Abstract
Objective
Previous research suggests that peripheral immune cells may play a role in the development of Alzheimer's disease (AD). Our study aims to determine if the composition of peripheral immune cells directly contributes to the occurrence of AD.
Methods
We utilized a two-sample Mendelian randomization (MR) approach to examine the association between peripheral immune cells and AD.The primary analysis method used was the inverse variance weighted (IVW) method, and we also conducted analyses using MR Egger, weighted median, simple mode, and weighted mode methods to ensure the accuracy of the results.Heterogeneity and horizontal pleiotropy were evaluated using Cochran's Q statistics and the MR Egger intercept, respectively.
Results
The study found a significant correlation between increased IgD + CD24- AC cells (Odds Ratio [OR] = 1.03, 95% Confidence Interval [CI] = 1.01–1.06, P = 0.0172), increased CD4 + %leukocyte (OR = 1.08, 95% CI = 1.02–1.14, P = 0.0086), and increased CD4 + CD8dim AC cells (OR = 1.06, 95% CI = 1.01–1.11, P = 0.0218), with an increased susceptibility to AD. Conversely, an increase in EM DN (CD4-CD8-) %T cells (OR = 0.95, 95% CI = 0.92–0.99, P = 0.0164) and an increase in DN (CD4-CD8-) AC cells (OR = 0.93, 95% CI = 0.88–0.99, P = 0.0145) were associated with a protective effect against AD.
Conclusion
Our findings establish a causal link between peripheral immune cells and AD. This study is the first to examine the relationship between peripheral immune cells and AD using MR, offering valuable insights for early diagnosis and treatment decisions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Alzheimer's disease (AD) is a specific disorder characterized by age-related cognitive and functional decline, ultimately leading to death (Alzheimer's Association, 2019) [1]. The prevalence of all-cause dementia is projected to increase from 50 million people in 2010 to 113 million by 2050 worldwide. AD imposes a significant burden on individuals and society [2]. Genetic factors account for approximately 70% of the risk of developing AD [3], Multiple large-scale genome-wide association studies (GWAS) and meta-analyses have identified over 40 genetic risk loci associated with AD, with the APOE ε4 allele remaining the strongest genetic risk factor for sporadic AD [4], Conversely, the APOE ε2 allele is the most robust genetic protective factor [5]. The pathology of AD is associated with synaptic loss [6], oxidative stress [7], abnormal mitochondrial structure and function [8], dysregulation of microRNAs [9], inflammatory response [10], neurodegeneration [11] and neurofibrillary tangles [12], accumulation of beta-amyloid (Aβ) [13] and phosphorylated tau (p-tau) protein accumulation [14]. The infiltration of peripheral immune cells into the brain is a prominent feature of aging and various neurodegenerative diseases, such AD [15]. In non-autoimmune conditions, alterations in the immune environment can occur, resulting in significant changes in the basic structure of the innate and adaptive immune systems within the brain parenchyma of AD due to local environmental changes and infiltration of immune cells from the blood and boundary regions [16]. the central and peripheral immune systems contribute to the clearance of Aβ peptide and immune dysfunction that lead to AD. This includes complement, microglia, and peripheral immune cells, such as monocytes and lymphocytes [17]. Increasing evidence supports the involvement of peripheral immune cells in the pathogenesis of AD [18]. Understanding the pathogenic mechanisms of AD remains challenging due to its complex etiology, chronic and progressive nature, long observational study periods, high research costs, and ethical limitations. Currently, there are no definitive therapeutic methods available to prevent or reverse the progression of AD, and there is a lack of effective treatment measures to validate or modulate factors associated with the disease's pathogenic mechanisms. Animal models have played a crucial role in disease research; however, finding a model that fully replicates the human pathological process of AD remains challenging. Despite significant advancements in research techniques over recent years, including proteomics, genomics, and imaging technologies, continuous improvement and development of more precise and sensitive techniques and methods are still needed to study the pathogenic mechanisms of AD. Therefore, we are searching for new research approaches to further clarify the pathogenic mechanisms of AD.
Mendelian randomization (MR) is an analytical method used to evaluate causal relationships by leveraging genetic variation. Based on the principles of Mendelian genetics, MR utilizes naturally occurring genetic variations to simulate the impact of randomization on investigating causal relationships. The core concept of MR is to link the exposure factor under investigation (such as a specific medication, lifestyle, or biomarker) to its associated genetic variants known as instrumental variables (IVs) in genetics and infer causal relationships through the influence of genetic variation. Genetic variation, as a mechanism of natural randomization, is unaffected by external environmental factors or individual selection, equivalent to the random allocation of different treatments in a randomized controlled trial. Compared to observational studies, MR reduces potential confounding factors and reverse causality, providing more reliable causal inference, minimizing the likelihood of reverse correlations, and avoiding ethical and practical limitations. In recent years, with the explosive growth of human genetic data, MR has emerged as a novel approach to examine causal relationships and has been proven as a valuable tool for studying disease risk factors.
In this study, we utilized large-scale GWAS summary statistics for blood-related traits to identify the pathophysiological role of the immune system in the onset and development of AD. Currently, there is limited research using MR to investigate causal relationships between circulatory immune cell counts and diseases, especially in relation to peripheral immune cells and AD. The objective of this paper is to determine the existence of a causal relationship between peripheral immune cells and the onset of AD. Furthermore, we aim to explore the mechanisms underlying AD pathogenesis and seek new therapeutic strategies for intervening and delaying the progression of AD.
Methods
Data sources
The data on peripheral immune cells is derived from a study published in 2020 by Orrù V et al. [19], investigating 731 immune cell features. The GWAS Catalog provides publicly available GWAS summary statistics for each immune trait (accession numbers from GCST90001391 to GCST90002121),The data can be obtained from the website: http://ftp.ebi.ac.uk/pub/databases/gwas/summary_statistics/. Covering a total of 731 immunophenotypes, this comprehensive analysis comprises absolute cell (AC) counts (n = 118), median fluorescence intensities (MFI) representing surface antigen levels (n = 389), morphological parameters (MP) (n = 32), and relative cell (RC) counts (n = 192). Specifically, the MFI, AC and RC features encompassed various immune cell types, including B cells, CDCs, mature stages of T cells, monocytes, myeloid cells, TBNK (T cells, B cells, natural killer cells), and Treg panels, while the MP feature encompassed CDC and TBNK panels. The original GWAS on immune traits was conducted using data from 3,757 European individuals without any overlapping cohorts. Approximately 22 million single nucleotide polymorphisms (SNPs) genotyped using high-density arrays were imputed with the Sardinian sequence-based reference panel [20] Associations were tested after adjusting for covariates such as sex, age, and age2.
The summary data for the GWAS of AD (ID: ieu-b-2, https://gwas.mrcieu.ac.uk/datasets/ieu-b-2/) was obtained from the International Genomics of Alzheimer's Project (IGAP) consortium. The GWAS data utilized in this study originated from a genetic meta-analysis of Alzheimer's disease conducted by Kunkle BW et al. in 2019. The analysis encompassed both male and female participants, comprising 21,982 individuals with Alzheimer's disease and 41,944 non-Hispanic White individuals as controls, leading to the identification of 10,528,610 SNPs [21]. To minimize potential bias in MR analyses caused by population stratification, the summary statistics were derived from individuals of European descent for both the exposure and outcome datasets.
Instrumental Variable
Instrumental variables (IVs) should meet three fundamental criteria: (1) They should be associated with peripheral immune cells. (2) They should be independent of confounding factors. (3) They should exclusively affect AD through peripheral immune cells. Consequently, in this study, SNPs exhibiting genome-wide significant differences (P < 1 × 10^-5) were chosen as IVs. The strength of the IVs was assessed using an F-statistic, with IVs having an F-statistic greater than 10 considered robust. The PhenoScanner (V2) website (http://www.phenoscanner.medschl.cam.ac.uk/) was utilized to identify SNPs displaying suggestive associations (p < 10^-5) with the risk factors.
To address linkage disequilibrium within SNPs and ensure the independence of IVs, the coefficient of linkage disequilibrium was set at r2 = 0.001 and kb = 10,000 using R software. Palindromic SNPs were removed to ensure that the same single nucleotide polymorphism (SNP) had identical alleles in both the immune cell group and the AD group. In instances where no immune cell-related SNPs were present in the GWAS data for AD, IVs without alternate loci were excluded to ensure the authenticity and accuracy of the results. The data analysis process is shown in Fig. 1.
Statistical Analysis
Validation of the Causal Relationship between Immune Cells and AD
Data statistical analysis was performed using R 4.3.1 and R packages (http://www.Rproject.org), including Two Sample MR、MR-PRESSO software package. This study primarily employed the inverse variance weighted analysis (IVW) to evaluate the association between circulating immune cells and Sepsis. The IVW results were considered as the main outcome measure of this study. If no heterogeneity was detected, a fixed-effects model was used; otherwise, a random-effects model was applied. Additionally, MR-Egger, weighted median, Weighted mode and simple mode analyses were conducted as supplementary analyses to further validate the reliability of the IVW results. The MR-Egger regression enables consistent estimation even in the presence of genetic pleiotropy across all instrumental variables (IVs). The main advantage of the weighted median method is its ability to provide consistent estimation of causal relationships even when more than 50% of the instrumental variables are invalid [22],Therefore, in this study, both the MR-Egger regression and the weighted median method were employed as complementary approaches. A significance level of P < 0.05 was considered statistically significant for detecting differences.
Sensitivity Analysis
We used MR-PRESSO to identify outliers that had a significant impact, and reanalyzed the data after excluding these outliers. This method can be employed using the MR-PRESSO software package. Sensitivity analysis using the "leave-one-out" approach was conducted by sequentially removing one SNP at a time to assess the individual SNP's impact on the results. Additionally, we used the intercept test in the MR-Egger regression to examine the presence of horizontal pleiotropy among SNPs. Cochran's Q test was performed to assess heterogeneity among the IVs, with a significance level set at P < 0.05 indicating the presence of heterogeneity. If heterogeneity among the IVs was detected, the IVW random-effects model was used to estimate the causal relationship. The significance level was set at α = 0.05, with P < 0.05 indicating statistically significant differences.
Results
Statistical analysis
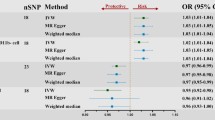
After rigorously selecting and harmonizing IVs, a total of 93 SNPs were utilized for cell count in the MR analysis. Importantly, all SNPs exhibited F statistics greater than 10, indicating their suitability as robust instruments. Comprehensive details of the harmonized data can be found in Supplementary Table 1. The causal estimate of peripheral immune cell count for AD risk is shown in Fig. 2. We detected five different cell phenotypes, including IgD + CD24- AC counts from the B cell panel and EM DN (CD4-CD8-) %T cell from the Maturation stages of T cell panel. Additionally, three cell phenotypes from the TBNK panel, namely DN (CD4-CD8-) AC, CD4 + %leukocyte, and CD4 + CD8dim AC were analyzed (Supplementary Table 2).
Using the IVW method and visualized through a forest plot, IgD + CD24- AC [odds ratio (OR),1.03;95% confidence interval (CI), 1.01–1.06; P = 0.0172], CD4 + %leukocyte [OR,1.08;95% CI, 1.02–1.14; P = 0.0086], and CD4 + CD8dim AC [OR,1.06;95% CI,1.01–1.11; P = 0.0218] were identified as potential risk factors promoting the occurrence of AD. On the other hand, EM DN (CD4-CD8-) %T cell [OR, 0.95; 95% CI, 0.92–0.99; P = 0.0164], Similarly, the Simple mode also supported these conclusions. DN (CD4-CD8-) AC [OR, 0.93; 95% CI, 0.88–0.99; P = 0.0145], the Weighted median also supported these conclusions. Both of these cellular phenotypes have a protective effect against AD. Except for EM DN (CD4 − CD8 −) %T cell with IVW and Simple mode, and DN (CD4 − CD −) AC with IVW and Weighted median supporting the same conclusion, the remaining cellular phenotypes only have support for the conclusion from the IVW method.
Horizontal pleiotropy and heterogeneity analysis
The p-value of MR-Egger intercept was found to be greater than 0.05, indicating no apparent presence of pleiotropy (Supplementary Table 3). Cochran's Q test revealed no heterogeneity in the present study (Supplementary Table 4). The funnel plot did not exhibit any evident directional pleiotropy. Sensitivity analysis using the "leave-one-out" approach showed that the inclusion of individual SNPs did not significantly affect the results, indicating the robustness of the findings. The results of the MR-PRESSO analysis indicated that the included SNP loci did not exhibit significant outliers.
Discussion
Investigating the causal relationship between peripheral immune cells and AD is crucial for understanding the disease's pathogenic mechanisms and developing effective prevention and treatment strategies. This study represents the first MR investigation into the association between peripheral immune cells and AD onset. Leveraging a vast array of publicly available genetic data, we explored the causal relationships between 731 immune cell types and AD. Among the three types of immune traits (BC, MT, TB) and five immunophenotypes (including IgD + CD24- AC from the B cell panel and EM DN (CD4-CD8-) AC from the Maturation stages of T cell panel, as well as DN (CD4-CD8-) AC, CD4 + %leukocyte, and CD4 + CD8dim AC from the TBNK panel), significant causal effects on AD were observed (adjusted for false discovery rate [FDR] < 0.05).
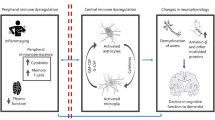
In recent years, there has been increasing research on the role of immune cells in the pathogenesis of AD. Genes and immune cells associated with innate immunity, especially peripheral ones, not only contribute positively to AD's neurodegenerative mechanisms but also display pathological effects. These cells can cross the blood–brain barrier, influencing AD's neurodegenerative processes. The study emphasizes the crucial role of interactions between adaptive and innate immune systems in both brain and peripheral contexts in AD onset and etiology [23]. Xu H et al. [24] observed characteristic changes in the proportions and gene expression patterns of peripheral blood immune cell subgroups in patients with AD, including alterations in CD4 + T cells, CD8 + T cells, B cells, natural killer cells, and monocytes. Additionally, Xiong LL et al. [25], using single-cell RNA-seq analysis, identified differential cell subgroups and genes in Peripheral Blood Mononuclear Cells between AD patients and healthy individuals, with the top 6 differing cell subgroups in AD patients being B cells, CD4 + T cells, CD8 + T cells, Hematopoietic Stem Cells, monocytes, and NK cells. Our study revealed an association between specific immune phenotypes within the TBNK panel, mature stages of T cell panel, and B cell panel with the onset of AD.
The most crucial characteristic of immune dysregulation is the activation and accumulation of T lymphocytes, the majority of which express both the α and β chains of the T-cell receptor (TCR), thus termed αβ T cells [26]. In αβ T cells, CD4 + T cells and CD8 + T cells are the most common subgroups, both exhibiting neurotoxicity by inducing substantial neuronal death through cell-to-cell contact mechanisms dependent on Fas ligand (FasL), lymphocyte function-associated antigen-1 (LFA-1), and CD40 [27]. In recent years, it has been found that the number of circulating activated CD4 + and CD8 + T cells increases in patients with AD [28]. Additionally, there is an increase in CD4 + and CD8 + T cells secreting IFN-γ in the circulation of AD patients [29]. Compared to relevant controls, AD patients and AD mouse models show a higher CD4 + /CD8 + T cell ratio in peripheral blood. Additionally, Slight expression of CD8 + and CD3 + [30]. Gate et al. [31] discovered an increased frequency of CD8 + effector memory T cells in the blood of AD patients, which showed a negative correlation with memory function. Peripheral CD8 + effector T cells infiltrate the central nervous system, undergo clonal expansion entering the cerebrospinal fluid, and participate in the formation of neuroinflammation. CD8 + T cells infiltrate the brain of APP-PS1 transgenic mice with AD and modulate the expression of genes related to synapses and neurons [18]. Jatin Machhi et al. [32] found that CD4 + effector T cells accelerate the progression of AD in mice. The mechanism may involve the conversion of disease-reactive Aβ-Th1 and Aβ-Th17 cells in the APP/PS1 mice, promoting a pro-inflammatory microenvironment and accelerating AD pathology. This study, through MR analysis, discovered that CD4 + %leukocyte and CD4 + CD8dim AC cells in the TBNK panel contribute to the exacerbation of AD onset. The current findings align with existing research conclusions, and this study provides more specific insights into the immune phenotype.
In addition, a small subset of αβ T cells that do not express CD4 and CD8 are referred to as "double-negative" T (DNT) cells [33], They have been found to play a role in the pathophysiological processes of certain autoimmune diseases [34].This type of cell constitutes approximately 1–3% of T lymphocytes in both mice and humans [35]. This study also identified that EM DN (CD4-CD8-) %T cells and DN (CD4-CD8-) AC cells in peripheral blood inhibit the progression of AD. Currently, there is limited research on the role of DNT cells in human and animal AD; however, existing studies have reported an association of this immune phenotype with AD. For example, in 3xTg-AD mice, the percentage of CD3 + CD4-CD8- (DN T) cells in the blood increases with age. Peripheral immune system abnormalities manifest earlier in male mice and are more pronounced in mice of the same age [36]. However, Chenyang Han et al. [37] discovered that double-negative T cells can exacerbate neuroinflammation and cognitive impairment in AD mice through TNF-α-NLRP3-mediated microglial M1 polarization. This finding contradicts our research results. Considering that mouse DN T cells and human DN T cells, while sharing similar functions, may not employ entirely similar mechanisms to inhibit their target cell populations [35]. For example, human DN T cells can effectively suppress the responses of both CD4 + and CD8 + T cells in vitro [38], and they exert cytotoxicity against CD8 + T cells in an antigen-specific manner [39]. On the other hand, unlike mouse DN T cells, the inhibition of CD4 + T cell proliferation by human DN T cells does not depend on the Fas/FasL pathway, perforin, and granzyme. Furthermore, DNT cells, similar to CD4 + T cells, are likely relatively unstable, and the reshaping cytokine environment may lead to their plastic fate, potentially having the ability to switch between anti-inflammatory and pro-inflammatory phenotypes [40], Therefore, this research conclusion requires further evidence for support.
The existing studies have shown controversial findings regarding the causal relationship between peripheral immune cells and AD. Therefore, this MR study provides theoretical support for previous observational studies. In this study, we utilized summary statistics from a recent large-scale GWAS of blood cell characteristics. Our findings, focusing on the European population, demonstrated that using genetic variations as causal tools and applying five MR analysis methods, the B cell panel IgD + CD24- AC and TBNK panel CD4 + %leukocyte, CD4 + CD8dim AC were associated with increased risk of AD. On the other hand, the maturation stages of T cell panel, specifically EM DN (CD4 − CD8 −) %T cell and DN (CD4 − CD8 −) AC, were found to have a protective effect against AD.
Furthermore, the current understanding of the distribution of peripheral blood immune cells in AD patients is limited to studies using flow cytometry, and the cell-type-specific functional states of immune cells, especially T and B cells, remain unclear. Additionally, there is currently no research reporting the existence of adaptive immune repertoire in the peripheral blood of AD patients [24]. Although previous observational studies have identified associations between certain immune cells and AD, the limitations of these observational studies (such as the inability to completely eliminate potential confounders, selection biases, and small sample sizes) and the expensive and time-consuming nature of randomized controlled trials present challenges in establishing a causal relationship between peripheral immune cells and AD. We did not observe any multi-collinearity or heterogeneity in our sensitivity analysis, suggesting the reliability of our results. Certain immune cell types and genetic susceptibility can serve as biomarkers for AD risk, potentially leading to earlier diagnosis and more effective treatment options. Therefore, investigating the mechanistic link between immune cells and the pathogenesis of AD at the genetic level is meaningful.
This study has several strengths. Firstly, it utilized data from the largest sample in GWAS studies. Sensitivity analyses using the MR-PRESSO, leave-one-out, MR-Egger, and Cochran Q tests were conducted to validate the IVW results. Secondly, by utilizing germline genetic variants as instrumental variables for exposure, random allocation was achieved, thereby avoiding reverse causation and confounding factors, with no ethical risks involved.
However, this study also has limitations. Firstly, among the five cell phenotypes in the conclusions of this study, the IVW method and the Simple mode supported the same conclusions for all cell phenotypes except for EM DN (CD4-CD8-) %T cell and DN (CD4 − CD −) AC, Only the IVW method supported the conclusions for the remaining cell phenotypes. This suggests that there may be limitations in the results. Possible reasons include the presence of multicollinearity leading to potential bias in the IVW method. Further studies are needed to validate the results, such as increasing the sample size and the number of instrumental variables and using more robust analysis methods to verify the findings. Subsequent research could consider communicating with the authors to supplement additional robust sensitivity analyses to support the results. Secondly, this study included only populations of European ancestry, which may introduce ethnic bias and limit the generalizability of the results. Thirdly, due to the unavailability of raw data, we could only estimate the approximate causal relationship between the variables and could not determine the more specific and detailed causal relationship between them. Lastly, through this method, the causal relationship between peripheral immune cells and the risk of AD can only be preliminarily assessed, and the underlying biological mechanisms between the two are not fully understood. Therefore, more research is needed to establish the relationship between them.
In summary, this study applies a two-sample MR method to infer the causal relationship between circulating immune cells and AD. The results suggest that IgD + CD24- AC, CD4 + %leukocyte, and CD4 + CD8dim AC may be associated with an increased risk of AD, while EM DN (CD4-CD8-) %T cell and DN (CD4-CD8-) AC may have a protective effect against AD. These findings may provide additional insights for early diagnosis and treatment decision-making, but further research is needed to support these conclusions.
Data Availability
The original contributions presented in the study are included in the article/Supplementary Material. Further inquiries can be directed to the corresponding authors.
References
Soria Lopez JA, González HM, Léger GC (2019) Alzheimer’s disease. Handb Clin Neurol 167:231–255. https://doi.org/10.1016/B978-0-12-804766-8.00013-3
Scheltens P, De Strooper B, Kivipelto M et al (2021) Alzheimer’s disease. Lancet 397:1577–1590. https://doi.org/10.1016/S0140-6736(20)32205-4
Silva MVF, de LouresMG, Alves LCV, C et al (2019) Alzheimer’s disease: risk factors and potentially protective measures. J Biomed Sci 26:33. https://doi.org/10.1186/s12929-019-0524-y
Veitch DP, Weiner MW, Aisen PS et al (2019) Understanding disease progression and improving Alzheimer’s disease clinical trials: Recent highlights from the Alzheimer’s Disease Neuroimaging Initiative. Alzheimers Dement 15:106–152. https://doi.org/10.1016/j.jalz.2018.08.005
Serrano-Pozo A, Das S, Hyman BT (2021) APOE and Alzheimer’s disease: advances in genetics, pathophysiology, and therapeutic approaches. Lancet Neurol 20:68–80. https://doi.org/10.1016/S1474-4422(20)30412-9
Gao L, Zhang Y, Sterling K, Song W (2022) Brain-derived neurotrophic factor in Alzheimer’s disease and its pharmaceutical potential. Transl Neurodegener 11:4. https://doi.org/10.1186/s40035-022-00279-0
Butterfield DA, Halliwell B (2019) Oxidative stress, dysfunctional glucose metabolism and Alzheimer disease. Nat Rev Neurosci 20:148–160. https://doi.org/10.1038/s41583-019-0132-6
Gowda P, Reddy PH, Kumar S (2022) Deregulated mitochondrial microRNAs in Alzheimer’s disease: Focus on synapse and mitochondria. Ageing Res Rev 73:101529. https://doi.org/10.1016/j.arr.2021.101529
Nagaraj S, Zoltowska KM, Laskowska-Kaszub K, Wojda U (2019) microRNA diagnostic panel for Alzheimer’s disease and epigenetic trade-off between neurodegeneration and cancer. Ageing Res Rev 49:125–143. https://doi.org/10.1016/j.arr.2018.10.008
Princiotta Cariddi L, Mauri M, Cosentino M et al (2022) Alzheimer’s Disease: From Immune Homeostasis to Neuroinflammatory Condition. Int J Mol Sci 23:13008. https://doi.org/10.3390/ijms232113008
Vasic V, Barth K, Schmidt MHH (2019) Neurodegeneration and Neuro-Regeneration-Alzheimer’s Disease and Stem Cell Therapy. Int J Mol Sci 20:4272. https://doi.org/10.3390/ijms20174272
Graff-Radford J, Yong KXX, Apostolova LG et al (2021) New insights into atypical Alzheimer’s disease in the era of biomarkers. Lancet Neurol 20:222–234. https://doi.org/10.1016/S1474-4422(20)30440-3
Jucker M, Walker LC (2023) Alzheimer’s disease: From immunotherapy to immunoprevention. Cell 186:4260–4270. https://doi.org/10.1016/j.cell.2023.08.021
Zhang H, Wei W, Zhao M et al (2021) Interaction between Aβ and Tau in the Pathogenesis of Alzheimer’s Disease. Int J Biol Sci 17:2181–2192. https://doi.org/10.7150/ijbs.57078
Altendorfer B, Unger MS, Poupardin R et al (2022) Transcriptomic Profiling Identifies CD8+ T Cells in the Brain of Aged and Alzheimer’s Disease Transgenic Mice as Tissue-Resident Memory T Cells. J Immunol 209:1272–1285. https://doi.org/10.4049/jimmunol.2100737
Chen X, Holtzman DM (2022) Emerging roles of innate and adaptive immunity in Alzheimer’s disease. Immunity 55:2236–2254. https://doi.org/10.1016/j.immuni.2022.10.016
Jevtic S, Sengar AS, Salter MW, McLaurin J (2017) The role of the immune system in Alzheimer disease: Etiology and treatment. Ageing Res Rev 40:84–94. https://doi.org/10.1016/j.arr.2017.08.005
Unger MS, Li E, Scharnagl L et al (2020) CD8+ T-cells infiltrate Alzheimer’s disease brains and regulate neuronal- and synapse-related gene expression in APP-PS1 transgenic mice. Brain Behav Immun 89:67–86. https://doi.org/10.1016/j.bbi.2020.05.070
Orrù V, Steri M, Sidore C et al (2020) Complex genetic signatures in immune cells underlie autoimmunity and inform therapy. Nat Genet 52:1036–1045. https://doi.org/10.1038/s41588-020-0684-4
Sidore C, Busonero F, Maschio A et al (2015) Genome sequencing elucidates Sardinian genetic architecture and augments association analyses for lipid and blood inflammatory markers. Nat Genet 47:1272–1281. https://doi.org/10.1038/ng.3368
Kunkle BW, Grenier-Boley B, Sims R et al (2019) Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Aβ, tau, immunity and lipid processing. Nat Genet 51:414–430. https://doi.org/10.1038/s41588-019-0358-2
Bowden J, Holmes MV (2019) Meta-analysis and Mendelian randomization: A review. Res Synth Methods 10:486–496. https://doi.org/10.1002/jrsm.1346
Jorfi M, Maaser-Hecker A, Tanzi RE (2023) The neuroimmune axis of Alzheimer’s disease. Genome Med 15:6. https://doi.org/10.1186/s13073-023-01155-w
Xu H, Jia J (2021) Single-Cell RNA Sequencing of Peripheral Blood Reveals Immune Cell Signatures in Alzheimer’s Disease. Front Immunol 12:645666. https://doi.org/10.3389/fimmu.2021.645666
Xiong L-L, Xue L-L, Du R-L et al (2021) Single-cell RNA sequencing reveals B cell-related molecular biomarkers for Alzheimer’s disease. Exp Mol Med 53:1888–1901. https://doi.org/10.1038/s12276-021-00714-8
Davis MM, Boniface JJ, Reich Z et al (1998) Ligand recognition by alpha beta T cell receptors. Annu Rev Immunol 16:523–544. https://doi.org/10.1146/annurev.immunol.16.1.523
Giuliani F, Goodyer CG, Antel JP, Yong VW (2003) Vulnerability of human neurons to T cell-mediated cytotoxicity. J Immunol 171:368–379. https://doi.org/10.4049/jimmunol.171.1.368
Lueg G, Gross CC, Lohmann H et al (2015) Clinical relevance of specific T-cell activation in the blood and cerebrospinal fluid of patients with mild Alzheimer’s disease. Neurobiol Aging 36:81–89. https://doi.org/10.1016/j.neurobiolaging.2014.08.008
Baglio F, Saresella M, Preti MG et al (2013) Neuroinflammation and brain functional disconnection in Alzheimer’s disease. Front Aging Neurosci 5:81. https://doi.org/10.3389/fnagi.2013.00081
St-Amour I, Bosoi CR, Paré I et al (2019) Peripheral adaptive immunity of the triple transgenic mouse model of Alzheimer’s disease. J Neuroinflammation 16:3. https://doi.org/10.1186/s12974-018-1380-5
Gate D, Saligrama N, Leventhal O et al (2020) Clonally expanded CD8 T cells patrol the cerebrospinal fluid in Alzheimer’s disease. Nature 577:399–404. https://doi.org/10.1038/s41586-019-1895-7
Machhi J, Yeapuri P, Lu Y et al (2021) CD4+ effector T cells accelerate Alzheimer’s disease in mice. J Neuroinflammation 18:272. https://doi.org/10.1186/s12974-021-02308-7
D’Acquisto F, Crompton T (2011) CD3+CD4-CD8- (double negative) T cells: saviours or villains of the immune response? Biochem Pharmacol 82:333–340. https://doi.org/10.1016/j.bcp.2011.05.019
Brandt D, Hedrich CM (2018) TCRαβ+CD3+CD4-CD8- (double negative) T cells in autoimmunity. Autoimmun Rev 17:422–430. https://doi.org/10.1016/j.autrev.2018.02.001
Hillhouse EE, Lesage S (2013) A comprehensive review of the phenotype and function of antigen-specific immunoregulatory double negative T cells. J Autoimmun 40:58–65. https://doi.org/10.1016/j.jaut.2012.07.010
Nava Catorce M, Acero G, Gevorkian G (2021) Age- and sex-dependent alterations in the peripheral immune system in the 3xTg-AD mouse model of Alzheimer’s disease: Increased proportion of CD3+CD4-CD8- double-negative T cells in the blood. J Neuroimmunol 360:577720. https://doi.org/10.1016/j.jneuroim.2021.577720
Han C, Sheng Y, Wang J et al (2022) Double-negative T cells mediate M1 polarization of microglial cells via TNF-α-NLRP3 to aggravate neuroinflammation and cognitive impairment in Alzheimer’s disease mice. J Cell Physiol 237:3860–3871. https://doi.org/10.1002/jcp.30839
Voelkl S, Gary R, Mackensen A (2011) Characterization of the immunoregulatory function of human TCR-αβ+ CD4- CD8- double-negative T cells. Eur J Immunol 41:739–748. https://doi.org/10.1002/eji.201040982
Fischer K, Voelkl S, Heymann J et al (2005) Isolation and characterization of human antigen-specific TCR alpha beta+ CD4(-)CD8- double-negative regulatory T cells. Blood 105:2828–2835. https://doi.org/10.1182/blood-2004-07-2583
Li H, Tsokos GC (2021) Double-negative T cells in autoimmune diseases. Curr Opin Rheumatol 33:163–172. https://doi.org/10.1097/BOR.0000000000000778
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical approval and Informed consent
It's important to note that this study solely utilized publicly available GWAS summary statistics and did not involve the use or identification of individual-level data, thus ethical approval was not required.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Glossary
- AD
-
Alzheimer's disease
- CI
-
confidence interval
- Aβ
-
beta-amyloid
- p-tau
-
phosphorylated tau
- MR
-
Mendelian randomization
- IVW
-
inverse variance weighted
- GWAS
-
Genome wide association study
- SNPs
-
single nucleotide polymorphisms
- IVs
-
instrumental variables
- DNT
-
Double negative T
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liao, J., Zhang, Y., Tang, Z. et al. Causal relationships between peripheral immune cells and Alzheimer’s disease: a two-sample Mendelian randomization study. Neurol Sci 45, 3117–3124 (2024). https://doi.org/10.1007/s10072-024-07324-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10072-024-07324-y