Abstract
Metarhizium spp. have emerged as an alternative to chemical pesticides for protecting crops from insect pest. Here, we investigated midgut microbial community and metabolites of Spodoptera litura at three different timepoints after infection with Metarhizium flavoviride. The innate immune system of S. litura was activated with levels of polyphenol oxidase, carboxylesterase, multifunctional oxidase, and glutathione S-transferase activity significantly increasing. Exposure to the fungal pathogen also altered bacterial abundance and diversity in host’s midgut, and these changes varied depending on the time elapsed since exposure. We identified more operational taxonomic units in the treated samples as compared to the control samples at all tested time points. A total of 372 metabolites were identified, and 88, 149, and 142 differentially accumulated metabolites (DAMs) were identified between the treatment and control groups at 3 timepoints after treatment, respectively. Based on the changes of DAMs in response to M. flavoviride infection at different timepoints and significantly enriched KEGG pathways, we speculated that “tyrosine metabolism,” “galactose metabolism,” “ATP-binding cassette transporters,” “neuroactive ligand-receptor interaction,” “purine metabolism,” “arginine and proline metabolism,” “beta-alanine metabolism,” “lysosome,” and “carbon metabolism” may participate in the metabolic-level defense response. An integrated pathway-level analysis of the 16S-rDNA and metabolomic data illustrated the connections and interdependencies between the metabolic responses of S. litura and the midgut microorganisms to M. flavoviride infection. This work emphasizes the value of integrated analyses of insect-pathogen interactions, provides a framework for future studies of critical microorganisms and metabolic determinants of these interactions, establishes a theoretical basis for the sustainable use of M. flavoviride.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Insect intestinal environments are symbiotic systems containing various microorganisms, including bacteria, fungi, viruses, and endophytic fungi (Hussa and Goodrich-Blair 2013). These microorganisms could have a mutually beneficial relationship with their insect host (Stouthamer et al. 1999; Hurst and Jiggins 2000; Oliver et al. 2003; Baumann 2005; Wu et al. 2006; Popa et al. 2012; Mercado et al. 2014) by providing a sanctuary for the microorganisms, which may in turn act as regulators of various life activities of the host (Shi et al. 2010). The symbiotic microbiome affects host physiology and evolution by participating in metabolic processes related to the anabolism of sterols, sugars, amino acids, digestive enzymes, and detoxifying enzymes (Crotti et al. 2010). Symbiotic microbial communities also help the host to detoxify harmful foreign substances and prevent the invasion of predators, parasites, and pathogens (Engel and Moran 2013; Kaltenpoth and Engl 2014). For example, the symbiotic bacteria of Acyrthosiphon pisum defend the insect host against pathogens, compensating for the host’s lack of relevant immune genes (Gerardo et al. 2010). Similarly, the infection rate of Metarhizium flavoviride ARSEF1764 in populations lacking Arsenophonus was significantly higher than in the carrying population (Chen 2013). In addition, gut fluids from monoassociated insects inhibited the germination of Metarhizium anisopliae conidia, while gut fluids from germ-free desert locusts (Schistocerca gregaria) did not (Dillon and Charnley 1995). However, gut populations of Wolbachia did not improve the resistance of Drosophila melanogaster to pathogenic bacteria; Wolbachia abundances associated with expression of key detoxifying genes, and correlated with insecticide susceptibility levels of Nilaparvata lugens (Zhang et al. 2021). Finally, symbiotic bacteria in Dendroctonus frontalis synthesize anti-pathogen substances, upregulate genes in specific defense pathways, and reduce the sensitivity of the host to pathogens (Oh et al. 2009). In insects, antioxidant enzyme activity reflects metabolic and stress-resistance activity; antioxidant enzyme activity is also an important index of the host’s pathological state (Ouedraogo et al. 2003; Boguś et al. 2007). Internal protective enzymes, such as superoxide dismutase (SOD), catalase (CAT), and other antioxidant enzymes, are activated by exogenous stress and could restore free-radical homeostasis and protect the host from threats (Liochev and Fridovich 2007).
In a recent revision of Metarhizium, Kepler (2014) expanded the definition of this genus to include most species previously assigned to the Metacordyceps, as well as the green-spored Nomuraea species and species in the more recently described genus Chamaeleomyces. Metarhizium spp. penetrate the insect cuticle and exoskeleton, consume host nutrients in the hemolymph, interfere with the host metabolism, and secrete destruxin toxins, which are insecticidal secondary metabolites isolated from Metarhizium spp. that alter cell morphology and destroy tissues, even at low doses (Fan et al. 2013). Therefore, Metarhizium spp. exert a remarkable effect on the pest host (Douglas 1998). Spodoptera litura is distributed worldwide and can severely damage more than 290 plant species belonging to 99 families (Xue et al. 2010; Koo et al. 2018). While most of the insect pests targeting various crops have previously been managed using pesticides, long-term pesticide use not only pollutes the environment and negatively affects human health, but also leads to the development of pesticide resistance. Metarhizium spp. have emerged as an alternative to chemical pesticides for protecting crops from S. litura. The infection rate of Metarhizium spp. is relatively slow and spends long time effecting, which are usually showing high selectivity against target pests, non-toxic to warm blood animals and have very fast degradation time in the air and light (Noor et al. 2019). Since host invasion by a fungus can be accompanied by changes in the host microbiome (Kikuchia et al. 2012), it is important to understand the effects of parasitization by M. flavoviride on S. litura, so that the use of M. flavoviride as a microbial control agent can be optimized. When penetrating the cuticle of S. litura, M. flavoviride causes trauma to the host and competes with the host for nutrients (Sánchez-Rodríguez et al. 2016). Hosts provide nutrition to symbiotes, which then use metabolic wastes to synthesize nutrients needed by the host, thus participating in the host metabolism (Douglas et al. 2001). Symbiotic microorganisms and exogenous pathogens compete for nutrients, and the symbiotic microorganisms could produce toxins to eliminate these competing pathogens (Kikuchia et al. 2012). Therefore, qualitative and quantitative analyses of the metabolites produced by the insect host under pathogen stress are needed to study the defense response of the insect metabolome to exogenous threats and identify useful target metabolites. Insect feces contain information about the host, the host’s intestinal microorganisms, and interactions among these microorganisms. These data could help to further clarify metabolic changes in the host and in the intestinal flora. Unlike other methods, fecal analyses are non-invasive and reveal changes in the complete metabolic profiles of the flora in response to different threats (Li et al. 2021).
Microbiomics and metabolomics are suitable choices for studying microbial community structure and metabolite composition since these techniques enable the in-depth examination of functional differences among microbial communities (Zhu et al. 2018). In this study, microbiome and metabolome analyses were used to explore the effects of M. flavoviride exposure on the midgut microbial community and metabolites of S. litura.
Materials and methods
Materials
S. litura populations that had not been exposed to any insecticide were collected from lotus plants (Nelumbo nucifera) near Wuhan, Hubei province, China, and were reared for one generation on lotus leaves (var. Taikong) in our laboratory under the following conditions: air temperature, 28 ± 2 °C; relative humidity (RH), 85%; and photoperiod, 14 h light:10 h dark.
M. flavoviride qc1401 cultures provided by the Institute of Subtropical Forestry, Chinese Academy of Forestry (Hangzhou, China), were grown on Petri plates (Diameter = 90 mm) containing potato dextrose agar (PDA) at 28 ± 2 °C and 93–98% RH for 10 days, after which the cultures had fully sporulated. Conidia were scraped into 10 mL of 0.05% Tween-80 to create a stock suspension, vortexed for 30 min, and then diluted to determine the solution concentration using a hemocytometer (XB. K. 25). The stock suspension was adjusted with 0.05% Tween-80 to create treatment concentrations of 108 conidia/ml. The control suspension contained 0.05% Tween-80 only.
Exposure of S. litura to M. flavoviride
Six sterile 200-mm plastic Petri dishes containing 500 s instar larvae of S. litura each (three replicate control and experimental plates) were uniformly sprayed with 10 mL of either the stock or control suspension using a hand-held sprayer (Choi et al. 2020). The larvae were held at 28 ± 2 °C, 60–85% RH, and a 14 h light:10 h dark cycle without diet for 24 h, to allow germination of conidia then the insects were transferred to new dishes with diet to follow the course of the infection. Larvae were sampled from each plate at 24 h, 48 h, and 96 h after spraying, and the resulting replicate samples were denoted as follows: Ck.24 h, Ck.48 h, and Ck.96 h, respectively, from the control plates. MF.24 h, MF.48 h, and MF.96 h, respectively, from the experimental plates. Insect feces were collected from each plate at 24 h, 48 h, and 96 h, and the resulting replicate samples were denoted as follows: Ck_24h, Ck_48h, and Ck_96h, respectively, from the control plates. MF_24h, MF_48h, and MF_96h, respectively, from the experimental plates.
Scanning electron microscopy
At 96 h after spraying, experimental and control S. litura larvae were fixed in 2.5% glutaraldehyde for 12 h at 4 °C. The fixed samples were rinsed four times with phosphate-buffered saline (PBS, 0.1 M, pH 7.2) and dehydrated using a graded ethanol series (30%, 50%, 70%, 80%, 90%, and 100%, v/v). After dehydration, the samples were dried with carbon dioxide in a critical point-dryer, mounted on stubs, coated with gold, and examined under a scanning electron microscope (JEOL JSM-6360 LV, Hitachi, Japan) at 15 kV. The colonization of S. litura by M. flavoviride was investigated using SEM to provide information about the colonization area and mode of the pathogenicity.
Enzyme activity assays
At 24 h, 48 h, and 96 h after spraying, 20 S. litura larvae were removed from each plate, ground whole bodies in liquid nitrogen., and 1 mL of sodium phosphate buffer (0.1 mol/L) was added, the pH values of the polyphenol oxidase (PPO), carboxylesterase (CarE), glutathione S-transferase (GST), and multifunctional oxidase (MFO) buffers were 7.8, 7.5, 7.0, and 7.8, respectively. After rapid vortexing and shaking, each mixture was centrifuged at 4 °C and 12,000 rpm for 15 min. Levels of PPO, CarE, MFO, and GST activity in the supernatant were measured using the corresponding assay kits (Suzhou Grace Biotechnology Co., Ltd., China) and following the manufacturer’s instructions. Absorbance was assessed using a microplate reader (SpectraMax 190, Molecular Devices, USA).
16S rRNA gene sequencing
Extraction of bacterial DNA
At 24 h, 48 h, and 96 h after spraying, 50 S. litura larvae were collected from each plate and dissected to obtain the midgut region of each larva. The collected midguts were rinsed with sterile water and stored at − 20 °C for DNA extraction. DNA was extracted using a bacterial DNA extraction kit (Omega Bio-Tek, USA) and dissolved in 200 μL of ddH2O. DNA concentration and purity were measured using 1% agarose gel electrophoresis and a nanodrop 1000 spectrophotometer (Saveen Werner ApS, Denmark), respectively. Extracted DNA was diluted to a concentration of 1 ng/μl and stored at − 20 °C. The diluted DNA was used as a template for the amplification of bacterial 16S rRNA genes using polymerase chain reactions (PCRs) with barcoded primers and Takara Ex Taq (Takara Biomedical Technology (Beijing) Co., Ltd.). To analyze bacterial diversity, the V3–V4 variable regions of the 16S rRNA sequence were amplified using the universal primers 343F (5′-TACGGRAGGCAGCAG-3′) and 798R (5′-AGGGTATCTAATCCT-3′).
Library construction
Amplicon quality was visualized using gel electrophoresis. Amplicons were purified using AMPure XP beads (Beckman Coulter) and re-amplified using PCR. After re-purification with AMPure XP beads, the final amplicons were quantified using Qubit dsDNA assay kits (Thermo Fisher). The purified amplicons were pooled in equal amounts for subsequent sequencing.
Bioinformatic analysis
The raw paired-end sequencing reads (in FASTAQ format) were preprocessed using Trimmomatic (Bolger et al. 2014) to detect and trim ambiguous bases (N) and to remove low-quality sequences (average quality score < 20) with a sliding-window trimming approach. After trimming, the paired-end reads were assembled using FLASH with the following parameters: minimum overlap, 10 bp; maximum overlap, 200 bp; and maximum mismatch rate, 20%. Sequences were further denoised by removing reads shorter than 200 bp, as well as ambiguous or homologous sequences. Reads where 75% of the bases had a quality score > 20 were retained. Then, reads with chimeras were detected and removed using QIIME (version 1.8.0) (Caporaso et al. 2010). After primer sequence removal, clean reads were clustered into operational taxonomic units (OTUs) using Vsearch with a 97% similarity cut-off (Edgar et al. 2011). The representative sequence for each OTU was selected using QIIME. All representative sequences were annotated and blasted against the Silva database using the RDP classifier with a confidence threshold of 70% (Wang et al. 2007). QIIME was used for alpha diversity analysis (Caporaso et al. 2010). The microbial diversity in samples was estimated using the alpha diversity that include Chao1 index and Shannon index. The unweighted Unifrac distance matrix performed by R package was used to analyze the significant differences between different groups using ANOVA/Kruskal–Wallis statistical test.
Metabolomic analysis
Gas chromatography-mass spectrometry
Fecal matter was collected from both the experimental and the control plates at 24 h, 48 h, and 96 h after spraying. We then transferred 60 mg of accurately weighed fecal matter from each plate to separate 1.5-mL Eppendorf tubes. Two small steel balls were added to each tube, along with 20 μL internal standard (2-chloro-l-phenylalanine in methanol, 0.3 mg/mL) and 600 μL of methanol/water extraction solvent (4/1, v/v). Samples were rapidly frozen at − 80 °C for 2 min and then ground to a powder at 60 HZ for 2 min using a grinder (Shanghai Jinxin Experimental Technology, China). We added 120 μL of chloroform to each sample, followed by vigorous vortexing. The vortexed samples were extracted at ambient temperature using an ultrasound (Shanghai Hannuo Experimental Technology, China) and then transferred to 4 °C for 10 min. The samples were then centrifuged at 12,000 rpm for 10 min at 4 °C. We then transferred 300 µL of each supernatant to a glass sampling vial; the quality-control sample was prepared by pooling equivalent aliquots of the supernatants of all samples. All samples were vacuum-dried at room temperature. After drying, 80 μL of methoxyamine hydrochloride (dissolved in pyridine, 15 mg/mL) was added to each sample, and the resulting mixture was vortexed vigorously for 2 min and incubated at 37 °C for 90 min. After incubation, 80 μL of N,O-bis (trimethylsilane) trifluoroacetamide (with 1% trimethylchlorosilane) and 20 μL n-hexane were added to each mixture. The mixtures were vortexed vigorously for 2 min and then derivatized at 70 °C for 60 min. The samples were incubated at ambient temperature for 30 min before GC–MS analysis, which was performed using an Agilent 7890A-5975C GC–MS system (Agilent, USA).
Raw MS data analysis
We used the Analysis Base File Converter (v2.74) to convert the raw MS data (in D format) to abf format. The abf files were imported into MD-DIAL for processing as follows: first, metabolites were annotated against the LUG database, which is an untargeted GC–MS database (Lumingbio, China). Second, data were aligned using the Statistic Compare module and converted into a text file containing a three-dimensional data array that included sample information, peak names (or retention time and m/z), and peak intensities. All internal standards and any known pseudo-positive peaks (caused by background noise, column bleed, or the BSTFA derivatization procedure) were removed from the data array. After interior labels with RSD values > 0.3 had been deleted, the strength (or area) of each peak was calculated by normalizing the multiple interior labels based on retention time and partition period.
Multivariate analysis
Data were log10 transformed, and the resulting data matrix was analyzed using the ropls R package as follows: after mean centering and unit variance scaling, principal component analysis (PCA), and (orthogonal) partial least-squares-discriminant analysis [(O)PLS-DA] were performed to visualize the metabolic differences between experimental groups. For both analyses, Hotelling’s T2 region, which is shown as an ellipse in score plots of the models and which defines the 95% confidence interval of the modeled variation, was calculated. We discriminated among groups using the variable importance in the projection (VIP) method, which ranks the overall contribution of each variable to the OPLS-DA model; variables with VIP > 1 are considered relevant for group discrimination. We used the default 7-round cross-validation, in which 1/7th of the samples were excluded from the mathematical model in each round to guard against overfitting. Metabolites that significantly differed between the experimental and control groups were identified based on both the VIP value in the OPLS-DA model and the p value of the difference between the normalized peak areas of the two groups, as calculated using two-tailed Student’s t tests. Metabolites with VIP value > 1.0 and p value < 0.05 were considered significantly different between the experimental and control groups.
Identification of differentially accumulated metabolites
Both multivariate and univariate statistical significance thresholds (VIP > 1.0 and P < 0.05) were used to identify metabolites that significantly differed with respect to accumulation between the groups. Metabolites were identified and annotated with ChromaTOF using several databases, including the NIST 11 standard MS databases, the Fiehn databases, and databases of reference standards available in our laboratory; metabolites were considered identified when similarity between the obtained mass spectrum and the spectrum of the reference standard was at least 70%.
Integration of the metabolomic and microbiomic (16S rDNA) data
The differentially accumulated metabolites and OTUs (P < 0.05) were selected and combined with the metabolite response data and the relative abundances of the OTUs. Then, Pearson correlations were used to determine the relationship between metabolite abundance and OTU abundance.
Results
Infection of S. litura by M. flavoviride conidia
Both spores and mycelia of M. flavoviride were present on the S. litura epidermis after treatment with the M. flavoviride suspension. The invasive behavior of M. flavoviride differed on different areas of the host epidermis. Infection always began on the host abdomen, possibly because the folds in the epidermis on the abdomen made invasion easier. After colonization of the abdomen, the fungus slowly spread to the folds and depressions on the host’s back, gradually infected the smooth part of the back, and surrounded the host body with mycelium (Fig. 1).
Invasion behavior of Metarhizium flavoviride on the epidermis of the host (Spodoptera litura). A S. litura 96 h after treatment with 10-mL 0.05% Tween 80 (control). B S. litura 96 h after treatment with 10-mL M. flavoviride (108 conidia/ml). Arrow indicates M. flavoviride mycelia. C Scanning electron microscope (SEM) image showing the cuticle of S. litura 96 h after treatment with 10-mL 0.05% Tween 80 (control). D SEM image showing the cuticle of S. litura 96 h after treatment with 10-mL M. flavoviride (108 conidia/mL). Arrow indicates M. flavoviride conidia
Enzyme activity in infected and uninfected S. litura
PPO and GST activity levels in the treatment groups were significantly higher than those in the control groups at 24, 48, and 96 h after spraying (Fig. 2A, 2B; TableS1, S2). CarE activity levels were also significantly greater in the treatment groups compared to the control groups at 24 and 48 h after spraying; there was no significant difference in CarE activity between the control and treatment groups at 96 h (Fig. 2C; Table S3). MFO activity levels in the treatment groups were significantly higher than that in the control group at 96 h after spraying only; at 24 and 48 h, there were no significant differences in MFO activity levels between the control and treatment groups (Fig. 2D; Table S4).
The activity levels of four enzymes in the treatment and control Spodoptera litura groups. Data in the figure are means ± SD; “*” and “**” above the bars indicate significant differences (P < 0.05) and extremely significant differences (P < 0.01), respectively, in enzyme activity levels between the two groups (Student’s t test). CK, control (suspension containing 0.05% Tween-80 only); MF, treatment (suspension containing 108 M. flavoviride conidia/ml). PPO, polyphenol oxidase; CarE, carboxylesterase; GST, glutathione S-transferase; MFO, multifunctional oxidase
16S rDNA sequencing results
Gut bacterial diversity
Across all midgut samples, Good’s coverage index was 0.99 (Table 1), indicating that the sequencing results were representative of the bacterial population. In general, Chao1 indices were greater for the treatment groups than the control groups, while the Shannon index varied little among groups (Table 1). The Shannon index fluctuated among groups, with the lowest value calculated for the CK.48 h group (4.95 ± 1.09) and the highest value calculated for the CK.24 h group (6.96 ± 0.94, Table 1). The observed species index was lowest for the CK.48 h group, followed by the CK.96 h group; this index was highest for the MF.96 h group (1538.88 ± 192.06).
Bacterial OTUs and taxonomic analysis
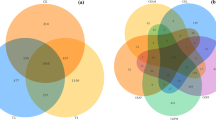
We identified 1136, 594, 834, 1335, 1260, and 1444 bacterial OTUs in groups CK.24 h, CK.48 h, CK.96 h, MF.24 h, MF.48 h, and MF.96 h, respectively (Fig. 3A). The number of OTUs shared between CK.24 h and MF.24 h was 178; the number shared between CK.48 h and MF.48 h was 133, and the number shared between CK.96 h and MF.96 h was 187 (Fig. 3B). In total, 93 OTUs were shared across all six groups. More OTUs were identified in all three treatment groups than in the corresponding controls, suggesting that the diversity of the midgut microbial community of S. litura could increase in response to treatment with M. flavoviride.
Across all six groups, bacteria from more than 30 classes were taxonomically annotated. The Gammaproteobacteria, Bacteroidia, and Clostridia had the highest relative abundance. The relative abundance of Gammaproteobacteria in the control group at 48 h after spraying (CK.48 h) was 74.99%, which was greater than the relative abundance of this taxon in any of the other control and treatment groups (CK.24 h: 40.00%, CK.96 h: 63.71%, MF.24 h: 55.50%, MF.48 h: 52.36%, and MF.96 h: 43.54%).
The composition of bacterial species differed between the control and treatment groups at all time points examined (Fig. 4). However, of the 10 most common species in each group, six were shared between CK.24 h and MF.24 h; seven were shared between CK.48 h and MF.48 h, and eight were shared between CK.96 h and MF.96 h (Fig. 4). The total abundance of the 10 most abundant species compared to the total abundance of all species identified also differed between the control and treatment groups at each time point: the total relative abundance of the top 10 species was 6.23% and 11.17% in groups Ck.24 h and MF.24 h, respectively; 9.15% and 3.10% in groups Ck.48 h and MF.48 h, respectively; and 9.01% and 3.11% in Ck.96 h and MF.96 h, respectively. As such, the total relative abundance of the top 10 species in the treatment group sharply decreased between 48 and 96 h after spraying but increased in the control group. Sphingobacterium multivorum was the most abundant bacterial species in both the control and treatment groups: S. multivorum abundance was 2.75%, 7.56%, and 6.23% in the control groups at 24, 48, and 96 h after spraying, respectively, but was 5.02%, 0.76%, and 1.17%, respectively, in the treatment groups.
Metabolomic analysis
Overview of metabolomic data corresponding to S. litura response to M. flavovirid
After the removal of unvalued peaks and the combination of peaks from the same metabolite, 372 metabolites remained. This analysis recovered the biological replicates of each group in tight clusters in the same hemisphere of each graph, suggesting high reproducibility. The treatment and control groups were separated along the predictive component of the OPLS-DA plot at each tested time point (Fig. 5), indicating that metabolite profiles substantially differed between the treatment and control groups.
Orthogonal projections to latent structures-discriminant analysis (OPLS-DA) and permutation tests of the OPLS-DA results, showing differences in metabolites between the treatment and control groups at A 24 h, B 48 h, and C 96 h after treatment. CK, control (suspension containing 0.05% Tween-80 only); MF, treatment (suspension containing 10.8 M. flavoviride conidia/ml)
Differentially accumulated metabolites
The metabolites that were differentially accumulated between the feces of the control and treatment groups also differed among the three tested time points. We identified 88 differentially abundant metabolites between the experimental and control feces at 24 h after spraying, 149 differentially abundant metabolites between the experimental and control feces at 48 h after spraying, and 142 differentially accumulated metabolites between the experimental and control feces at 96 h after spraying (Fig. 6). At 24 h post-spraying, the differentially accumulated metabolites were primarily benzene and substituted derivatives, organooxygen compounds, carboxylic acids, and derivatives. Twelve differentially accumulated metabolites were carboxylic acids and derivatives (Fig. 6A). At 48 h post-spraying, 14 differentially accumulated metabolites were carboxylic acids and derivatives, while 16 differentially accumulated metabolites were organooxygen compounds (Fig. 6B). At 96 h post-spraying, the differentially accumulated metabolites were organic acids and derivatives, lipids and lipid-like molecules, benzenoids, organic oxygen compounds, phenylpropanoids, and polyketides; 32 differentially accumulated metabolites were organic acids and derivatives (Fig. 6C). The number of differentially accumulated metabolites between the control and treatment groups increased with the length of time after spraying. However, the species of metabolites differentially accumulated between the control and treatment groups were similar in the 24 h and 48 h treatment groups.
Heatmap of metabolites that differed between the feces of control Spodoptera litura and the feces of S. litura treated with Metarhizium flavoviride at different time. A 24 h after M. flavoviride treatment, B 48 h after M. flavoviride treatment, and C 96 h after M. flavoviride treatment, a tree showing the sample clustering pattern is shown above each heatmap, and metabolite groups are shown along the y-axis. Redder cells correspond to more accumulated metabolites, while bluer cells correspond to fewer accumulated metabolites. CK, control (suspension containing 0.05% Tween-80 only); MF, treatment (suspension containing 10.8 M. flavoviride conidia/ml)
Pathways enriched in the differentially abundant metabolites
KEGG metabolic pathway analysis indicated that differentially accumulated metabolites between the control and experimental groups at 24, 48, and 96 h post-spraying were enriched in 17, 18, and 26 metabolic pathways, respectively. The 10 pathways most significantly enriched in the differentially abundant metabolites were the tyrosine metabolism, the galactose metabolism, the ABC transporters, and neuroactive ligand-receptor interactions at 24 h post-spraying (P < 0.05); the purine metabolism, the ABC transporters, and neuroactive ligand–receptor interactions at 48-h post-spraying (P < 0.05); and the ABC transporters, the arginine and proline metabolism, the purine metabolism, the beta-alanine metabolism, lysosome, and the carbon metabolism at 96-h post-spraying (P < 0.05; Fig. 7).
The 10 KEGG pathways most significantly enriched in the differentially abundant metabolites at A 24-h post-spraying, B 48-h post-spraying, and C 96-h post-spraying. P values in the metabolic pathways correspond to the significance of the enrichment of that pathway. Red lines indicate a p value of 0.01, while blue lines indicate a p value of 0.05. A bar above the blue line indicates that the signaling pathway is significant
Combined metabolomic and 16S analysis
Across the three time points tested, there were no strong correlations among the 20 most differentially accumulated metabolites and the 20 most differentially abundant gut microbes (Fig. 8). For example, melezitose and galactose were among the 20 most differentially abundant metabolites both 24 h and 48 h post-spraying, but were correlated with different microorganisms at the two time points. That is, at 24 h post-spraying, melezitose was positively correlated with the microbes Bacteroides fragilis, uncultured Alistipes sp., uncultured gamma proteobacterium, and uncultured beta proteobacterium (Fig. 8A); at 48 h post-spraying, melezitose was negatively correlated with Weissella confusa and bacterium enrichment culture clone BBMC-5 (Fig. 8B). Two metabolites appeared at both 24 and 96 h post-spraying (catechin and glycerol-alpha-phosphate), while six metabolites were present at both 48 and 96 h post-spraying (UDP-glucuronic acid, oxamic acid, urea, 4',5-dihydroxy-7-glucosyloxyflavanone, methylamine, and 5-methyluridine). However, these shared differentially abundant metabolites were correlated with different microbes at each time point. S. multivorum, the most abundant bacterium in both the control and treatment groups (Fig. 6), was negatively correlated with most of the 20 most differentially abundant metabolites except for chenodeoxycholic acid, methylamine, and galactose at 48 h post-spraying. This bacterium was negatively correlated with all differential metabolites at 96 h post-spraying.
Discussion
The intestine possesses the largest mucosal surface in the insect body; the intestinal mucosa digests and absorbs nutrients into the haemocoel (Moor 2018; Haber 2017). Intestinal microorganisms participate in insect digestive processes by synthesizing digestive enzymes; producing mucins, hormones, and antimicrobial molecules; assisting in the degradation of toxic substances (Zheng et al. 2018), improving host defense and building a physical and chemical barrier that protects the host against pathogenic microbes. Metabolomic analyses found that the honeybee gut flora produces organic acids and metabolizes flavonoids and pollen exine (Kesnerova et al. 2017). In addition, honeybees degrade plant polymers with assistance of the intestinal flora, which also metabolize amino acids and vitamins for the host (Zheng et al. 2017). In D. melanogaster, previous studies have shown that several metabolic pathways (including amino acid metabolism, glycolysis, purine metabolism, pyrimidine metabolism, and lipid metabolism) changed after permethrin treatment, and tryptophan catabolism played an important role in the formation of host insecticide resistance (Brinzer et al. 2015). Many previous studies have shown that the intestinal microbiota can independently metabolize compounds in the intestine, we concluded that the host metabolic process would be altered after the disruption of the host intestinal microbiota during the stress-resistance response. This study explores changes in the microbial community structure of the midgut, metabolite abundance, and activated metabolic pathways in S. litura in response to M. flavoviride stress.
The insect immune defense system metabolizes foreign toxic substances. For example, multifunctional oxidases and glutathione transferases decompose fat-soluble and toxic pesticides before further metabolism occurs (Grant and Matsumura 1989). The metabolic resistance of insect results from the increased metabolic activity of these enzymes (Adeyi et al. 2015). CarE, a member of the esterase supergene family, enhances the metabolic activity of pesticides through the upregulation and point mutation of carboxylesterase genes. These genes are crucial in the metabolism of organophosphorus and carbamate pesticides (Zhang et al. 2010). In this study, CarE activity significantly increased in response to M. flavoviride treatment, suggesting that infection by pathogenic fungi activated a similar immune system response to that triggered by exposure to chemical insecticides. Polyphenol oxidase (PPO) could induce melanization under pathogen stress (Zhao et al. 2007; Cerenius et al. 2010), and the PPO cascade affects the resistance of the mosquito to Beauveria bassiana (Yassine et al. 2012; Ashida et al. 1990). PPO activity in the treatment group was significantly higher than in the control group at 24, 48, and 96 h after infection, indicating that PPO plays a role in the resistance of S. litura to M. flavoviride.
In this study, exposure to M. flavoviride altered the abundance and diversity of the bacterial microorganisms in the midgut of S. litura, and these changes differed with time post-exposure. In general, exposure to M. flavoviride increased bacterial abundance in the host midgut, but alpha diversity fluctuated across time points. Microbial symbiont assemblages with consistent relative abundance and diversity indicate a stable community or ecosystem (Glasl et al. 2016). The overall increases in bacterial abundance and diversity in the midgut of S. litura after M. flavoviride treatment suggested that M. flavoviride infection destabilized the microecological environment of the host. After treatment with M. flavoviride, bacterial species composition in the midgut of S. litura was altered, suggesting that this fungal infection affected bacterial community structure. In particular, the dominant bacterial species in the S. litura midgut shifted after treatment, with substantial decreases in the abundances of W. confusa and S. multivorum. Indeed, the relative total abundance of the 10 most abundant species in the midgut decreased significantly between 24 and 48 h, 96 h post-spraying. The high relative abundance of W. confuse at 24 h after spraying was likely associated with the host immune system response to pathogen stress, as this species colonizes the human and animal intestinal tract (Jeong et al. 2007; Harlan et al. 2011), plays a prebiotic role in regulating the balance of the intestinal flora (Zhang et al. 2011), inhibits Helicobacter pylori (Nam et al. 2002), and synthesizes exopolysaccharides with potential pharmacological properties, including anti-inflammatory, anti-tumor, and immunosuppressive effects (Wang et al. 2022). The further colonization of the pathogen modulated the host gut biome and which made the relative abundance of Weissella confuse decreased between 24 and 48, 96 h after spraying.
S. multivorum is associated with the degradation of pesticides, especially nicotine, imidacloprid, and pentachlorophenol (Ye et al. 2006). In Wuhan, where the S. litura population used in this study was collected, S. litura populations are 14–229-fold more resistant to organophosphates and 12–227-fold more resistant to pyrethroids compared with a susceptible Lab-BJ strain (Hong et al. 2013). Whether the high relative abundance of S. multivorum at 24 h after spraying is consistent with the ability of this bacterium to degrade pesticides needs to be further. However, with the further colonization of the host by the pathogen, the relative abundance of S. multivorum decreased, and S. multivorum was negatively correlated with almost all metabolites. ABC transporters use the energy generated by ATP hydrolysis to transport various bound substances, such as alkanes, amino acids, and antibiotics (Lane et al. 2016). ABC transporters play important physiological roles in prokaryotes as well as higher animals and plants (Kathawala et al. 2015; Ponte 2007). In this study, we found that the ABC transporter pathway was significantly enriched in metabolites that were differentially abundant in the S. litura population after M. flavoviride treatment, suggesting an increased nutrient demand in the part of the host, possibly for immune defense. In this study, as the time post-spraying increased, the number of differentially abundant metabolites between the control and treatment groups increased. Several metabolic pathways related to anabolism (e.g., the galactose metabolism, the purine metabolism, the beta-alanine metabolism, and the carbon metabolism) were also significant enriched in the differentially accumulated metabolites. Further study is necessary to identify significantly enriched metabolites associated with improvements in host immunity or resistance to M. flavoviride.
Neuroactive ligand receptors participate in the intracellular and extracellular signal pathways on the plasma membrane (Lauss et al. 2007). Several exogenous substances downregulate multiple receptor genes (e.g., chrm3, DRD5, and HTR1B) in these pathways, affecting the host’s physiological rhythms and endocrine system (Zhuang et al. 1999; Svob et al. 2008). The changes in the expression levels of related genes in response to pathogen stress might lead to the enrichment of neuroactive ligand receptors in the differentially abundant metabolites at 24 and 48 h after spraying.
The enrichment of the tyrosine metabolism pathway in the significantly differentially abundant metabolites in the feces at 48 h after spraying may not only have been related to host nutritional requirements, but could also berelated to host immune defense against M. flavoviride. The tyrosine metabolism pathway serves as a starting point for the production of various structurally diverse natural compounds, such as tocopherols, plastoquinone, ubiquinone, betalains, salidroside, and benzylisoquinoline alkaloids (Schenck and Maeda 2018). Some of these, including tyrosine-derived metabolites, tocopherols, plastoquinone, and ubiquinone, are essential for survival (Xu et al. 2019). Hydroxylation of tyrosine at the 3-position leads to the formation of 3,4-dihydroxy-l-phenylalanine (l-DOPA); this reaction can be catalyzed by the tyrosinase (EC1.14.18.1) activity of PPOs, which play important roles in insect development, wound healing, and immunity (Tran et al. 2012). In a previous study, recolonization of the gut biome by Klebsiella michiganensis removed Bactrocera dorsalis, significantly upregulated the arginine and proline metabolic pathway (10- to 13-fold), maintained the structure and function of mitochondria, and improved host resistance to stress (Muhammad et al. 2020). Thus, enrichment of the arginine and proline metabolism pathway in the significantly differentially abundant metabolites in S. litura feces at 96 h after spraying could have been related to the stress resistance function of this pathway. Further studies are needed to detect intestinal flora related to the upregulation of the arginine and proline metabolic pathway.
Lysosomes are essential intracellular digestive stations that are indispensable for energy and metabolic homeostasis (Safig and Klumperman 2009). Lysosomes are also involved in signal transduction, organelle recycling (Lawrence and Zoncu 2019), and other physiological and pathological processes (Ravikumar. 2010; Saxton and Sabatini 2017). Lysosome dysfunction may contribute to lysosomal storage diseases as well as the progression of certain types of neurodegenerative disorders (Platt et al. 2018; Zhu et al. 2021). The significant enrichment of the lysosomal metabolic pathway in the differentially accumulated metabolites in the S. litura gut at 96 h after spraying was probably also related to the response of host immune system to pathogenic fungus.
The results showed that the structure of the intestinal flora, the metabolic pathways, and the metabolites of S. litura were altered in response to M. flavoviride challenge. The evidence gathered in this study indicated that the ultimate purpose of such changes is to protect the host by competing for nutrients and even killing invaders. The using of M. flavoviride as a substitute for chemical pesticides may also brings the risk of developing resistance in the field. Further research is needed to confirm the roles of highly abundant strains, significantly enriched metabolic pathways, and significantly differentially abundant metabolites in host immune defense, in order to provide a theoretical basis for the sustainable use of M. flavoviride in pest control.
Data availability
The datasets generated for this study are available upon request to the corresponding author. Clean data were deposited in the NCBI Short Read Archive (SRA) database (http://www.ncbi.nlm.nih.gov/Traces/sra/). The accession number of 16S rRNA Gene sequencing data was PRJNA862281; the access link is http://www.ncbi.nlm.nih.gov/bioproject/862281.
References
Ashida M, Kinoshita K, Brey PT (1990) Studies on propheno-loxidase activation in the mosquito Aedes aegypti L. Eur J Biochem 188:507–515. https://doi.org/10.1111/j.1432-1033.1990.tb15429.x
Adeyi AO, Akozi GO, Adeleke MA (2015) Induction and activity of glutathione S-transferases extracted from Zonocerus variegatus (Orthoptera: Pyrgomorphidae) exposed to insecticides. Int J Trop Insect Sci 35(1):27–33. https://doi.org/10.1017/S174275841400054X
Boguś MI, Kędra E, Bania J, Szczepanik M, Czygier M, Jabłoński P, Pasztaleniec A, Samborski J, Mazgajska J, Polanowski A (2007) Different defense strategies of Dendrolimus pini, Galleria mellonella, and Calliphora vicina against fungal infection. J Insect Physiol 53(9):909–922. https://doi.org/10.1016/j.jinsphys.2007.02.016
Brinzer RA, Henderson L, Marchiondo AA (2015) Metabolomic profiling of permethrin ⁃ treated Drosophila melanogaster identifies a role for tryptophan catabolism in insecticide survival. Insect Biochem Mol Biol 67:74–86. https://doi.org/10.1016/j.ibmb.2015.09.009
Baumann P (2005) Biology of bacteriocyte-associated endosymbionts of plant sap-sucking insects. Annu Rev Microbiol 59:155–189. https://doi.org/10.1146/annurev.micro.59.030804.121041
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30(15):2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Crotti E, Rizzi A, Chouaia B, Ricci I, FaviaA G, Alberto A (2010) Acetic acid bacteria, newly emerging symbionts of insects. Appl Environ Microb 76:6963–6970. https://doi.org/10.1128/AEM.01336-10
Caporaso J, Gregory KJ, Stombaugh J (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336. https://doi.org/10.1038/nmeth.f.303
Choi CJ, Lee JY, Woo RM (2020) An effective entomopathogenic fungus Metarhizium anisopliae for the simultaneous control of Aedes albopictus and Culex pipiens mosquito adults. J Asia Pac Entomol 23(2):585–590. https://doi.org/10.1016/j.aspen.2020.04.007
Chen Y (2013) The biological function of symbiotic bacteria Arsenophonus for the brown planthopper, Nilaparvata lugens (Stål). Chines academy of agricultural sciences, Beijing, pp 30–33
Cerenius L, Babu R, Soderhall K (2010) In vitro effects on bacterial growth of phenol oxidase reaction products. J Invertebr Pathol 103:21–23. https://doi.org/10.1016/j.jip.2009.09.006
Dillon RJ, Charnley AK (1995) Chemical barriers to gut infection in the desert locust: in vivo production of antimicrobial phenols associated with the bacterium Pantoea agglomerans. J Invertebr Pathol 66:72–75. https://doi.org/10.1006/jipa.1995.1063
Douglas AE, Minto LB, Wilkinson TL (2001) Quantifying nutrient production by the microbial symbionts in an aphid. J Exp Biol 204:349–358. https://doi.org/10.1242/JEB.204.2.349
Douglas AE (1998) Nutritional interactions in insect-microbial symbioses: aphids and their symbiotic bacteria Buchnera. Annu Revi Entomol 43:17–37. https://doi.org/10.1146/annurev.ento.43.1.17
Engel P, Moran NA (2013) The gut microbiota of insects: diversity in structure and function. FEMS Microbiol 37:699–735. https://doi.org/10.1111/1574-6976.12025
Edgar RC, Haas BJ, Clemente JC (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27(16):2194–2200. https://doi.org/10.1093/bioinformatics/btr381
Fan JQ, Chen XR, Hu QB (2013) Effect of destruxins A on hemocytes morphology of Bombyx mori. J Integr Agr 12(6):1042–1048. https://doi.org/10.1016/S2095-3119(13)60324-X
Gerardo NM, Altincicek B, Anselme C, Atamian H, Barribeau SM, Duncan EJ (2010) Immunity and other defenses in pea aphids Acyrthosiphon Pisum. Genome Biol 11(2):R21. https://doi.org/10.1186/gb-2010-11-2-r21
Grant DF, Matsumura F (1989) Glutathione S-transferase 1 and 2 in susceptible and insecticide resistant Aedes aegypt. Pestic Biochem Physiol 33(2):132–143. https://doi.org/10.1016/0048-3575(89)90004-7
Glasl B, Herndl GJ, Frade PR (2016) The microbiome of coral surface mucus has a key role in mediating holobiont health and survival upon disturbance. ISME J 10:2280–2292. https://doi.org/10.1038/ismej.2016.9
Hussa EA, Goodrich-Blair H (2013) It takes a village: ecological and fitness impacts of multipartite mutualism. Annu Rev Microbiol 67:161–178. https://doi.org/10.1146/annurev-micro-092412-155723
Hurst GD, Jiggins FM (2000) Male-killing bacteria in insects: mechanisms, incidence, and implications. Emerg Infec Dis 6:329–336. https://doi.org/10.3201/eid0604.000402
Haber AL, Biton M, Rogel N, Herbst RH, Shekhar K, Smillie C (2017) A singlecell survey of the small intestinal epithelium. Nature 551:333–339. https://doi.org/10.1038/nature24489
Hong T, Qi S, Zhou XM, Bai LY (2013) Field resistance of Spodoptera litura (Lepidoptera: Noctuidae) to organophosphates, pyrethroids, carbamates and four newer chemistry insecticides in Hunan, China. J Pest Sci 86:599–609. https://doi.org/10.1007/s10340-013-0505-y
Harlan NP, Kempker RR, Parekh SM (2011) Weissella confusa bacteremia in a liver transplant patient with hepatic artery thrombosis. Transpl Infect Dis 13:290–293. https://doi.org/10.1111/j.1399-3062.2010.00579.x
Jeong HS, Doo IK, Hye RK (2007) Severe infective endocarditis of native valves caused by Weissella confusa detected incidentally onechocardiography. J Infect 54:149–151. https://doi.org/10.1016/j.jinf.2006.09.009
Kaltenpoth M, Engl T (2014) Defensive microbial symbionts in Hymenoptera. Funct Ecol 28:315–327. https://doi.org/10.1111/1365-2435.12089
Kathawala RJ, Gupta P, Ashby CR (2015) The modulation of ABC transporter-mediated multidrug resistance in cancer: a review of the past decade. Drug Resist Update 18:1–17. https://doi.org/10.1016/j.drup.2014.11.002
Kepler RM, Humber RA, Bischoff JF (2014) Clarification of generic and species boundaries for Metarhizium and related fungi through multigene phylogenetics. Mycologia 106(4):811–829. https://doi.org/10.3852/13-319
Kikuchia Y, Hayatsuc M, Hosokawad T, Nagayamae A, Tagoc K, Fukatsuduk T (2012) Symbiont-mediated insecticide resistance. PNAS 109:8618–8622. https://doi.org/10.1073/pnas.1200231109
Kesnerova L, Mars R, Ellegaard KM (2017) Disentangling metabolic functions of bacteria in the honey bee gut. PLoS Biol 15(12):e2003467. https://doi.org/10.1371/journal.pbio.2003467
Koo HN, Yun SH, Kim HK, Kim GH (2018) Elucidation of molecular expression associated with abnormal development and sterility caused by electron beam irradiation in Spodoptera litura (F.) (Lepidoptera: Noctuidae). Int J Radiat Biol 95:1–8. https://doi.org/10.1080/09553002.2019.1552376
Liochev SI, Fridovich I (2007) The effects of superoxide dismutase on H2O2 formation. Free Radical Bio Med 42(10):1465–1469. https://doi.org/10.1016/j.freeradbiomed.2007.02.015
Lawrence RE, Zoncu R (2019) The lysosome as a cellular centre for signalling, metabolism and quality control. Nat Cell Biol 21:133–142. https://doi.org/10.1038/s41556-018-0244-7
Lauss M, Kriegner A, Vierlinger K (2007) Characterization of the drugged human genome. Pharmacogenomics 8(8):1063–1073. https://doi.org/10.2217/14622416.8.8.1063
Lane TS, Rempe CS, Davitt J, Staton ME, Peng YH, Soltis DE (2016) Diversity of ABC transporter genes across the plant kingdom and their potential utility in biotechnology. BMC Biotechnol 16(1):47. https://doi.org/10.1186/s12896-016-0277-6
Li JC, Rayko H, Dapeng L, Christian P, Haichao S, Sven H, Shu QX, Ian TB (2021) Controlled hydroxylations of diterpenoids allow for plant chemical defense without autotoxicity. Science 371(6526):255–260. https://doi.org/10.1126/science.abe4713
Mercado JE, Hofstetter RW, Reboletti DM, Negron JF (2014) Phoretic symbionts of the mountain pine beetle (Dendroctonus ponderosae Hopkins). Forest Sci 60:512–526. https://doi.org/10.5849/FORSCI.13-045
Muhammad FR, Wang YC, Cai ZH, Bai S, Yao ZC, Umar A (2020) Gut microbiota promotes host resistance to low-temperature stress by stimulating its arginine and proline metabolism pathway in adult Bactrocera dorsalis. Plos Pathog 16(4):e1008441. https://doi.org/10.1371/journal.ppat.1008441
Moor AE, Harnik Y, Ben-Moshe S, Massasa EE, Rozenberg M, Eilam R (2018) Spatial reconstruction of single enterocytes uncovers broad zonation alongthe intestinal villus axis. Cell 175:1156–1167. https://doi.org/10.1016/j.cell.2018.08.06
Nam H, Ha M, Bae O (2002) Effect of Weissella confusa strain PL9001 on the adherence and growth of Helicobacter pylori. Appl Environ Microbiol 68:4642–4645. https://doi.org/10.1128/AEM.68.9.4642-4645.2002
Noor HR, Suzana Y, Armando TQ, Khairiraihana J, Benjamin WBK (2019) Optimization of saponin extracts using microwave-assisted extraction as a sustainable biopesticide to reduce Pomacea canaliculata population in paddy cultivation. Sustain Chem Pharm 11:23–35
Ouedraogo RM, Cusson M, Goettel MS, Brodeur J (2003) Inhibition of fungal growth in thermoregulating locusts, Locusta migratoria, infected by the fungus Metarhizium anisopliae var acridum. J Invertebr Pathol 82(2):103–109. https://doi.org/10.1016/S0022-2011(02)00185-4
Oliver KM, Russell JA, Moran NA (2003) Facultative bacterial symbionts in aphids confer resistance to parasitic wasps. PNAS 100:1803–1807. https://doi.org/10.1073/pnas.0335320100
Oh DC, Scott JJ, Currie CR (2009) Mycangimycin, A polyene peroxide from a mutualist Streptomyces sp. Org Lett 11(3):633–636. https://doi.org/10.1021/ol802709x
Popa V, Deziel E, Lavallee R, Baucec E, Guertina C (2012) The complex symbiotic relationships of bark beetles with microorganisms: a potential practical approach for biological control in forestry. Pest Manag Sci 68:963–975. https://doi.org/10.1002/ps.3307
Platt FM, Davidson BL, Neufeld EF, Tif CJ (2018) Lysosomal storage diseases. Nat Rev Dis Primers 4:27–51. https://doi.org/10.1038/s41572-018-0025-4
Ponte SA (2007) Availability and applications of ATP-binding cassette (ABC) transporter blockers. Appl Microbiol Biot 76(2):279–286. https://doi.org/10.1007/s00253-007-1017-6
Ravikumar B (2010) Regulation of mammalian autophagy in physiology and pathophysiology. Physiol Rev 90:1383–1435. https://doi.org/10.1152/physrev.0030.2009
Stouthamer R, Breeuwer JAJ, Hurst GDD (1999) Wolbachia pipientis: microbial manipulator of arthropod reproduction. Annu Rev Microbiol 53:71–102. https://doi.org/10.1146/annurev.micro.53.1.71
Saxton RA, Sabatini DM (2017) mTOR signaling in growth, metabolism and disease. Cell 168:960–976. https://doi.org/10.1016/j.cell.2017.03.035
Svob SD, Vlainic J, Jazvinscak JM (2008) Differential effects of diazepam treatment and withdrawal on recombinant GABA receptor expression and functional coupling. Brain Res 1246:29–40. https://doi.org/10.1016/j.brainres.2008.09.093
Shi W, Syrenne R, Sun JZ, Yuan JS (2010) Molecular approaches to study the insect gut symbiotic microbiota at the ‘omics’ age. Insect Sci 17:199–219. https://doi.org/10.1111/j.1744-7917.2010.01340.x
Schenck CA, Maeda HA (2018) Tyrosine biosynthesis, metabolism, and catabolism in plants. Phytochemistry 149:82–102. https://doi.org/10.1016/j.phytochem.2018.02.003
Safig P, Klumperman J (2009) Lysosome biogenesis and lysosomal membrane proteins: trafcking meets function. Nat Rev Mol Cell Biol 10:623–635. https://doi.org/10.1038/nrm2745
Sanchez-Rodriguez AR, Barron V, Campollo MCD, Quesada-Moraga OE (2016) The entomopathogenic fungus Metarhizium brunneum: a tool for alleviating Fe chlorosis. Plant Soil 406:295–310. https://doi.org/10.1007/s11104-016-2887-0
Tran LT, Taylor JS, Constabel CP (2012) The polyphenol oxidase gene family in land plants: lineage-specific duplication and expansion. BMC Genom 13:395. https://doi.org/10.1186/1471-2164-13-395
Wu D, Daugherty SC, Van SE (2006) Metabolic complementarity and genomics of the dual bacterial symbiosis of sharpshooters. Plos Biol 4:1079–1092. https://doi.org/10.1371/journal.pbio.0040188
Wang Q, Garrity GM, Tiedje JM (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microb 73(16):5261–5267. https://doi.org/10.1128/AEM.00062-07
Wang WQ, Li SP, Heng X, Chu WH (2022) Weissella confusa CGMCC 19,308 strain protects against oxidative stress, increases lifespan, and bacterial disease resistance in Caenorhabditis elegans. Probiotics Antimicro 14:121–129. https://doi.org/10.1007/s12602-021-09799-z
Xu JJ, Fang X, Chen YL, Lei Y, Chen XY (2019) General and specialized tyrosine metabolism pathways in plants. aBiotech 1:97–105. https://doi.org/10.1007/s42994-019-00006-w
Xue M, Pang YH, Li QL, Liu TX (2010) Efects of four host plants on susceptibility of Spodoptera litura (Lepidoptera: Noctuidae) larvae to five insecticides and activities of detoxifcation esterases. Pest Manag Sci 66:1273–1274. https://doi.org/10.1002/ps.2005
Ye YE, Min H, Lu ZM (2006) Effect of herbicide mefenacet pollution on antioxidant enzyme and ATPase of Sphingobacterium multivorum. Acta Sci Circum 26(1):151–156. https://doi.org/10.3321/j.issn:0253-2468.2006.01.025
Yassine H, Kamareddine L, Osta MA (2012) The mosquito melanization response is implicated in defense against the entomopathogenic fungus Beauveria bassiana. PloS Pathog 8:e1003029. https://doi.org/10.1371/journal.ppat.1003029
Zhuang X, Gross C, Santarelli L (1999) Altered emotional states in knockout mice lacking 5-HT1A or 5-HT1B receptors. Neuropsychopharmacol 21(2 Suppl):52–60. https://doi.org/10.1016/S0893-133X(99)00047-0
Zhao PC, Li JJ, Wang Y (2007) Broad-spectrum antimicrobial activity of the reactive compounds generated in vitro by Manduca sexta phenoloxidase. Insect Biochem MolBiol 37:952–959. https://doi.org/10.1016/j.ibmb.2007.05.001
Zhang L, Shi J, Shi XY (2010) Quantitative and qualitative changes of the carboxylesterase associated with beta-cypermethrin resistance in the housefly, Musca domestica (Diptera: Muscidae). Comp Biochem Physiol b: Biochem Mol Biol 156(1):6–11. https://doi.org/10.1016/j.cbpb.2010.01.011
Zhu LM, Zhou Y, Li XN, Zhao JM, Guo N, Xing H (2018) Metabolomics analysis of soybean hypocotyls in response to phytophthora sojae infection. Front Plant Sci 9. https://doi.org/10.3389/fpls.2018.01530
Zhu HY, Li QQ, Liao TP (2021) Metabolomic profiling of single enlarged lysosomes. Nat Methods 18:788–798. https://doi.org/10.1038/s41592-021-01182-8
Zheng H, Steele MI, Leonard SP (2018) Honey bees as models for gut microbiota research. Lab Anim 47(11):317–325. https://doi.org/10.1038/s41684-018-0173-x
Zheng H, Powell JE, Steele MI (2017) Honeybee gut microbiota promotes host weight gain via bacterial metabolism and hormonal signaling. PNAS 114(18):4775–4780. https://doi.org/10.1073/pnas.1701819114
Zhang JC, Guo Z, Sun ZH (2011) The effect of probiotics on intestinal flora-Take Lactobacillus casei Zhang study as an example. J Chin Inst Food Sci Technol 11:58–68 ((in Chinese))
Zhang YH, Cai TW, Cai YF, Shu RH, Yuan MJ, Wan H (2021) Microbiome variation correlates with the insecticide susceptibility in different geographic strains of a significant agricultural pest, Nilaparvata Lugens. NPJ Biofilms Microbi 9:2. https://doi.org/10.1038/s41522-023-00369-5
Funding
This work was supported by Zhejiang Province science and technology plan projects (Grant No.2022SNJF016) and China Agriculture Research System (Grant No. CARS-24-G-07). Zhang juefeng has received research support from Zhejiang Province Department of Agriculture and Rural Affairs, Chen jianming has received CARS-24-G-07 support from Ministry of Agriculture of the PRC. The authors declare that no funds, grants, or other support were received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
We confirm that all authors have approved the manuscript and agree with submission to the Journal. Zhang Juefeng carried out the experiments and drafted the manuscript. Li Fang, Zhong Haiying and Liu Liwei helped with experimental procedures and manuscript preparation. Chen Jianming designed the study and critically revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethical approval
The insects involved in the experiment were all artificially reared in the experimental base owned by our laboratory (Institute of Plant Protection and Microbiology, Zhejiang Academay of Agricultural Sciences). All the experimental insects were collected with permission from local authorities of agricultural department. All the experimental research on insects complied with relevant guidelines and regulations.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Juefeng, Z., Fang, L., Haiying, Z. et al. Integrated microbiome and metabolomic analysis of Spodoptera litura under Metarhizium flavoviride qc1401 stress. Int Microbiol (2024). https://doi.org/10.1007/s10123-024-00574-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10123-024-00574-y