrieved from P2CS database (Ortet et al. 2015). Finally, synonymous codon usage order (SCUO) for each CDS was computed using CodonO (Angellotti et al. 2007). Studies have shown that a correlation exists between codon usage biases, and protein and transcript levels in different species (Gupta et al. 2004; Sahoo and Das 2014; Zhou et al. 2016; Das et al. 2017; Uddin 2017). Hence, in this analysis, the SCUO values were used as proxies to estimate gene expression levels. The CDSs with higher SCUO based on statistical analysis were thus assumed to be potentially highly expressed. The terms CDS, protein-coding sequence, ORF (open-reading frame), and protein (translated CDS) were used interchangeably for brevity and clarity, while genes refer to both protein-coding sequences and other features such as transfer and ribosomal RNA genes.
Phylogenetics and Comparative Genomics
A small subunit (SSU/16S) rRNA gene was identified from the assembly using RNAmmer (Lagesen et al. 2007). Homologous sequences (n = 12) were retrieved from the Ribosomal Database Project (RDP) (Cole et al. 2009). Phylogenetic analysis was performed using MEGA (Kumar et al. 2016) and MrBayes (Huelsenbeck and Ronquist 2001) following multiple sequence alignment (MSA), removal of flanking 5′ and 3′ ends by manual trimming, model testing, and tree building. The phylogenetic tree was inferred from the multiple sequence alignment using Bayesian Markov Chain Monte Carlo (MCMC) method and Kimura-2 parameter (K2P) G + I as the best nucleotide substitution model with 1,000,000 generations. The same workflow was also employed for the RNA polymerase Beta subunit (rpoB) protein using Bayesian MCMC method employing model jumping between fixed rate amino acid models. The analysis was iteratively run until convergence at 100,000 generations. The operational taxonomic units (OTUs) included in the rpoB analysis used in the analysis were based on the list of Spring et al. (2015). Complete or draft genomes of related taxa based on 16S rRNA gene tree were searched for each taxon at the genus or species level. As not all taxa included in the analysis have complete or draft genomes available, only those genus or species with available genome data were included in the phylogenomic analysis. Gene (CDS) features were extracted from respective GenBank files as multi-FASTA amino acid (.faa) files. Species identification tool (specI) (Mende et al. 2013) was used to extract all universal single-copy genes (40 in the specI COG database, Supplementary Table 2) from the list of identified taxa. Then, a common set of marker genes were identified leaving 38 genes for comparison (Supplementary Table 2). The phylogenomic analysis using concatenated marker genes was performed following the same workflow as described previously for SSU rRNA. The evolutionary history was inferred using the neighbor-joining (NJ) method where the evolutionary distances were computed using the JTT matrix-based method with 1000 bootstrap resamplings. In silico DNA-DNA hybridization was simulated based on pairwise genome comparisons between AOL6 with Teredinibacter turnerae T7901 and with Saccharophagus degradans 2-40 using JSpecies (Richter and Rosselló-Móra 2009). In this analysis, the genome of AOL6 was randomly divided to 1020 bp where the resulting fragments were used as query for reference mapping.
Comparative genomics with respect to phylogenetically related carbohydrate degraders Teredinibacter turnerae T7901 (NC_012997.1) and Saccharophagus degradans 2-40 (NC_007912.1) was performed based on COGs using OrthoVenn (Wang et al. 2015) and CAZyme/carbohydrate-binding module profiles and metabolic pathways.
CAZymes—κ-Carrageenases
Computational analysis was performed to search for κ-carrageenases (κCa). An approach combining similarity- and profile-based methods, i.e., BLASTp search against GenBank NR, trEMBL and Swiss-Prot databases and HMM search (Eddy 1998; Finn et al. 2011) based on annotated 18 κCas (protein) sequences from UniProt (The UniProt Consortium 2014), and HMM search based on 120 SEED sequences for GH 16 superfamily was implemented. The intersection of the results, containing potential κCa-encoding sequences, was identified from the consolidated results.
Detection and Measurement of κ-Carrageenase Activity
Protein isoforms were detected from the culture medium (supplemented with 0.2% κ-carrageenan in marine broth) of AOL6 using zymography-based assay as described by Smith et al. (2005). The enzyme activity from the culture medium was measured using 3,5-dinitrosalicylic acid reducing sugar assay following the protocol of Tayco et al. (2013) against agar, alginate, ι-carrageenan, λ-carrageenan, and starch. This assay was performed in triplicates. The carrageenan (κ, ι, λ), alginate and starch powder used in this assay were obtained from Sigma-Aldrich. Agar was obtained from Conda Pronadisa.
Results
Morphological and Biochemical Characteristics of the Isolate
Using standard microbiological procedures, isolate AOL6 was found to be Gram-negative, aerobic, mesophilic, pigmented, rod-shaped, motile (Supplementary Fig. 1), and carrageenolytic. Details of the phenotypic and biochemical characteristics are described in a separate paper (David et al., manuscript in preparation).
Genome Assembly Metrics and Features
The sequencing run generated 166,418 single-end reads with mean base length per read of 446.95 bases. These reads are equivalent to a total of 74,380,448 bases. The Newbler-generated assembly consisted of 55 contigs covering a total of 4,129,655 bases, with an N50 of 193,405 bases and a mean coverage of 18×. The genome size of the bacterium was estimated to be 4.4 Mb. Thus, the sequencing experiment has covered 93.86% of the expected genome size. On the other hand, k-mer counting methods estimated the genome size to be 4.04 Mb. The assembly metrics, detailed in Table 1, indicated that majority of the sequence reads were used in the assembly leaving only a miniscule portion (0.0012% reads, 0.093% total bases) as singletons—generating 55 contiguous sequences/contigs. On the other hand, the general features of the genome showed a GC composition of 52%, 1 copy of small subunit (SSU/16S) rRNA gene, 1 copy of large subunit (LSU/23S) rRNA gene, 1 copy of 5S rRNA gene, 40 tRNA genes (Supplementary Table 1), and 1 transfer-messenger RNA (tmRNA). The genome was observed to encode 3659 protein-coding genes (CDS).
Phylogenetic Affiliation
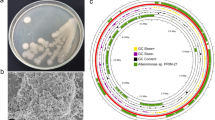
Phylogenetic analysis of 16S rRNA gene sequences and rpoB protein sequences extracted from the assembly allowed the identification of species closely related to isolate AOL6 (Fig. 1a). Based on the topology of the trees (Fig. 1a, b, and c), AOL6 was found to be most closely related to known carbohydrate-active bacteria Teredinibacter turnerae T7901 (Ttu T7901) and Saccharophagus degradans 2-40 (Sde 2-40) and to a free-living marine bacterium Eionea nigra 17X/A02/237 (Fig. 1a). Together, they form a distinct clade under family Cellvibrionaceae, affiliated to phylum Proteobacteria and class Gammaproteobacteria. Of these three related taxa, only Ttu T7901 and Sde 2-40 have genome data available for comparison. Hence, whole genome comparisons were limited to these two-related species. Phylogenomics based on 38 universal single-copy marker genes (Supplementary Table 2) produced a tree with similar topology (Fig. 1c), placing AOL6 adjacent to but distinct from Ttu T7901 and Sde 2-40 species. The phylogenetic affiliation of AOL6 with respect to these two carbohydrate degraders was in congruence with in silico DNA-DNA hybridization results (Fig. 1d). The average nucleotide identity (ANI, BLAST and MUMmer—ANIb and ANIm) and tetra (tetranucleotide signatures) values between the taxa were lower than the required threshold for species circumscription. Traditional DNA-DNA hybridization values recommend the species borders to be set at 60–70% which can be equated to ANI (average nucleotide identity) range of 95–96% and tetra value of ≥ 0.99 (Richter and Rosselló-Móra 2009).
a Phylogenetic affiliation of AOL6 with other marine bacteria based on 16S (SSU) rDNA gene sequences and b rpoB protein sequences reconstructed using Bayesian MCMC method. Numbers at the nodes indicate posterior probabilities based on 1,000,000 (16S) and 100,000 (rpoB) generations. Bar (0.05) indicates nucleotide/amino acid substitutions per site. Accession numbers are indicated before the name of each taxon. AOL6 and closely related bacteria are grouped as a clade under Cellvibrionaceae. c Neighbor-joining (NJ) tree based on 38 single-copy marker genes (proteins, Supplementary Table 2) where bar (0.05) and numbers at the nodes indicate amino acid substitutions per site and bootstrap support, respectively. Accession numbers are indicated before the name of each taxon. d In silico DNA-DNA hybridization with respect to T. turnerae T7901 (Ttu T7901) and S. degradans (Sde 2-40) genomes
Comparative Genome Analysis
Comparative analysis of AOL6 with respect to Ttu T7901 and Sde 2-40 highlighted key similarities and differences between the species, including differences in metabolic pathways (Supplementary Tables 4 and 5). Table 2 shows a comparison of the general features of the genomes of AOL6 and the closely related Cellvibrionaceae species Ttu T7901 (Yang et al. 2009), Sde 2-40 (Weiner et al. 2008) and Cellvibrio japonicus Ueda107 ((DeBoy et al. 2008), GenBank Accession No. CP000934). The genome of AOL6 is the smallest in this group (20% smaller compared to Ttu T7901 and Sde 2-40, respectively), and contained the fewest ribosomal RNA operon (rrn) copy number (1 vs. 2 or 3) and fewer CDS regions (3659 vs. 4105 and 4065 of Ttu T7901 and Sde 2-40, respectively). It is important to take note, however, that the exact rrn copy number of AOL6 cannot be definitively determined using the current genome assembly.
COG category assignment of AOL6, Ttu T7901, and Sde 2-40 shows that 86, 88, and 89% annotation coverage of respective CDSs have been achieved (Table 3). Considering that CDSs may be assigned to multiple COG categories and in order to get an accurate representation of COG categories, CDS assigned to multiple COG categories was thus counted individually and separately. Hence, the number of COG category assignments would not necessarily reflect the number of CDSs/proteins annotated. The number of “S, unknown function” category was excluded from downstream analyses involving total number of COG category assignments as its sheer overrepresentation would potentially mask important recognizable features as presented in Fig. 2 and Table 9.
Functional classification of COGs appears to be similar among the three bacteria (Fig. 2). A genome-wide comparative clustering analysis of homologous proteins among AOL6, Ttu T7901, and Sde 2-40 indicated they have 2432, 2497 and 2584 clusters, respectively (Fig. 3). Together these species formed 2912 clusters, 2759 orthologous clusters (which comprise at least two species), and 1782 single-copy clusters. From these clusters, there are 34 paralogous clusters in AOL6. Gene ontology term enrichment (GO enrichment) assigned these clusters to 12 GO terms which comprised 48 proteins. These proteins belong to 12 distinct names representing specific biological process, molecular function, and cellular component (Table 4). Noteworthily, some of these proteins are putatively involved in carbohydrate processing specifically κ-carrageenan hydrolysis and other carbohydrate modification enzymes.
The proportion of CAZymes in the genome (CAZome) of AOL6 was analyzed and compared to CAZomes of known carbohydrate-degrading marine bacteria as shown in Table 5. The proportion of CAZymes in the genome of AOL6 is 5.82% which consist of the hydrolytic enzymes glycoside hydrolases, polysaccharide lyases, carbohydrate esterases, glycosyl transferases, and auxiliary REDOX enzymes. This proportion exceeded that of known and previously identified carbohydrate degraders except Saccharophagus degradans 2-40 (Mann et al. 2013). In-depth analysis of more species-specific CAZymes and CBMs with respect to Ttu T7901 and Sde 2-40 revealed distinct patterns that differentiate one from the other species (Supplementary Fig. 2), such as in terms of the number and relative abundance of each CAZyme and CBMs in the genome. The difference in the composition and abundance of carbohydrate-binding modules (CBMs) in the genomes of AOL6, Ttu T7901, and Sde 2-40 was apparent, where each genome appeared to have a unique CBM profile (Fig. 4). Proteins containing a CBM but without a catalytic domain were also observed in the genome of AOL6 (Supplementary Table 3). For instance, it was observed that CBM8 and CBM61 are only found in AOL6 (Fig. 4). Interestingly, these two CBMs have never been observed to be associated or to co-exist with any catalytic domains of CAZymes identified in AOL6.
Comparative analysis of the carbohydrate-binding modules (CBM) of AOL6 including T. turnerae T7901 and S. degradans 2-40 using dbCAN (database for carbohydrate-active enzyme annotation) as a reference/database. CBMs marked with asterisk (*) are those not found to be in the same protein with CAZyme catalytic domains (in AOL6 only) (left panel). Top 10 CBMs per species are shaded in gray (right panel)
κ-Carrageenases and Polysaccharide-Degrading Activities
Three potential κCa-encoding ORFs were identified in the genome of AOL6 designated as κCa02330, κCa02332, and κCa10164 (Table 6). κCa02330 has a predicted molecular weight of 51.15 kDa similar to previously identified κ-carrageenases (Potin et al. 1991; Zhou et al. 2008; Sun et al. 2010; Liu et al. 2011, 2013; Zhu and Ning 2016). κCa02332 and κCa10164 were predicted to be relatively larger with molecular weights of 86.98 and 95.98 kDa, respectively, but these are smaller than the 128-kDa κ-carrageenase identified by Khambhaty et al. (2007) from Pseudomonas elongata. A zymography assay (which would reveal the combined effect of a mixture of active enzymes secreted by the bacteria) using κ-carrageenan as substrate detected at least 3 isozymes (Fig. 5). This result was consistent with the result of the computational analysis which identified 3 potential κ-carrageenase-encoding genes (Table 6). Assays were also carried out to determine the enzymatic activity of crude extract of AOL6 (grown in κ-carrageenan-supplemented marine broth) against κ-carrageenan and other polysaccharide substrates. Figure 6 revealed high enzyme activity against κ-carrageenan at 80.22 U/ml where 1 unit of enzyme is equivalent to the protein that produces 1 μg of D-galactose per minute (significantly lower activities were detected against the other polysaccharides, including other carrageenans, which ranged from 16 to 20 U/ml).
Bacterial Secretion System
Homologs of most genes/proteins involved in the bacterial type ii secretion system (T2SS) were found in the genome of isolate AOL6 including Sec and Tat general secretion systems Fig. 7). These proteins included GspD, GspC, GspF, GspG, GspI, GspJ, GspK, GspL, GspM, GspE, SecD/F, SecE, SecG, SecY, YajC, YidC, SecA, FtsY, SecB, ffh, TatA, TatB, and TatC. Approximately 12% (434 of 3659) of the total predicted proteins from the genome of AOL6 (Table 7) maintain an N-terminal signal peptide sequence where 16% (71 of 434) of these are putative CAZymes comprised by GH (glycoside hydrolases), PL (polysaccharide lyases), CE (carbohydrate esterases), GT (glycosyl transferases), and AA (auxiliary enzymes or REDOX enzymes). Based on the number of predicted CAZymes including CBMs (carbohydrate-binding modules), it is therefore predicted that about 34% (81 of 239) are possible substrates of or secreted extracellularly through a T2SS.
Two-Component Signal Transduction Systems
Bioinformatics analysis based on P2RP (Barakat et al. 2013) identified several putative TCS proteins. From this analysis, 101 TCS proteins were identified and classified to 49 histidine kinases (HKs), 49 response regulators (RRs), and 3 phosphotransferases (PPs) (Table 8). Based on a 200-bp distance cut-off (Williams and Whitworth 2010) for functionally-linked TCS proteins, the proportion of paired TCS genes in the genome of AOL6 was 54% (54/101) while the remaining fraction was classified as either complex (8%, 8/101) or orphaned (39%, 39/101). This number is much higher than in Teredinibacter turnerae T7901 and Saccharophagus degradans 2-40, which have 20% paired-50% orphaned and 24% paired-46% orphaned TCS, respectively (Ortet et al. 2015). On the other hand, in E. coli, a distant relative in Gammaproteobacteria, a large majority (71.5%) of TCS proteins are organized in pairs while orphaned and complex TCS account for only 14.5 and 14.5%, respectively (Mizuno 1997). While majority of the prokaryotic TCS proteins (62%, e.g., Firmicutes) are encoded in pairs at loci within genomes, some organisms appear to favor encoding TCS proteins as orphan genes (e.g., Xanthomonas) (Williams and Whitworth 2010). Comparison of the number of TCS proteins among carbohydrate-degrading representatives of phylum Proteobacteria and Bacteriodetes (Table 8 and Fig. 8) revealed interesting patterns of composition and abundance of RRs; the distribution of RRs appeared to be clade-specific regardless of the habitat of the bacteria.
Discussion
AOL6 Represents a Novel Copiotrophic, Carrageenolytic Marine Bacterium
Molecular phylogenetic analysis based on 16S rRNA gene sequences and rpoB protein sequences identified the bacterial species most closely related to isolate AOL6 (Fig. 1a, b), Teredinibacter turnerae T7901 (Ttu T7901) and Saccharophagus degradans 2-40 (Sde 2-40). However, the 16S rRNA and rpoB sequences of AOL6 were only 93 and 90% identical to the corresponding sequences in Ttu T7901 and Sde 2-40, respectively, hence indicating that AOL6 is a distinct species. Phylogenomic and in silico DNA-DNA hybridization analyses (Fig. 1c, d) also supported the same conclusion. The results of the analyses in fact suggest that AOL6 represents not only a novel species but also a new genus in the family Cellvibrionceae. AOL6 is a carrageenolytic marine bacterium isolated from a diseased thallus of a farmed rhodophyte Kappaphycus alvarezii (Gigartinales, Rhodophyta). Ttu T7901, a closely related species, is a cellulolytic, diazotrophic, and Gram-negative marine bacterium isolated from the gills of a broad range of wood-boring marine bivalves of the family Teredinidae while Sde 2-40 is a free-living Gram-negative bacterium isolated from decaying seagrass (Weiner et al. 2008; Yang et al. 2009). Eionea nigra 17X/A02/237, a member of the same group, is a Gram-negative bacterium isolated from the coastal waters of the north-western Mediterranean Sea (Urios et al. 2011). The carbohydrate-degrading activity of Eionea nigra remains to be confirmed. The phylogenetic affinity of AOL6 places it as potential member of the emerging Microbulbifer/Teredinibacter/Saccharophagus group of marine carbohydrate degraders (Weiner et al. 2008). Spring et al. (2015) identified the members of the family Cellvibrionaceae and Microbulbiferaceae that comprise a copiotrophic group of marine bacteria with special ability to utilize complex polysaccharides as substrates. Based on rpoB sequences (Fig. 1b), AOL6 belongs to a group of bacteria known as r-strategists, characterized by preference for nutrient-rich environments (Spring et al. 2015). Furthermore, the definition of prokaryotic K-strategists and r-strategists can be based on the copy number of rRNA operons, rpoB sequence delineation in reference to Fig. 1b, and the number of COG category assignments in Table 9 (Spring et al. 2015). This inferred ecological adaptation based on rpoB protein sequence (r-selection) was also consistent with the overrepresentation of COG categories namely motility, defense mechanisms, signal transduction mechanisms, and transcription (Table 9). Finally, on the basis of genome size and number of protein-coding sequences (Giovannoni et al. 2005), AOL6 appeared unlikely to be an obligate symbiont and is therefore free-living or host-associated. On the other hand, ribosomal RNA operon (rrn) copy number in a bacterial genome can be used to infer the ecological and reproductive strategies of an organism in response to specific ecological niche where copiotrophs generally have more than 1 copy of rrn while oligotrophs have only 1 copy of rrn (Klappenbach et al. 2000; Lauro et al. 2009; Roller et al. 2016). In AOL6, the exact rrn copy number (Table 2) cannot be conclusively determined as the average read length of pyrosequencing introduces an inherent limitation in resolving tandemly repeated copies of ribosomal RNA genes. The sequencing depth spanning the predicted 16S rRNA gene (170×) far exceeds that of the average sequencing depth for the draft genome (18×). Therefore, this raises the possibility of a mis-assembly leading to a generalized repeat collapse and expansion (Phillippy et al. 2008) and which can only be addressed if the reads are long enough to span this tandemly repetitive elements (Sohn and Nam 2018). Notwithstanding the general trend of rrn copy number with respect to trophic strategies, it has been shown that counterintuitively large genomes of Planctomycetes (a feature of copiotrophs) possess only a single copy of rrn (a feature of oligotrophs) (Lauro et al. 2009).
Carrageenan-degrading or carrageenolytic marine bacteria belong to two distantly related phyla Proteobacteria and Bacteriodetes, although most of the isolates belong to the former group (Chauhan and Saxena 2016). Most of these isolates are Gram-negative, including Pseudoalteromonas, Cellulophaga, Pseudomonas, Cytophaga, Tamlana, Vibrio, Catenovolum, Microbulbifer, Zobellia, and Alteromonas (Michel et al. 2001; Mou et al. 2004; Sun et al. 2010; Hatada et al. 2011; Li et al. 2013; Liu et al. 2013; Yao et al. 2013; Ziayoddin et al. 2014; Zhu and Ning 2016). Like most carrageenolytic marine bacteria, AOL6 is a Gram-negative bacterium that belongs to phylum Proteobacteria. Some Gram-positive bacteria such as Bacillus sp. (Kang and Kim 2015) have also been reported to produce carrageenases. While several carrageenolytic bacteria have already been identified and characterized, AOL6 is among the relatively few bacteria whose genome was sequenced and characterized.
Comparative Genomics Reveals Shared and Distinct Features with Respect to Closely Related Carbohydrate-Degrading Species T. turnerae T7901 and S. degradans 2-40
Comparative genomic analysis with respect to Ttu T7901 and Sde 2-40 identifies the key features that differentiate AOL6 from the two species. GO enrichment analysis (Fig. 2) identified 34 paralogous clusters in AOL6 that are involved in carbohydrate processing, and these included κ-carrageenase activity. Ttu T7901 and Sde 2-40 are carbohydrate-active bacteria that degrade cellulosic materials (Weiner et al. 2008; Yang et al. 2009). Therefore, this indicated that the bacteria can be differentiated based on carbohydrate metabolism, e.g., carbohydrate specificity (carbohydrate-binding modules, CBM) and utilization (CAZymes). Analysis of more species-specific CAZymes and CBMs revealed distinct patterns that differentiate one from the other (Fig. 4 and Supplementary Fig. 2). This analysis revealed a unique profile for each species highlighting the number and relative abundance of each CAZyme and CBM in the genome. Specifically, it was observed that CBM8 and CBM61 are only found in AOL6 (Fig. 4 and Supplementary Table 3). These two CBMs are observed neither to be associated nor to co-exist with any catalytic domain in CAZymes. CBM8 is a cellulose-binding module from a cellulase of the slime mold Dictyostelium discoideum while CBM61 is a binding module for β-1,4-galactan and is found to co-occur with catalytic domains in GH16, GH30, GH31, GH43, GH53, and GH66 (Lombard et al. 2014). CBMs are non-catalytic in nature and are usually found in association with the catalytic domains of CAZymes. CBM is as a contiguous amino acid sequence within a CAZyme with a discrete fold having carbohydrate-binding activity. The presence of CBM in an enzyme confers substrate specificity to the corresponding CAZyme (Boraston et al. 2004; Lombard et al. 2014). In the case of cellulases, CBMs comprise part of a scaffolding subunit that organizes the catalytic subunits into a cohesive multienzyme complex known as cellulosome (Shoseyov et al. 2006).
Critical to the assembly of cellulosomes are two groups of structural modules, namely, cohesins and dockerins which mediate the interaction of different enzymes to form molecular assemblies that degrade plant biomass (Bayer et al. 2004). Interestingly, a search for these modules in the genome of AOL6 revealed that neither cohesins nor dockerins were found in the genome. A similar search in the genomes of Teredinibacter turnerae T7901 or Saccharophagus degradans 2-40 found that either cohesins or dockerins, but not both, were encoded in the genome of these two species. The implications of these findings remain to be investigated.
CAZome Analysis Reveals CAZyme Abundance Comparable to Other Carbohydrate-Degrading Marine Bacteria: a Fraction of Cell Wall-Degrading Enzymes Is Predicted to Be Secreted Through the T2SS Pathway
An organism may be described by the number of CAZymes in its genome. In the study conducted by Barbeyron et al. (2016), carbohydrate catabolism of marine heterotrophic bacteria (MHB) is influenced by phylogeny, as well as habitat, and as a consequence, CAZymes can be used as a predictive tool to assess a bacterial species’ ecological niche, in particular its role in the carbon cycle. Therefore, the varying proportion of CAZymes in the genome can be used as an indication of bacterial behavior or lifestyle. The CAZome of free-living organisms typically correspond to 1–5% of the predicted coding sequences. Extremely reduced CAZomes are characteristic of species with a strict intracellular parasitic lifestyle (Lombard et al. 2014). For instance, microorganism that degrade plant cell walls synthesizes an extensive portfolio of hydrolytic enzymes that display complex molecular structures (DeBoy et al. 2008). In AOL6, the proportion of CAZymes in the genome is 5.82%—possibly indicating a specialized function to degrade a range of polysaccharides. This value exceeds that of most known and previously identified carbohydrate degraders (Table 5) (Mann et al. 2013). The number of CAZymes in the genome of AOL6 suggests that it does not only live and thrive in symbiosis with carrageen-rich algae but is also capable of degrading and utilizing other polysaccharide substrates—suggesting a promiscuous lifestyle.
Protein secretion plays a central role in bacterial response to environmental conditions or stimuli. In particular, symbiotic bacteria, whether pathogenic, commensal, or mutualistic, rely on a variety of secreted proteins in its interaction with their host organisms (Tseng et al. 2009a). The presence of genes involved in the bacterial T2SS in the genome of AOL6 (Fig. 7) but not of other types of secretion systems was an interesting finding. T2SS, a protein secretion machinery found in various Gram-negative bacteria and is one of the known six protein secretory systems (Tseng et al. 2009b), enables the transport of cytoplasmic proteins that are generally associated with the breakdown of host tissues and therefore important in causing the symptoms associated with bacterial infection. The secreted proteins have a range of biological functions, but they are generally enzymes. They include proteases, lipases, phosphatases, and CAZymes where the site of action is primarily extracellular (Korotkov et al. 2012). In contrast to AOL6, the genome of the endosymbiont Teredinibacter turnerae T7901 apparently encodes types III, IV, and VI (T3SS, T4SS, and T6SS) secretion systems apart from T2SS (Yang et al. 2009). The presence of such secretion systems is believed to be the hallmarks of intracellular symbionts (Lee and Schneewind 2001), and therefore, their absence in the genome of AOL6 suggests that this species is unlikely to be an intracellular pathogen or symbiont. On the other hand, the genome of Saccharophagus degradans 2-40 encodes T1SS and T2SS secretion systems (Weiner et al. 2008).
Genome analysis (Table 7) shows that approximately 12% (432/3659) of the total predicted proteins from the genome of AOL6 maintain an N-terminal signal peptide sequence where 16% (71/434) of these are CAZymes comprised by GH (glycoside hydrolases), PL (polysaccharide lyases), CE (carbohydrate esterases), GT (glycosyl transferases), and AA (auxiliary enzymes or REDOX enzymes). Based on the number of predicted CAZymes including CBMs (carbohydrate-binding modules), it is proposed that about 34% (81/239) of these are possible substrates of or secreted extracellularly through the type II secretion system.
Aside from the apparent exclusive use of T2SS for CAZymes secretion, the analysis of the AOL6 genome also revealed how CAZymes (and perhaps other hydrolytic or degradative enzymes) may be delivered to their target efficiently by this species. Codon usage preference analysis can be used to predict the highly expressed genes in a genome (Gupta et al. 2004; Angellotti et al. 2007; Sahoo and Das 2014; Zhou et al. 2016; Das et al. 2017; Uddin 2017). From this strategy, pilZ gene is identified to be potentially highly expressed. The product of this gene is believed to be involved in type IV fimbrial biogenesis promoting adhesion to host cells during colonization and are associated with a form of surface translocation called twitching motility (Alm et al. 1996; Han et al. 2007; Yamagata et al. 2012; Nivaskumar and Francetic 2014; Green and Mecsas 2016). Conceivably, the putative high expression pilZ gene may play a role in secreting CAZymes efficiently during colonization of algal hosts.
The Collection of TCS Proteins in AOL6 Genome Putatively Reflects Role in Coordinating Complex and Dynamic Interactions Between the CAZyme-Rich Bacterium and Its Polysaccharide-Rich Host
The TCSs are one of the primary means by which bacteria regulate their behavior in response to external or environmental stimuli (Borland et al. 2015). They play important roles in a broad range of physiological mechanisms such as virulence, chemotaxis, motility, and metabolism which are essential for plant-associated bacteria (Gao and Stock 2009; Leonard et al. 2017). Coordination of virulence mechanisms involves several TCSs (Leonard et al. 2017). Therefore, identification of TCSs may provide valuable insights into the behavior and virulence mechanism of a bacterium. Table 8 shows that majority of the predicted TCSs in AOL6 are involved in bacterial chemotaxis and surface colonization, which could mediate the infection process. Diseases in seaweeds such as the “ice-ice” disease observed in Kappaphycus species result from infection by opportunistic pathogens promoted by unfavorable environmental conditions (Largo et al. 1999). Bacteria with polysaccharide-degrading abilities tend to be opportunistic pathogens or saprophytes rather than commensal or mutualistic macroalgal symbionts (Egan et al. 2013). Initial events in most plant-microbe interactions include bacterial chemotaxis toward plant exudates and bacterial attachment on the plant surface (Yao and Allen 2006; Antúnez-Lamas et al. 2009; Heindl et al. 2014 cited in Leonard et al. 2017). It is surmised that regulation of expression of chemotaxis and surface colonization proteins, as well as the synthesis of extracellular enzymes, is facilitated by the TCS proteins coordinating complex and dynamic host-bacterial interactions. In the case of AOL6, its colonization of its algal hosts would conceivably involve its rich repertoire of predicted response regulators putatively involved in bacterial chemotaxis and surface colonization, in concert with its T2SS that enable type IV fimbrial biogenesis followed by extrusion of cell wall-degrading enzymes, including κ-carrageenases. Investigation of this hypothesis would be a step toward a deeper mechanistic understanding of how bacteria cause diseases in seaweeds.
Comparative investigation reveals a pattern of TCS protein composition and abundance among carbohydrate-degrading marine bacteria at different taxonomic lineages and habitats. The repertoire of TCS proteins appears to be phylogeny-dependent rather than habitat-dependent (Table 8 and Fig. 8) suggesting that bacteria belonging to a specific taxonomic rank could alternate from one lifestyle to another and assume different lifestyles in response to habitat condition. T. turnerae is an endosymbiont from the gills of a broad range of wood-boring bivalves of family Teredinae (shipworms) (Distel et al. 2002; Yang et al. 2009). S. degradans 2-40 was isolated from a dead, decaying seagrass. By far, S. degradans 2-40 has the largest repertoire of CAZymes and has been demonstrated to degrade an array of complex polysaccharides (Ekborg et al. 2005; Weiner et al. 2008). Microbulbifer agarilyticus GP101 (genome accession no. NZ_CP019650.1), isolated from ocean sediments, possessed both agarolytic and carrageenolytic activities (Miyazaki et al. 2008). Alteromonas macleodii ATCC 27126 (genome accession no. NC_018632.1) is a copiotrophic and planktonic marine bacteria (Ivars-Martinez et al. 2008). Zobellia galactanivorans DSIJ (genome accession no. NC_015844.1) is a seaweed-associated (rhodophyte) marine bacterium that possesses both agarolytic and carrageenolytic activities and is currently a model for studying algal polysaccharide bioconversions (Barbeyron et al. 2001; Barbeyron et al. 2016).
Conclusion
In this study, the genome of a newly discovered seaweed-associated, carrageenolytic, and culturable marine bacterium designated as isolate AOL6 was sequenced and analyzed. Phylogenetic and phylogenomic analyses suggest that this bacterium represents not only a novel species but also a novel genus in the family Cellvibrionaceae. Analysis of its genome revealed several interesting features. Similar to its relatives in the family Cellvibrionaceae, such as T. turnerae T7901 and S. degradans 2-40, AOL6 has an abundance of CAZyme-encoding genes in the genome. Unlike Ttu T7901 and Sde 2-40, however, it includes carrageenases in its CAZyme repertoire and apparently relies exclusive on T2SS for secreting proteins, presumably including the cell wall-degrading enzymes polysaccharide lyases, carbohydrate esterases, and glycoside hydrolases. In addition, majority of its putative TCSs are predicted to be involved in bacterial chemotaxis and surface colonization. Its pilZ gene, whose product is presumably involved in cell adhesion and CAZyme secretion, also appears to be potentially actively expressed based on codon usage bias analysis. These characteristics suggest that AOL6 is an opportunistic, potentially pathogenic, r-strategist, adapted to colonizing polysaccharide-rich hosts, such as seaweeds.
References
Aldridge P, Paul R, Goymer P, Rainey P, Jenal U (2003) Role of the GGDEF regulator PleD in polar development of Caulobacter crescentus. Mol Microbiol 47:1695–1708
Alm RA, Bodero AJ, Free PD, Mattick JS (1996) Identification of a novel gene, pilZ, essential for type 4 fimbrial biogenesis in Pseudomonas aeruginosa. J Bacteriol 178:46–53
Altschul S, Gish W, Miller W, Myers E, Lipman D (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Angellotti MC, Bhuiyan SB, Chen G, Wan XF (2007) CodonO: codon usage bias analysis within and across genomes. Nucleic Acids Res 35:W132–W136
Antúnez-Lamas M, Cabrera-Ordóñez E, López-Solanilla E, Raposo R, Trelles-Salazar O, Rodríguez-Moreno A, Rodríguez-Palenzuela P (2009) Role of motility and chemotaxis in the pathogenesis of Dickeya dadantii 3937 (ex Erwinia chrysanthemi 3937). Microbiology 155:434–442
Barakat M, Ortet P, Whitworth DE (2013) P2RP: a web-based framework for the identification and analysis of regulatory proteins in prokaryotic genomes. BMC Genomics 14:269
Barbeyron T, L’Haridon S, Corre E, Kloareg B, Potin P (2001) Zobellia galactanovorans gen. nov., sp. nov., a marine species of Flavobacteriaceae isolated from a red alga, and classifica- tion of [Cytophaga] uliginosa (Zo Bell and Upham 1944) Reichenbach 1989 as Zobellia uliginosa gen. nov., comb. nov. Int J Syst Evol Microbiol 51(51):985–997
Barbeyron T, Thomas F, Barbe V, Teeling H, Schenowitz C, Goesmann A, Leblanc C, Glöckner FO, Czjzek M, Michel G et al (2016) Habitat and taxon as driving forces of carbohydrate catabolism in marine heterotrophic bacteria: example of the model algae-associated bacterium Zobellia galactanivorans DsijT. Environ Microbiol 18:4610–4627
Bayer EA, Belaich J-P, Shoham Y, Lamed R (2004) The cellulosomes: multienzyme machines for degradation of plant cell wall polysaccharides. Annu Rev Microbiol 58:521–554. http://www.annualreviews.org/doi/10.1146/annurev.micro.57.030502.091022. Accessed 30 May 2018
Bischoff M, Berger-Bachi B (2001) Teicoplanin stress-selected mutations increasing B activity in Staphylococcus aureus. Antimicrob Agents Chemother 45:1714–1720. http://www.ncbi.nlm.nih.gov/pubmed/11353616. Accessed 20 Oct 2017
Boraston AB, Bolam DN, Gilbert HJ, Davies GJ (2004) Carbohydrate-binding modules: fine-tuning polysaccharide recognition. Biochem J 382:769–781. http://biochemj.org/lookup/doi/10.1042/BJ20040892. Accessed 29 Aug 2017
Borland S, Oudart A, Prigent-combaret C, Brochier-armanet C, Wisniewski-dyé F (2015) Genome-wide survey of two-component signal transduction systems in the plant growth-promoting bacterium Azospirillum. BMC Genomics 16:1–17
Chauhan P, Saxena A (2016) Bacterial carrageenases: an overview of production and biotechnological applications. 3 Biotech 6:146
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ, Kulam-Syed-Mohideen AS, McGarrell DM, Marsh T, Garrity GM et al (2009) The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37:D141–D145
Cousens D, Clarke P, Drew R (1987) The amidase regulatory gene (amiR) of Pseudomonas aeruginosa. J Gen Microbiol 133:2041–2052
Das S, Chottopadhyay B, Sahoo S (2017) Comparative analysis of predicted gene expression among crenarchaeal genomes. Genomics Inform 15:38–47
De Ruiter G, Rudolph B (1997) Carrageenan biotechnology. Trends Food Sci Technol 8:389–395
DeBoy RT, Mongodin EF, Fouts DE, Tailford LE, Khouri H, Emerson JB, Mohamoud Y, Watkins K, Henrissat B, Gilbert HJ et al (2008) Insights into plant cell wall degradation from the genome sequence of the soil bacterium Cellvibrio japonicus. J Bacteriol 190:5455–5463
Delcher A, Harmon D, Kasif S, White O, Salzberg S (1999) Improved microbial gene identification with GLIMMER. Nucleic Acids Res 27:4636–4641
Deorowicz S, Kokot M, Grabowski S, Debudaj-Grabysz A (2015) KMC 2: fast and resource-frugal k-mer counting. Bioinformatics 31:1569–1576. https://academic.oup.com/bioinformatics/article-lookup/doi/10.1093/bioinformatics/btv022. Accessed 26 Apr 2018
Distel D, Morrill W, Maclaren-Toussaint N, Franks D, Waterbury J (2002) Teredinibacter turnerae gen. nov., sp. nov., a dinitrogen-fixing, cellulolytic, endosymbiotic γ-proteobacterium isolated from the gills of wood-boring molluscs (Bivalvia : Teredinidae). Int J Syst Evol Microbiol 52:2261–2269
Dons L, Eriksson E, Jin Y, Rottenberg ME, Kristensson K, Larsen CN, Bresciani J, Olsen JE (2004) Role of flagellin and the two-component CheA/CheY system of Listeria monocytogenes in host cell invasion and virulence. Infect Immun 72:3237–3244. http://www.ncbi.nlm.nih.gov/pubmed/15155625. Accessed 20 Oct 2017
Eddy SR (1998) Profile hidden Markov models. Bioinformatics 14:755–763
Egan S, Harder T, Burke C, Steinberg P, Kjelleberg S, Thomas T (2013) The seaweed holobiont: understanding seaweed–bacteria interactions. FEMS Microbiol Rev 37:462–476. https://academic.oup.com/femsre/article-lookup/doi/10.1111/1574-6976.12011. Accessed 21 May 2017
Ekborg NA, Gonzalez JM, Howard MB, Taylor LE, Hutcheson SW, Weiner RM (2005) Saccharophagus degradans gen. nov., sp. nov., a versatile marine degrader of complex polysaccharides. Int J Syst Evol Microbiol 55:1545–1549
Ewann F, Jackson M, Pethe K, Cooper A, Mielcarek N, Ensergueix D, Gicquel B, Locht C, Supply P (2002) Transient requirement of the PrrA-PrrB two-component system for early intracellular multiplication of Mycobacterium tuberculosis. Infect Immun 70:2256–2263. http://www.ncbi.nlm.nih.gov/pubmed/11953357. Accessed 20 Oct 2017
Finn RD, Clements J, Eddy SR (2011) HMMER web server: interactive sequence similarity searching. Nucleic Acids Res 39:W29–W37. http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=3125773&tool=pmcentrez&rendertype=abstract. Accessed 28 Sep 2013
Galperin MY (2006) Structural classification of bacterial response regulators: diversity of output domains and domain combinations. J Bacteriol 188:4169–4182
Gao R, Stock A (2009) Biological insights from structures of two-component proteins. Annu Rev Microbiol 63:133–154
Giovannoni S, Tripp J, Givan S, Podar M, Vergin K, Baptista D, Bibbs L, Eads J, Richardson T, Noordewier M et al (2005) Genome streamlining in a cosmopolitan oceanic bacterium. Science 309:1242–1245
Green ER, Mecsas J (2016) Bacterial secretion systems: an overview. In: Kudva I, Cornick N, Plummer P, Zhang Q, Nicholson T, Bannantine J, Bellaire B (eds) Virulence mechanisms of bacterial pathogens, 5th edn, vol 4. ASM Press, Washington, DC, pp 215–239
Gupta SK, Bhattacharyya TK, Ghosh TC (2004) Synonymous codon usage in lactococcus lactis: mutational bias versus translational selection. J Biomol Struct Dyn 21:527–535
Han X, Kennan RM, Parker D, Davies JK, Rood JI (2007) Type IV fimbrial biogenesis is required for protease secretion and natural transformation in Dichelobacter nodosus. J Bacteriol 189:5022–5033
Hatada Y, Mizuno M, Li Z, Ohta Y (2011) Hyper-production and characterization of the ι-Carrageenase useful for ι-carrageenan oligosaccharide production from a deep-sea bacterium, Microbulbifer thermotolerans JAMB-A94 T, and insight into the unusual catalytic mechanism. Mar Biotechnol 13:411–422
Hehemann JH, Boraston AB, Czjzek M (2014) A sweet new wave: structures and mechanisms of enzymes that digest polysaccharides from marine algae. Curr Opin Struct Biol 28:77–86. http://www.sciencedirect.com/science/article/pii/S0959440X14000815. Accessed 17 Aug 2015
Heindl JE, Wang Y, Heckel BC, Mohari B, Feirer N, Fuqua C (2014) Mechanisms and regulation of surface interactions and biofilm formation in agrobacterium. Front Plant Sci 5:176
Hu X, Jiang X, Aubree E, Boulenguer P, Critchley A (2006) Preparation and in vivo antitumor activity of kappa-carrageenan oligosaccharides. Pharm Biol 44:646–650
Huelsenbeck JP, Ronquist F (2001) MR BAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17:754–755
Huerta-Cepas J, Szklarczyk D, Forslund K, Cook H, Heller D, Walter MC, Rattei T, Mende DR, Sunagawa S, Kuhn M, Jensen LJ, von Mering C, Bork P (2016) EGGNOG 4.5: a hierarchical orthology framework with improved functional annotations for eukaryotic, prokaryotic and viral sequences. Nucleic Acids Res 44:D286–D293
Huson DH, Auch AF, Qi J, Schuster SC (2007) MEGAN analysis of metagenomic data. Genome Res 17:377–386. http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=1800929&tool=pmcentrez&rendertype=abstract. Accessed 9 Jul 2014
Iuchi S, Lin EC (1987) The narL gene product activates the nitrate reductase operon and represses the fumarate reductase and trimethylamine N-oxide reductase operons in Escherichia coli. Proc Natl Acad Sci U S A 84:3901–3905
Ivars-Martinez E, Martin-Cuadrado A-B, D’Auria G, Mira A, Ferriera S, Johnson J, Friedman R, Rodriguez-Valera F (2008) Comparative genomics of two ecotypes of the marine planktonic copiotroph Alteromonas macleodii suggests alternative lifestyles associated with different kinds of particulate organic matter. ISME J 2:1194–1212. http://www.ncbi.nlm.nih.gov/pubmed/18670397. Accessed 20 Dec 2017
Kalitnik AA, Byankina Barabanova AO, Nagorskaya VP, Reunov AV, Glazunov VP, Solov’eva TF, Yermak IM (2012) Low molecular weight derivatives of different carrageenan types and their antiviral activity. J Appl Phycol 25:65–72. http://springerlink.bibliotecabuap.elogim.com/10.1007/s10811-012-9839-8. Accessed 31 Oct 2014
Kanehisa M, Sato Y, Morishima K (2016) BlastKOALA and GhostKOALA: KEGG tools for functional characterization of genome and metagenome sequences. J Mol Biol 428:726–731
Kang S, Kim JK (2015) Reuse of red seaweed waste by a novel bacterium, Bacillus sp. SYR4 isolated from a sandbar. World J Microbiol Biotechnol 31:209–217. http://springerlink.bibliotecabuap.elogim.com/10.1007/s11274-014-1778-x. Accessed 15 Jan 2017
Khambhaty Y, Mody K, Jha B (2007) Purification and characterization of κ-carrageenase from a novel γ-proteobacterium,Pseudomonas elongata (MTCC 5261) syn. Microbulbifer elongatus comb. Nov. Biotechnol Bioprocess Eng 12:668–675. http://springerlink.bibliotecabuap.elogim.com/10.1007/BF02931084. Accessed 28 Mar 2015
Klappenbach JA (2001) rrndb: the ribosomal RNA operon copy number database. Nucleic Acids Res 29:181–184
Klappenbach JA, Dunbar JM, Thomas M, Schmidt TM (2000) rRNA operon copy number reflects ecological strategies of bacteria. Appl Environ Microbiol 66:1328–1333
Knobloch JK-M, Jager S, Horstkotte MA, Rohde H, Mack D (2004) RsbU-dependent regulation of Staphylococcus epidermidis biofilm formation is mediated via the alternative sigma factor B by repression of the negative regulator gene icaR. Infect Immun 72:3838–3848. http://www.ncbi.nlm.nih.gov/pubmed/15213125. Accessed 20 Oct 2017
Korotkov KV, Sandkvist M, Hol WGJ (2012) The type II secretion system: biogenesis, molecular architecture and mechanism. Nat Rev Microbiol 10:336–351
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Lagesen K, Hallin P, Rødland EA, Staerfeldt H-H, Rognes T, Ussery DW (2007) RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res 35:3100–3108. http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=1888812&tool=pmcentrez&rendertype=abstract. Accessed 25 Nov 2014
Lane D (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. John Wiley & Sons, Inc., New York, pp 115–176
Largo DB, Fukami K, Nishijima T (1999) Time-dependent attachment mechanism of bacterial pathogen during ice-ice infection in Kappaphycus alvarezii (Gigartinales, Rhodophyta). J Appl Phycol 11:129–136. http://springerlink.bibliotecabuap.elogim.com/article/10.1023/A%3A1008081513451. Accessed 25 Feb 2015
Laslett D, Canback B (2004) ARAGORN, a program to detect tRNA genes and tmRNA genes in nucleotide sequences. Nucleic Acids Res 32:11–16
Lauro FM, McDougald D, Thomas T, Williams TJ, Egan S, Rice S, DeMaere MZ, Ting L, Ertan H, Johnson J et al (2009) The genomic basis of trophic strategy in marine bacteria. Proc Natl Acad Sci 106:15527–15533
Lee VT, Schneewind O (2001) Protein secretion and the pathogenesis of bacterial infections. Genes Dev 15:1725–1752
Lee SH, Angelichio MJ, Mekalanos JJ, Camilli A (1998) Nucleotide sequence and spatiotemporal expression of the Vibrio cholerae vieSAB genes during infection. J Bacteriol 180:2298–2305
Leonard S, Hommais F, Nasser W, Lyon CB (2017) Plant–phytopathogen interactions: bacterial responses to environmental and plant stimuli. Environ Microbiol 19:1689–1716
Leung E, Huang A, Cadag E, Montana A, Soliman JL, Zhou CLE (2016) Protein sequence annotation tool (PSAT): a centralized web-based meta-server for high-throughput sequence annotations. BMC Bioinformatics 17:1–7
Li S, Jia P, Wang L, Yu W, Han F (2013) Purification and characterization of a new thermostable κ-carrageenase from the marine bacterium Pseudoalteromonas sp. QY203. J Ocean Univ China 12:155–159. http://springerlink.bibliotecabuap.elogim.com/10.1007/s11802-013-1994-2. Accessed 2 Nov 2014
Liu G-L, Li Y, Chi Z, Chi Z-M (2011) Purification and characterization of κ-carrageenase from the marine bacterium Pseudoalteromonas porphyrae for hydrolysis of κ-carrageenan. Process Biochem 46:265–271. http://linkinghub.elsevier.com/retrieve/pii/S1359511310003326. Accessed 2 Nov 2014
Liu Z, Li G, Mo Z, Mou H (2013) Molecular cloning, characterization, and heterologous expression of a new κ-carrageenase gene from marine bacterium Zobellia sp. ZM-2. Appl Microbiol Biotechnol 97:10057–10067
Liu G, Wu S, Jin W, Sun C (2016) Amy63, a novel type of marine bacterial multifunctional enzyme possessing amylase, agarase and carrageenase activities. Sci Rep 6:18726.
Lombard V, Golaconda Ramulu H, Drula E, Coutinho PM, Henrissat B (2014) The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res 42:D490–D495. http://nar.oxfordjournals.org/content/42/D1/D490.long. Accessed 10 Jul 2014
Makarova KS, Wolf YI, Snir S, Koonin EV (2011) Defense Islands in bacterial and archaeal genomes and prediction of novel defense systems. J Bacteriol 193:6039–6056
Mann AJ, Hahnke RL, Huang S, Werner J, Xing P, Barbeyron T, Huettel B, Stüber K, Reinhardt R, Harder J, Glöckner FO, Amann RI, Teeling H (2013) The genome of the alga-associated marine flavobacterium Formosa agariphila KMM 3901Treveals a broad potential for degradation of algal polysaccharides. Appl Environ Microbiol 79:6813–6822
Marçais G, Kingsford C (2011) A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27:764–770. https://academic.oup.com/bioinformatics/article-lookup/doi/10.1093/bioinformatics/btr011. Accessed 29 Apr 2018
Marchant J, Wren B, Ketley J (2002) Exploiting genome sequence: predictions for mechanisms of campylobacter chemotaxis. Trends Microbiol 10:155–159
Mende DR, Sunagawa S, Zeller G, Bork P (2013) Accurate and universal delineation of prokaryotic species. Nat Methods 10:881–884
Mendoza A, Leija A, Martínez-Romero E, Hernández G, Mora J (1995) The enhancement of ammonium assimilation in Rhizobium etli prevents nodulation of Phaseolus vulgaris. Mol Plant-Microbe Interact 8:584–592. http://www.ncbi.nlm.nih.gov/pubmed/8589413. Accessed 20 Oct 2017
Michel G, Chantalat L, Duee E, Barbeyron T, Henrissat B, Kloareg B, Dideberg O, Teissier PG, Cedex R (2001) The k-carrageenase of P. carrageenovora features a tunnel-shaped active site: a novel insight in the evolution of clan-B glycoside hydrolases. Structure 9:513–525
Michel G, Nyval-Collen P, Barbeyron T, Czjzek M, Helbert W (2006) Bioconversion of red seaweed galactans: a focus on bacterial agarases and carrageenases. Appl Microbiol Biotechnol 71:23–33. http://www.ncbi.nlm.nih.gov/pubmed/16550377. Accessed 2 Nov 2014
Miyazaki M, Nogi Y, Ohta Y, Hatada Y, Fujiwara Y, Ito S, Horikoshi K (2008) Microbulbifer agarilyticus sp. nov. and Microbulbifer thermotolerans sp. nov., agar- degrading bacteria isolated from deep-sea sediment. Int J Syst Evol Microbiol 58:1128–1133
Mizuno T (1997) Compilation of all genes encoding two-component phosphotransfer signal transducers in the genome of Escherichia coli. DNA Res 4:161–168
Mizuno T, Kato M, Jo YL, Mizushima S (1988) Interaction of OmpR, a positive regulator, with the osmoregulated ompC and ompF genes of Escherichia coli. Studies with wild-type and mutant OmpR proteins. J Biol Chem 263:1008–1012. http://www.ncbi.nlm.nih.gov/pubmed/3275653. Accessed 23 Oct 2017
Mou H, Jiang X, Liu Z, Guan H (2004) Structural analysis of kappa-carrageenan oligosaccharides released by carrageenase from marine Cytophaga MCA-2. J Food Biochem 28:245–260
Nivaskumar M, Francetic O (2014) Type II secretion system: a magic beanstalk or a protein escalator. Biochim Biophys Acta Mol Cell Res 1843:1568–1577
Nucifora G, Chu L, Silver S, Misra TK (1989) Mercury operon regulation by the merR gene of the organomercurial resistance system of plasmid pDU1358. J Bacteriol 171:4241–4247
Ortet P, Whitworth DE, Santaella C, Achouak W, Barakat M (2015) P2CS: updates of the prokaryotic two-component systems database. Nucleic Acids Res 43:D536–D541
Patriarca EJ, Riccio A, Colonna-Romano S, Defez R, Iaccarino M (1994) DNA binding activity of NtrC from Rhizobium grown on different nitrogen sources. FEBS Lett 354:89–92
Petersen TN, Brunak S, von Heijne G, Nielsen H (2011) SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods 8:785–786
Phillippy AM, Schatz MC, Pop M (2008) Genome assembly forensics: finding the elusive mis-assembly. Genome Biol 9:1–13. http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=2397507&tool=pmcentrez&rendertype=abstract. Accessed 31 Jan 2013
Potin P, Sanseau A, Gall YLE, Rochas C, Kloareg B (1991) Purification and characterization of a new k-carrageenase from a marine Cytophaga-like bacterium. Eur J Biochem 247:241–247
Rhein-Knudsen N, Ale MT, Meyer AS (2015) Seaweed hydrocolloid production: an update on enzyme assisted extraction and modification technologies. Mar Drugs 13:3340–3359
Rice P, Longden I, Bleasby A (2000) EMBOSS: the European Molecular Biology Open Software Suite. Trends Genet 16:276–277
Richter M, Rosselló-Móra R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci U S A 106:19126–19131
Roller B, Stoddard S, Schmidt T (2016) Exploiting rRNA operon copy number to investigate bacterial reproductive strategies. Nat Microbiol 1–7:16160
Sahoo S, Das S (2014) Analyzing gene expression and codon usage Bias in diverse genomes using a variety of models. Curr Bioinforma 9:102–112. http://www.eurekaselect.com/openurl/content.php?genre=article&issn=1574-8936&volume=9&issue=2&spage=102. Accessed 30 May 2018
Salzberg SL, Delcher a L, Kasif S, White O (1998) Microbial gene identification using interpolated Markov models. Nucleic Acids Res 26:544–548
Schmieder R, Edwards R (2011) Quality control and preprocessing of metagenomic datasets. Bioinformatics 27:863–864
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30:2068–2069. http://bioinformatics.oxfordjournals.org/content/30/14/2068. Accessed 10 Jul 2014
Shoseyov O, Shani Z, Levy I (2006) Carbohydrate binding modules: biochemical properties and novel applications. Microbiol Mol Biol Rev 70:283–295
Slater H, Alvarez-Morales A, Barber CE, Daniels MJ, Dow JM (2000) A two-component system involving an HD-GYP domain protein links cell-cell signalling to pathogenicity gene expression in Xanthomonas campestris. Mol Microbiol 38:986–1003. http://www.ncbi.nlm.nih.gov/pubmed/11123673. Accessed 2017 Oct 20
Smith J, Mountfort D, Falshaw R (2005) A zymogram method for detecting carrageenase activity. Anal Biochem 347:336–338. http://www.ncbi.nlm.nih.gov/pubmed/16259937. Accessed 2 Nov 2014
Sohn J, Nam J (2018) The present and future of de novo whole-genome assembly. Brief Bioinform 19:23–24
Spring S, Scheuner C, Göker M, Klenk H-P (2015) A taxonomic framework for emerging groups of ecologically important marine gammaproteobacteria based on the reconstruction of evolutionary relationships using genome-scale data. Front Microbiol 6:1–17
Stephens BB, Loar SN, Alexandre G (2006) Role of CheB and CheR in the complex chemotactic and aerotactic pathway of Azospirillum brasilense. J Bacteriol 188:4759–4768
Sun F, Ma Y, Wang Y, Liu Q (2010) Purification and characterization of novel κ-carrageenase from marine Tamlana sp. HC4. Chin J Oceanol Limnol 28:1139–1145. http://springerlink.bibliotecabuap.elogim.com/10.1007/s00343-010-9012-7. Accessed 2 Nov 2014
Tayco C, Tablizo F, Regalia R, Lluisma A (2013) Characterization of a κ-Carrageenase-producing marine bacterium, isolate ALAB-001. Philipp J Sci 142:45–54
The UniProt Consortium (2014) UniProt: a hub for protein information. Nucleic Acids Res 43:D204–D212. http://nar.oxfordjournals.org/content/43/D1/D204. Accessed 29 Oct 2014
Tischler AD, Camilli A (2004) Cyclic diguanylate (c-di-GMP) regulates Vibrio cholerae biofilm formation. Mol Microbiol 53:857–869
Trudeau K, Ward M, Zusman DR (1996) Identification and characterization of FrzZ, a novel response regulator necessary for swarming and fruiting-body formation in Myxococcus xanthus. Mol Microbiol 20:645–655
Tseng T-T, Tyler BM, Setubal JC (2009a) Protein secretion systems in bacterial-host associations, and their description in the Gene Ontology. BMC Microbiol 9(Suppl 1):S2
Tseng T-T, Tyler BM, Setubal JC (2009b) Protein secretion systems in bacterial-host associations, and their description in the Gene Ontology. BMC Microbiol 9(Suppl 1):S2. http://www.ncbi.nlm.nih.gov/pubmed/19278550. Accessed 29 Oct 2014
Uddin A (2017) Codon usage bias: a tool for understanding molecular evolution. J Proteomics Bioinform 10:1000e32
Urios L, Intertaglia L, Lesongeur F, Lebaron P (2011) Eionea nigra gen. nov., sp. nov., a gammaproteobacterium from the Mediterranean Sea. Int J Syst Evol Microbiol 61:1677–1681. http://www.ncbi.nlm.nih.gov/pubmed/20729310. Accessed 21 Aug 2017
Venturi V, Otten M, Korse V, Brouwer B, Leong J, Weisbeek P (1995) Alginate regulatory and biosynthetic gene homologs in Pseudomonas putida WCS358: correlation with the siderophore regulatory gene pfrA. Gene 155:83–88. http://www.ncbi.nlm.nih.gov/pubmed/7698672. Accessed 2017 Oct 20
Wang W, Zhang P, Yu G-L, Li C-X, Hao C, Qi X, Zhang L-J, Guan H-S (2012) Preparation and anti-influenza A virus activity of κ-carrageenan oligosaccharide and its sulphated derivatives. Food Chem 133:880–888. http://linkinghub.elsevier.com/retrieve/pii/S0308814612001604. Accessed 13 Nov 2014
Wang Y, Coleman-Derr D, Chen G, Gu YQ (2015) OrthoVenn: a web server for genome wide comparison and annotation of orthologous clusters across multiple species. Nucleic Acids Res 43:W78–W84. https://oup.silverchair-cdn.com/oup/backfile/Content_public/Journal/nar/43/W1/10.1093_nar_gkv487/2/gkv487.pdf?Expires=1493294814&Signature=IHz9Ch1yvjV4cpE4iUuLAqC1p6cKj9HqJQKMkCrz3MEBnkdFEPwl2JPuWhEbHP0gDbpTV-edCGGniATALrwxGMKebLywzH9NrzotOJHIU9z4QL99FQBE. Accessed 23 Apr 2017
Weiner RM, Taylor LE, Henrissat B, Hauser L, Land M, Coutinho PM, Rancurel C, Saunders EH, Longmire AG, Zhang H, Bayer EA, Gilbert HJ, Larimer F, Zhulin IB, Ekborg NA, Lamed R, Richardson PM, Borovok I, Hutcheson S (2008) Complete genome sequence of the complex carbohydrate-degrading marine bacterium, Saccharophagus degradans strain 2-40T. PLoS Genet 4:1–13
Williams R, Whitworth D (2010) The genetic organization of prokaryotic two-component system signalling pathways. BMC Genomics 11:720
Wu S, Chai W, Lin J (1999) General nitrogen regulation of nitrate assimilation regulatory gene nasR expression in Klebsiella oxytoca M5al general nitrogen regulation of nitrate assimilation regulatory gene nasR expression in Klebsiella oxytoca M5al. J Bacteriol 181:7274–7284
Yamagata A, Milgotina E, Scanlon K, Craig L, Tainer JA, Donnenberg MS (2012) Structure of an essential type IV pilus biogenesis protein provides insights into pilus and type II secretion systems. J Mol Biol 419:110–124
Yang JC, Madupu R, Durkin AS, Ekborg NA, Pedamallu CS, Hostetler JB, Radune D, Toms BS, Henrissat B, Coutinho PM, Schwarz S, Field L, Trindade-Silva AE, Soares CAG, Elshahawi S, Hanora A, Schmidt EW, Haygood MG, Posfai J, Benner J, Madinger C, Nove J, Anton B, Chaudhary K, Foster J, Holman A, Kumar S, Lessard PA, Luyten YA, Slatko B, Wood N, Wu B, Teplitski M, Mougous JD, Ward N, Eisen JA, Badger JH, Distel DL (2009) The complete genome of Teredinibacter turnerae T7901: an intracellular endosymbiont of marine wood-boring bivalves (shipworms). PLoS One 4:1–17
Yao J, Allen C (2006) Chemotaxis is required for virulence and competitive fitness of the bacterial wilt pathogen Ralstonia solanacearum. J Bacteriol 188:3697–3708
Yao R, Burr DH, Guerry P (1997) CheY-mediated modulation of Campylobacter jejuni virulence. Mol Microbiol 23:1021–1031. http://www.ncbi.nlm.nih.gov/pubmed/9076738. Accessed 2017 Oct 20
Yao Z, Wang F, Gao Z, Jin L, Wu H (2013) Characterization of a κ-carrageenase from marine Cellulophaga lytica strain N5-2 and analysis of its degradation products. Int J Mol Sci 14:24592–24602
Yin Y, Mao X, Yang J, Chen X, Mao F, Xu Y (2012) DbCAN: a web resource for automated carbohydrate-active enzyme annotation. Nucleic Acids Res 40:W445–W451
Yuan H, Zhang W, Li X, Lü X, Li N, Gao X, Song J (2005) Preparation and in vitro antioxidant activity of kappa-carrageenan oligosaccharides and their oversulfated, acetylated and phosphorylated derivatives. Carbohydr Res 340:685–692
Yuan H, Song J, Li X, Li N, Dai J (2006a) Immunomodulation and antitumor activity of kappa-carrageenan oligosaccharides. Cancer Lett 243:228–234. http://www.ncbi.nlm.nih.gov/pubmed/16410037. Accessed 21 Nov 2014
Yuan H, Song J, Zhang W, Li X, Li N, Gao X (2006b) Antioxidant activity and cytoprotective effect of kappa-carrageenan oligosaccharides and their different derivatives. Bioorg Med Chem Lett 16:1329–1334. http://www.ncbi.nlm.nih.gov/pubmed/16337378. Accessed 21 Nov 2014
Zhou M, Ma J, Li J, Ye H, Huang K, Zhao X (2008) A κ-carrageenase from a newly isolated pseudoalteromonas-like bacterium, WZUC10. Biotechnol Bioprocess Eng 13:545–551. http://springerlink.bibliotecabuap.elogim.com/10.1007/s12257-007-0200-z. Accessed 2 Nov 2014
Zhou Z, Dang Y, Zhou M, Li L, Yu C-H, Fu J, Chen S, Liu Y (2016) Codon usage is an important determinant of gene expression levels largely through its effects on transcription. Proc Natl Acad Sci U S A 113:E6117–E6125. http://www.ncbi.nlm.nih.gov/pubmed/27671647.Accessed 29 May 2018
Zhu B, Ning L (2016) Purification and characterization of a new k-carrageenase from the marine Bacterium Vibrio sp. NJ-2. J Microb Biotechnol 26:255–262
Ziayoddin M, Lalitha J, Shinde M (2014) Optimization of Agrase production by alkaline Pseudomonas aeruginosa ZSL-2 using Taguchi experimental design. ILNS 12:194–207
Funding
This study was funded by a research grant of the Philippine Department of Science and Technology (DOST)—Philippine Council for Agriculture, Aquatic and Natural Resources Research and Development (PCAARRD) to AOL.
Author information
Authors and Affiliations
Contributions
DATB performed the genomics and bioinformatics experiments and prepared the manuscript. AMED isolated the bacterium, performed microbial characterization, and assisted in the preparation of the manuscript. AOL conceptualized the project, assisted in the analysis of data and in the preparation of the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Electronic Supplementary Material
ESM 1
(DOCX 210 kb)
Rights and permissions
About this article
Cite this article
Boncan, D.A.T., David, A.M.E. & Lluisma, A.O. A CAZyme-Rich Genome of a Taxonomically Novel Rhodophyte-Associated Carrageenolytic Marine Bacterium. Mar Biotechnol 20, 685–705 (2018). https://doi.org/10.1007/s10126-018-9840-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10126-018-9840-6