Abstract
The application of molecular parameters in the World Health Organization classification of central nervous system tumors has advanced remarkably in this field. Large-scale genomic DNA analyses, including gene expression profiling, genome-wide association studies, and single-nucleotide polymorphism analysis, have revealed differences between tumors with the same pathological features. Because mutated genes and their signaling pathways can be targets for therapy, categorizing tumors by molecular parameters facilitates the selection of optimal therapeutic methods. Many genes are either oncogenes or tumor suppressor genes, and many of them are also involved in normal development, such as neural stem cell maintenance and neural differentiation. Moreover, genetic engineering has enabled the generation of tumors that phenocopy human tumors in mice. Here, I will discuss key molecular parameters, mechanisms of neural differentiation, isocitrate dehydrogenases, 1p36/19q13, and p53 in gliomagenesis. Because future therapeutic methods will be determined by the molecular mechanisms of tumors, identification of new parameters is still needed for further classification of glioma.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
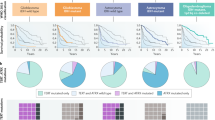
The main change in the 2016 World Health Organization (WHO) classification of central nervous system (CNS) tumors is the use of molecular parameters for diagnosis [1]. This change is a significant advance, because genome-wide gene expression analyses of CNS tumors have revealed that tumors diagnosed in the same group (e.g., glioblastoma; GBM) have different gene expression profiles, which were recently categorized as classical, neural, pro-neural, and mesenchymal types [2]. In addition, therapeutic targets are largely dependent on the molecular mechanisms of tumors. Therefore, classification of CNS tumors by molecular parameters is appropriate.
The development of glioma involves many factors. Three signaling pathways, namely, p53, retinoblastoma (RB), and receptor tyrosine kinase (RTK), play crucial roles in GBM development [3, 4]. Mutations in isocitrate dehydrogenase 1 and 2 (IDH1/2), which reduce their enzymatic activities, have been the center of attention in gliomagenesis, because patients with IDH mutations have a better outcome than those with wild-type IDHs [5, 6]. Chromosomes 1p36 and 19q13 are frequently deleted in oligodendroglioma, although the genes encoded in these chromosomes have not been identified yet [7–9]. It has been further demonstrated that factors, which regulate stem cell maintenance and inhibit differentiation, are aberrantly activated in glioma [10–14]. These factors are not only diagnostic markers and therapeutic targets, but can also be used to establish glioma models with the pathological features of human tumors [2–15].
Based on these findings, I will summarize how these factors are involved in gliomagenesis and discuss future insights for glioma therapy.
Molecular mechanisms of glial differentiation
Neural stem cells (NSCs) self-renew and give rise to neurons, astrocytes, and oligodendrocytes [10, 11]. Many extrinsic and intrinsic factors have been demonstrated to regulate NSC self-renewal and differentiation in rodent systems (Fig. 1). Basic fibroblast growth factor (bFGF) and epidermal growth factor (EGF) are used to maintain NSCs in culture [16, 17]. Sox2, Bmi1, Hairy-and-enhancer-split (Hes), and Inhibitor-of-differentiation (Id) are involved in NSC maintenance by preventing the functions of basic helix-loop-helix (bHLH)-type differentiation inducers, such as Mash1 and neurogenin, and the cell cycle inhibitor Ink4a/ARF [18–21].
Bone morphogenic proteins (BMPs) and leukemia inhibitory factor (LIF)/ciliary neurotrophic factor (CNTF) induce astrocytic differentiation through activation of Smads and STAT3 transcription factors, respectively [22, 23], whereas BMPs block neuronal and oligodendrocyte differentiation by induction of Ids [24, 25]. Many other factors also play dual functions in a similar manner. Notch signaling maintains NSCs and induces astrocytiv differentiation, but it blocks neuronal and oligodendrocyte differentiation by induction of Hes transcription factors. Oligodendrocyte lineage transcription factor (Olig) 2, which is induced by hedgehog (Hh)-Gli signaling, is not only involved in oligodendrocyte differentiation by forming a heterodimer with the homeobox transcription factor Nkx2.2, but also prevents astrocytic differentiation by blocking STAT3-p300 histone acetyltransferase association [26]. In addition, both Wnt and Hh promote oligodendrocyte differentiation and NSC self-renewal. Oligodendrocyte differentiation is also regulated by platelet-derived growth factor AA (PDGFAA), thyroid hormone (TH), and retinoic acid (RA) [27–34]. Fyn kinase regulates oligodendrocyte morphology through activation of the integrin α6/β1 complex [35]. Although many factors have been identified to exclusively regulate NSC maintenance and glial differentiation, it remains to evaluate whether the same factors regulate human NSC maintenance and their differentiation.
Functions of IDHs and their mutants
Whole genome sequences of human GBMs have revealed IDH1 and IDH2 mutations at amino acid residues 132 (mainly arginine to histidine) and 172 (arginine to lysine, methionine, or glycine), respectively [5, 6]. Surprisingly, IDH1 mutations have been found in more than 70% of low-grade astrocytoma and oligodendroglioma [6]. Over 80% of secondary GBMs also contain the same mutation, whereas less than 5% of primary GBMs contain the mutation [5]. In addition, the Cancer Genome Atlas has shown that the average ages of GBM patients with wild-type or mutant IDH1 are 53.3 and 33.2 years, respectively [6]. Taken together, these data suggest that IDH1 is primarily mutated in gliomagenesis.
Wild-type IDH1 converts isocitrate to α-ketoglutarate (α-KG) by oxidation of nicotinamide adenine dinucleotide phosphate (NADP+) to NADPH, whereas mutant IDH1 catalyzes reduction of α-KG to 2-hydroxyglutarate (2-HG), which is called an oncometabolite, with production of NADP+ from NADPH (Fig. 2) [36]. Moreover, 2-HG promotes tumorigenesis through multiple mechanisms. First, 2-HG competitively inhibits the activity of α-KG-dependent Jumonji C domain containing histone demethylases (JHDMs), thereby maintaining the repressive histone methylation of certain chromosomal domains, such as trimethylation of H3K9 and H3K27, and blocking cell differentiation [37]. Second, 2-HG blocks α-KG-dependent prolyl hydroxylase (PHD) activity, which induces degradation of hypoxia-inducible factor 1α (HIF1α) through the proteasome pathway, and promotes activation of HIF1α downstream factors including nuclear factor-κB (NF-κB) [38]. Third, 2-HG blocks mature collagen formation by α-KG-dependent PHD activity [39]. Because precise collagen formation is essential for the proper function of various types of tissues and cytoplasmic apparatus, abnormal collagen, such as misfolding, might contribute to tumorigenesis by induction of necrosis and tumor angiogenesis. Finally, decreased levels of NADPH by mutant IDHs prevent the conversion of glutathione disulfide to glutathione, a major antioxidant, causing an increase of reactive oxygen species that induce DNA damage and genetic instability [40]. Thus, mutant IDH1 likely prepares tumorigenesis in multiple manners.
Metabolic regulation in mutant IDH-bearing cancer cells. Mutant IDH1 converts α-ketoglutarate (α-KG), which activates Jumonji-C domain histone demethylases (JHDMs) and TET DNA demethylase 2 (TET2), to 2-hydroxyglutarate (2-HG). In turn, 2-HG not only competes with α-KG for JHDM binding, but also inhibits demethylases. Reduction of α-KG suppresses the Krebs cycle and increases HIF1α expression. Eventually, these events induce the expression of stemness- and cancer-related genes
Genes encoded on human 1p36 and 19q13
Chromosomes 1p36 and 19q13 are frequently deleted in oligodendroglioma, thereby making this co-deletion a surrogate marker. Whole genome sequence analysis has revealed that these chromosomal regions contain many important genes involved in tumorigenesis and oligodendrocyte differentiation, including Notch-related factors (HES2-5, MINDBOMB2, and DLL3), WNT factors (WNT4, DVL1, and GSK3A), oncogenes/proto-oncogenes/tumor suppressors (p73, CHD5, SKI, HKR1, AKT2, TGFB1, ARHGAP35, and FOSB), apoptosis-related factors (DFFA, DFFB, CASPASE9, and BAX), and cancer-related factors [mTOR (mammalian target of Rapamycin), CEACAM1, PLAUR, RELB, and DYRK1B] (Fig. 3). The fact that 1p36 encodes important apoptosis regulators DFFA, DFFB, and CASPASE 9 suggests that oligodendroglioma cells with 1p36 loss may not exhibit typical apoptotic phenotypes in their death.
Among these genes, p73 and Arhgap35 (also known as p190RhoGAP) are involved in oligodendrocyte differentiation. Overexpression of a dominant negative form of p73 inhibits differentiation of oligodendrocyte precursor cells (OPCs) into mature oligodendrocytes in culture [41]. Phosphorylated Arhgap35 by Fyn kinase induces morphological changes in differentiating oligodendrocytes [42], whereas its overexpression induces cell cycle arrest and enhances process extension [43]. In addition, Arhgap35 is involved in cell motility and metastasis [44]. These findings suggest that co-deletion of 1p36 and 19q13 might maintain OPCs in the undifferentiated state, although it is unknown why co-deletion is specific to oligodendroglioma.
Bagchi et al. identified chromodomain helicase DNA binding domain 5 (CHD5) encoded on 1p36 and demonstrated that CHD5 controls cell proliferation, apoptosis, and senescence through the cyclin-dependent kinase inhibitor (CDKI) p14ARF-p53 pathway, thereby identifying CHD5 as a new tumor suppressor gene [45]. This finding indicates that p53 functions should decrease in oligodendroglioma with 1p36 loss, although 1p36 loss and p53 mutation are exclusive in glioma.
It is notable that 1p36 and 19q13 encode many differentiation inhibitors and oncogenes/proto-oncogenes. Among them, both AKT2 and mTOR are key effectors in gliomagenesis in multiple mechanisms [46, 47]. CEACAM1, the plasminogen activator system, and non-canonical NF-κB pathway also play crucial roles in GBM-initiating cells [48–50]. These findings might explain the reason why 1p36 and 19q13-co-deleted oligodendrogliomas are susceptible to therapy.
p53 regulates multiple functions
p53 was originally found as an essential tumor suppressor preventing cell cycle progression and inducing apoptosis [51]. External and internal stress signals, including DNA damage, oncogene activation, hypoxia, and nutrient stress, induce p53 expression at transcriptional and post-transcriptional levels and regulate its activation through various kinds of modification, including phosphorylation, ubiquitination, acetylation, methylation, glycosylation, sumoylation, and ADP ribosylation (Fig. 4) [52, 53]. It is well known that ubiquitination of p53 by mouse double minute gene 2 (Mdm2) induces its degradation via a proteasome pathway, whereas phosphorylation at the N-terminal blocks the interaction of p53 with Mdm2 and induces cell cycle arrest and senescence by increasing Cdki p21/Cip1 and plasminogen activator inhibitor 1 (Pai1) [54]. Acetylation of p53 at lysine 120 activates p21/Cip1 expression by binding with monocytic and promyelocytic leukemia zinc finger proteins or triggers cell death by inducing p53-upregulated modulator of apoptosis (Puma) and cofactors in a cell context-dependent manner [55]. In addition, p53 induces the expression of other apoptosis-related factors, such as Bcl2-associated X protein (Bax), Noxa, Fas, death receptor (DR) 4 and DR5.
It is evident that p53 supports the oxidative phosphorylation pathway and blocks glycolysis at multiple steps. For example, p53 inhibits expression of glucose transporters Glut1 and 4, glycolytic enzyme phosphoglycerate mutase (Pgm), and pyruvate dehydrogenase kinase 2 (Pdk2) that prevent acetyl-CoA production from pyruvate. Moreover, p53 induces expression of Tiger, which reduces glycolysis indirectly, glutaminase 2 (Gls2) that produces glutamate from glutamine, another source of α-KG (Fig. 2), and apoptosis-inducing factor (Aif) that is essential for both the electron transport chain in mitochondria and DNA fragmentation in apoptosis.
There is increasing evidence that p53 induces the expression of autophagy-related genes, including DNA-damaged-regulated autophagy-modulator 1 (Dram1) and UNC51-like autophagy-activating kinase 1 (Ulk1). Conversely, autophagy reduces various types of stress, such as DNA damage and damaged organelles, which activate p53. Furthermore, p53 binds to autophagy-related factors, such as autophagy protein 7 (ATG7) and RB1 inducible coiled-coil 1 (RB1CC1), and inhibits cell cycle progression and autophagy [56]. It would be of interest to further analyze the relationship between p53 and autophagy in tumorigenesis.
Taken together, these findings indicate that p53 blocks tumorigenesis by regulating multiple pathways, DNA repair, senescence, cell death, glycolysis, mitochondrial functions, and autophagy.
Conclusion and future perspective
There is no doubt that the molecular parameters introduced in the new WHO classification of CNS tumors will facilitate diagnosis of certain tumors with difficult pathological classification. Simultaneously, this raises the interesting question of why some tumors with different molecular parameters, such as GBM with wild-type or mutant IDH, show the same characteristics pathologically. Diagnostic parameters and other mutations in p53, RB, and RTK pathways may share targets or complement essential oncogenic signals [3, 4]. For example, HIF1α is activated by mutant IDH-dependent reduction of α-KG, while activation of RTKs indicues HIF1α, indicating that HIF1α is commonly activated in GBM with wild-type or mutant IDH in different manners [57]. The shared regulators are likely primary therapeutic targets. Furthermore, such parameter-specific factors can be useful as selective targets.
1p36/19q13 loss and p53 mutation are exclusive markers for oligodendroglioma and astrocytoma, respectively. Nonetheless, the evidence that CHD5, a functional activator of p53, is encoded on 1p36, indicates that p53 functions may decrease in 1p36-deleted oligodendroglioma as well as p53-mutated astrocytoma. Billon et al. have shown that p53 is also involved in oligodendrocyte differentiation, suggesting that p53 mutations found in astrocytoma may prevent oligodendrocyte differentiation [41]. Thus, it is essential to identify factors regulating glioma phenotypes, oligodendroglioma, and astrocytoma on 1p36/19q13. In summary, further investigations of molecular mechanisms involved in glioma may be applied to diagnosis, new molecular classification, and therapy in the future.
References
Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK, Ohgaki H, Wiestler OD, Kleihues P, Ellison DW (2016) The 2016 World Health Organization Classification of Tumors of the central nervous system: a summary. Acta Neuropathol 131:803–820
Verhaak RG, Hoadley KA, Purdom E, Wang V, Qi Y, Wilkerson MD, Miller CR, Ding L, Golub T, Mesirov JP, Alexe G, Lawrence M, O’Kelly M, Tamayo P, Weir BA, Gabriel S, Winckler W, Gupta S, Jakkula L, Feiler HS, Hodgson JG, James CD, Sarkaria JN, Brennan C, Kahn A, Spellman PT, Wilson RK, Speed TP, Gray JW, Meyerson M, Getz G, Perou CM, Hayes DN; Cancer Genome Atlas Research Network (2010) Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17:98–110
Cancer Genome Atlas Research Network (2008) Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455:1061–1068
Parsons DW, Jones S, Zhang X, Lin JC, Leary RJ, Angenendt P, Mankoo P, Carter H, Siu IM, Gallia GL, Olivi A, McLendon R, Rasheed BA, Keir S, Nikolskaya T, Nikolsky Y, Busam DA, Tekleab H, Diaz LA Jr, Hartigan J, Smith DR, Strausberg RL, Marie SK, Shinjo SM, Yan H, Riggins GJ, Bigner DD, Karchin R, Papadopoulos N, Parmigiani G, Vogelstein B, Velculescu VE, Kinzler KW (2008) An integrated genomic analysis of human glioblastoma multiforme. Science 321:1807–1812
Yan H, Parsons DW, Jin G, McLendon R, Rasheed BA, Yuan W, Kos I, Batinic-Haberle I, Jones S, Riggins GJ, Friedman H, Friedman A, Reardon D, Herndon J, Kinzler KW, Velculescu VE, Vogelstein B, Bigner DD (2009) IDH1 and IDH2 mutations in gliomas. N Engl J Med 360:765–773
Dimitrov L, Hong CS, Yang C, Zhuang Z, Heiss JD (2015) New developments in the pathogenesis and therapeutic targeting of the IDH1 mutation in glioma. Int J Med Sci 12:201–213
Reifenberger J, Reifenberger G, Liu L, James CD, Wechsler W, Collins VP (1994) Molecular genetic analysis of oligodendroglial tumors shows preferential allelic deletions on 19q and 1p. Am J Pathol 145:1175–1190
Ueki K (2005) Oligodendroglioma: impact of molecular biology on its definition, diagnosis and management. Neuropathology 25:247–253
Wesseling P, van den Bent M, Perry A (2015) Oligodendroglioma: pathology, molecular mechanisms and markers. Acta Neuropathol 129:809–827
Gage F (2000) Mammalian neural stem cells. Science 287:1433–1438
Temple S (2001) The development of neural stem cells. Nature 414:112–117
Singh SK, Clarke ID, Hide T, Dirks PB (2004) Cancer stem cells in nervous system tumors. Oncogene 23:7267–7273
Kondo T (2006) Brain cancer stem-like cells. Eur J Cancer 42:1237–1242
Vescovi AL, Galli R, Reynolds BA (2006) Brain tumour stem cells. Nat Rev Cancer 6:425–436
Hide T, Takezaki T, Nakatani Y, Nakamura H, Kuratsu J, Kondo T (2009) Sox11 prevents tumorigenesis of glioma-initiating cells by inducing neuronal differentiation. Cancer Res 69:7953–7959
Reynolds BA, Weiss S (1992) Generation of neurons and astrocytes from isolated cells of the adult mammalian central nervous system. Science 255:1707–1710
Vescovi AL, Reynolds BA, Fraser DD, Weiss S (1993) bFGF regulates the proliferative fate of unipotent (neuronal) and bipotent (neuronal/astroglial) EGF-generated CNS progenitor cells. Neuron 11:951–966
Norton JD (2000) ID helix-loop-helix proteins in cell growth, differentiation and tumorigenesis. J Cell Sci 113:3897–3905
Graham V, Khudyakov J, Ellis P, Pevny L (2003) SOX2 functions to maintain neural progenitor identity. Neuron 39:749–765
Bruggeman SW, Valk-Lingbeek ME, van der Stoop PP, Jacobs JJ, Kieboom K, Tanger E, Hulsman D, Leung C, Arsenijevic Y, Marino S, van Lohuizen M (2005) Ink4a and Arf differentially affect cell proliferation and neural stem cell self-renewal in Bmi1-deficient mice. Genes Dev 19:1438–1443
Molofsky AV, He S, Bydon M, Morrison SJ, Pardal R (2005) Bmi-1 promotes neural stem cell self-renewal and neural development but not mouse growth and survival by repressing the p16Ink4a and p19Arf senescence pathways. Genes Dev 19:1432–1437
Bonni A, Sun Y, Nadal-Vicens M, Bhatt A, Frank DA, Rozovsky I, Stahl N, Yancopoulos GD, Greenberg ME (1997) Regulation of gliogenesis in the central nervous system by the JAK-STAT signaling pathway. Science 278:477–483
Nakashima K, Yanagisawa M, Arakawa H, Kimura N, Hisatsune T, Kawabata M, Miyazono K, Taga T (1999) Synergistic signaling in fetal brain by STAT3-Smad1 complex bridged by p300. Science 284:479–482
Lyden D, Young AZ, Zagzag D, Yan W, Gerald W, O’Reilly R, Bader BL, Hynes RO, Zhuang Y, Manova K, Benezra R (1999) Id1 and Id3 are required for neurogenesis, angiogenesis and vascularization of tumour xenografts. Nature 401:670–677
Kondo T, Raff M (2000) The Id4 HLH protein and the timing of oligodendrocyte differentiation. EMBO J 19:1998–2007
Fukuda S, Kondo T, Takebayashi H, Taga T (2004) Negative regulatory effect of an oligodendrocytic bHLH factor OLIG2 on the astrocytic differentiation pathway. Cell Death Differ 11:196–202
Noble M, Murray K, Stroobant P, Waterfield MD, Riddle P (1988) Platelet-derived growth factor promotes division and motility and inhibits premature differentiation of the oligodendrocyte/type-2 astrocyte progenitor cell. Nature 333:560–562
Raff MC, Lillien LE, Richardson WD, Burne JF, Noble MD (1988) Platelet-derived growth factor from astrocytes drives the clock that times oligodendrocyte development in culture. Nature 333:562–565
Richardson WD, Pringle N, Mosley MJ, Westermark B, Dubois-Dalcq M (1988) A role for platelet-derived growth factor in normal gliogenesis in the central nervous system. Cell 53:309–319
Rodriguez-Pena A (1999) Oligodendrocyte development and thyroid hormone. J Neurobiol 40:497–512
Rowitch DH, S-Jacques B, Lee SM, Flax JD, Snyder EY, McMahon AP (1999) Sonic hedgehog regulates proliferation and inhibits differentiation of CNS precursor cells. J Neurosci 19:8954–8965
Mekki-Dauriac S, Agius E, Kan P, Cochard P (2002) Bone morphogenetic proteins negatively control oligodendrocyte precursor specification in the chick spinal cord. Development 129:5117–5130
Lai K, Kaspar BK, Gage FH, Schaffer DV (2003) Sonic hedgehog regulates adult neural progenitor proliferation in vitro and in vivo. Nat Neurosci 6:21–27
Shimizu T, Kagawa T, Wada T, Muroyama Y, Takada S, Ikenaka K (2005) Wnt signaling controls the timing of oligodendrocyte development in the spinal cord. Dev Biol 282:397–410
Krämer-Albers EM, White R (2011) From axon-glial signaling to myelination: the integrating role of oligodendroglial Fyn kinase. Cell Mol Life Sci 68:2003–2012
Zhao S, Lin Y, Xu W, Jiang W, Zha Z, Wang P, Yu W, Li Z, Gong L, Peng Y, Ding J, Lei Q, Guan KL, Xiong Y (2009) Glioma-derived mutations in IDH1 dominantly inhibit IDH1 catalytic activity and induce HIF-1alpha. Science 324:261–265
Lu C, Ward PS, Kapoor GS, Rohle D, Turcan S, Abdel-Wahab O, Edwards CR, Khanin R, Figueroa ME, Melnick A, Wellen KE, O’Rourke DM, Berger SL, Chan TA, Levine RL, Mellinghoff IK, Thompson CB (2012) IDH mutation impairs histone demethylation and results in a block to cell differentiation. Nature 483:474–478
Wang G, Sai K, Gong F, Yang Q, Chen F, Lin J (2014) Mutation of isocitrate dehydrogenase 1 induces glioma cell proliferation via NK-κB activation in a hypoxia-inducible factor 1-α dependent manner. Mol Med Rep 9:1799–1805
Sasaki M, Knobbe CB, Itsumi M, Elia AJ, Harris IS, Chio II, Cairns RA, McCracken S, Wakeham A, Haight J, Ten AY, Snow B, Ueda T, Inoue S, Yamamoto K, Ko M, Rao A, Yen KE, Su SM, Mak TW (2012) D-2-hydroxyglutarate produced by mutant IDH1 perturbs collagen maturation and basement membrane function. Genes Dev 26:2038–2049
Bonner MY, Arbiser JL (2012) Targeting NADPH oxidases for the treatment of cancer and inflammation. Cell Mol Life Sci 69:2435–2442
Billon N, Terrinoni A, Jolicoeur C, McCarthy A, Richardson WD, Melino G, Raff M (2004) Roles for p53 and p73 during oligodendrocyte development. Development 131:1211–1220
Wolf RM, Wilkes JJ, Chao MV, Resh MD (2001) Tyrosine phosphorylation of p190 RhoGAP by Fyn regulates oligodendrocyte differentiation. J Neurobiol 49:62–78
Wolf RM, Draghi N, Liang X, Dai C, Uhrbom L, Eklöf C, Westermark B, Holland EC, Resh MD (2003) p190RhoGAP can act to inhibit PDGF-induced gliomas in mice: a putative tumor suppressor encoded on human chromosome 19q13.3. Genes Dev 17:476–487
Bartolomé RA, Wright N, Molina-Ortiz I, Sánchez-Luque FJ, Teixidó J (2008) Activated G(alpha)13 impairs cell invasiveness through p190RhoGAP-mediated inhibition of RhoA activity. Cancer Res 68:8221–8230
Bagchi A, Papazoglu C, Wu Y, Capurso D, Brodt M, Francis D, Bredel M, Vogel H, Mills AA (2007) CHD5 is a tumor suppressor at human 1p36. Cell 128:459–475
Chautard E, Ouédraogo ZG, Biau J, Verrelle P (2014) Role of Akt in human malignant glioma: from oncogenesis to tumor aggressiveness. J Neurooncol 117:205–215
Li X, Wu C, Chen N, Gu H, Yen A, Cao L, Wang E, Wang L (2016) PI3K/Akt/mTOR signaling pathway and targeted therapy for glioblastoma. Oncotarget 7:33440–33450
Kaneko S, Nakatani Y, Takezaki T, Hide T, Yamashita D, Ohtsu N, Ohnishi T, Terasaka S, Houkin K, Kondo T (2015) Ceacam1L Modulates STAT3 signaling to control the proliferation of glioblastoma-initiating cells. Cancer Res 75:4224–4234
Yamashita D, Kondo T, Ohue S, Takahashi H, Ishikawa M, Matoba R, Suehiro S, Kohno S, Harada H, Tanaka J, Ohnishi T (2015) miR340 suppresses the stem-like cell function of glioma-initiating cells by targeting tissue plasminogen activator. Cancer Res 75:1123–1133
Ohtsu N, Nakatani Y, Yamashita D, Ohue S, Ohnishi T, Kondo T (2016) Eva1 maintains the stem-like character of glioblastoma-initiating cells by activating the noncanonical NF-κB signaling pathway. Cancer Res 76:171–181
Kruiswijk F, Labuschagne CF, Vousden KH (2015) p53 in survival, death and metabolic health: a lifeguard with a licence to kill. Nat Rev Mol Cell Biol 16:393–405
Hollstein M, Hainaut P (2010) Massibely regulated genes: the example of TP53. J Pathol 220:164–173
Meek DW, Anderson CW (2009) Posttranslational modification of p53: cooperative integrators of function. Cold Spring Harb Perspect Biol 1:a000950
Salama R, Sadaie M, Hoare M, Narita M (2014) Cellular senescence and its effector programs. Genes Dev 28:99–114
Reed SM, Quelle DE (2014) p53 acetylation: regulation and consequences. Cancer 7:30–69
Tang J, Jiehui D, Cao H, Bai J, Zheng J (2015) p53-mediated autophagic regulation: A prospective strategy for cancer therapy. Cancer Lett 363:101–107
Nilsson MB, Zage PE, Zeng L, Xu L, Cascone T, Wu HK, Saigal B, Zweidler-McKay PA, Heymach JV (2010) Multiple receptor tyrosine kinases regulate HIF-1alpha and HIF-2alpha in normoxia and hypoxia in neuroblastoma: implications for antiangiogenic mechanisms of multikinase inhibitors. Oncogene 29:2938–2949
Acknowledgements
T.K. was supported, in part, by a research program of the Project for Development of Innovative Research on Cancer Therapeutics (P-DIRECT), Ministry of Education, Culture, Sports, Science and Technology of Japan.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kondo, T. Molecular mechanisms involved in gliomagenesis. Brain Tumor Pathol 34, 1–7 (2017). https://doi.org/10.1007/s10014-017-0278-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10014-017-0278-8