Abstract
High-altitude cold habitats of the Karakoram are rarely explored for their bacterial community characterization and metabolite productions. In the present study, bacterial communities in ice, water, and sediments of Batura Glacier were investigated using culture-dependent and culture-independent methods. Twenty-seven cold-adapted bacterial strains (mostly psychrotrophic) were isolated using R2A, Tryptic Soy Agar (TSA), and Luria-Bertani (LB) media, at 4 °C and 15 °C. Most of the isolates exhibited growth at a wide range of temperature (4–35 °C), pH (5–12), and salinity (1–6%). Among the bacterial isolates, 52% were identified as Gram-positive and the remaining 48% represented as Gram-negative. The results of phylogenetic analysis indicated that all the culturable bacteria belonged to 3 major phylogenetic groups, i.e., Actinobacteria (48%), Bacteroidetes (26%), and Proteobacteria (22%), while Flavobacterium (26%), Arthrobacter (22%), and Pseudomonas (19%) were represented as the dominant genera. Similarly, Illumina amplicon sequencing of 16S rRNA genes after PCR amplification of DNA from the whole community revealed dominance of the same phylogenetic groups, Proteobacteria, Actinobacteria, and Bacteroidetes, while Arthrobacter, Mycoplana, Ochrobactrum, Kaistobacter, Janthinobacterium, and Flavobacterium were found as the dominant genera. Among the culturable isolates, 70% demonstrated activity for cellulases, 48% lipases, 41% proteases, 41% DNases, and only 7% for amylases. Most of the glacial isolates demonstrated antimicrobial activity against other microorganisms including the multiple-drug-resistant strains of Candida albicans, Klebsiella pneumoniae, Acinetobacter sp., and Bacillus sp. 67% of Gram-negative while 46% of Gram-positive glacial bacteria were resistant to trimethoprim/sulfamethoxazole. Resistance against methicillin and vancomycin among the Gram-positive isolates was 23% and 15%, respectively, while 11% of the Gram-negative isolates exhibited resistance against both colistin sulfate and nalidixic acid.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
More than 80% of the biosphere is constantly at temperatures under 5 °C. The deep sea represents a major fraction of these cold environments as the oceans cover 71% of the planet and 90% of their volume is below 5 °C (Casanueva et al. 2010; Rodrigues and Tiedje 2008). Additionally, glaciers account for 10% of the land surface (Russell 1990; Margesin and Miteva 2011). The cold biosphere harbors a diverse group of cold-adapted microorganisms called psychrophiles or psychrotrophs, inhabiting terrestrial and aquatic environments in the Arctic and Antarctic polar regions as well as the deep ocean, high mountains of alpine regions, upper atmosphere, and the cave systems. Cold-adapted bacteria have successfully colonized these environments through a vast array of adaptation strategies including maintenance of membrane fluidity and structural flexibility of their enzymes, expression of the cold shock proteins, and production of antifreeze proteins and compatible solutes (Collins and Margesin 2019; Tribelli and López 2018; Barria et al. 2013).
The concept of glacial ecology has gained the interest of researchers in recent years, mainly focusing on the two key glacier ecosystems: the supraglacial and subglacial (Hodson et al. 2008). The supraglacial ecosystems represent the outer surface of the glaciers and were inhabited by a group of complex microbial communities including algae, bacteria, phytoflagellates, viruses, archaea, fungi, and rotifers (Hodson et al. 2008). The subglacial ecosystems represent the ice-bed interface and are dominated by the heterotrophs including bacteria and fungi (Hodson et al. 2008). Cold-adapted microorganisms are involved in key ecological functions in freezing environments where photoautotrophy is the basis of food webs in the proglacial and chemoautotrophy dominates the subglacier environments (Boetius et al. 2015). Glacier ice with extremely low temperature is considered one of the harsh environments for living organisms on the biosphere. Apart from the extremely low temperatures, other limitations include low nutrient and water availability, frequent freeze-thaw events, and increased darkness (Margesin and Miteva 2011). However, despite all these limitations, it is estimated that glaciers and ice sheets around the world contain as many as 1029 microbial cells (Irvine-Fynn and Edwards 2014). Studies on the structure of microbial communities and their possible ecological role in cold environments have been reported from various glaciers of the polar and non-polar regions (Segawa et al. 2011; Shivaji et al. 2011; Miteva et al. 2004).
Extremophiles are organisms that can thrive in environments that are hostile or inhabitable to other forms of life (Horikoshi et al. 2010). Psychrophiles represent the most abundant group of extremophiles in terms of distribution, diversity, and biomass (D’Amico et al. 2006). Apart from their ecological role, cold-adapted microorganisms have shown tremendous potential for biotechnological applications ranging from enzymes and antimicrobial productions to environmental bioremediations (Cavicchioli et al. 2011; Sánchez et al. 2009; Gratia et al. 2009). Nature is considered the best reservoir for the discovery of industrially important metabolites. The less explored extreme environments are seen as new avenues for the bioprospecting of novel bioactive molecules including enzymes and antimicrobial compounds (Sánchez et al. 2009; Cavicchioli et al. 2002).
The north-most territories of Pakistan consist of the greatest mountain ranges in the world including the western Himalayas, Karakoram, and Hindukush. These ranges host at least 5000 glaciers in the Pakistani geographical region (Rasul et al. 2011). Some of the notable glaciers of the Karakoram includes Siachen Glacier (∼ 75 km long), Biafo Glacier (∼ 68 km), Baltoro Glacier (∼ 62 km), and Batura Glacier (∼ 59 km) (Hewitt 2014). Interestingly, glaciers of the Karakoram are stable or even growing contrary to the ice bodies worldwide that are receding due to global warming (Rankl et al. 2014). This unusual behavior of the Karakoram glaciers is known as the “Karakorum Anomaly” (Hewitt 2005). However, glaciers in the Karakoram are the least explored in terms of their bacterial diversity. It would be interesting to study the bacterial community of these glaciers that have exhibited resilience in climate change when other glaciers in the world are retreating. Previously, genome sequencing of the glacial isolate Pseudomonas sp. BGI-2 unveiled 11 exopolysaccharide (EPS)-producing genes, which are not present in the closely related mesophilic Pseudomonas species (Ali et al. 2019a). Also, BGI-2 produced high yield of a cryoprotective EPS at low temperatures and demonstrated ability to withstand freeze-thaw stress (Ali et al. 2020). Another glacial isolate BGI-11 demonstrated the ability to grow at higher salinity and utilized hexadecane as carbon source (Ali et al. 2019b).
The aim of the present study was to characterize the bacterial community of an unexplored glacier in the Karakoram Range of Pakistan and explore some of their metabolic potential for possible industrial applications. To the best of our knowledge, this is the first study where culture-independent method was used to characterize the bacterial community in any glacier of the Karakoram in the Pakistani geographical region.
Materials and methods
Sampling, bacterial count, and isolation
Description of the sampling site and collection of samples
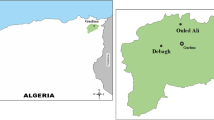
Sampling was done in Batura Glacier located in the Passu village of the upper Hunza valley in the northern Pakistan (Fig. 1). Hunza valley is located at the base of the Karakoram Range along the Karakoram Highway. Batura Glacier (36° 32′ N, 74° 40′ E) is one of the longest glacier outside the Polar regions with the highest elevation around 7795 m in the West and the lowest nearly 2570 m in the East (Hewitt 2014). Batura Glacier basin is ∼ 48% glacierized and surrounded by several major peaks with altitude > 7000 m including, Batura Muztagh ∼ 7795 m (Hodson et al. 2002).
Triplicate samples of ice, water, and sediment were collected aseptically in sterile 500-mL wide-mouth polypropylene bottles. Handling of samples was performed according to standard microbiological techniques. Sterilized tools were used for sampling of glacial ice and sediments. Surface ice and sediment up to 10–13 cm were removed and discarded. Underlying samples were collected into sterile bottles, temperature, and pH recorded at the time of sampling. The tubes were sealed, placed in an isothermal box, and transported to Applied, Environmental and Geomicrobiology Laboratory, at the Department of Microbiology, Quaid-i-Azam University Islamabad, and stored at − 20 °C. Geographic coordinates were recorded at the time of sampling using a GPS device.
Direct bacterial count using epifluorescence microscopy
Direct bacterial count from the three glacier samples performed using the method used by Zhang et al. (2008) and Chen et al. (2001) with minor modifications. About 2 to 3 mL of glacier samples (melted ice and water) were filtered through 0.2-μm polycarbonate membrane filter (Millipore) under gentle vacuum pressure. CYBR Gold (Invitrogen) working solution (2 ×) was prepared from an original stock (10,000 ×) using TE buffer (10 mM Tris-Cl; 1 mM EDTA, pH 7.4–7.6). Bacterial cells on the filters were stained with 300 μL of the CYBR Gold solution (2 ×) for 15 min in the dark and the residual stain was removed by applying vacuum. Ten microliters of mounting solution (50% glycerol and 50% TE buffer) was placed on the slides and stained filters were mounted on top of the drop with cover slips placed over it. The samples were examined under blue-green light excitation (480–495 nm) using fluorescent microscope (Axioplan, Zeiss). The number of bacterial cells per milliliter or gram for each sample was calculated based on count of at least 20 randomly selected microscopic fields and the volume of each filtered sample.
Enumeration, isolation, and viable bacterial count
Enumeration of culturable heterotrophic bacteria was carried out through the plate count method. Three general purpose culture media including Tryptic Soy agar (Oxoid), Luria Bertani agar (Miller), and R2A agar (Difco) were used to isolate psychrophilic/psychrotrophic bacteria from the samples. About 100 μL aliquots of the three glacier samples, or a 10-fold dilution of them, was spread onto agar plates. These plates were incubated at two different temperatures (4 °C and 15 °C) for up to 4 weeks and observed daily for colony appearance. Plates were also incubated at 4 °C for an extended period of up to several months until growth of new colonies was no longer detected. Morphologically different colonies from each plate were sub-cultured on separate fresh plates to obtain pure cultures. All the viable counts were performed in replicates and the results presented are the mean of duplicate assays.
For ice samples, 100 μL of melted ice was directly plated on tryptic soy agar, LB agar, and R2A agar, and incubated at 4 °C for 9 months and 15 °C for 6 months. Enrichment was also carried out by adding 10 mL of melted ice sample to 90 mL of liquid culture media (R2A broth, TSB and LB broth). Enrichment samples were incubated in shaker incubators (150 rpm) at two different temperatures (4 °C and 15 °C). Samples were then visualized for bacterial growth (media turbidity) every day up to 2 to 3 weeks. One hundred microliters of diluted and undiluted samples from each enrichment culture was plated on their corresponding solid media for separate colonial growth. Morphologically different colonies observed were streaked on separate plates to obtain pure cultures. A similar procedure was used for enrichment of water and sediment samples for maximum recovery of bacterial isolates.
Identification of culturable bacterial strains
Biochemical identification
The bacterial isolates were differentiated and identified initially based on colony morphology (size, elevation, margin, and pigmentation), cell morphology (cocci, coccobacilli, or rods), and biochemical characteristics (sugar fermentation, H2S production, catalase, oxidase, nitrate reduction, and citrate utilization), while later on subjected to molecular identification through 16S rRNA gene sequencing.
Identification through 16S rRNA gene sequencing and phylogenetic analysis
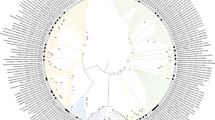
DNA was extracted from each isolate using a Genomic DNA Isolation Kit (Norgen Biotek Corporation, Canada). The 16S rRNA gene was amplified using universal primers 27F and 1492R. The PCR conditions for 16S rRNA gene amplification were as follows: initial denaturation at 96 °C for 5 min followed by 35 cycles of denaturation at 96 °C for 30 s, annealing at 55 °C for 30 s, and elongation at 72 °C for 30 s. The final elongation step was 10 min at 72 °C. Sequencing of 16S rRNA was done by Macrogen Inc., Seoul, Korea. The ambiguous low-quality aligned regions from sequences were removed using Chromas (version 2.6.6). 16S rRNA gene sequences for all the isolates were analyzed using EzBioCloud database (Yoon et al. 2017). 16S rRNA gene sequences for all the isolates were deposited in the NCBI GenBank database and accession numbers were obtained. A phylogenetic tree was constructed by neighbor-joining method with bootstrap value > 50 using the MEGA 7.0 software (Fig. 2) (Saitou and Nei 1987).
Phylogenetic tree of isolates recovered from the three glacier samples using neighbor-joining method with a bootstrap value (%) greater than 50 from 1000 replicates. Numbers in the brackets are GenBank accession numbers for the 16S rRNA sequences and E.coli ATCC 11775T was used as an outgroup to root the tree
Characterization of isolated bacteria
Optimization of physico-chemical culture conditions
Growth of the isolates was checked at different temperatures (4, 5, 10, 15, 20, 25, 30, 35, 40, and 45 °C) and pH (3.0, 4.0, 5.0, 6.0, 7.0, 8.0, 9.0, 10.0, 11.0, 12.0, and 13.0) on R2A agar plates (Zhang et al. 2013). Isolates were checked for their ability to grow under halophilic conditions by streaking them on R2A agar plates supplemented with NaCl at concentration ranges from 2 to 10% (w/v). Growth medium was also optimized by inoculating on to different media named R2A (Difco), tryptic soy agar (Oxoid), nutrient agar (Oxoid), LB agar (Oxoid), Sabouraud dextrose agar (Oxoid), mannitol salt agar (Oxoid), MacConkey agar (Oxoid), and Mueller Hinton agar (Oxoid). For all the above tests, the result was analyzed after incubating plates at 15 °C for 5 days in duplicates. These tests were performed according to the methods as previously described (Zhang et al. 2013).
Antibiotic resistance
Antimicrobial susceptibility was determined on R2A and Mueller-Hinton agar (MHA) plates using disc diffusion method (Bauer et al. 1966). All the isolates were grown in R2A broth at 15 °C for 48 h. Inoculums were diluted to turbidity equivalent to 0.5 McFarland standard and swabbed on the surface of agar plates (R2A and MHA) using sterile cotton swabs. Seven antibiotics representing different classes were used including methicillin (10 μg), imipenem (10 μg), ofloxacin (5 μg), vancomycin (5 μg), colistin sulfate (10 μg), nalidixic acid (30 μg), and trimethoprim sulfamethoxazole (30 μg). All the plates were incubated at 15 °C and results were interpreted after 72–96 h. The susceptibility of strains to each antibiotic was determined by measuring zone of inhibition (Fig. 3a).
a Antibiotic susceptibility test for Okibacterium sp. BGS-13 shows resistance to methicillin, imipenem, and trimethoprim sulfamethoxazole. b Cellulolytic enzyme activity by the Bacillus sp. BGI-4 on carboxymethylcellulose (CMC) plate. c Antimicrobial activities by Pseudomonas sp. BGI-1 and Pseudomonas sp. BGI-2 against Candida albicans by spot on lawn assay using Mueller Hinton agar plate
Extracellular enzyme activity
Isolates were screened for their ability to produce extracellular enzymes. Starch hydrolysis test was performed for amylase production (Priest 1977). Lipase activity was checked by streaking isolates on basal medium supplemented with 1% Tween-40 along with 0.001% rhodamine B and 0.01% CaCl2, according to the method as previously described (Booth 1971). Cellulase activity was demonstrated on carboxymethylcellulose (CMC) agar plates according to the methods from Kasana et al. (2008). Zone of clearance around the isolates indicated a positive response for cellulase production (Fig. 3b). Protease activity was checked by the method as previously described by Priest (1977). For DNase test, isolates were spot inoculated on surface of DNase agar and plates were incubated at 15 °C for 5 days. Plates were then flooded with 1 N hydrochloric acid. Hydrolysis of DNA was indicated by a clear zone around the growth of tested organism.
Antimicrobial activity
Antimicrobial activity of all the glacial isolates was tested against a number of microorganisms including the multidrug resistance strains of Candida albicans, Klebsiella pneumoniae, Acinetobacter sp., and Bacillus sp. and two ATCC strains Pseudomonas aeruginosa (ATCC27853) and Staphylococcus aureus (ATCC6538). Antimicrobial activity was determined through spot on lawn assay (Rafiq et al. 2017). Briefly, R2A agar and Mueller-Hinton agar plates were swabbed with overnight culture of indicator organisms and isolates were spot inoculated on plates. The plates were incubated at 15 °C for 3 to 5 days and observed for a clear zone around the isolates (Fig. 3c). The multiple drug resistance strains were provided by the Medical Microbiology Laboratory, in the Department of Microbiology, Quaid-i-Azam University, Islamabad.
High-throughput sequencing of bacterial community 16S rRNA gene sequencing
Extraction of community DNA and 16S rRNA gene sequencing
DNA from ice, water, and sediment samples was extracted using DNeasy® PowerWater Kit (Qiagen GmbH, Hilden, Germany) and DNeasy® PowerSoil® Kit (Qiagen GmbH, Hilden, Germany). One hundred milliliters each of water and ice samples and 50 g sediment sample were used for community DNA extraction. Quality and quantity of the extracted DNA were determined using NanoDrop 2000 spectrophotometer (Thermo Scientific). 16S rRNA genes for the three samples were amplified using the forward primer (5′ GTGCCAGCMGCCGCGGTAA) and reverse primer (5′ GGACTACHVGGGTWTCTAAT) designed for the V3 and V4 regions (Klindworth et al. 2013). Size of the PCR amplicons was checked through 1% agarose gel electrophoresis. Sequencing was done using MiSeq (Illumina, San Diego, CA) at the BioAnalytical Laboratory, Institute of Marine and Environmental Technology (IMET), University of Maryland Center for Environmental Sciences (UMCES). The libraries were prepared using the Nextera XT kit (Illumina) and sequencing was done using 2 × 300 cycle run using the MiSeq reagent kit v.3. The 16S rRNA gene dataset consisted of 189,890 raw sequence reads for the ice sample, 451,614 for water, and 225,910 reads for the sediment sample.
16S rRNA gene data and operational taxonomic unit classification
16S rRNA gene MiSeq sequencing reads for the three glacier samples were assembled and classified using QIIME software v 1.9.1 (Caporaso et al. 2010). Alignment of the unique sequences was performed with the ‘align.seqs’ command against a Greengenes database. Quality filtered sequences were clustered into operational taxonomic units (OTUs) according to sequence similarity using a 97% similarity threshold against the respective reference databases (Marzorati et al. 2008). The remaining sequences were clustered into de novo OTUs with UCLUST within QIIME.
RDP classifier in QIIME was used for the assignment of taxonomy to these sequences.
Nucleotide sequence accession numbers
The 16S rRNA gene accession numbers for isolates from different samples were designated as follows: bacteria from ice (MK522040-MK522046), from water (MK558827-MK558837), and from sediment (MK558838-MK558845). The bacterial 16S rRNA gene raw data from MiSeq sequencing for the ice, water, and sediment samples was deposited into the Sequence Read Archive (SRA) of the National Center for Biotechnology Information (NCBI) under the BioProject accession number PRJNA533649.
Results
Bacterial count, isolation, and identification
Microscopic and viable count
Direct bacterial count using epifluorescence microscopy revealed the presence of high number of bacterial cells in all three samples. The highest bacterial count was observed in sediment (3.3 × 106 cell g−1), followed by water (2.8 × 106 cells mL−1) and ice (6.4 × 105 cells mL−1) (Table 1).
For viable count in the ice sample, no bacterial colony was observed on any of the three media used (R2A, TSA, LBA) even after 9 months of incubation at 4 °C and 6 months of incubation at 15 °C. Viable bacterial count from the water and sediment samples varied depending on the incubation temperature and the type of medium used. The maximum viable count for water (2.0 × 103 CFU mL−1) and sediment (1.8 × 104 CFU g−1) samples was recorded when R2A medium was used at 15 °C (Table 1).
Isolation of bacteria
A total of 27 bacterial isolates were recovered from three glacier samples, including 11 from water and 8 from each ice and sediment samples. Most of the isolates were recovered on the oligotrophic medium R2A, whereas the nutrient-rich media (TSA and LBA) yielded less colonies. Most of the colonies appeared after 2 to 3 weeks of incubation (4 °C and 15 °C) and only few appeared in the first week. However, the same isolates formed visible colonies within 2 to 5 days on subsequent re-streaking. Initial identification was done based on the colony and cell morphologies, pigmentation, and growth characteristics.
Direct plating of the melted ice sample on agar media (R2A, TSA, and LBA) did not yield colonies on any of the plates even after 9 and 6 months of incubation at 4 °C and 15 °C, respectively. On the other hand, in enriched with liquid media (R2B, TSB, and LB), growth was observed in the second week at 15 °C as indicated by turbidity in the media. When inoculated on the respective solid medium, colonies appeared within the first to the 3rd week of incubation at 4 °C and 15 °C. A total of 8 different bacterial isolates were obtained after plating the liquid enrichment culture on agar media; six colonies from R2B and one each from TSB and LB enrichments were recovered.
Gram staining revealed Batura Glacier is inhabited by representatives from both Gram-positive (14 isolates) and Gram-negative (13 isolates), with majority of the isolates exhibited coccobacillary cell morphologies (Table 2). Almost half of the isolates showed pigmented colonies. Ten of the isolates exhibited yellowish pigmentation while one each orange and pink (Table 2). Strains isolated from the glacier ice, water, and sediment were named with the prefix BGI (Batura Glacier Ice), BGW (Batura Glacier Water), and BGS (Batura Glacier Sediment).
Taxonomic classification of culturable isolates based on 16S rDNA gene sequences
Molecular identification based on 16S rDNA gene sequences revealed Batura Glacier is inhabited by a diverse group of bacteria. The isolates belong to 3 major phylogenetic groups: Actinobacteria (48%), Bacteroidetes (26%), and Proteobacteria (22%) (Fig. 2). The dominant genera were Flavobacterium (26%), Arthrobacter (22%), and Pseudomonas (19%). A phylogenetic tree was constructed to determine the affiliation of 27 isolates with other bacteria in the NCBI GenBank database (Fig. 2). The majority of 16S rRNA sequences was similar to previously determined sequences with 98% or above identical values (Table 3). Some of the isolates had 98 to 99% 16S rRNA gene sequence similarity with psychrophilic bacteria isolated from various cold environments. BGI-14 was closely related to Cryobacterium psychrotolerans (> 99%), BGW-1/BGW-17/BGW-18 to Flavobacterium psychrolimnae (> 99%), BGW-2/BGW-7/BGW-12 to Flavobacterium glaciei (> 98%), and BGW-5 to Pseudomonas antarctica (> 99%) (Table 3).
Characterization of bacterial isolates
Growth characteristics of the isolated bacteria
Phenotypic characteristics including temperature, pH, and salinity ranges for growth were determined. Results revealed that most of the isolates could grow at a wide range of temperature (4–35 °C) and pH (5–12) (Table 2). A few strains, including BGI-10 and BGI-11, demonstrated the ability to grow at the extremes of pH range (3 and 13). The majority of the isolates were either slight halophiles or halotolerant. BGI-10 and BGI-11 were able to grow at salt concentrations of up to 8% (w/v); however, none of the isolates demonstrated growth at 10% NaCl. Most of the isolates preferred the oligotrophic medium R2A and also demonstrated growth at nutrient-rich media (Table S1). BGI-10 and BGI-11 were able to grow in all the 8 media used. Five Pseudomonas species BGI-1, BGI-2, BGW-4, BGW-5, and BGW-10 demonstrated growth on MacConkey’s agar and could not ferment lactose (Table S1). BGS-6, BGI-10, and BGI-11 demonstrated good growth on mannitol salt agar with high salt concentration (7.5% w/v) and deemed halotolerant.
Extracellular enzymes and antimicrobial activities
Isolates were screened for enzymatic and antimicrobial activities. Most of the isolates demonstrated the ability to produce one or more extracellular enzymes (Table S2). Enzymatic assays revealed that 70% of the strains produced cellulases, 48% lipases, 41% proteases, 41% DNases, and only 7% amylases. Some isolates demonstrated remarkable ability to produce 3 or more different enzymes including strains BGS-9, BGI-1, BGI-2, BGS-2, BGS-12, BGW-4, BGW-5, and BGW-15. Flavobacterium sp. BGW-15 and Flavobacterium sp. BGW-17 were the only strains able to produce amylases (Table S2).
The majority of the glacier isolates demonstrated antimicrobial activity against one or more test microorganisms: 33% against C. albicans, 30% against Bacillus sp., 26% against P. aeruginosa, 26% against S. aureus, 22% against Acinetobacter sp., and only 4% against K. pneumoniae. Some strains exhibited broad-spectrum activity including BGI-2, BGW-4 and BGW-5, BGI-1, BGW-8, and BGW-10 (Table S2).
Prevalence of antibiotic resistance in the glacier isolates
Antibiotic resistance to commercial antibiotics was found among the glacial isolates (Table S3). As many as 67% of the Gram-negative and 46% of the Gram-positive bacteria demonstrated resistance to trimethoprim/sulfamethoxazole (30 μg) (Table 4). Among the Gram-negative bacteria, 11% were resistant to colistin sulfate (10 μg) and nalidixic acid (30 μg) each. Resistance against methicillin (10 μg) and vancomycin (5 μg) among the Gram-positive isolates was 23% and 15%, respectively. All the Gram-negative bacteria were sensitive to the broad-spectrum antibiotics imipenem (10 μg) and ofloxacin (5 μg). Only one Gram-positive isolate exhibited resistance to imipenem (10 μg) and all 27 isolates were sensitive to ofloxacin (5 μg) (Table S3).
Culture-independent characterization of bacterial community
The results from Illumina amplicon sequencing of 16S rRNA genes revealed Proteobacteria, Actinobacteria, and Bacteroidetes as major phyla from the three glacier samples (Fig. 4a). Proteobacteria dominated in all the three samples, followed by Actinobacteria and Bacteroidetes. A small fraction of the bacterial communities was also represented by the Saccharibacteria and Firmicutes and a considerably large fraction of the sequences (20%) in the sediment sample was unassigned.
The sequences in glacier samples were assigned to 24 genera (> 0.5%) and the majority of them could not be refined to the genus level and were then grouped under the category “Bacteria (others).” The genera included the following: Arthrobacter, Flavobacterium, Pseudomonas, Kaistobacter, Mycoplana, Ochrobactrum, Devosia, Rhodococcus, Rhodobacter, Janthinobacterium, Acinetobacter, Segetibacter, Flavisolibacter, Methylotenera, Polaromonas, Sphingomonas, Novosphingobium, Agrobacterium, Methylobacterium, Lysinibacillus, Chryseobacterium, Salinibacterium, Mycetocola, and Microbacterium (Fig. 4b).
Mycoplana (16%) and Arthrobacter (12%) represented the major genera in the ice sample. The genera Kaistobacter (20%) and Arthrobacter (16%) dominated in sediment sample; Ochrobactrum (30%), Janthinobacterium (24%), and Flavobacterium (11%) dominated in ice sample. Arthrobacter was found predominantly in all the three samples. Graphs (Fig. 4a and b) depict the phyla and genera with operational taxonomic units (OTUs) that have abundance greater than 0.5%.
Discussion
In the present study, we examined the possibility of using a glacier in the Karakoram Range of Pakistan as source for isolation of novel cold-adapted bacteria, and identification and exploration of some of their metabolic potential to be used for biotechnological or industrial applications. The glaciers of the Karakoram, particularly in the Pakistani geographical region, are not commonly explored in terms of their bacterial community characterization and metabolite production.
Viable count was the highest on R2A among the three media and the majority of the isolates were recovered using this medium. R2A is a low nutrient medium designed for the recovery of bacteria from oligotrophic samples (Reasoner and Geldreich 1985). Many studies have previously been reported where R2A was considered the medium of choice for isolation of psychrophilic or psychrotrophic bacteria from cold environments (Christner et al. 2001; Segawa et al. 2011; Tam et al. 2015). The maximum viable counts for water and sediment samples were measured at 15 °C on R2A agar. A similar range of viable counts has been reported previously for samples from the cold habitats (Zhang et al. 2013; Segawa et al. 2011). In our study, isolates from the ice were recovered through culture enrichment in liquid media, which is in accordance with a previous study (Antony et al. 2012). Only a small fraction of microorganisms in any environment is culturable and a large fraction remains uncultivable and extreme environments make it even harder for the isolation of bacteria; their nutritional requirements are limited and somewhat unknown. Other studies have also reported extremely low viable bacterial counts from the glacier ice samples (Zhang et al. 2008; Christner et al. 2001).
Another feature, which was predominant in the glacial isolates, was the formation of pigmented colonies. More than half of the isolates formed pigmented colonies that appeared yellowish and orange. Studies showing pigmented bacteria dominating bacterial communities have been reported from glaciers and other cold habitats (Shen et al. 2018; Christner et al. 2000; Antony et al. 2012). Carotenoid pigmentation is considered a strategy to withstand environmental stresses including low temperatures (Chattopadhyay and Jagannadham 2001; Dieser et al. 2010). Fong et al. (2001) found correlation between temperature and carotenoid production, and a decrease in cultivation temperature resulted in a concomitant increase in carotenoid production, which is speculated to contribute to membrane stabilization at low temperatures.
All the isolated bacteria belonged to 3 major phyla including Actinobacteria, Proteobacteria, and Bacteroidetes. The predominance of these groups in glaciers and other cold environments has been previously reported (Silva et al. 2018; Shen et al. 2018; Liu et al. 2018; Cheng and Foght 2007). Flavobacterium, Arthrobacter, and Pseudomonas were the dominant genera in overall the three glacier samples which conforms to the previous reports from cold habitats (Thakur et al. 2018; Reddy et al. 2009; Cheng and Foght 2007). All the isolated bacteria from sediment samples were members of Actinobacteria with 63% representing the genus Arthrobacter. Zhao et al. (2018) studied the culturable bacteria in soil along an altitudinal gradient in the Tianshan Mountains and found that almost 50% represented the genus Arthrobacter.
16S rRNA gene meta-barcoding also revealed Proteobacteria, Actinobacteria, and Bacteroidetes as the dominant taxa in the glacier samples. Proteobacteria accounted for more than 50% of relative abundance in all the three glacier samples. Proteobacteria, Actinobacteria, and Bacteroidetes have been previously reported as the dominant phyla from glacier environments using culture-independent methods (Lopez et al. 2018; Venkatachalam et al. 2015). Mycoplana, Arthrobacter, Kaistobacter, Ochrobactrum, Janthinobacterium, and Flavobacterium represented the major genera in the glacier samples. Arthrobacter was consistently found as one of the dominant genera in terms of the relative abundance in all the three glacier samples. Arthrobacter has been previously reported as the dominant genus from cold environments (Thakur et al. 2018; Zhao et al. 2018). Similarly, Mycoplana (Yang et al. 2016), Kaistobacter (Yang et al. 2016), Janthinobacterium (Rassner et al. 2016), Ochrobactrum (Liu et al. 2009), and Flavobacterium (Segawa et al. 2011) have also been reported previously from glacial environments. Interestingly, Ochrobactrum was the dominant genus among all in the ice sample and accounted for 30% of the relative abundance. Members of the genus Ochrobactrum have rarely been reported from glaciers and other cold environments. Ochrobactrum species are metabolically versatile and produce a variety of enzymes to degrade recalcitrant compounds including pesticides, phenols, and petroleum hydrocarbons, making them potential candidates for bioremediation technologies (Ermakova et al. 2017; Arulazhagan and Vasudevan 2011).
According to Moyer and Morita (2007) definitions of cold-adapted bacteria, Batura Glacier is mainly dominated by psychrotrophic bacteria as the majority of the isolates exhibited a wide growth temperature range (4–30 °C or 4–35 °C). Flavobacterium isolates BGW-15, BGW-17, and BGW-18 could not grow above 20 °C and were determined to be representatives of psychrophilic bacteria. According to Russell (1990), psychrophiles are found in permanently cold environments and psychrotrophs dominate habitats which experience more thermal fluctuations. This is the case for non-polar glaciers like Batura Glacier in the Karakoram, as it experiences wide seasonal temperature fluctuations. Therefore, psychrotrophic bacteria are expected to dominate over their psychrophilic counterparts. The dominance of psychrotrophic bacteria compared to true psychrophilic bacteria has been previously reported in similar cold environments (Liu et al. 2018; Thakur et al. 2018; Reddy et al. 2009; Steven et al. 2007).
Antibiotic resistance patterns in the glacier isolates revealed resistance against some commercial antibiotics. High resistance was documented against trimethoprim/sulfamethoxazole among the Gram-negative and Gram-positive bacteria. However, resistance against the broad-spectrum antibiotics (ofloxacin and imipenem) was the least observed. Imipenem is a β-lactam antibiotic and resistant bacteria produce β-lactamases to inactivate the drug. In a recent study, Van Goethem et al. (2018) found a very low abundance of β-lactamase genes in the metagenome of a pristine cold habitat of Antarctica. Nevertheless, antibiotic resistance genes and organisms have been reported from non-clinical and natural environments including the ocean and polar region (Hatosy and Martiny 2015; Van Goethem et al. 2018; Segawa et al. 2013). The resistance found in the remote pristine regions with no or less human impact suggests a natural process that is acquired by these microorganisms over time through the process of evolution and the passing off through generations. Nevertheless, Batura Glacier is located close to living populations and anthropogenic transfer through migratory birds and airborne bacteria could be a possibility (Segawa et al. 2013).
Most of the isolates in our study exhibited antimicrobial activities against one or more of the test organisms. All cultured Pseudomonas species demonstrated broad-spectrum antimicrobial activities. Cold-adapted Pseudomonas species with wide-spectrum antimicrobial activities have been reported previously from Antarctica (Silva et al. 2018; Tedesco et al. 2016; O'Brien et al. 2004). Isolated strains from Batura Glacier also exhibited abilities to produce a variety of extracellular enzymes including cellulases, proteases, lipases, DNases, and amylases. Cold-adapted bacteria with enzyme activities have been reported previously from the cold environments (Reddy et al. 2009; Margesin et al. 2003). The majority of the isolates demonstrated cellulolytic activity (70%), similar to a recent study by Thakur et al. (2018); their result revealed 71% of cellulolytic bacteria in their strains using carboxymethyl cellulose (CMC) plates. Overall, Pseudomonas species dominated among the glacier isolates in terms of enzymatic activities. Pseudomonads have been reported previously as producers of lipases (Ganasen et al. 2016), proteases (Zeng et al. 2003), and cellulases (Menéndez et al. 2015).
Conclusions
Results of the culture-dependent and culture-independent study revealed bacterial community from the glacier samples belonged to 3 major phylogenetic groups including Actinobacteria, Bacteroidetes, and Proteobacteria. The range of temperature for growth of the isolates indicated that Batura Glacier is mainly inhabited by psychrotrophic bacteria rather than the true psychrophiles. The majority of the isolates were either slight halophiles or halotolerant. Our results also indicated that cold environments in the Karakoram Range are promising sources of cold-adapted bacteria with enormous potential for biotechnological exploitation. Most of the isolates produced one or more extracellular enzymes and exhibited antimicrobial activity against a number of microorganisms including multidrug-resistant pathogenic strains. Almost all the isolates were sensitive to the broad-spectrum antibiotics (ofloxacin and imipenem), while varying degree of resistance was found against the narrow spectrum antibiotics.
References
Ali P, Shah AA, Hasan F, Cai H, Sosa A, Chen F (2019a) Draft genome sequence of a cold-adapted Pseudomonas sp. strain, BGI-2, isolated from the ice of Batura Glacier, Pakistan. Microbiol Resour Announc 8:e00320–e00319. https://doi.org/10.1128/mra.00320-19
Ali P, Hasan F, Khan S, Badshah M, Shah AA (2019b) Cold-adapted halotolerant Rhodococcus sp. BGI-11, candidate for biotechnological applications. Int J Biosci 15:475–489. https://doi.org/10.12692/ijb/15.2.475-489
Ali P, Shah AA, Hasan F, Hertkorn N, Gonsior M, Sajjad W, Chen F (2020) A glacier bacterium produces high yield of cryoprotective exopolysaccharide. Front Microbiol 10:3096. https://doi.org/10.3389/fmicb.2019.03096
Antony R, Krishnan KP, Laluraj CM, Thamban M, Dhakephalkar PK, Engineer AS, Shivaji S (2012) Diversity and physiology of culturable bacteria associated with a coastal Antarctic ice core. Microbiol Res 167:372–380. https://doi.org/10.1016/j.micres.2012.03.003
Arulazhagan P, Vasudevan N (2011) Biodegradation of polycyclic aromatic hydrocarbons by a halotolerant bacterial strain Ochrobactrum sp. VA1. Mar Pollut Bull 62:388–394. https://doi.org/10.1016/j.marpolbul.2010.09.020
Barria C, Malecki M, Arraiano CM (2013) Bacterial adaptation to cold. Microbiology 159:2437–2443. https://doi.org/10.1099/mic.0.052209-0
Bauer AW, Kirby WMM, Sherris JC, Turck M (1966) Antibiotic susceptibility testing by a standardized single disk method. Am J Clin Pathol 45:493–496. https://doi.org/10.1093/ajcp/45.4_ts.493
Boetius A, Anesio AM, Deming JW, Mikucki JA, Rapp JZ (2015) Microbial ecology of the cryosphere: sea ice and glacial habitats. Nat Rev Microbiol 13:677–690. https://doi.org/10.1038/nrmicro3522
Booth C (1971) Methods in microbiology (Vol. 4). Academic Press
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena AG, Goodrich JK, Gordon JI, Huttley GA (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336. https://doi.org/10.1038/nmeth.f.303
Casanueva A, Tuffin M, Cary C, Cowan DA (2010) Molecular adaptations to psychrophily: the impact of ‘omic’ technologies. Trends Microbiol 18:374–381. https://doi.org/10.1016/j.tim.2010.05.002
Cavicchioli R, Charlton T, Ertan H, Omar SM, Siddiqui KS, Williams TJ (2011) Biotechnological uses of enzymes from psychrophiles. Microb Biotechnol 4:449–460. https://doi.org/10.1111/j.1751-7915.2011.00258.x
Cavicchioli R, Siddiqui KS, Andrews D, Sowers KR (2002) Low-temperature extremophiles and their applications. Curr Opin Biotechnol 13:253–261. https://doi.org/10.1016/S0958-1669(02)00317-8
Chattopadhyay MK, Jagannadham MV (2001) Maintenance of membrane fluidity in Antarctic bacteria. Polar Biol 24:386–388. https://doi.org/10.1007/s003000100232
Chen F, Lu JR, Binder B, Hodson RE (2001) Enumeration of viruses in aquatic environments using SYBR gold stain: application of digital image analysis and flow cytometer. Appl Environ Microbiol 67:539–545. https://doi.org/10.1128/AEM.67.2.539-545.2001
Cheng SM, Foght JM (2007) Cultivation-independent and-dependent characterization of bacteria resident beneath John Evans Glacier. FEMS Microbiol Ecol 59:318–330. https://doi.org/10.1111/j.1574-6941.2006.00267.x
Christner BC, Mosley-Thompson E, Thompson LG, Reeve JN (2001) Isolation of bacteria and 16S rDNAs from Lake Vostok accretion ice. Environ Microbiol 3:570–577. https://doi.org/10.1046/j.1462-2920.2001.00226.x
Christner BC, Mosley-Thompson E, Thompson LG, Zagorodnov V, Sandman K, Reeve JN (2000) Recovery and identification of viable bacteria immured in glacial ice. Icarus 144:479–485. https://doi.org/10.1006/icar.1999.6288
Collins T, Margesin R (2019) Psychrophilic lifestyles: mechanisms of adaptation and biotechnological tools. Appl Microbiol Biotechnol 103:2857–2871. https://doi.org/10.1007/s00253-019-09659-5
D’Amico S, Collins T, Marx JC, Feller G, Gerday C (2006) Psychrophilic microorganisms: challenges for life. EMBO Rep 7:385–389. https://doi.org/10.1038/sj.embor.7400662
Dieser M, Greenwood M, Foreman CM (2010) Carotenoid pigmentation in Antarctic heterotrophic bacteria as a strategy to withstand environmental stresses. Arct Antarct Alp Res 42:396–405. https://doi.org/10.1657/1938-4246-42.4.396
Ermakova IT, Shushkova TV, Sviridov AV, Zelenkova NF, Vinokurova NG, Baskunov BP, Leontievsky AA (2017) Organophosphonates utilization by soil strains of Ochrobactrum anthropi and Achromobacter sp. Arch Microbiol 199:665–675. https://doi.org/10.1007/s00203-017-1343-8
Fong N, Burgess M, Barrow K, Glenn D (2001) Carotenoid accumulation in the psychrotrophic bacterium Arthrobacter agilis in response to thermal and salt stress. Appl Microbiol Biotechnol 56:750–756. https://doi.org/10.1007/s002530100739
Ganasen M, Yaacob N, Rahman RNZRA, Leow ATC, Basri M, Salleh AB, Ali MSM (2016) Cold-adapted organic solvent tolerant alkalophilic family I.3 lipase from an Antarctic Pseudomonas. Int J Biol Macromol 92:1266–1276. https://doi.org/10.1016/j.ijbiomac.2016.06.095
Gratia E, Weekers F, Margesin R, D’Amico S, Thonart P, Feller G (2009) Selection of a cold-adapted bacterium for bioremediation of wastewater at low temperatures. Extremophiles 13:763–768. https://doi.org/10.1007/s00792-009-0264-0
Hatosy SM, Martiny AC (2015) The ocean as a global reservoir of antibiotic resistance genes. Appl Environ Microbiol 81:7593–7599. https://doi.org/10.1128/AEM.00736-15
Hewitt K (2005) The Karakoram anomaly? Glacier expansion and the ‘elevation effect’ Karakoram Himalaya. Mt Res Dev 25:332–340. https://doi.org/10.1659/0276-4741(2005)025[0332:TKAGEA]2.0.CO;2
Hewitt K (2014) Glaciers of the Karakoram Himalaya: glacial environments, processes, hazards and resources. Springer, Netherlands. https://doi.org/10.1007/978-94-007-6311-1
Hodson A, Anesio AM, Tranter M, Fountain A, Osborn M, Priscu J, Laybourn-Parry J, Sattler B (2008) Glacial ecosystems. Ecol Monogr 78:41–67. https://doi.org/10.1890/07-0187.1
Hodson A, Porter P, Lowe A, Mumford P (2002) Chemical denudation and silicate weathering in Himalayan glacier basins: Batura Glacier, Pakistan. J Hydrol 262:193–208. https://doi.org/10.1016/S0022-1694(02)00036-7
Horikoshi K, Antranikian G, Bull AT, Robb FT, Stetter KO (2010) Extremophiles handbook. Springer, Berlin
Kasana RC, Salwan R, Dhar H, Dutt S, Gulati A (2008) A rapid and easy method for the detection of microbial cellulases on agar plates using Gram’s iodine. Curr Microbiol 57:503–507. https://doi.org/10.1007/s00284-008-9276-8
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, Glöckner FO (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41:e1–e1. https://doi.org/10.1093/nar/gks808
Liu Y, Priscu JC, Yao T, Vick-Majors TJ, Michaud AB, Sheng L (2018) Culturable bacteria isolated from seven high-altitude ice cores on the Tibetan Plateau. J Glaciol 65:29–38. https://doi.org/10.1017/jog.2018.86
Liu Y, Yao T, Jiao N, Kang S, Xu B, Zeng Y, Huang S, Liu X (2009) Bacterial diversity in the snow over Tibetan Plateau Glaciers. Extremophiles 13:411–423. https://doi.org/10.1007/s00792-009-0227-5
Lopez EG, Rodriguez-Lorente I, Alcazar P, Cid C (2018) Microbial communities in coastal glaciers and tidewater tongues of Svalbard Archipelago, Norway. Front Mar Sci 5:512. https://doi.org/10.3389/fmars.2018.00512
Margesin R, Miteva V (2011) Diversity and ecology of psychrophilic microorganisms. Res Microbiol 162:346–361. https://doi.org/10.1016/j.resmic.2010.12.004
Margesin R, Gander S, Zacke G, Gounot AM, Schinner F (2003) Hydrocarbon degradation and enzyme activities of cold-adapted bacteria and yeasts. Extremophiles 7:451–458. https://doi.org/10.1007/s00792-003-0347-2
Marzorati M, Wittebolle L, Boon N, Daffonchio D, Verstraete W (2008) How to get more out of molecular fingerprints: practical tools for microbial ecology. Environ Microbiol 10:1571–1581. https://doi.org/10.1111/j.1462-2920.2008.01572.x
Menéndez E, Ramírez-Bahena MH, Fabryová A, Igual JM, Benada O, Mateos PF, Peix A, Kolařík M, García-Fraile P (2015) Pseudomonas coleopterorum sp. nov., a cellulase-producing bacterium isolated from the bark beetle Hylesinus fraxini. Int J Syst Evol Microbiol 65:2852–2858. https://doi.org/10.1099/ijs.0.000344
Miteva VI, Sheridan PP, Brenchley JE (2004) Phylogenetic and physiological diversity of microorganisms isolated from a deep Greenland glacier ice core. Appl Environ Microbiol 70:202–213. https://doi.org/10.1128/aem.70.1.202-213.2004
Moyer CL, Morita RY (2007) Psychrophiles and psychrotrophs. John Wiley and Sons, Ltd. eLS. https://doi.org/10.1002/9780470015902.a0000402.pub2
O'Brien A, Sharp R, Russell NJ, Roller S (2004) Antarctic bacteria inhibit growth of food-borne microorganisms at low temperatures. FEMS Microbiol Ecol 48:157–167. https://doi.org/10.1016/j.femsec.2004.01.001
Priest FG (1977) Extracellular enzyme synthesis in the genus Bacillus. Bacteriol Rev 41:711–753
Rafiq M, Hayat M, Anesio AM, Jamil S, Hassan N, Shah AA, Hasan F (2017) Recovery of metallo-tolerant and antibiotic resistant psychrophilic bacteria from Siachen glacier, Pakistan. PLoS One 12:e0178180. https://doi.org/10.1371/journal.pone.0178180
Rankl M, Kienholz C, Braun M (2014) Glacier changes in the Karakoram region mapped by multimission satellite imagery. Cryosphere 8:977–989. https://doi.org/10.5194/tc-8-977-2014
Rassner SM, Anesio AM, Girdwood SE, Hell K, Gokul JK, Whitworth DE, Edwards A (2016) Can the bacterial community of a high Arctic glacier surface escape viral control? Front Microbiol 7:956. https://doi.org/10.3389/fmicb.2016.00956
Rasul G, Chaudhry QZ, Mahmood A, Hyder KW, Dahe Q (2011) Glaciers and glacial lakes under changing climate in Pakistan. Pakisan J Meteorol 8:1–8
Reasoner DJ, Geldreich EE (1985) A new medium for the enumeration and subculture of bacteria from potable water. Appl Environ Microbiol 49:1–7
Reddy PVV, Rao SSSN, Pratibha MS, Sailaja B, Kavya B, Manorama RR, Singh SM, Srinivas TNR, Shivaji S (2009) Bacterial diversity and bioprospecting for cold-active enzymes from culturable bacteria associated with sediment from a melt water stream of Midtre Lov´ enbreen glacier, an Arctic glacier. Res Microbiol 160:538–546. https://doi.org/10.1016/j.resmic.2009.08.008
Rodrigues DF, Tiedje JM (2008) Coping with our cold planet. Appl Environ Microbiol 74:1677–1686. https://doi.org/10.1128/AEM.02000-07
Russell NJ (1990) Cold adaptation of microorganisms. Philos Trans R Soc Lond Ser B Biol Sci 326:595–611. https://doi.org/10.1098/rstb.1990.0034
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Sánchez LA, Gómez FF, Delgado OD (2009) Cold-adapted microorganisms as a source of new antimicrobials. Extremophiles 13:111–120. https://doi.org/10.1007/s00792-008-0203-5
Irvine-Fynn TD, Edwards A (2014) A frozen asset: the potential of flow cytometry in constraining the glacial biome. Cytometry Part A 85:3–7. https://doi.org/10.1002/cyto.a.22411
Segawa T, Takeuchi N, Rivera A, Yamada A, Yoshimura Y, Barcaza G, Shinbori K, Motoyama H, Kohshima S, Ushida K (2013) Distribution of antibiotic resistance genes in glacier environments. Environ Microbiol Rep 5:127–134. https://doi.org/10.1111/1758-2229.12011
Segawa T, Yoshimura Y, Watanabe K, Kanda H, Kohshima S (2011) Community structure of culturable bacteria on surface of Gulkana Glacier, Alaska. Polar Sci 5:41–51. https://doi.org/10.1016/j.polar.2010.12.002
Shen L, Liu Y, Wang N, Jiao N, Xu B, Liu X (2018) Variation with depth of the abundance, diversity and pigmentation of culturable bacteria in a deep ice core from the Yuzhufeng Glacier, Tibetan Plateau. Extremophiles 22:29–38. https://doi.org/10.1007/s00792-017-0973-8
Shivaji S, Pratibha MS, Sailaja B, Kishore KH, Singh AK, Begum Z, Anarasi U, Prabagaran SR, Reddy GSN, Srinivas TNR (2011) Bacterial diversity of soil in the vicinity of Pindari glacier, Himalayan mountain ranges, India, using culturable bacteria and soil 16S rRNA gene clones. Extremophiles 15:1–22. https://doi.org/10.1007/s00792-010-0333-4
Silva TR, Duarte AW, Passarini MR, Ruiz ALT, Franco CH, Moraes CB, de Melo IS, Rodrigues RA, Fantinatti-Garboggini F, Oliveira VM (2018) Bacteria from Antarctic environments: diversity and detection of antimicrobial, antiproliferative, and antiparasitic activities. Polar Biol 41:1505–1519. https://doi.org/10.1007/s00300-018-2300-y
Steven B, Briggs G, McKay CP, Pollard WH, Greer CW, Whyte LG (2007) Characterization of the microbial diversity in a permafrost sample from the Canadian high Arctic using culture-dependent and culture-independent methods. FEMS Microbiol Ecol 59:513–523. https://doi.org/10.1111/j.1574-6941.2006.00247.x
Tam HK, Wong CMVL, Yong ST, Blamey J, González M (2015) Multiple-antibiotic-resistant bacteria from the maritime Antarctic. Polar Biol 38:1129–1141. https://doi.org/10.1007/s00300-015-1671-6
Tedesco P, Maida I, Palma Esposito F, Tortorella E, Subko K, Ezeofor C, Zhang Y, Tabudravu J, Jaspars M, Fani R, de Pascale D (2016) Antimicrobial activity of monoramnholipids produced by bacterial strains isolated from the Ross Sea (Antarctica). Mar Drugs 14:83. https://doi.org/10.3390/md14050083
Thakur V, Kumar V, Kumar S, Singh D (2018) Diverse culturable bacterial communities with cellulolytic potential revealed from pristine habitat in Indian trans-Himalaya. Can J Microbiol 64:798–808. https://doi.org/10.1139/cjm-2017-0754
Tribelli P, López N (2018) Reporting key features in cold-adapted bacteria. Life 8:8. https://doi.org/10.3390/life8010008
Van Goethem MW, Pierneef R, Bezuidt OK, Van De Peer Y, Cowan DA, Makhalanyane TP (2018) A reservoir of ‘historical’ antibiotic resistance genes in remote pristine Antarctic soils. Microbiome 6:40. https://doi.org/10.1186/s40168-018-0424-5
Venkatachalam S, Gowdaman V, Prabagaran SR (2015) Culturable and culture-independent bacterial diversity and the prevalence of cold-adapted enzymes from the Himalayan mountain ranges of India and Nepal. Microb Ecol 69:472–491. https://doi.org/10.1007/s00248-014-0476-4
Yang GL, Hou SG, Le Baoge R, Li ZG, Xu H, Liu YP, Du WT, Liu YQ (2016) Differences in bacterial diversity and communities between glacial snow and glacial soil on the chngce ice cap, west Kunlun Mountains. Sci Rep 6:36548. https://doi.org/10.1038/srep36548
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y, Seo H, Chun J (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol 67:1613–1617. https://doi.org/10.1099/ijsem.0.001755
Zeng R, Zhang R, Zhao J, Lin N (2003) Cold-active serine alkaline protease from the psychrophilic bacterium Pseudomonas strain DY-A: enzyme purification and characterization. Extremophiles 7:335–337. https://doi.org/10.1007/s00792-003-0323-x
Zhang DC, Brouchkov A, Griva G, Schinner F, Margesin R (2013) Isolation and characterization of bacteria from ancient Siberian permafrost sediment. Biology 2:85–106. https://doi.org/10.3390/biology2010085
Zhang XF, Yao TD, Tian LD, Xu SJ, An LZ (2008) Phylogenetic and physiological diversity of bacteria isolated from Puruogangri ice core. Microb Ecol 55:476–488. https://doi.org/10.1007/s00248-007-9293-3
Zhao Y, Song C, Dong H, Luo Y, Wei Y, Gao J, Wu Q, Huang Y, An L, Sheng H (2018) Community structure and distribution of culturable bacteria in soil along an altitudinal gradient of Tianshan Mountains, China. Biotechnol Biotechnol Equip 32:397–407. https://doi.org/10.1080/13102818.2017.1396195
Acknowledgments
We gratefully acknowledge the International Research Support Initiative Program–Higher Education Pakistan (IRSIP–HEC) for providing fellowship to conduct this research in the Institute of Marine and Environmental Technology (IMET), University of Maryland Center for Environmental Sciences (UMCES) Maryland, USA.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(DOCX 33 kb)
Rights and permissions
About this article
Cite this article
Ali, P., Chen, F., Hassan, F. et al. Bacterial community characterization of Batura Glacier in the Karakoram Range of Pakistan. Int Microbiol 24, 183–196 (2021). https://doi.org/10.1007/s10123-020-00153-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10123-020-00153-x