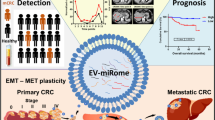
Abstract
Background
Peritoneum, liver and lymph node are the most common metastatic sites of gastric cancer (GC). Biomarkers for GC’s organo-tropic metastasis remained largely unknown, which was investigated in this study from the perspective of small extracellular vesicle (sEV)-derived miRNAs.
Methods
Plasma from treatment-naïve GC patients including no metastasis (M0), peritoneal metastasis (PM), hepatic metastasis (HM) and distant lymph node metastasis (dLNM)) were divided into one discovery (N = 40), one training (N = 40) and one validating cohort (N = 86), then assessed by sEV-miRNA-sequencing and sEV-miRNA-qPCR. Functional explorations were also performed for verification.
Results
The expression profiles of sEV-miRNAs varied greatly across different metastatic patterns. Based on logistic regression models, we constructed signatures for M0 (hsa-miR-186-5p/hsa-miR-200c-3p/hsa-miR-429/hsa-miR-5187-5p/hsa-miR-548ae-5p), PM (hsa-miR-200c-3p/hsa-miR-429), HM (hsa-miR-200c-3p/hsa-miR-429) and dLNM (hsa-miR-324-5p/hsa-miR-374a-5p/hsa-miR-429/hsa-miR-548ae-5p). These signatures vigorously characterized organo-tropic metastasis (all displaying AUC > 0.8, consistency ≥ 75%), and effectively conjectured the risk of future metastasis within 5 years (accuracy 45.5% for occurrence, 70% for organotropism, P = 0.002 for prognostic diversity). Additionally, we explored these seven biomarker miRNAs’ impact on GC’s in vitro motility and discussed their potential involvement in cancer-related biological processes and pathways.
Conclusions
Our work highlighted that plasma sEV-miRNAs powerfully characterized and predicted the organo-tropic metastasis of GC and provided new insight into the applications of sEV-based liquid biopsy in clinical practice.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As one of the most representative hallmarks of cancer, metastasis accelerated malignant progression through facilitating the dissemination of cancer cells and is thus regarded as the greatest contributors to cancer death [1]. The fact that metastasis relies on the interplay between cancer cells (the “seeds”) and appropriate microenvironment (the “soil”) for attachment, colonization and re-dissemination emphasized that the organ/site-specific distribution of distant metastasis, or metastatic organotropism, was shaped by diverse molecular bases and could be characterized by specific biomarkers [2, 3]. Metastatic tumors usually generate new genetic alterations during evolvement and develop biological traits different from primary tumors [4]. Thus, in order to select appropriate clinical management for patients, it is crucial to develop biomarkers specifically predicting metastatic organotropism of cancer.
Gastric cancer (GC), the fifth most prevalent type of cancer and the third leading cause of cancer death around the world [5], is often diagnosed at advanced stages, bears high metastatic risk, and exhibits a poor prognosis [6]. GC’s most prevalent metastatic patterns were peritoneal metastasis (PM), hepatic metastasis (HM) and distant-lymph node metastasis (dLNM) [7,8,9]. Frequently observed in abdominal-occupying malignancies, peritoneal metastasis accounts for around 40% of GC metastasis and is often accompanied by ascites and occult lesions that lead to cancer death [10]. Liver, connected with stomach by hepatic portal vein, is the organ where hematogenous metastasis of GC takes place, which shared a proportion of around 40% in all GC metastasis [11]. Scattered all over the human body, lymph nodes distant to stomach are also usually found to be nested with metastatic lesions of GC [12]. GC patients with each of the three representative forms of metastasis lose the chance of radical resection and have a poor survival [9, 13, 14]. Subsequent to systemic chemotherapy, the development of targeted therapy and immunotherapy ignited new hope for the precision medication of advanced GC, yet the standardized management for unresectable metastatic GC still achieves unsatisfying responses [15, 16]. Therefore, the timely detection of metastatic organotropism will provide patients with more therapeutic options and better outcomes. However, due to the complicated anatomy of the stomach, current detection methods (such as computed tomography, CT) for GC metastasis are insufficient to fulfill the need of real-time diagnosis and are limited in characterizing occult lesions. Consequently, it is important to develop new methods to distinguish the organo-tropic metastatic modes of GC.
Small extracellular vesicles (sEVs, also known as exosomes) are membrane bounding, nanosized particles released by cells. sEVs are detected in almost all body fluids and contain multiple bioactive substances including DNAs/mRNAs/noncoding RNAs/proteins [17]. Travelling along with local or systemic blood/lymph circulation, sEV transfers specific cargos and directed cell–cell or cell–environment communications [18]. Recent studies pointed out the miRNAs are enriched in cancer-associated sEVs and play key roles in mediating malignant transformation, drug resistance, and metastasis [19], suggesting that sEV-derived miRNAs can be effectively used as metastatic biomarkers in liquid biopsy [20]. However, sEV-derived biomarkers addressing the metastatic organotropism of GC have not yet been systemically reported. In this study, we screened the expressional profiles of the sEV-derived miRNAs in patient plasma and explored their correlations with the most representative metastatic organotropism of GC.
Materials and methods
Specimen collection and information
Patients diagnosed with unresectable or metastatic GEJ (gastroesophageal junction) or non-GEJ GC during Aug 2014 to Sep 2019 by the Department of Gastrointestinal Oncology, Peking University Cancer Hospital & Institute were proceeded for selection. To rule out the interference of therapeutic management on plasma sEV’s quantity and content, enrolled patients must be naïve for surgery or any anti-cancer medical therapies. Patients harboring GEJ/non-GEJ combined with other cancer types, histopathologically diagnosed as neuroendocrine tumor or squamous type of tumor, or diagnosed with active hepatitis B were excluded. Metastatic locations of patients were comprehensively assessed by experienced radiologists/pathologists with CT/laparoscopy/biopsy or peritoneal lavage cytology, and diagnosed by referring to the Guidelines of Chinese Society of Clinical Oncology (CSCO), Gastric Cancer (2020 edition). A total amount of 166 patients were enrolled in this study, including 42 M0, 31 PM, 34 HM, 34 dLNM and 25 MM cases (10 PM + dLNM, 10 HM + dLNM, 5 PM + HM + dLNM), then randomly allocated into one 40-case discovery cohort, one 40-case training cohort and one 86-case validating cohort. The procedures of patient selection and organotropism classification were demonstrated in Supplementary Fig. S1.
To ensure the accuracy of metastasis organotropism, the characterized metastatic status must remain unchanged for at least 3 months after blood sampling in order for each patient to be selected. Each peripheral whole blood specimen was collected from the donor after the definite diagnosis of metastasis status and before the administration of subsequent anti-cancer therapies. Paired clinical information was obtained from the medical records of patients.
Experimental and analytical procedures
Details for experimental and analytical procedures, including plasma isolation, sEV extraction, nanoparticle tracking analysis (NTA), transmission electron microscopy (TEM), sEV miRNA extraction/quantification and real-time qPCR, library preparation and sequencing, processing of sequencing data, degradation assay, western blot assay, cell lines, miRNA transfection, wound healing assay, invasion assay and viability assay, are provided in Supplementary Materials and Methods section. Primer and probe sequences are listed in Supplementary Table S2.
Statistical analysis and formatting
Details for statistical analysis and formatting are provided in Supplementary Materials and Methods section.
Results
Characterization of patients and sEV-extracted fractions
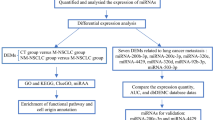
As summarized by an overall workflow (Fig. 1), a 40-patient discovery cohort (including 10 M0/10 PM/10 HM/10 dLNM cases) was assessed by sEV-related miRNA sequencing, while a 40-patient training cohort (10 M0/10 PM/10 HM/10 dLNM) and an 86-patient validating cohort (22 M0/11 PM/14 HM/14 dLNM/25 MM (mixed-site metastasis, including 10 PM + dLNM/10 HM + dLNM/5 PM + HM + dLNM)) were assessed by sEV-related miRNA qPCR. Patients in all three cohorts displayed comparable demographic characteristics (Table 1), which minimized the confounding bias across different cohorts.
The workflow of the whole study. miRNA biomarkers for organo-tropic metastasis were screened with miRNA-sequencing from a 40-case discovery cohort, then analyzed with miRNA RT-qPCR (with three replicates) in a 40-case training cohort and an 86-case validating cohort, then proceeded for additional analysis
Plasma-borne sEVs for all samples were isolated with SEC method. As inspected with TEM and NTA methods, isolated sEVs were clear-shaped ovoid or spheres with a diameter that ranged between 50 and 200 nm (Supplementary Fig. S2A–S2B). sEV markers (Alix/CD9/CD63) were enriched while the cellular marker calnexin was absent in the extracted fractions compared with HEK293T cell lysate/total plasma (Supplementary Fig. S2C), suggesting isolated sEVs were of good quality and purity. An average sEV concentration (18.22 ng/mL) was quantified across all cohorts, in which M0/PM/HM/dLNM groups displayed comparable concentrations, while mixed-site metastasis group displayed higher concentrations of sEV than single-metastasis groups (Supplementary Fig. S2D).
The expressional landscape of sEV-miRNAs across GC’s organo-tropic metastasis
Plasma-borne sEV specimens in the 40-patient discovery cohort were assessed by miRNA sequencing. With an average clean read of 13.74 M for each sample, a total of 1997 miRNAs (1636 known/361 novel) were identified across the cohort. In order to minimize the systemic bias induced by unevenly distributed miRNA expressions, only those miRNAs with median TPM ≥ 10 were included for further analysis. We noticed through t-SNE analysis that sEV miRNAs shared similar expressional profiles between M0 and PM or between HM and dLNM groups (Supplementary Fig. S3A), which emphasized that peritoneal metastasis had more involvement with the local tumor microenvironment in abdominal cavity, while hepatic metastasis and distant-lymph node metastasis were more likely to follow a hematogenous manner [3]. With fold changes (FC) ≥ 1.5 and ≤ 1/1.5 and Mann–Whitney U test P ≤ 0.01 as the threshold, differentially expressed miRNAs (DEMs) were screened for all metastatic sites. The amount of upregulated/downregulated DEMs were 8/47 for M0 vs others (PM/HM/dLNM), 12/3 for PM vs others (M0/HM/dLNM), 13/2 for HM vs others (M0/PM/dLNM), 6/0 for dLNM vs others (M0/PM/HM) and 9/33 for HM/dLNM vs M0/PM (Supplementary Fig. S3B, Table S1). The phenomenon that M0 possessed more specifically downregulated DEMs while other groups possessed more specifically upregulated DEMs suggested an enrichment of upregulated sEV miRNAs after the occurrence of distant metastasis.
By filtering DEMs sharing a high correlation (R2 > 0.5) with the rest miRNAs in the same group, 23 candidate miRNAs (hsa-miR-127-3p/hsa-miR-1277-5p/hsa-miR-141-3p/hsa-miR-148b-3p/hsa-miR-17-5p/hsa-miR-186-5p/hsa-miR-191-5p/hsa-miR-193a-5p/hsa-miR-200a-3p/hsa-miR-200b-3p/hsa-miR-200c-3p/hsa-miR-203a-3p/hsa-miR-22-3p/hsa-miR-30e-5p/hsa-miR-324-5p/hsa-miR-3613-5p/hsa-miR-374a-5p/hsa-miR-429/hsa-miR-485-3p/hsa-miR-5187-5p/hsa-miR-548ae-5p/hsa-miR-548b-5p/hsa-miR-574-3p) were identified. t-SNE analysis validated these 23 miRNAs also shared approximate expressional profiles between M0 and PM or between HM and dLNM (Supplementary Fig. S4A), while hierarchical clustering emphasized these 23 miRNAs could outline M0/PM/HM/dLNM metastatic patterns in training cohort (Supplementary Fig. S4B). The specificity of these candidate DEMs for each metastatic group were compared (Supplementary Fig. S4C).
Selection of sEV-derived biomarker miRNAs for metastatic organotropism
After removing three highly homologous candidates (hsa-miR-200a-3p/hsa-miR-200b-3p/hsa-miR-548b-5p), the remaining 20 sEV-derived DEMs were selected as the representative miRNAs across M0/PM/HM/dLNM groups. In order to concise the panel and to improve its feasibility in identifying organo-tropic metastasis, we enrolled another 40 GC patients (including 10 M0/10 PM/10 HM/10 dLNM) as the training cohort. After plasma-sEV isolation and miRNA extraction, all specimens in this cohort were quantified with RT-qPCR for the 20 panel DEMs. Since U6 (the most commonly used miRNA internal control) may not be well-functioning for miRNA quantification in terms of sEV, hsa-miR-140-5p (the most stably expressed sEV miRNA across all metastatic classifications in discovery cohort) was compared with U6 for the efficacy as internal control in the training cohort (Fig. 2A). hsa-miR-140-5p generated more significant overall variances of the 20 DEMs than U6 as the internal control (Fig. 2B); thus we measured the expressions of the 20 miRNAs with hsa-miR-140-5p as the internal control while cel-miR-39 as the external control. The group-specificity for the adjusted expressional profiles of 20 DEMs was measured with Shapiro test and Bartlett test paired with ANOVA test, or Shapiro test and Bartlett test paired with Kruskal test. Seven miRNAs (hsa-miR-186-5p/hsa-miR-200c-3p/hsa-miR-324-5p/hsa-miR-374a-5p/hsa-miR-429/hsa-miR-5187-5p/hsa-miR-548ae-5p) were found displaying the top-half of variances (reflected by the Final P-value) across different metastatic groups in training cohort (Table 2), simultaneously included in one or more organo-tropic combinations exported by a step logistic regression model (described in following parts). These 7 miRNAs also shared similar expressional profiles between discovery and training cohorts (Supplementary Fig. S5); thus they were selected as the biomarker miRNAs for GC’s metastatic organotropism (Fig. 2C).
Selection of biomarker miRNAs for metastatic organotropism. A The average Ct value of 20 candidate miRNAs and 3 reference miRNAs in training cohort were measured by qPCR. B The overall variance of 20 candidate miRNAs was adjusted with three referenced miRNAs and measured by ANOVA and Kruskal P value. C The hsa-miR-140-5p and cel-miR-39-adjusted expression profile of 20 miRNAs was displayed, in which 7 miRNAs (red mark) with the highest variance were selected as organo-tropic biomarkers
Selected biomarker miRNAs were predominantly originated from sEV
Since plasma miRNAs could be in both sEV-capsulated and non-SEV forms, it is vital to clarify their predominant forms of existence in order for potential diagnostic applications. Degradation assay was performed to verify the origination of the 7 biomarker miRNAs. Total plasma or isolated sEV samples from M0/PM/HM/dLNM patients were mixed and treated with proteinase K/RNase A, then processed for RNA/protein quantification. Both RNA concentration and protein density were reduced for total plasma samples while isolated sEV samples were largely unaffected (Fig. 3A, B), confirming that sEV-originated RNA/protein contents were protected from degradation.
Identifying the sEV-origination of organo-tropic biomarker miRNAs. For representative samples, changes of A RNA concentration and B protein contents in total plasma or isolated sEV were assessed after degradation assay. C Expression of 7 biomarker miRNAs as well as 2 control miRNAs was quantified by qPCR (with three replicates) after degradation assay. PK proteinase K; RA RNase A; p total plasma; s sEV
Total plasma or isolated sEV samples pretreated with degradation assay were then quantified for the 7 biomarker miRNAs as well as two controls (hsa-miR-21-5p/hsa-miR-30a-5p) specifically expressed in circulating forms. Compared with controls, hsa-miR-186-5p/hsa-miR-200c-3p/hsa-miR-324-5p/hsa-miR-374a-5p/hsa-miR-5187-5p displayed higher degraded proportions in total plasma than in sEV, suggesting they had higher concentrations of sEV-capsulated form than non-sEV form in plasma, while only a small fraction of hsa-miR-429/hsa-miR-548ae-5p were degraded in both total plasma and sEV (Fig. 3C), indicating they existed majorly in sEV-capsulate form. These findings verified all 7 biomarker miRNAs were predominantly originated from sEV.
sEV-miRNA-based signatures effectively characterized metastatic organotropism of GC
ROC analysis showed that the AUC for 7 miRNAs in correlation with M0/PM/HM/dLNM were relatively low (between 0.49–0.8) and non-organo-tropic (Supplementary Fig. S6A). For improvement, we applied a step logistic regression model and referred to the AIC (Akaike information criterion) to evaluate miRNA combinations. We found that the combination of hsa-miR-186- 5p/hsa-miR-200c-3p/hsa-miR-429/hsa-miR-5187-5p/hsa-miR-548ae-5p displayed M0 specificity, the combinations of hsa-miR-200c-3p/hsa-miR-429 (with different coefficients) displayed PM and HM specificity, while the combination of hsa-miR-324-5p/hsa-miR-374a-5p/hsa-miR-429/hsa-miR-548ae-5p displayed dLNM specificity. We defined each combination as an organotropic signature and evaluated their efficacy in representing corresponding metastatic patterns. According to ROC analysis, the AUC for M0/PM/HM/dLNM signatures were 0.91/0.81/0.8567/0.9567 (Fig. 4B), displaying high effectiveness in specifying respective metastatic organotropism.
miRNA signatures efficiently characterized GC’s metastatic organotropism. The power of M0, PM, HM and dLNM signatures in characterizing corresponding organo-tropic metastasis was measured by ROC analysis in A discovery cohort, B training cohort and C validating cohort. The consistency between practical and signature-predicted metastatic organotropism were compared in D discovery cohort, E training cohort and F validating cohort
We then introduced a validating cohort of GC patients for verification, which contained 22 M0/11 PM/14 HM/14 dLNM and 25 MM cases (10 PM + dLNM, 10 HM + dLNM, 5 PM + HM + dLNM). Similar to training cohort, the 7 individual miRNAs were insufficient to characterize (Supplementary Fig. S6B), while combined signatures were powerful to characterize organo-tropic metastasis in validating cohort (the AUC for M0/PM/HM/dLNM signatures were 0.9033/0.9273/0.8328/0.9635, Fig. 4C). These signatures also displayed high efficacy in characterizing organo-tropic metastasis in discovery cohort (the AUC for M0/PM/HM/dLNM signatures were 0.8833/0.9067/0.8767/0.9133, Fig. 4A).
Additionally, metastatic organotropism predicted by sEV-derived miRNA signatures shared high consistency with the practical metastatic organotropism in discovery (77.5%, Fig. 4D), training (75%, Fig. 4E) and validating cohorts (75.41% for single-metastasis cases, 76% for multi-metastatic cases, Fig. 4F), indicating that these signatures could help locating untraceable lesions or occult metastasis for GC patients.
sEV miRNA signatures conjectured metastatic risk and organotropism for M0 patients
Apart from characterizing existing metastasis, it is equally important to predict the risk of future distant metastasis for M0 patients. In this study, 42 M0 patients were recruited across discovery/training/validating cohorts. Using a logistic regression model, we compared the signature values of PM/HM/dLNM to measure each M0 case’s risk to develop peritoneal/hepatic/distant lymph node metastasis during 5-year follow-up. According to the rank of these three signature scores, 22 patients were classified as high-risk while another 20 were classified as low-risk for future distant metastasis (Fig. 5A, upper panel). Practically, 45.5% (10/22) of high-risk patients developed metastasis during the following 5 years, while no future metastasis (0%, 0/20) was observed for low-risk patients (Fig. 5A, lower panel). High-risk patients also displayed a significantly degenerated DMFS (distant metastasis-free survival) than low-risk patients (Fig. 5B). High concordances were achieved by comparing predicted and practical metastatic organotropism (70% (7/10) for all patients that developed distant metastasis, including 100% (3/3) for PM, 50% (1/2) for HM and 60% (3/5) for dLNM) (Fig. 5A). CT images for 3 representative M0 cases before/after the development of PM/HM/dLNM were exhibited (Fig. 5C).
Organo-tropic miRNA signatures conjectured future metastatic risk for M0 patients. For 42 M0 patients across all three cohorts, A the metastatic risk and organotropism were predicted based on miRNA signatures (upper panel) and compared with the clinical practices in following 5 years (lower panel). B The 5-year DMFS of 42 M0 patients were stratified by signature-based metastatic risk scores. C Representative CT images for three M0 cases before and after developed PM/HM/dLNM. Metastatic lesions in M1 images and the corresponding location in M0 images were marked with yellow arrow
Since two cases in 42 have follow-up plasma samples collected at/after dLNM, we also assessed the time course of sEV miRNA expressions as well as organo-tropic metastatic risk for these two cases. The expression profile of 7 biomarker miRNAs displayed similar trend of changes from M0 to dLNM for both cases (Supplementary Fig. S7A). The risk for dLNM was predicted in case 17,684 at baseline (M0) and remained unchanged at/after practical dLNM, while although case 11,165 was inadequately diagnosed as with HM risk at baseline, it was correctly defined as with dLNM risk at/after practical dLNM (Supplementary Fig. S7B). These findings emphasized that sEV derived miRNA signatures estimate future metastatic risk and metastatic patterns for GC.
In addition, among the 42 patients that underwent prediction of metastasis, 23 received neoadjuvant chemotherapy (SOX/POS/XELOX) plus surgery, 16 received chemotherapy (SOX/POS/XELOX), while 3 received immunotherapy plus chemotherapy after the sampling of peripheral blood. The DMFS (Supplementary Fig. S8) and predicted/practical metastasis concordance (Fig. 5A) for treatment-stratified subgroups were similar to the previous conclusions for all 42 M0 patients, indicating that anti-cancer therapies have minimal impact on the prediction of GC metastasis. Collectively, our sEV miRNA signatures displayed considerable potential in conjecturing organo-tropic metastasis.
Functional implications of biomarker miRNAs
We first introduced 20 plasma samples from healthy people to compare the qPCR-quantified plasma-sEV expression profiles of the 7 biomarker miRNAs between GC (combining training/validating cohorts) and healthy control, the result of which suggested that hsa-miR-374a-5p/hsa-miR-429/hsa-miR-5187-5p/hsa-miR-548ae-5p displayed higher cancer-specificity (Supplementary Fig. S9). We then explored the functional implications of the 7 biomarker miRNAs by assessing their impact on GC phenotypes. GC cell lines AGS/HGC27 were transfected with mimics of the 7 miRNAs (Fig. 6A, B) and then evaluated by wound healing/invasion/viability assay. Transfecting hsa-miR-374a-5p elicited an elevation of wound healing capability/invasiveness/proliferation, while transfecting other miRNAs impaired wound healing/invasiveness/proliferation on diverse degrees (Fig. 6C, D, Supplementary Fig. S10A–S10B), suggesting that in addition to being metastatic biomarkers, these miRNAs were also involved in the modulation of GC’s malignancy. We referred to GO/KEGG databases to explore their mechanistic implications. On individual level, 5 miRNAs (hsa-miR-186-5p/hsa-miR-324-5p/hsa-miR-429/hsa-miR-5187-5p/hsa-miR-548ae-5p) were significantly enriched to certain target genes and pathways predominantly involved in metabolism and estrogen receptor signaling (Fig. 7A, B). Also, M0, PM, HM and dLNM signatures were also dominantly enriched in metabolic terms (Fig. 7C) and pathways (Fig. 7D), suggesting that these miRNAs might be involved in GC metastasis in a metabolism-associated manner. Nevertheless, their functional insights into GC metastasis need to be addressed by further studies.
Discussion
Specific molecular features for the metastatic organotropism of GC deserved to be clarified for the sake of early diagnosis and treatment selection. Possessing the advantage of non-invasive and dynamic sampling, omics-powered liquid biopsy exhibited promising applications for the diagnosis and decision-making of cancer. Current study of liquid biopsy majorly focused on circulating tumor DNA (ctDNA) and circulating tumor cells (CTCs), while the application of sEV contents in liquid biopsy remained largely unexplored, especially in GC [21, 22]. sEV carries crucial biological substances, such as DNAs/RNAs/proteins [23]. Compared with ctDNA, sEV miRNAs provide expressional information and thus expand the potential for subsequent mechanistic investigations. Additionally, sEV miRNAs are much more stable than cell and protein contents due to their nature as small nucleotides and thus maintain higher availability than CTCs for retrospective study of samples that underwent long-term or nonstandard storage [24]. Therefore, we believed that sEV-derived miRNA testing is an ideal option of liquid biopsy for clinical diagnosis and surveillance.
In this study, by referring to the plasma specimens from treatment naïve GC patients, we analyzed the sEV-derived miRNA expressional spectrums for representative metastatic patterns (M0/PM/HM/dLNM). Both single- and multi-metastatic patients were enrolled for investigation. In order to minimize the bias from regional lymph node metastasis, only non-regional distant lymph node metastases were included for the dLNM or dLNM-related groups. Across all plasma samples, mixed-site metastasis group displayed higher sEV concentrations than single metastasis groups. Since multiple-metastatic patients were usually at advanced stages, their heavy tumor burden may contribute to the high concentration of plasma sEV. Differentially expressed miRNAs across metastatic groups were selected by fulfilling the following three additional criteria: 1. abundantly expressed in plasma; 2. with significant fold-changes between one metastatic group and others; 3. with a high expressional correlation with the rest of differentially expressed miRNAs. We noticed that the expressional profiles of sEV miRNA were similar between M0 and PM or between HM and dLNM, in accordance with the current clinical consensus, which could be due to the mechanistic diversity for different metastatic patterns. Peritoneum is anatomically adjacent to stomach, enriched in lymphatic vessels, and was thus easily invaded and adhered by regionally disseminated GC cells, while hepatic or distant-lymph node metastasis more likely originated from circulating tumor cells in a hematogenous manner [3, 25]. We selected hsa-miR-140-5p instead of U6 as qPCR control for more effective quantification of sEV miRNAs and selected 7 sEV miRNAs as representative metastatic biomarkers by further considering intergroup variance.
The majority of these 7 miRNAs were associated with cancer development, among which hsa-miR-186-5p/hsa-miR-324-5p/hsa-miR-429 were tumor suppressive, hsa-miR-374a-5p was oncogenic, while both tumor-suppressive and -promoting roles were reported for hsa-miR-200c-3p. hsa-miR-186-5p was reported to be prognostic and playing a tumor-suppressive role in multiple malignancies. It inhibited lung cancer proliferation by inducing G1–S checkpoint arrest [26] and maintained sensitivity to chemotherapy agents in ovarian cancer [27]. In GC, has-miR-186-5p suppressed aerobic glycolysis and PD-L1 expression by targeting HIF-1α [28]. hsa-miR-324-5p induced apoptosis, inhibited viability/in vivo tumorigenesis of GC through regulating TSPAN8 [29] or suppressed migration/invasion via targeting WNT2B in nasopharyngeal carcinoma [30]. It was also found playing a tumor-suppressive role in colorectal and breast cancer [31]. hsa-miR-429 targeted ZEB1 and exerted tumor-suppressive effects in osteosarcoma and pancreatic cancer [32, 33]. In GC, hsa-miR-429 inhibited Bcl-2-mediated tumor survival, restored apoptosis and alleviated chemoresistance [34]. hsa-miR-374a-5p was implicated as an oncogenic miRNA in colorectal, non-small cell lung and breast cancers [35,36,37]. Similarly, in GC, hsa-miR-374a-5p promoted proliferation/migration/invasion by targeting SRCIN1 [38], predicted poor prognosis and promoted chemoresistance through inhibiting Neurod1 [39]. A series of studies pointed out that hsa-miR-200c-3p was associated with poor prognosis in GC patients [40], while another study suggested that hsa-miR-200c-3p was a favorable prognostic factor in GC, according to which overexpressing hsa-miR-200c-3p targeted DNMT3A/DNMT3B and led to a decrease in global DNA methylation [41]. It was reported that through suppressing ZEB1/2, hsa-miR-200c-3p inhibited TGF-β-induced-EMT and restored GC’s sensitivity to Trastuzumab [42]. hsa-miR-200c-3p was also reported to reduce GC motility and eliminate chemoresistance [43]. Apart from the five cancer-related miRNAs mentioned above, another two biomarker miRNAs were seldom studied in cancer. hsa-miR-5187-5p was downregulated in the patient plasma of unprotected left main coronary artery disease (uLMCAD) and can thus be used as a powerful diagnostic biomarker [44]. It was also found to be downregulated in stroke patients and upregulated in endometriosis patients [45]. However, its correlation with cancer has not yet been addressed. On the other hand, the biological roles of hsa-miR-548ae-5p remained largely uncategorized. Collectively, through carrying out preliminary phenotypic investigation in GC cell lines, we verified that the impacts of these 7 miRNAs on cancer motility/invasiveness/proliferation were generally consistent with published data. Nonetheless, to better understand the mechanisms of GC’s metastatic organotropism, functional details of these biomarker miRNAs as well as their roles in sEV-directed tumor-microenvironment crosstalks remained to be explored by future studies.
It has been reported the contents of sEV-derived miRNAs were not precisely correlated with total circulating miRNAs [46]. Consequently, it needs to be verified whether these biomarker miRNAs mainly originated from sEV-capsulated or non-sEV forms in order to ensure the validity of sEV-based miRNA as metastatic biomarkers. Through degradation assay, we verified these 7 biomarker miRNAs existed in plasma, mainly in sEV-capsulated form, suggesting their future clinical applications in liquid biopsy should be based on sEV-targeted assays. Since the robustness of 7 miRNAs in individually characterizing metastatic sites were literally low and lack organotropism, we referred to a logistic regression model and combined these miRNAs as signatures, which effectively optimized the characterizing power to 0.81–0.96. On the other hand, a major problem in clinical practice of GC is that current imaging diagnosis cannot provide real-time surveillance of metastasis and has a relatively low efficiency in distinguishing occult lesions. Signature-predicted metastasis displayed ≥ 75% consistency with practical metastasis in all three cohorts, suggesting our signatures provided a chance to help locating the metastatic sites for GC patients with unknown or unrecognizable metastasis lesions, which could be a potential complement for imaging diagnosis. It also should be noted that although PM and HM signatures were constructed with the same combination of biomarker miRNAs (hsa-miR-429/hsa-miR-200c-3p) and thus shared the same enrichment profile of functions/pathways, it did not support that the modes or biomarkers of peritoneal metastasis and hepatic metastasis were similar. Instead, different logistic regression coefficients were applied to generate PM and HM signatures, and the comparison of consistency showed that only a very small fraction of cases were inconsistently paired between PM and HM groups, verifying that PM and HM signatures had distinct specificity and were competent for the characterization of respective organotropism. Furthermore, these signatures were used to calculate metastatic risk scores and also displayed robustness in predicting future metastasis for M0 patients. Around 45% of patients stratified into high-risk group developed distant metastasis within the next 5 years, while an overall high accuracy (70%) was achieved in predicting the metastatic organotropism. Thus, for the sake of disease prevention and control, it is valuable for non-metastatic GC patients to pay extra attention to monitoring those sEV-derived miRNA signatures.
A series of methods have been applied to surveillant cancer metastasis, yet they showed disadvantages in different ways. Circulating biomarkers, such as CEA/CA199/CA72.4, were powerful indexes to predict GC progression, but were insufficient to distinguish metastasis and relevant organotropism [47]. Imaging methods hold promise for tumor staging and detecting distant metastasis, but frequently missed occult lesions in abdominal cavity and peritoneal regions [48, 49]. In this study, through identifying sEV-derived miRNA signatures robustly predicting metastatic organotropism of GC, we brought new clues to screen lesions unrecognizable/untraceable for endoscopic or imaging methods. Compared with other biomarker studies, our work offered a more efficient and specific option to diagnose and surveillant metastasis. Since our signatures were promising tools in facilitating clinical decision making, it worth to be expanded to larger patient populations for further verification.
In conclusion, our work provided a clinical insight into the metastatic correlation of plasma-borne sEVs in GC and verified sEV-derived miRNA signatures outlining GC’s representative organo-tropic metastasis. These findings expanded the current understandings to the metastatic modes of GC and provided potential reference for clinical applications.
Data availability
Data have been deposited into NCBI Gene Expression Omnibus (GEO) as series GSE178283.
References
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
Gupta GP, Massagué J. Cancer metastasis: building a framework. Cell. 2006;127:679–95.
Gao Y, Bado I, Wang H, et al. Metastasis organotropism: redefining the congenial soil. Dev Cell. 2019;49:375–91.
Turajlic S, Swanton C. Metastasis as an evolutionary process. Science (New York, NY). 2016;352:169–75.
Bray F, Ferlay J, Soerjomataram I, et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424.
Yang D, Hendifar A, Lenz C, et al. Survival of metastatic gastric cancer: significance of age, sex and race/ethnicity. J Gastrointestinal Oncol. 2011;2:77–84.
Thomassen I, van Gestel YR, van Ramshorst B, et al. Peritoneal carcinomatosis of gastric origin: a population-based study on incidence, survival and risk factors. Int J Cancer. 2014;134:622–8.
Jo J-C, Ryu M-H, Koo D-H, et al. Serum CA 19–9 as a prognostic factor in patients with metastatic gastric cancer. Asia Pac J Clin Oncol. 2013;9:324–30.
Bernards N, Creemers GJ, Nieuwenhuijzen GAP, et al. No improvement in median survival for patients with metastatic gastric cancer despite increased use of chemotherapy. Ann Oncol. 2013;24:3056–60.
Chen Y, Zhou Q, Wang H, et al. Predicting peritoneal dissemination of gastric cancer in the era of precision medicine: molecular characterization and biomarkers. Cancers. 2020;12:2236.
Kodera Y, Fujitani K, Fukushima N, et al. Surgical resection of hepatic metastasis from gastric cancer: a review and new recommendation in the Japanese gastric cancer treatment guidelines. Gastric Cancer. 2013;17:206–12.
Deng J, Liang H. Clinical significance of lymph node metastasis in gastric cancer. World J Gastroenterol. 2014;20:3967–75.
Strong V, Wu A, Selby L, et al. Differences in gastric cancer survival between the US and China. J Surg Oncol. 2015;112:31–7.
Li P, Huang C, Zheng C, et al. Comparison of gastric cancer survival after R0 resection in the US and China. J Surg Oncol. 2018;118:975–82.
Gadde R, Tamariz L, Hanna M, et al. Metastatic gastric cancer (MGC) patients: Can we improve survival by metastasectomy? A systematic review and meta-analysis. J Surg Oncol. 2015;112:38–45.
Shen L, Shan Y-S, Hu H-M, et al. Management of gastric cancer in Asia: resource-stratified guidelines. Lancet Oncol. 2013;14:e535–47.
Witwer K, Buzás E, Bemis L et al. Standardization of sample collection, isolation and analysis methods in extracellular vesicle research. J Extracell Vesicle 2013; 2.
Xu R, Rai A, Chen M, et al. Extracellular vesicles in cancer—implications for future improvements in cancer care. Nat Rev Clin Oncol. 2018;15:617–38.
Murillo O, Thistlethwaite W, Rozowsky J, et al. exRNA atlas analysis reveals distinct extracellular RNA cargo types and their carriers present across human biofluids. Cell. 2019;177:463-477.e415.
Jiang C, Chen X, Alattar M, et al. MicroRNAs in tumorigenesis, metastasis, diagnosis and prognosis of gastric cancer. Cancer Gene Ther. 2015;22:291–301.
Bardelli A, Pantel K. Liquid biopsies, what we do not know (yet). Cancer Cell. 2017;31:172–9.
Siravegna G, Marsoni S, Siena S, Bardelli A. Integrating liquid biopsies into the management of cancer. Nat Rev Clin Oncol. 2017;14:531–48.
Palmirotta R, Lovero D, Cafforio P, et al. Liquid biopsy of cancer: a multimodal diagnostic tool in clinical oncology. Ther Adv Med Oncol. 2018;10:175883591879463.
Thind A, Wilson C. Exosomal miRNAs as cancer biomarkers and therapeutic targets. J Extracell Vesicle. 2016;5:31292.
Pretzsch E, Bösch F, Neumann J, et al. Mechanisms of metastasis in colorectal cancer and metastatic organotropism: hematogenous versus peritoneal spread. J Oncol. 2019;2019:7407190.
Cai J, Wu J, Zhang H, et al. miR-186 downregulation correlates with poor survival in lung adenocarcinoma, where it interferes with cell-cycle regulation. Can Res. 2013;73:756–66.
Zhu X, Shen H, Yin X, et al. miR-186 regulation of Twist1 and ovarian cancer sensitivity to cisplatin. Oncogene. 2016;35:323–32.
Liu L, Wang Y, Bai R, et al. MiR-186 inhibited aerobic glycolysis in gastric cancer via HIF-1α regulation. Oncogenesis. 2016;5:e224–e224.
Lin H, Zhou A-J, Zhang J-Y, et al. MiR-324-5p reduces viability and induces apoptosis in gastric cancer cells through modulating TSPAN8. J Pharm Pharmacol. 2018;70:1513–20.
Liu C, Li G, Yang N et al. miR-324-3p suppresses migration and invasion by targeting WNT2B in nasopharyngeal carcinoma. Cancer Cell Int. 2017; 17.
Kuo W, Yu S, Li S, et al. MicroRNA-324 in human cancer: miR-324-5p and miR-324-3p have distinct biological functions in human cancer. Anticancer Res. 2016;36:5189–96.
Liu X, Liu Y, Wu S, et al. Tumor-suppressing effects of miR-429 on human osteosarcoma. Cell Biochem Biophys. 2014;70:215–24.
Shen J, Hong L, Yu D, et al. LncRNA XIST promotes pancreatic cancer migration, invasion and EMT by sponging miR-429 to modulate ZEB1 expression. Int J Biochem Cell Biol. 2019;113:17–26.
Zhu P, Zhang J, Zhu J, et al. MiR-429 induces gastric carcinoma cell apoptosis through Bcl-2. Cell Physiol Biochem. 2015;37:1572–80.
Wang Y, Zhang X, Zhang B, et al. Initial study of microRNA expression profiles of colonic cancer without lymph node metastasis. J Dig Dis. 2010;11:50–4.
Wang Y, Xia H, Zhuang Z, et al. Axl-altered microRNAs regulate tumorigenicity and gefitinib resistance in lung cancer. Cell Death Dis. 2014;5: e1227.
Cai J, Guan H, Fang L, et al. MicroRNA-374a activates Wnt/β-catenin signaling to promote breast cancer metastasis. J Clin Investig. 2013;123:566–79.
Xu X, Wang W, Su N, et al. miR-374a promotes cell proliferation, migration and invasion by targeting SRCIN1 in gastric cancer. FEBS Lett. 2015;589:407–13.
Ji R, Zhang X, Gu H, et al. miR-374a-5p: a new target for diagnosis and drug resistance therapy in gastric cancer. Mol Therapy Nucl Acids. 2019;18:320–31.
Huang Z-S, Guo X-W, Zhang G, et al. The diagnostic and prognostic value of miR-200c in gastric cancer: a meta-analysis. Dis Markers. 2019;2019:1–9.
Tang H, Deng M, Tang Y, et al. miR-200b and miR-200c as prognostic factors and mediators of gastric cancer cell progression. Clin Cancer Res. 2013;19:5602–12.
Zhou X, Men X, Zhao R, et al. miR-200c inhibits TGF-β-induced-EMT to restore trastuzumab sensitivity by targeting ZEB1 and ZEB2 in gastric cancer. Cancer Gene Ther. 2018;25:68–76.
Ghasabi M, Majidi J, Mansoori B, et al. The effect of combined miR-200c replacement and cisplatin on apoptosis induction and inhibition of gastric cancer cell line migration. J Cell Physiol. 2019;234:22581–92.
Zhu L, Chen T, Ye W, et al. Circulating miR-182-5p and miR-5187-5p as biomarkers for the diagnosis of unprotected left main coronary artery disease. J Thorac Dis. 2019;11:1799–808.
Modak JM, Roy-O’Reilly M, Zhu L, et al. Differential microribonucleic acid expression in cardioembolic stroke. J Stroke Cerebrovasc Dis. 2019;28:121–4.
Min L, Zhu S, Chen L, et al. Evaluation of circulating small extracellular vesicles derived miRNAs as biomarkers of early colon cancer: a comparison with plasma total miRNAs. J Extracell Vesicle. 2019;8:1643670.
Lin J-X, Wang W, Lin J-P, et al. Preoperative tumor markers independently predict survival in stage III gastric cancer patients: should we include tumor markers in AJCC staging? Ann Surg Oncol. 2018;25:2703–12.
Wang Y, Liu W, Yu Y, et al. CT radiomics nomogram for the preoperative prediction of lymph node metastasis in gastric cancer. Eur Radiol. 2019;30:976–86.
Kamiya S, Takeuchi H, Nakahara T, et al. Auxiliary diagnosis of lymph node metastasis in early gastric cancer using quantitative evaluation of sentinel node radioactivity. Gastric Cancer. 2015;19:1080–7.
Acknowledgements
We deeply appreciate Dr. Sai Ge and Dr. Jiajia Yuan (Department of Gastrointestinal Oncology, Peking University Cancer Hospital & Institute) for suggestions on patient selection, and Dr. Yuezong Bai (Medical Affairs, 3D Medicines Inc, Shanghai, China) and Lingyu Wang (SIP LifeLink Oncology Research Institute, Suzhou, P. R. China) for suggestions on bioinformatic analysis.
Funding
This work was supported by the National Natural Science Foundation of China (No. 81872341 and No. 81802327), the third round of public welfare development and reform pilot projects of Beijing Municipal Medical Research Institutes (Beijing Medical Research Institute, 2019-1), and the Major Program of National Natural Science Foundation of China (No. 91959205).
Author information
Authors and Affiliations
Contributions
CZ, JG and LS designed and supervised the project. CZ, JY, YC and FJ performed the research. HL, XZ and JL collected biospecimens and clinical information. XL, YW and GK integrated the data and instructed statistical analysis.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
Written informed consents were acquired from all donors or their legal guardians. Patient samples and relevant clinical data were approved by the Institutional Ethics Committee, Peking University Cancer Hospital & Institute for experimental applications.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary Fig. S1
Procedures and criteria for patient selection. Treatment-naïve GC patients were grouped according to their metastatic sites. A total of 166 (including 42 M0, 31 PM, 34 HM, 34 dLNM and 25 MM) cases were selected and distributed into three cohorts. ■, inclusive criteria. ×, exclusive criteria. Supplementary file4 (TIF 1198 KB)
Supplementary Fig. S2
Isolation and identification of plasma EV. Representative (A) morphology, (B) diameter and (C) purity of isolated sEV were demonstrated. (D) The concentrations of sEV RNA were mutually compared among M0, PM, HM and dLNM groups, then pairwise compared with MM group. ns, not significant. *,P ≤ 0.05.. Supplementary file5 (TIF 6966 KB)
Supplementary Fig. S3
Differentially expressed sEV miRNA profiles across metastatic modes. (A) t-SNE analysis for all sEV miRNA profiles across all metastatic modes. (B) The landscape of miRNAs differentially expressed for all 4 metastatic modes. Supplementary file6 (TIF 2046 KB)
Supplementary Fig. S4
Candidate sEV miRNAs outlined the four metastatic modes. (A) t-SNE analysis for candidate sEV miRNAs across all metastatic modes. (B) Hierarchical clustering of candidate sEV miRNAs efficiently represented M0, PM, HM and dLNM groups. (C) The intersectional part of differentially expressed candidate miRNAs across all metastatic modes. Supplementary file7 (TIF 1210 KB)
Supplementary Fig. S5
Expression of the 7 biomarker miRNAs in discovery cohort and training cohort. In discovery cohort (upper panel) and training cohort (lower panel), expressions of 7 miRNAs were quantified by qPCR for M0 vs nonM0, PM vs nonPM, HM vs nonHM and dLNM vs nondLNM comparisons.Supplementary file8 (TIF 1671 KB)
Supplementary Fig. S6
The power of 7 miRNAs in individually characterizing GC’s metastatic organotropism. The AUC for 7 miRNAs in correlation to M0, PM, HM and dLNM in (A) discovery cohort and (B) training cohort were demonstrated. Supplementary file9 (TIF 1581 KB)
Supplementary Fig. S7
The time course of sEV miRNA expressions as well as metastatic risk for these two cases. (A) The time course of 7 sEV miRNAs’ expression profile for two cases were assessed with qPCR. (B) The time course of organo-tropic metastatic risk for two cases. Supplementary file10 (TIF 1046 KB)
Supplementary Fig. S8
The 5-year DMFS for M0 patients subsequently received diverse anti-cancer therapies. For M0 patients received neoadjuvant chemotherapy+surgery (left panel) or chemotherapy (right panel), the 5-year DMFS were stratified by signature-based metastatic risk scores. Supplementary file11 (TIF 689 KB)
Supplementary Fig. S9
The sEV expression profiles of the 7 biomarker miRNAs between gastric cancer patients and healthy control. The sEV expression profiles of 7 biomarker miRNAs in 20 plasma samples from healthy people (N) were quantified with qPCR, then compared with those in training cohort combined validating cohort (GC). Statistic comparisons were performed with t-test. ns, not significant. *, P≤0.05. Supplementary file12 (TIF 999 KB)
Supplementary Fig. S10
Relevance of biomarker miRNAs to GC invasiveness/proliferation. The 7 biomarker miRNAs were overexpressed in gastric cancer cell lines, then administrated with (A) invasion assay and (B) proliferation assay. ns, not significant. *,P ≤ 0.05. Supplementary file13 (TIF 6286 KB)
Rights and permissions
About this article
Cite this article
Zhang, C., Yang, J., Chen, Y. et al. miRNAs derived from plasma small extracellular vesicles predict organo-tropic metastasis of gastric cancer. Gastric Cancer 25, 360–374 (2022). https://doi.org/10.1007/s10120-021-01267-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10120-021-01267-5