Abstract
Rare missense variants play a crucial role in amyotrophic lateral sclerosis (ALS) pathophysiology. We report rare/novel missense variants from 154 Indian ALS patients, identified through targeted sequencing of 25 ALS-associated genes. As pathogenic variants could explain only a small percentage of ALS pathophysiology in our cohort, we investigated the frequency of tolerated and benign novel/rare variants, which could be potentially ALS susceptible. These variants were identified in 5.36% (8/149) of sporadic ALS (sALS) cases; with one novel variant each in ERBB4, SETX, DCTN1, and MATR3; four rare variants, one each in PON2 and ANG and two different rare variants in SETX. Identified variants were either absent or present at extremely rare frequencies (MAF < 0.01) in large population databases and were absent in 50 healthy controls sequenced through Sanger method. Furthermore, an oligogenic basis of ALS was observed in three sALS, with co-occurrence of intermediate-length repeat expansions in ATXN2 and a rare/novel variant in DCTN1 and SETX genes. Additionally, molecular dynamics and biochemical functional analysis of an angiogenin variant (R21G) identified from our cohort demonstrated loss of ribonucleolytic and nuclear translocation activities. Our findings suggest that rare variants could be potentially pathogenic and functional studies are warranted to decisively establish the pathogenic mechanisms associated with them.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Amyotrophic lateral sclerosis (ALS) is a neurodegenerative disorder characterized by the selective degeneration of both upper and lower motor neurons. This wasting of motor neurons ultimately leads to progressive paralysis and death of patients from respiratory failure, usually within 3–5 years of symptom onset [1]. Primarily, ALS is a sporadic disease although ~ 10% of patients have an associated family history.
The etiology of sALS is a complex interplay between genetic and environmental factors [2]. Recent studies suggest that although genetic factors may not be directly involved in disease manifestation, yet these could significantly increase the risk of ALS in the presence of conducive environmental factors. Through next-generation sequencing (NGS), a substantial number of novel genes, mutations, disease modifiers, and susceptibility factors have been identified in sALS. However, a notable fraction of genetic variations contributing to ALS still remains unknown and poses a problem in defining the genetic paradigm of ALS. Recent studies have led to the identification of rare variants in both familial and sALS [3, 4]. Since the missing genetic basis of sALS could lie in these rare variants, and may act as disease modifiers and susceptibility factors, the role of tolerated and benign mutations in ALS manifestation cannot be ruled out [5, 6]. However, only a few functional studies have assessed the disease-causing role of these variants. For example, functional characterization of angiogenin (ANG), RNASE4, and MATR3 has shown that certain variants that are predicted to be tolerated/benign (Table 1) could be pathogenic due to perturbation in their normal cellular functions [7,8,9,10, 12]. Recent studies have implicated the role of two new genes namely TBK1 and NEK1 in the ALS manifestation through the loss of kinase function or through induction of DNA damage, respectively [13, 14]. Another study employs extensive use of molecular dynamics (MD) simulations to explain the loss of enzymatic functions underlying the DAO mutations [15].
Our previous study could explain the genetic basis of approximately ~ 13% of patients in our cohort [16, 17]. Therefore, we analyzed our cohort for rare/novel variants that are predicted to be tolerated and benign in silico but could possibly play a role in ALS susceptibility and manifestation. Since previous reports suggest that tolerated and benign ANG variants could lead to ALS due to loss of ribonucleolytic or nuclear translocation or both of these activities (Table 1), we specifically carried out a comprehensive investigation of loss-of-function mechanisms of a rare R21G (ANG) variant identified in one sALS case using MD simulations, followed by functional assay experiments. Our findings show that variants predicted as tolerated and benign could also be potentially pathogenic and hence highlights the need to carefully examine such variants in other populations to uncover their roles in ALS pathophysiology.
Material and methods
ALS patients
A total of 154 ALS patients (5 familial and 149 sporadic ALS) described previously were studied. We have included 5 fALS and 149 sALS patients visiting the neurology clinic at the All India Institute of Medical Sciences, a tertiary care academic hospital in New Delhi, India. Among these, 124 were classified as definite ALS, 29 were classified as probable ALS, and 1 was classified as clinically possible ALS based on the revised El Escorial Criteria. Additionally, ALS functional rating score was evaluated for all the patients to confirm diagnosis (Table 2) [17]. We also sequenced 50 healthy controls (Supplementary Table 1) for the rare/novel variants identified in our patients. Informed written consent was obtained from all patients and controls before the blood sample collection. The age, gender, and ethnicity of these patients are provided in Table 2. This study was conducted with the approval of and in accordance with the guidelines of the Institutional Ethics Committee, All India Institute of Medical Sciences, New Delhi.
Targeted sequencing and bioinformatics analysis
As described previously, our targeted NGS panel consisted of 25 ALS-associated genes and was tested on 154 patient samples. Sequence data was analyzed using the Variant Studio software (Illumina, v2.2) [17]. The variants were filtered by the following criteria: read depth 100×, variant type SNV, variant quality scores 30, variant consequence, alternate variant frequency ≥ 40, variant frequency in 1000 genomes and Exome Variant Server (EVS), and functional impact predicted through SIFT [18] and PolyPhen 2.0 [19]. All nonsynonymous variants predicted to be tolerated and benign through variant analysis were reconfirmed by Sanger sequencing (Supplementary Fig. 1). The variants that passed Sanger validation were selected for further analysis. Since the number of healthy controls was limited, we used allele frequency data of 60,706 individuals from the ExAC database for all statistical analysis [20].
Molecular dynamics simulations and analyses
The crystal structure of human ANG was obtained from the Protein Data Bank (PDB entry: 1B1I), and the structure of R21G variant was modeled in silico by mutating the target residue with the desired amino acid while keeping the secondary structure intact (Supplementary Fig. 2). The heteroatoms (crystallographic waters and cofactor, CIT) were removed from the structure before simulation, followed by the addition of hydrogen atoms using the X-leap tool of AMBER 14 and subsequent solvation in an octahedral box of TIP3P water with ~ 10 Å between the protein surface and the box boundary. Each system was then electrostatically neutralized, and simulations were carried out using AMBER 14 software package. Topology and parameter files for the proteins were generated using the “ff14SB” force field. A minimization step employing 2500 steps of steepest descent followed by 1000 steps of conjugate gradient was performed. The SANDER module of AMBER was used for all the MD simulations. A heating step, in which each system is gradually heated from 0 to 300 K in 200 ps, was followed by constant temperature equilibration at 300 K for 1000 ps. Following the equilibration phase, 100 ns MD simulations were carried out with periodic boundary conditions in the NPT ensemble, at a temperature of 300 K with Berendsen temperature coupling and a constant pressure of 1 atm with isotropic molecule-based scaling. SHAKE algorithm was applied to fix all covalent bonds containing hydrogen atoms. The particle mesh-Ewald (PME) method was used to treat the long-range electrostatic interactions and a 10-Å cutoff was applied to treat nonbonding short-range electrostatics and van der Waals interactions. The MD-generated trajectory files were analyzed using the CPPTRAJ module of AMBER. Coordinates of the trajectories were sampled every 1 ps for analysis of energy stabilization and root mean square deviations (RMSD) of the proteins. The H-bonds plugin incorporated in visual molecular dynamics (VMD) was used for the calculation of percentage hydrogen bond occupancy of wild type and R21G variant [21, 22]. All figures for representing ANG structures were generated using PyMOL and VMD.
Cloning, expression, and purification of ANG
The wild type and R21G variant were cloned in the BamHI and EcoRI restriction sites of the Escherichia coli expression vector pGEX-6P-2 (GE-Healthcare) with a C-terminal hexa-histidine His-tag as described previously [18]. Briefly, the missense variant, R21G, was generated through standard site-directed mutagenesis technique, verified by Sanger’s sequencing, followed by transformation into competent cells. Protein expression was induced in E. coli BL21 cells at 37 °C by addition of 0.5 mM isopropyl b-d-1-thiogalactopyranoside (IPTG) (HiMedia), at an A600 of 0.6. Cells were harvested after 4 h of induction, and cell pellets were disrupted by the addition of lysozyme (MP Biomedical), followed by sonication (Branson Sonifier 250, Netherlands). Followed by centrifugation at 8000 rpm for 1 h, the wild type and R21G variant were recovered from the soluble fraction. Purification of wild type and R21G variant was carried out through standard Ni-nitrilotriacetic acid (Ni-NTA) affinity-based purification (Qiagen), followed by dialysis and desirable concentration. The concentration of wild type and R21G was determined at 595 nm using Bradford assay.
Secondary structure assessment
Circular dichroism (CD) spectroscopic studies of wild type and R21G proteins were performed on a peltier-attached JASCO J-815 spectropolarimeter (Tokyo, Japan). The purified proteins were diluted in phosphate-buffered saline (PBS) to yield a concentration of 0.5 mg/ml, and measurements were taken in a 1-mm path length quartz cell at a scan speed of 100 nm/min. For each sample, three spectra were recorded, averaged, and plotted after subtracting the buffer baseline.
Ribonucleolytic activity assay
The ribonucleolytic activity of wild type and R21G variant proteins was determined using yeast transfer RNA (t-RNA) as substrate. Proteins were added in the concentration range of 0.05–0.5 mg/ml to a 150-μl assay mixture containing 0.3 mg yeast t-RNA (Sigma), 15 mg RNase-free bovine serum albumin (HiMedia), 30 mM 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES) at pH 6.8 (MP Biomedicals), and 30 mM sodium chloride (HiMedia), and the assay mixtures were incubated for 2 h at 37 °C. Subsequently, 350 μl of 3.4% pre-chilled, ice-cold perchloric acid (Sigma) was added to the assay mixtures to terminate the reactions. The mixtures were then rigorously vortexed, incubated in ice for 15 min, followed by centrifugation at 14,000 rpm for 12 min at 4 °C. The supernatants were collected and diluted by 3-fold, and their absorbance was measured at 260 nm. The percentage loss of ribonucleolytic activity was calculated by considering the difference in the optical density of wild type and variant at a particular concentration of protein (0.3 mg/ml). All the experiments were done in triplicates.
Nuclear translocation assay
The nuclear translocation ability of wild type and R21G variant was examined in HeLa cells. These cells were seeded at a density of 3 × 105 cells on cover slip placed in 35 mm culture dishes in Dulbecco’s modified Eagle’s medium (DMEM) (Cell Clone) supplemented with 10% fetal bovine serum (FBS) (Cell Clone) and 1% penicillin-streptomycin solution (Cell Clone). Subsequently, cells were treated with 2 μg/ml of wild type or R21G proteins and incubated for 30 min at 37 °C. Cells were then fixed with 2% paraformaldehyde (Sigma) for 10 min at 4 °C and then permeabilized with 0.2% Triton X-100 (Sigma) at room temperature. Cells were then incubated in 2% bovine serum albumin (HiMedia) to prevent nonspecific binding. The fixed cells were subsequently treated with mouse anti-ANG monoclonal antibody (Abcam), at a concentration of 2.5 μg/ml for 3 h, and Alexa Fluor 555 goat anti-mouse IgG (H + L) (Molecular Probes) for 1 h at room temperature. The nuclei were counter-stained with 300 nM 4′,6-diamidino-2-phenylindole (DAPI) dihydrochloride (Molecular Probes), mounted, and visualized using Olympus fluorescence microscope through a × 60 objective. All experiments were done in duplicates.
Angiogenesis assay
Angiogenic activities of the wild type and R21G variant were determined by assessing endothelial cell tube formation. Angiogenesis assay was carried out in a 96-well plate, which was coated with Gibco™Geltrex™ LDEV-Free Reduced Growth Factor Basement Membrane Matrix by loading 50 μl of matrix in each well and allowing the matrix to solidify at 37 °C. Subsequently, HUVEC cells were seeded at a density of 1 × 104 cells per well in endothelial growth medium from Lonza (EGM™-2 BulletKit™) and incubated with 0.2 μg/ml of wild type or R21G-ANG at 37 °C under humidified 5% CO2 [7]. The cells were imaged at three time points: 1 h (t1), after 3 h (t3), and 6 h (t6). The images from these experiments were then analyzed for tube formation properties using the angiogenesis plugin of the ImageJ software. All experiments were done in duplicates.
Statistical analyses
All data are presented as mean ± the standard deviation. The differences in ribonucleolytic activity between wild type and variant were determined by Student’s t test (two-tailed). Significant differences were observed with p < 0.05, n = 3. The strength of association between the identified tolerated and benign rare/novel variants and ALS was determined through odds ratio (OR) analysis by utilizing the allele frequency from the ExAC database. The significance of OR analysis was determined by using the Fisher exact test. The p values were adjusted for multiple testing using Benjamini-Hochberg method (Table 3).
Results
Novel and rare susceptibility variants
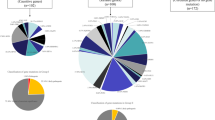
Using targeted NGS analysis, we identified a total of eight susceptibility variants (four rare and four novel variants) in eight patients (5.36%) from our cohort (Table 3) that are predicted to be tolerated and benign by in silico analysis through SIFT and PolyPhen 2.0. Of these, four variants were present at MAF < 0.01 in ExAC; we refer to these four as rare variants. Four variants identified in our cohort were absent in ExAC and hence referred to as novel variants (Table 3). The novel variants we identified include E738G (MATR3), Y1306C (SETX), E69V (ERBB4), and L1207P (DCTN1), and the rare variants include Q2588H (SETX), N226S (PON2), R45G (ANG), and I591V (SETX). Statistical analysis shows that the adjusted p value (for multiple testing) was significant for seven of these variants and was not significant for the PON2 variant, although the minor allele frequency was equal to 0.001. As reported previously, the pathogenic role of mutations associated with ALS can be defined by their absence in large-scale public databases and when the minor allele frequency is < 0.01 [6]. Our study further extends this finding by the detection of such variants in Indian ALS patients, which are tolerated and benign but are present at rare/novel variant frequency.
Repeat expansions in Indian ALS patients with tolerated and benign variants
We had previously reported ATXN2 polyQ repeats of intermediate length in three of these ten ALS patients [16]. Interestingly, among these three patients with ATXN2 polyQ repeats of intermediate length (27–32 repeats), we found that two patients additionally harbored rare/novel variants in the SETX gene (Q2588H and I591V) and one had a variant in the DCTN gene (L1207P), which were identified through this study. Taken together, these findings suggest that about a third of Indian ALS patients with tolerated and benign variants in ALS-associated genes may be carriers of ATXN2 intermediate-length repeat expansions.
Analysis of a rare ANG variant
We functionally characterized the rare ANG variant (R45G representing R21G in mature ANG after the removal of signal peptide sequence of 21 residues) identified in a patient from our cohort. Prior to the functional analysis, we predicted the nature of the R21G variant using our previously developed ANGDelMut web-tool [8]. Preliminary findings from this suggested that the R21G variant may exhibit loss of functions. To validate the computational predictions, we carried out extensive MD simulations of the R21G variant followed by biochemical functional assay experiments.
Functional characterization of the R21G-ANG variant
It has been established that ANG variants cause ALS through loss of either ribonucleolytic activity or nuclear translocation activity or both of these functions caused by a structural and dynamic change in the catalytic triad residues His13, Lys40, and His114 and nuclear localization signal 29IMRRRGL35, respectively. To analyze the pathogenicity of the R21G variant, we generated glutathione S-transferase (GST)-fusion constructs of wild type ANG and the R21G variant with a C-terminus His-tag, then expressed and purified the functional proteins to > 95% homogeneity through Ni-NTA affinity chromatography as described previously [18] (Supplementary Fig. 3a). Biochemical characterization of the purified wild type and R21G variant using CD spectroscopy showed that the far-UV spectra of the R21G was largely similar to that of wild type ANG (Supplementary Fig. 3b), indicating that the R21 to G21 variation did not significantly affect the secondary structure, overall stability, or folding. The root mean square deviation (RMSD) profile of the R21G variant (Supplementary Fig. 3c) also supports our findings from CD spectroscopy, suggesting that the structural stability of the variant was similar to that of wild type ANG.
Conformational switching of a catalytic histidine causes loss of ribonucleolytic activity in the R21G-ANG variant
We visualized the structural and dynamic changes of the catalytic residues from the trajectories of the wild type and R21G using VMD and observed that the R21G variant exhibited a characteristic conformational switching of catalytic residue His114 from its native position, while wild type ANG did not demonstrate any alteration within the catalytic triad from 100 ns MD simulation trajectory (Fig. 1A). This conformational switching of His114 in the R21G variant was supported by the quantitative HA-CA-CB-CG dihedral angle measurement over the course of simulations, where the dihedral angle of the R21G variant changed from the mean − 80° position to − 179° during simulations (Fig. 1B). Our previous results have indicated that this characteristic conformational switching of His114 is detrimental for ribonucleolytic activity of ANG as His114 in its altered conformation is unable to form hydrogen bond and salt-bridge interactions with NCI-65828 (a known inhibitor of the ribonucleolytic activity of ANG) [18]. Thus, based on the MD simulations of the wild type ANG and the R21G variant, we predict that the R21G variant is possibly linked to loss of ribonucleolytic activity.
Ribonucleolytic activity of wild type ANG and R21G variant. A Snapshot of MD simulations for wild type ANG and R21G variant exhibiting loss of ribonucleolytic activity. MD simulation of (a, b) wild type ANG showing stable and native conformation of catalytic residue (c, d) Conformational switching of His114 observed from the MD simulation of R21G variant. B Change in dihedral angle of catalytic residue His114 from MD simulation as a function of time. R21G variant shows change in dihedral angle of His114, indicating its role in loss of ribonucleolytic activity whereas wild type ANG maintained a stable His114 conformation. C. Absorbance and resulting ribonucleolytic activity of wild type (black) and R21G variant (red) proteins measured using yeast t-RNA as substrate
We next performed ribonucleolytic assay using yeast transfer RNA as substrate to validate MD simulation predictions. Our quantitative analysis showed that the R21G variant had only 59% of the ribonucleolytic activity as compared with the wild type (considered 100%) (Fig. 1C).
Local folding of nuclear localization signal abrogates nuclear translocation of the R21G-ANG variant
Next, we investigated the nuclear translocation activity of the R21G variant from MD simulations. Our 100 ns simulation showed that the 31RRR33 residues undergo local folding and becomes less accessible to solvent in R21G as compared with the wild type (Fig. 2A). This resulted in a reduced solvent accessible surface area (SASA) in the R21G variant than the wild type that retained an open conformation and loose packing of 31RRR33 residues during the simulation. Since the 31RRR33 residues are known to play a pivotal role in the nuclear translocation of ANG among the complete nuclear localization signal 29IMRRRGL35, we predicted that a decrease in SASA due to the local folding and close packing of 31RRR33 residues will probably lead to the loss of nuclear translocation activity in the R21G variant (Fig. 2B).
Local folding of nuclear localization signal residues in R21G variant. A Wild type (a, b): MD simulation profile showing open and loose conformation of nuclear localization signal residues 31RRR33, R21G (c, d): MD simulation of variant showing closed and tight packing of 31RRR33residues, indicating loss of nuclear translocation activity. B Computed solvent accessible surface area from MD simulations of nuclear localization signal residues 31RRR33 for wild type ANG and R21G variant
To verify our MD simulation-based prediction that the R21G variant would indeed result in loss of nuclear translocation activity, we performed nuclear translocation assay. We observed that while wild type ANG translocated into the nucleus and the R21G variant remained primarily in the cytoplasm with decreased nuclear translocation (Fig. 3). Taken together, our results demonstrate that the R21G-ANG variant identified in our ALS patient, though predicted to be tolerated and benign in silico, is a loss-of-function variant exhibiting loss of both ribonucleolytic and nuclear translocation activities.
Nuclear translocation activity of wild type ANG and R21G variant. Nuclear translocation assay demonstrating the activity of wild type ANG and R21G variant. Individual and merge channels are shown. All images were taken at × 60 magnification and experiments were done in duplicates. The images are representative of cells from at least three areas (containing 20–25 cells) from two independent experiments
Loss of angiogenic property
To evaluate the role of the R21G-ANG variant on its angiogenic ability, we performed the in vitro angiogenesis assay using HUVEC cells and subsequently assessing its tube formation characteristic on the Matrigel. It was observed that while the wild type ANG formed tubular structures as expected upon the treatment of angiogenic factor, the R21G variant does not show the formation of these structures, thereby indicating loss of angiogenic activity (Fig. 4). It is interesting to note that in the wild type ANG, the formation of tubular structures was consistently higher until 3- and 6-h time points compared with the R21G variant. This further confirmed that the R21G-ANG is a loss-of-function variant.
Angiogenic activity of wild type ANG and R21G variant proteins. Endothelial cell tube formation assay performed on human umbilical vein endothelial cells (HUVECs) were cultured on serum-free Matrigel and imaged at three time points t1,t3, and t6 to assess the formation of tubular structures. The difference between the angiogenic activity of wild type and R21G was found to be significant p < 0.05 for the properties (total length, total branching length, total segment lengths, mesh index, mean mesh size, and the path) of HUVEC tubes determined through Angiogenesis plugin of ImageJ software. Images shown are representative of two independent experiments. Original magnification, × 10
Discussion
In this study, we report eight novel and rare variants predicted to be tolerated and benign in silico in ALS-associated genes of Indian patients using targeted sequencing. For variant classification as novel and rare, we mined the allele frequency data from the ExAC database to determine the presence of these variants across individuals of different ethnicities. These patients were also screened for 25 ALS-associated genes [17] in addition to the repeat expansions in the ATXN2 and C9orf72 [16]. Interestingly, we also observed that certain Indian ALS patients with rare/novel variations also carry intermediate-length repeat expansions in ATXN2 gene, thereby supporting the proposition of increased ALS risk in individuals carrying multiple genetic aberrations [23]. Intermediate-length repeat expansions (27–33 CAG) in the ATXN2 have been previously identified as a potential risk factor for ALS [24]. The presence of aberrations in more than one ALS-associated genes further supports the premise that a fraction of sALS could be influenced by such multiple genetic perturbations, which in turn could exert a synergistic deleterious effect leading to neurodegeneration [3].
We also searched the ALSoD [25] and ALSdb (http://alsdb.org/) databases for all the missense mutations reported in ANG, DCTN1, ERBB4, MATR3, PON2, and SETX genes. We then predicted the nature of these mutations using SIFT and PolyPhen 2.0 (Table 4) and observed that several mutations that are predicted as tolerated/benign have previously been found in ALS patients and constitute around 50% of the total mutations reported in ALSoD and ALSdb. Additional literature searches also revealed that some of these predicted tolerated and benign mutations such as reported in ANG [22] and MATR3 [10] have shown to be pathogenic through functional studies (Table 1).
We detected a heterozygous missense mutation, E738G (MATR3), in a young female sALS patient with bulbar onset. Earlier reports show that mutations in MATR3 have predisposed at a frequency of 0.2, 1.6, and 1.8% in Taiwanese and Chinese, Italian, and Canadian sALS cases, respectively [11, 26,27,28]. The frequency of variation carrier in this gene is 0.67% (1/149) in our cohort and is slightly higher as compared with that reported in ALS cases in other Asian countries including Taiwan and China [25].
DCTN1 plays an important role in maintenance of cytoskeletal dynamics and is involved in the retrograde axonal transport [29]. Missense mutations in DCTN1 have been proposed as a plausible risk factor for ALS and have been reported in Chinese, American, Irish, and Japanese ALS cases at a frequency of 0.39–2.87% in sALS cases [30,31,32,33]. We identified a novel variant L1207P in one sALS patient in this gene, who also harbored intermediate-length repeat in ATXN2 gene.
We identified a novel E69V variant in the ERBB4 gene in one (0.67%) sALS case from our cohort. Mutation in ERBB4 was initially identified in a Japanese fALS and subsequently in a Canadian family [34].
Polymorphisms in paraoxonase family of genes (PON1, PON2, and PON3) have been reported as risk factors for developing sALS in Caucasians [35]. This family of genes has a crucial role in enzymatic breakdown of nerve toxins and plays a role in cellular response to oxidative stress [36]. Mutations in this gene have been associated with sALS manifestation in Polish, North American, French, Canadian, and Swedish populations [37, 38]. In this study, we identified a rare variant in PON2 in one sALS patient from our cohort.
SETX emerged to be an important gene in our panel as we identified three variants (one novel and two rare) in three sALS cases, each carrying one variant. We found variants in this gene in 2.01% (3/149) of our adult-onset sALS cases. Interestingly, two of these variant carriers (one novel and one rare) had intermediate-length expansion in ATXN2. Previously, mutations in this gene were associated with juvenile-onset sALS [39, 40]. Importantly, our study substantiates the link between SETX and adult-onset ALS, as all three SETX variant carriers in our cohort were adult ALS patients.
It is now well established that nonsynonymous variations in ANG manifest ALS through loss-of-function mechanisms. A recent study in Hungarian ALS patients further lays emphasis on the role of ANG mutations in ALS manifestation by identifying this mutation in 2.9% of the cases [41]. Previous studies reported by us [21, 22] and others [7, 9] have demonstrated that ANG variants cause ALS through loss of either ribonucleolytic or nuclear translocation or both of these activities. The substitution in ANG (R45G; R21G in mature ANG protein) occurred in only 1 ALS patient, and this variation was present at rare heterozygous state in only 2 of 60,705 individuals in ExAC database. Preliminary analysis by ANGDelMut web-tool [8] suggested that the R21G variant would exhibit loss of functions. Therefore, this variant was a good candidate for further computational and functional analyses for determining its possible association with ALS.
In order to confirm the pathogenicity of rare ANG variant, we employed extensive MD simulations and found that although the G21 is distally located from the catalytic triad, it induced a characteristic conformational switching of catalytic residue His114 during the simulations. The conformational change of the His114 residue is important because it has previously been shown that a mutation of this residue could cause a 3300-fold reduction in the ribonucleolytic activity of ANG. The R21G variant had only 59% of ribonucleolytic activity compared with the wild type ANG, thus completely corroborating our MD simulation predictions with functional assays and thereby suggesting that the conformational alteration of catalytic residue His114 is indeed responsible for loss of ribonucleolytic activity.
Next, our investigation on nuclear translocation activity showed that the R21G variant could adversely affect nuclear translocation ability of the variant, as it exhibited a local folding of nuclear localization signal residues 31RRR33 during the simulations resulting in reduction of SASA. The nuclear translocation assay in HeLa cells clearly demonstrated that while the wild type ANG translocated into the nucleus, the R21G variant had decreased localization in the nucleus and remained predominantly in the cytoplasm. Moreover, angiogenesis assay revealed that the R21G variant would lose its angiogenic activity in the HUVEC cells. As the loss of ribonucleolytic, angiogenic, and nuclear translocation activities in ANG has been shown to be critical for abolishment of neurite outgrowth, path finding, and several other crucial biological functions, our combined simulation and functional assay results suggest that the novel R21G variant identified in our Indian sALS patient could impair its major neuroprotective functions, leading to ALS origin and progression.
Conclusion
In sum, our findings highlight the significance of novel and rare variants in manifestation of ALS. Our computational and experimental results for the R21G variant of ANG show that it exhibits a significant loss-of-function characteristic. Hence, variants in ALS-associated genes predicted to be tolerated/benign can play a role in the pathogenesis of ALS. This work further emphasizes the need for prioritizing functional studies of ALS-associated genetic variants regardless of the predicted functional impact.
References
Renton AE, Chiò A, Traynor BJ (2014) State of play in amyotrophic lateral sclerosis genetics. Nat Neurosci
Al-Chalabi A, Hardiman O (2013) The epidemiology of ALS: a conspiracy of genes, environment and time. Nat Rev Neurol. https://doi.org/10.1038/nrneurol.2013.203
Cady J, Allred P, Bali T et al (2015) Amyotrophic lateral sclerosis onset is influenced by the burden of rare variants in known amyotrophic lateral sclerosis genes. Ann Neurol. https://doi.org/10.1002/ana.24306
Pang SYY, Hsu JS, Teo KC et al (2017) Burden of rare variants in ALS genes influences survival in familial and sporadic ALS. Neurobiol Aging. https://doi.org/10.1016/j.neurobiolaging.2017.06.007
Krüger S, Battke F, Sprecher A et al (2016) Rare variants in neurodegeneration associated genes revealed by targeted panel sequencing in a German ALS cohort. Front Mol Neurosci. https://doi.org/10.3389/fnmol.2016.00092
Marangi G, Traynor BJ (2015) Genetic causes of amyotrophic lateral sclerosis: new genetic analysis methodologies entailing new opportunities and challenges. Brain Res
Wu D, Yu W, Kishikawa H et al (2007) Angiogenin loss-of-function mutations in amyotrophic lateral sclerosis. Ann Neurol. https://doi.org/10.1002/ana.21221
Padhi AK, Vasaikar SV, Jayaram B, Gomes J (2014) ANGDelMut—a web-based tool for predicting and analyzing functional loss mechanisms of amyotrophic lateral sclerosis-associated angiogenin mutations. F1000Research. https://doi.org/10.12688/f1000research.2-227.v3
Thiyagarajan N, Ferguson R, Subramanian V, Acharya KR (2012) Structural and molecular insights into the mechanism of action of human angiogenin-ALS variants in neurons. Nat Commun. https://doi.org/10.1038/ncomms2126
Boehringer A, Garcia-Mansfield K, Singh G et al (2017) ALS associated mutations in Matrin 3 alter protein-protein interactions and impede mRNA nuclear export. Sci Rep. https://doi.org/10.1038/s41598-017-14924-6
Marangi G, Lattante S, Doronzio PN et al (2017) Matrin 3 variants are frequent in Italian ALS patients. Neurobiol Aging. https://doi.org/10.1016/j.neurobiolaging.2016.09.023
Padhi AK, Narain P, Dave U et al (2019) Insights into the role of ribonuclease 4 polymorphisms in amyotrophic lateral sclerosis. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2017.1419147
de Majo M, Topp SD, Smith BN et al (2018) ALS-associated missense and nonsense TBK1 mutations can both cause loss of kinase function. Neurobiol Aging. https://doi.org/10.1016/j.neurobiolaging.2018.06.015
Higelin J, Catanese A, Semelink-Sedlacek LL et al (2018) NEK1 loss-of-function mutation induces DNA damage accumulation in ALS patient-derived motoneurons. Stem Cell Res. https://doi.org/10.1016/j.scr.2018.06.005
Padhi AK, Hazra S (2019) Insights into the role of d-amino acid oxidase mutations in amyotrophic lateral sclerosis. J Cell Biochem. https://doi.org/10.1002/jcb.27529
Narain P, Gomes J, Bhatia R et al (2017) C9orf72 hexanucleotide repeat expansions and Ataxin 2 intermediate length repeat expansions in Indian patients with amyotrophic lateral sclerosis. Neurobiol Aging. https://doi.org/10.1016/j.neurobiolaging.2017.04.011
Narain P, Pandey A, Gupta S et al (2018) Targeted next-generation sequencing reveals novel and rare variants in Indian patients with amyotrophic lateral sclerosis. Neurobiol Aging. https://doi.org/10.1016/j.neurobiolaging.2018.05.012
Ng PC, Henikoff S (2003) SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res
Adzhubei IA, Schmidt S, Peshkin L et al (2010) A method and server for predicting damaging missense mutations. Nat Methods. https://doi.org/10.1038/nmeth0410-248
Lek M, Karczewski KJ, Minikel EV et al (2016) Analysis of protein-coding genetic variation in 60,706 humans. Nature. https://doi.org/10.1038/nature19057
Padhi AK, Kumar H, Vasaikar SV et al (2012) Mechanisms of loss of functions of human angiogenin variants implicated in amyotrophic lateral sclerosis. PLoS One. https://doi.org/10.1371/journal.pone.0032479
Padhi AK, Banerjee K, Gomes J, Banerjee M (2014) Computational and functional characterization of angiogenin mutations, and correlation with amyotrophic lateral sclerosis. PLoS One. https://doi.org/10.1371/journal.pone.0111963
Al-Chalabi A, Van Den Berg LH, Veldink J (2017) Gene discovery in amyotrophic lateral sclerosis: implications for clinical management. Nat Rev Neurol
Sproviero W, Shatunov A, Stahl D et al (2017) ATXN2 trinucleotide repeat length correlates with risk of ALS. Neurobiol Aging. https://doi.org/10.1016/j.neurobiolaging.2016.11.010
Abel O, Powell JF, Andersen PM, Al-Chalabi A (2012) ALSoD: a user-friendly online bioinformatics tool for amyotrophic lateral sclerosis genetics. Hum Mutat. https://doi.org/10.1002/humu.22157
Lin KP, Tsai PC, Liao YC et al (2015) Mutational analysis of MATR3 in Taiwanese patients with amyotrophic lateral sclerosis. Neurobiol Aging. https://doi.org/10.1016/j.neurobiolaging.2015.02.008
Xu L, Li J, Tang L et al (2015) MATR3 mutation analysis in a Chinese cohort with sporadic amyotrophic lateral sclerosis. Neurobiol Aging. https://doi.org/10.1016/j.neurobiolaging.2015.11.023
Leblond CS, Gan-Or Z, Spiegelman D et al (2016) Replication study of MATR3 in familial and sporadic amyotrophic lateral sclerosis. Neurobiol Aging. https://doi.org/10.1016/j.neurobiolaging.2015.09.013
Brown RH, Al-Chalabi A (2017) Amyotrophic lateral sclerosis. Prog Med Chem. https://doi.org/10.1016/bs.pmch.2018.12.001
Couthouis J, Raphael AR, Daneshjou R, Gitler AD (2014) Targeted exon capture and sequencing in sporadic amyotrophic lateral sclerosis. PLoS Genet. https://doi.org/10.1371/journal.pgen.1004704
Kenna KP, McLaughlin RL, Byrne S et al (2013) Delineating the genetic heterogeneity of ALS using targeted high-throughput sequencing. J Med Genet. https://doi.org/10.1136/jmedgenet-2013-101795
Maruyama H, Morino H, Ito H et al (2010) Mutations of optineurin in amyotrophic lateral sclerosis. Nature. https://doi.org/10.1038/nature08971
Nakamura R, Sone J, Atsuta N et al (2016) Next-generation sequencing of 28 ALS-related genes in a Japanese ALS cohort. Neurobiol Aging. https://doi.org/10.1016/j.neurobiolaging.2015.11.030
Takahashi Y, Fukuda Y, Yoshimura J et al (2013) Erbb4 mutations that disrupt the neuregulin-erbb4 pathway cause amyotrophic lateral sclerosis type 19. Am J Hum Genet. https://doi.org/10.1016/j.ajhg.2013.09.008
Zhang G, Li W, Li Z et al (2013) Association between paraoxonase gene and stroke in the Han Chinese population. BMC Med Genet. https://doi.org/10.1186/1471-2350-14-16
Valdmanis PN, Kabashi E, Dyck A et al (2008) Association of paraoxonase gene cluster polymorphisms with ALS in France, Quebec, and Sweden. Neurology. https://doi.org/10.1212/01.wnl.0000324997.21272.0c
Saeed M, Siddique N, Hung WY et al (2006) Paraoxonase cluster polymorphisms are associated with sporadic ALS. Neurology. https://doi.org/10.1212/01.wnl.0000227187.52002.88
Slowik A, Tomik B, Wolkow PP et al (2006) Paraoxonase gene polymorphisms and sporadic ALS. Neurology. https://doi.org/10.1212/01.wnl.0000219565.32247.11
Chen Y-Z, Bennett CL, Huynh HM et al (2004) DNA/RNA helicase gene mutations in a form of juvenile amyotrophic lateral sclerosis (ALS4). Am J Hum Genet. https://doi.org/10.1086/421054
Hirano M, Quinzii CM, Mitsumoto H et al (2011) Senataxin mutations and amyotrophic lateral sclerosis. Amyotroph Lateral Scler. https://doi.org/10.3109/17482968.2010.545952
Tripolszki K, Danis J, Padhi AK et al (2019) Angiogenin mutations in Hungarian patients with amyotrophic lateral sclerosis: clinical, genetic, computational, and functional analyses. Brain Behav. https://doi.org/10.1002/brb3.1293
Acknowledgments
Helpful suggestions from Prof B. Jayaram (IIT Delhi) are gratefully acknowledged.
Funding
The authors gratefully acknowledge the funding received from the Kusuma Trust UK and the financial and infrastructure support of the Indian Institute of Technology, Delhi for carrying out this research. DM, PN, and UD were supported by Research Fellowship from IIT Delhi, India.
Author information
Authors and Affiliations
Contributions
PN, AKP, VP, and JG conceived and designed the experiments. PN performed the experiments. AKP performed molecular dynamics simulations and analyses. PN, UD, and DM handled protein purification and assays. PN, AKP, UD, DM, VP, and JG analyzed the data. RB handled patient recruitment and clinical evaluation. PN, AKP, RB, VP, and JG contributed to the writing of the manuscript.
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Narain, P., Padhi, A.K., Dave, U. et al. Identification and characterization of novel and rare susceptible variants in Indian amyotrophic lateral sclerosis patients. Neurogenetics 20, 197–208 (2019). https://doi.org/10.1007/s10048-019-00584-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10048-019-00584-3