Abstract
Binding of histamine to the G-protein coupled histamine H1 receptor plays an important role in the context of allergic reactions; however, no crystal structure of the resulting complex is available yet. To deduce the histamine binding site, we performed unbiased molecular dynamics (MD) simulations on a microsecond time scale, which allowed to monitor one binding event, in which particularly the residues of the extracellular loop 2 were involved in the initial recognition process. The final histamine binding pose in the orthosteric pocket is characterized by interactions with Asp1073.32, Tyr1083.33, Thr1945.43, Asn1985.46, Trp4286.48, Tyr4316.51, Phe4326.52, and Phe4356.55, which is in agreement with existing mutational data. The conformational stability of the obtained complex structure was subsequently confirmed in 2 μs equilibrium MD simulations, and a metadynamics simulation proved that the detected binding site represents an energy minimum. A complementary investigation of a D107A mutant, which has experimentally been shown to abolish ligand binding, revealed that this exchange results in a significantly weaker interaction and enhanced ligand dynamics. This finding underlines the importance of the electrostatic interaction between the histamine ammonium group and the side chain of Asp1073.32 for histamine binding.
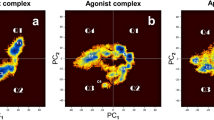
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Histamine is an endogenous tissue hormone and a key molecule for the regulation of arousal, inflammatory, and allergic reactions [1, 2]. Being extensively studied by medical research [1], it has been shown to stimulate lymphocyte activity, to facilitate the migration of immune cells, and to influence the behavior of granulocytes and mast cells [3, 4]. Histamine is known to be directly involved in the genesis of key symptoms observed in the context of allergic reactions such as an excessive production of mucus, sneezing, and pruritus [4]. The detection of histamine by a cell is mediated via four different G-protein coupled receptors (GPCRs): H1, H2, H3, and H4. All of them belong to the family of class-A GPCRs [5]. The histamine H1 receptor is expressed in many different cell types, e.g., immune cells, neurons, as well as the smooth muscle cells of respiratory or intestinal epithelium as well as vascular endothelial cells [2]. It has been described to play a major role in type I hypersensitivity reactions: histamine released from mast cells binds to the receptor and leads to its activation [6], which is followed by a signal transduction via a Gq protein [5]. The activated heterotrimeric G-protein hydrolyses GTP to GDP and dissociates into its α and a dimeric (β + γ) subunit. The α subunit binds to the enzyme phospholipase C (PLC), which is thereby triggered to catalyze the reaction of phosphoinositide-3,4-bisphosphate (PIP2) to diacyl glycerol (DAG) and inositol 1,4,5-trisphosphate (IP3). IP3 leads to a release of the second messenger Ca2+ from the endoplasmatic reticulum, whereas DAG activates the protein kinase C (PKC) [7]. Both mechanisms convey changes in cellular metabolism and behaviour, which occur in the context of allergic processes. Due to its particular role for hypersensitivity reactions, the histamine H1 receptor is a main target for anti-allergic drugs [8], which are used for the treatment of hay fever, conjunctivitis, allergic rhinitis, or anaphylactic shock [9]. The most relevant substances applied in therapy are inverse agonists of the H1 receptor such as loratadine, olopatadine, or cetirizine [8].
In 2011, Shimamura et al. published a crystal structure of the histamine H1 receptor bound to the small antagonist doxepin (PDB code 3RZE) [10]. However, there is to date no structure of a complex with the physiological ligand histamine available. The binding mode of histamine is so far more or less known due to a site-directed mutagenesis study by Ohta et al., which indicates that the residues Asp1073.32, Thr1945.43, and Asn1985.46 (superscript numbers indicate Ballesteros–Weinstein numbering [11] of the respective residues) might be involved in ligand binding [12]. Before the crystal structure of the H1 receptor became available, a first computational prediction for the histamine binding pocket was proposed based on a homology model [13]. Besides the three residues proposed by Ohta et al., this work suggested an involvement of Tyr1083.33, Ser1113.36, and Lys1915.40. After the crystal structure was published, Panula et al. performed a docking study and suggested interactions of histamine with Asp1073.32, Lys1915.40, Thr1945.43, Asn1985.46, Tyr4316.51, and Phe4356.55 [14]. Taken together, these findings suggest that histamine binds to the orthosteric pocket of the H1 receptor in a similar fashion like the antagonist doxepin.
To obtain further details of the histamine-H1 receptor interaction, we applied a set of different molecular dynamics simulations: For the initial deduction of the histamine binding site, we performed three unbiased molecular dynamics (MD) simulations starting from an unbound ligand, one of which succeeded in placing the ligand in the orthosteric pocket. The obtained binding pose was subsequently validated by 2 μs equilibrium MD simulations and by a thermodynamic analysis based on metadynamics simulations. These simulations demonstrate the conformational stability of the detected binding mode, which also represents a minimum in the energy landscape. Moreover, we conducted analogous simulations of a D107A mutant, which reveal the role of a saltbridge between Asp1073.32 and the histamine amino group for a stable binding position. Taken together, the results from our simulations allowed to refine the histamine binding mode and to identify the key interacting residues of the H1 receptor. In the future, this information may be helpful for drug development or further functional studies of histamine receptors.
Methods
Simulations performed
To elucidate the binding site of histamine in the H1 receptor, three simulations of at least 1 μs were performed, in which a single histamine molecule was placed ≈ 10 Å above the GPCR. The binding mode was afterwards verified by two independent 2-μs MD simulations. In addition, a D107A mutant was generated from the same starting structure and also simulated in two 2-μs simulation runs. An overview of the systems investigated is given in Table 1.
Preparation of histamine for MD simulations
Coordinates for histamine were downloaded from the PubChem database [15]. For the correct physiological protonation state (pKa= 9.7 for amino group, pKa= 5.8 for imidazole ring [16]), a third hydrogen atom was added to the amino group using Avogadro 1.1 [17]. gaff [18] atom types were assigned with the AmberTool antechamber. Partial charges for histamine were derived from a RESP/ESP fit with the RESP/ESP Charge Derive Server [19] using Firefly 7.1 [20, 21] and the base set RESP-C2 (HF/6-31G*//HF/6-31G*). The resulting final prep file is provided as the supplementary information of this paper (Online Resource 5). With tleap from AmberTools 17 [22], Amber coordinate and topology files for histamine were generated and then converted to the respective Gromacs formats with amb2gmx.pl as described in [23].
Preparation of the H1 receptor starting structure
The structure of the histamine H1 receptor was taken from PDB entry 3RZE [10]. The ligand doxepin and the T4 lysozyme used for crystallization were removed. The resulting gap between Cys221 and Leu405 was closed by an eight-residue spacer (sequence GSGSGSGS) using ModLoop [24, 25]. Additionally, the unresolved residues between His167 and Arg175 were completed with the native sequence. Since there were some missing terminal residues, an N-terminal acetyl and a C-terminal N-methyl capping group were added to the structure using Sybyl7.3 [26] to avoid artificial charges at the termini. The setup of the structure for the subsequent MD simulations was performed with tleap. Missing hydrogen atoms were added, the disulfide bridges mentioned in the PDB file were created, and ff99SB [27] parameters were assigned to the protein. The structure was then temporarily solvated in a TIP3P [28] waterbox (capped octahedron, minimum distance of 8 Å from the solute to the borders) with Cl− counter ions for electrical neutralization. This was done to perform an initial energy minimization prior to membrane embedding in order to reduce steric tension in the starting structure. The energy minimization was conducted with sander from Amber17 and comprised 500 steps of steepest descent as well as 4500 steps of the conjugate gradient algorithm. Afterwards, water and ions were removed and tleap was run to generate new Amber coordinate and topology files for the minimized protein structure using again parameters from the ff99SB force field. As described for histamine, the resulting files were then converted into Gromacs file formats with amb2gmx.pl. Based on the entry 3RZE from the OPM database [29], the structure was then overlaid with a pre-equilibrated dioleoylphosphatidylcholine (DOPC) bilayer (gaff force field) [30] and solvated in SPC water [31]. This was done using an in-house Perl script, which minimized the sum of the squared z distances between the DOPC C1 atoms and the pseudoatom entries from the OPM entry by which the position of the extracellular and intracellular membrane layer are coded. Coordinates and topology of the solvated bilayer and the protein were then combined and the resulting structure was embedded into the membrane using the gmx membed [32] functionality of Gromacs 2016.5 [33]. An electrical neutralization of the system was performed by addition of Cl− ions with gmx genion.
Then, a three-step energy minimization of the system was conducted, which comprised three steps in total. First, only water molecules and ions were minimized whereas all other atoms were restrained by the use of harmonic potentials with a force constant of 1000 kJ⋅mol− 1 ⋅nm− 2 in x, y, and z direction. In the second step, the DOPC membrane and most of the H1 receptor were minimized as well, only the Cα atoms were still restrained with the same force constant. Finally, the complete system including the Cα atoms was kept unrestrained. Every minimization phase was subdivided in a first part with the steepest descent algorithm and a second part with the conjugate gradient algorithm. The minimization was performed using gmx mdrun and terminated as soon as machine precision reached (≈ a few thousands of steps).
In order to equilibrate the GPCR in the membrane, a series of 300 consecutive MD simulations (see Section “Molecular dynamics simulations” for details) of 100 ps length was performed. After each simulation, water molecules that had diffused between receptor and membrane were deleted using a self-written Perl script. During these equilibration simulations, position restraints with a force constant of 1000 kJ⋅mol− 1 ⋅nm− 2 in x, y, and z direction were imposed on the protein backbone. After the 300 restrained simulations, an additional 2 μs free equilibration simulation of the system was performed to remove structural artifacts that might be due to the co-crystallization with the T4 lysozyme and the antagonist doxepin. For the histamine binding simulations, a single histamine molecule was added about 10 Å above the receptor on the extracellular side and an additional Cl− ion was added to ensure electrical neutrality.
For the simulation of the D107A mutant, the Asp1073.32 residue was exchanged for alanine with the Chimera swapaa command. Then, a new topology was created with tleap and converted into the Gromacs file format as described above.
Molecular dynamics simulations
All unbiased MD simulations were performed with Gromacs 2016.5 [33]. Periodic boundary conditions were applied in x, y, and z direction. The simulations were run at constant pressure and temperature (NpT ensemble) with surface-tension coupling. The reference surface tension was set to 1.1 nm⋅bar and the reference z pressure to 1 bar. The temperature was constantly held at 310 K with temperature coupling being achieved by a Berendsen thermostat [35] in three separate coupling groups for (i) solvent and ions, (ii) protein and ligand, and (iii) the DOPC membrane. A time step of 2 fs was used because bonds involving hydrogen atoms were constrained with the LINCS algorithm [36].
The analysis of the simulations was performed with cpptraj [37] from AmberTools 17. Contacts were defined between any atom pair (including hydrogen atoms) with a maximum distance of 5 Å, as described previously [38]. Plots were created with gnuplot [39], structure visualization was done with UCSF Chimera [34].
To assess the energy landscape of the obtained binding mode and to investigate the energetic effect of the D107A mutant, a well-tempered multiple walker metadynamics simulation was performed for both wild type and mutant according to a strategy established by Saleh et al. [40,41,42]. The metadynamics simulation was conducted using Gromacs 2016.3 [33] with the plumed 2.3.1 plugin [43]. As a collective variable (CV), we chose the z component of the distance between the conserved Trp4286.48 Cα atom at the bottom of the orthosteric binding pocket and the histamine amino nitrogen atom. Methodical details are given in the supplementary information to this paper (Online Resource 1). Briefly, we initially conducted two metadynamics runs (one run for the wild type and one for the D107A mutant), which started from the bound state of histamine, in order to obtain a rapid ligand unbinding. From these initial simulations, 32 starting conformations were selected that were equidistantly distributed between the bound and the totally unbound state of histamine. Subsequently, for both wild type and mutant, multiple walker metadynamics simulations with a simulation time of ≈ 1500 ns (32×48 ns) were conducted. The integration of the free energy landscape was performed with the plumed shell command.
Results and discussion
Determination of the histamine binding site
Molecular dynamics (MD) simulations, either in combination with enhanced sampling methods such as metadynamics or adaptive biasing force calculations or without external bias, have been successfully applied in the past in GPCR research to investigate ligand binding [41, 42, 44,45,46,47]. Therefore, we decided to use microsecond unbiased MD simulations for the identification of the histamine binding site in the present study. We performed a total of three simulations (1 μs each), in which histamine was placed above the H1 receptor. Since only one of these simulations led to successful binding, the other two runs were extended to 3 μs. However, no binding event could be observed in those simulations despite this prolongation. Therefore, we will only discuss the successful run in more detail. As a measurement for the progress of binding, we chose two different parameters: (i) the z component of the distance between the histamine amino nitrogen atom and the Trp4286.48 Cα atom (Fig. 1a), which is located directly below the binding pocket as described by Saleh et al. [40], and (ii) the number of contacts between histamine and the receptor (Fig. 1b). The binding process (movie in Online Resource 2) started after a simulation time of 600 ns and was complete at about 900 ns, taking 300 ns in total. As can be seen from Fig. 1b, it may be subdivided into three consecutive steps (visible as three plateaus). First, a small number of initial contacts (< 100) was established mainly with charged residues of the ECL2 (Glu177, Asp178, Lys179, Glu181) (Fig. 1c). Then, a rapid increase to an intermediate phase of 300–400 contacts occurred when the ligand moved to the vestibule of the binding pocket. During this second step of the binding process, the ligand maintained interactions with residues of the ECL2 (Asp178, Lys179, Thr182, Tyr185) that form a kind of lid over the pocket (Fig. 1c). In addition, it contacted residues (such as Leu1043.29, Lys1915.40, His4507.34, and Ile4547.38), which belong to the outer parts of the transmembrane helices 3, 5, and 7. At about 900 ns of simulation time, the ligand descended further into the deep orthosteric binding pocket and established stable interactions with Asp1073.32, Tyr1083.33, Thr1945.43, Asn1985.46, Trp4286.48, Tyr4316.51, Phe4326.52, and Phe4356.55 (Fig. 1c).
Histamine binding to the H1 receptor in an unbiased MD simulation. a Distance between the Trp4286.48 Cα atom at the bottom of the orthosteric binding pocket and the histamine amino nitrogen atom as a function of simulation time. b Number of contacts between histamine and the H1 receptor as a function of simulation time. c Structure of the upper part of the H1 receptor. The backbone is depicted in white, all residues which are contacted by histamine during the binding process are shown as sticks. The color code indicates at what point in time they interact with histamine: first magenta, then green, light blue, and violet in the end. For better orientation, the backbone of the ECL2 is highlighted in light green. Histamine is shown as black sticks in a position at the beginning of the binding process. Its further movement during the following binding process is shown as a black line (one point every 10 ns)
This final binding mode (Fig. 2a, PDB file in Online Resource 6) is especially characterized by a saltbridge between one of the Asp1073.32 Oδ atoms and the histamine ammonium group. Moreover, a hydrogen bond to Tyr4316.51 was observed. Besides electrostatic interactions with the different tyrosine residues, hydrophobic interactions of the two phenylalanines Phe4326.52 and Phe4356.55 with the carbon atoms of the imidazole ring of histamine play a role as well.
Binding mode of histamine to the H1 receptor from the last frame of the binding MD simulation. The H1 receptor is shown from the extracellular side. a Overview of main interacting residues. The backbone is colored in white and the main residues which interact with histamine are displayed as grey sticks. Histamine is visualized as black sticks. The saltbridge between Asp1073.32 and the histamine ammonium group as well as a hydrogen bond with Tyr4316.51 are illustrated as dashed red lines. b Overlay of the histamine binding mode and the doxepin-bound crystal structure of the histamine H1 receptor (PDB code 3RZE) [10]. The backbone of the crystal structure is colored in blue, doxepin (Z isomer from PDB 3RZE) and Asp1073.32 are displayed as violet sticks. The backbone of the histamine-bound structure is colored in green, histamine and Asp1073.32 are shown as orange sticks
When we analyzed the pathway from our initial binding simulation, we detected that especially the residues of the ECL2 formed contacts with histamine in the first phase of the binding process (Fig. 1c). This finding is in good agreement with the fact that this loop is already known to be particularly relevant for ligand recognition and ligand selectivity of class-A GPCRs [45, 48]. The entry of histamine happened through the crevice between the ECL2 and the transmembrane helices 5, 6, and 7 in a manner quite comparable to a binding pathway that was elucidated in an MD study for alprenolol at the β2 adrenergic receptor [45].
Having reached the vestibule of the binding pocket, histamine interactions with the residues of the ECL2 remained important for a certain period of time until the ligand moved further inward into the orthosteric binding site (Fig. 1c). One role of the ECL2 for ligand binding is thus to catch the free ligand by means of a variety of charged residues and then to guide the ligand to the binding site as described in previous publications [49]. Such a role of the ECL2 as an enclosure for bound ligands has been discussed before [50]. Thus, our study is in line with previous publications [45, 48, 50] demonstrating that the ECL2 can be of high functional relevance and that its role is not limited on simply connecting two transmembrane helices with each other.
In summary, the comparison with the experimental data suggests that the present binding MD simulation has sampled a plausible entrance pathway to the orthosteric binding pocket. However, we are aware that a comprehensive investigation of ligand binding pathways, which will allow to assess the detailed role of individual residues in the binding process, requires the observations of more successful binding events. For example, Dror et al. performed 82 simulations with 21 binding events to get a statistically sound data basis for such an analysis [45]. Thus, we consider our approach mainly as an alternative strategy to docking or enhanced sampling simulations to obtain a candidate binding mode for the ligand, which requires subsequent verification. We have performed this further verification by performing extended equilibrium MD simulations (Section “Stability of the histamine binding mode and role of D107 for the interaction”), by thermodynamic analyses (Section “Thermodynamics of the receptor–ligand interaction”), and by a comparison to previous experimental and computational data for the H1 receptor (Section “Conclusions”).
Comparison of the histamine binding mode with the doxepin-bound crystal structure
To date, the only available crystal structure for the H1 receptor is a complex with the antagonist doxepin [10]. To investigate if our obtained binding mode involves similar interacting residues as this crystal structure, we performed an overlay of both structures (Fig. 2b). The result shows that the overall ligand position within the binding pocket is the same for histamine and doxepin. The two ligands have a similar orientation with their amino/amine nitrogen atom directing towards Asp1073.32. To further characterize the similarities between the two binding modes, we compiled a table (Table 2), which lists all interacting amino acids for the two ligands.
Besides Asp1073.32, common interactions occurred with Tyr1083.33, Ser1113.36, Lys1915.40, Thr1945.43, Ala1955.44, Asn1985.46, Trp4286.48, Tyr4316.51, the two phenylalanines Phe4326.52 and Phe4356.55 as well as with Ile4547.38 and Tyr4587.42. Just one single residue, Ala1103.35, was found to interact exclusively with histamine and not with doxepin. However, there were a couple of residues that were contacted by doxepin, but not involved in the binding of histamine (e.g., Thr1123.37, Ile1153.40, Trp1584.57, Lys179, Phe1995.47, and Phe4246.44). This is most probably due to the fact that doxepin is larger compared to histamine, allowing it to reach residues that are located more distant from the center of the binding pocket. We also added information about mutagenesis studies with an effect on histamine or doxepin binding to Table 2 based on the information available from the GPCRdb [51]. These data will be discussed in more detail in Section “Conclusions”.
Stability of the histamine binding mode and role of D107 for the interaction
In order to test if the obtained binding mode remains stable and to investigate the relevance of the observed saltbridge (Fig. 2a), we performed subsequent 2 μs simulations of the wild-type receptor and a D107A mutant, in which the saltbridge is absent (two runs for each system). In case of the wild-type trajectories, we found that histamine remained constantly deep inside the binding pocket (movie in Online Resource 3, final structures from the two MD runs as PDB files in Online Resources 7, 8). This is also visible from the fact that the z distance between the histamine amino nitrogen atom and the Trp4286.48 Cα atom remained constant over the entire trajectory (Fig. 3a). The number of contacts changed only slightly over the simulation time apart from some fluctuations and a small increase, which indicates a further enhancement of binding (Fig. 3b). In contrast, the runs of the D107A mutant showed both a lot more flexibility in the z distance of histamine from the bottom of the binding pocket (Fig. 3a). In run2, there was already at the beginning of the simulation (from about 150 to 400 ns) a phase where histamine moved from the orthosteric binding pocket towards the vestibule, which is visible as a plateau at about 10–15 Å. The ligand returned then into its initial position. However, in this second run, the z distance increased again at about 1 μs of simulation time reaching the same plateau before a complete dissociation of histamine occurred after about 1600 ns (movie in Online Resource 4). This becomes also evident from the fact that the number of contacts between histamine and the receptor decreased to zero at this point of simulation time (Fig. 3b). Subsequently, the unbound ligand moved through the solvent very quickly, forming only transient contacts with the protein (predominantly with the ECL2) until a re-association took place about 100 ns later. In the simulated time range, histamine went only back into the vestibule and did not re-enter the deep orthosteric region of the binding pocket although the remaining simulation time was longer than the time span observed for this descent in the initial binding simulation with the wild-type receptor.
Stability of the detected histamine binding mode for the wild-type H1 receptor and the D107A mutant. a Distance between the Trp4286.48 Cα atom at the bottom of the orthosteric binding pocket and the histamine amino nitrogen atom as a function of simulation time. b Number of contacts between histamine and the H1 receptor as a function of simulation time
The different dynamics of histamine between the wild-type and the mutant H1 receptor can be visualized by an overlay of snapshots at different time points of the trajectories (Fig. 4). For the two wild-type runs (Fig. 4a, b), the general orientation of the ligand remained always the same with the amino group directing towards the side chain of Asp1073.32. A distance analysis revealed that the saltbridge between both charged groups persists over the majority (87 %) of the simulation time (Fig. 5a). The pronounced conformational stability becomes also apparent from a plot of the histamine root mean square deviation as a function of simulation time (Fig. 5b). In contrast, in case of the D107A mutant (Fig. 4c, d), histamine sometimes completely reversed its orientation, so that the amino group pointed towards the extracellular direction even if the ligand stayed within the orthosteric binding site (run1, Fig. 4c). In run2, there were because of the dissociation of histamine also snapshots in which the ligand was located in the vestibule or in the solvent outside the binding pocket.
Histamine flexibility within the binding pocket. Structure of the upper part of the H1 receptor with an overlay of the histamine position for every 200 ns of simulation time. The backbone of the receptor is colored in white, and the residues Asp1073.32/Ala107 are shown as yellow sticks. Histamine is shown as black sticks with nitrogen atoms colored in blue. The three protons of the NH3+ group are highlighted in violet to allow for an easier recognition of the orientation within the binding pocket. a Wild type, run1. b Wild type, run2. c D107A mutant, run1. d D107A mutant, run2
Dynamics of histamine in the wild-type equilibrium MD simulations over time. a Saltbridge between the Asp1073.32 Oδ atoms and the histamine amino nitrogen atom. To take into account the chemical equivalency of the two aspartate Oδ atoms, for every frame, the minimum distance out of the two possible atom combinations was calculated. b Root mean square deviation of the histamine heavy atoms. A translational and rotational fit was performed to the protein backbone for the analyzed frames to adjust the protein moiety
To investigate the differences in histamine binding between wild-type and D107A mutant receptor in more detail, we performed an analysis of the average number of contacts between histamine and all protein residues, as shown in Fig. 6. Their localization in the 3D structure of the receptor is visualized in Fig. 7 with a color code for the relevance of the individual residues. The two wild-type runs (Fig. 6a, b) are very similar to each other regarding the involved residues. Similar to the results from the end of the binding simulation (Fig. 2a), most contacts are observed with Tyr1083.33, Asp1073.32, Tyr4316.51, Trp4286.48 at the bottom of the binding pocket, and the two phenylalanines Phe4326.52 and Phe4356.55. The sum of all occurring interactions yields between 580 (run2) and 590 (run1) contacts per frame. For the D107A mutant runs (Fig. 6c, d), the main contacts are formed with the same residues. The alanine residue, by which Asp1073.32 is replaced, forms about 40 contacts with the ligand compared to about 70 that were detected for the wild type. In fact, the numbers of contacts per residue are in general slightly lower for the mutant runs leading to a sum of 490 contacts in run1 and only 440 contacts in run2 where the transient dissociation occurred. Moreover, it becomes apparent that histamine interacts in case of the mutant receptor with a larger variety of different residues due to its higher flexibility within the binding pocket. This includes some minor interactions with the residues Val1093.34, Phe184, and Phe1905.39 in run1. For run2, there are some contacts with side chains from the ECL2 (residues 177-185, depicted in Fig. 7b), which arise when histamine leaves the orthosteric binding pocket, e.g., during the dissociation from the receptor and the subsequent re-association to the vestibule.
Number of contacts between histamine and the H1 receptor, dissected into individual amino acid residues. Averages per frame ± standard deviations. The diagrams contain all side chains of residues where on average at least one contact was detected per frame in at least one of the performed simulations. a Wild type, run1. b Wild type, run2. c D107A mutant, run1. d D107A mutant, run2
H1 receptor residues contacted by histamine. Structure of the H1 receptor shown from the extracellular side. The backbone is colored in white. Histamine is shown as black sticks, all residues that interact with histamine are also depicted as sticks and colored according to the number of contacts they formed during the MD simulations: magenta> orange > yellow > green >cyan. For a better orientation, the backbone of the ECL2 is highlighted in light green. a Wild type, average over both runs. b D107A mutant, average over both runs
To verify if the contacts between histamine and the H1 receptor remained stable for the wild-type simulations, we plotted the contact numbers as a function of the simulation time (Fig. 8a, b). The graphs demonstrate that there is only very limited variability: Most interactions are constantly present from the beginning until the end of the trajectory. For the D107A mutant, the same kind of analysis (Fig. 8c, d) shows that the contacts are much less stable. There are more breaks indicating that the ligand is not as tightly bound as in the wild-type receptor. For run2, it can be seen in Fig. 8d that the contacts, which occur with the residues of the ECL2, were not exclusively formed during the dissociation and re-association processes after 1500 ns of simulation time. In fact, there was already at the beginning (about 150-400 ns) a phase where the ECL2 was frequently contacted because histamine moved upward into the direction of the vestibule (Fig. 3a). Then, the ligand re-entered the orthosteric binding site so that no contacts with the ECL2 were formed for about 1 μs until the second movement to the vestibule and the consecutive dissociation took place. Interestingly, the ECL2 was not only contacted directly before the dissociation but also in an earlier phase followed by a return of histamine to the orthosteric binding pocket (Figs. 8d and 4d). This observation suggests that the ECL2, which forms a lid over the binding pocket, hampers the dissociation process holding the ligand back inside. The ECL2 was also important for the following re-association, especially Asp178 and Lys179. Besides these residues, the re-association was characterized by several predominantly hydrophobic interactions with the outer parts of transmembrane helix 6 and 7 such as Phe4326.52, Phe4346.54, Phe4356.55, Ile4386.58 and His4507.34. Together, these results indicate that the histamine amino group is strongly bound to Asp1073.32 which also explains that the overall orientation of histamine in the binding pocket remains stable compared to the much higher dynamics in case of the D107A mutant (Fig. 4).
Thermodynamics of the receptor–ligand interaction
After the analysis of purely structural properties, we performed a complementing metadynamics simulation to investigate whether the binding mode from our unbiased MD study represents an energetic minimum. In order to explore how the energy landscape of the binding process is influenced by the D107A mutation, both the wild-type and the D107A mutant were studied. As a collective variable (CV), we chose in accordance with Saleh et al. [40] again the z component of the distance between the conserved Trp4286.48 Cα atom at the bottom of the orthosteric binding pocket and the histamine amino nitrogen atom. Using the multiple walker technique with 32 different starting conformations between the bound and the totally unbound state (for details see methods section and Online Resource 1), we reached a cumulative simulation time of approximately 1500 ns (48 ns per walker) per system (wild-type or D107A mutant).
As shown in Fig. 9, the free energy landscape of the wild type has a pronounced global minimum at a CV value between 0.4 and 0.5 nm. This result is in good agreement with the distance observed in our long unbiased MD simulations (Fig. 3a), which suggests that the obtained binding position actually represents an energetic minimum. The D107A mutant, in contrast, displays only a weak local minimum at this CV value. This effect can most probably be attributed to the loss of electrostatic interactions between histamine and the charged Asp1073.32 residue. The difference in the energy landscape of the D107A mutant also offers an energetic explanation for the higher dynamics of histamine within the binding pocket of the mutant GPCR and the observed dissociation in one of the two unbiased MD runs.
Conformational stability of the H1 receptor
Since the crystal structure we used for our MD study is stabilized by an antagonist and thus represents an inactive conformation, we also investigated the type of structural changes that occur upon histamine binding. In addition, we checked whether the D107A mutation has effects on the structure of the H1 receptor. Therefore, we performed an analysis of the backbone root mean square deviation (RMSD) for the initial binding simulations (Fig. 10a) and the 2 μs simulations with bound histamine (Fig. 10b). The RMSD was generally rather low with about 2-3 Å at the end of the simulations. The final RMSD values for the D107A mutant are only marginally increased compared to the wild type (Fig. 10b), suggesting that no large structural rearrangements are triggered by the exchange of Asp1073.32 for alanine. A structural overlay of frames at different time points of the trajectory (Fig. 11) shows that the general position of the transmembrane helices is rather stable for both wild type and mutant whereas the loops (especially the artificially constructed ICL 3) are more flexible.
In the context of agonist binding to an inactive GPCR, the question arises if there should be signs of receptor activation. Some activation mechanisms, which have been reported for class A GPCRs in the literature, include the disruption of the so-called ionic lock between helices 3 and 6, an outward rotation of transmembrane helix 6, as well as a downward movement of the Tyr7.53 side chain (belongs to the NPxxY motif) [59,60,61]. The histamine H1 receptor, however, lacks the ionic lock already in its inactive crystal structure [10], which makes it impossible to use it as an indicator for activation. From a structural overlay of the crystal structure and the two final structures of the wild-type MD runs with bound histamine (Fig. 12a), only a very slight outward rotation of helix 6 can be seen. However, we observed a downward movement of Tyr4687.53 in helix 7 (Fig. 12b), which might be a first sign of a beginning activation process. The fact that there is no stronger evidence for receptor activation in our MD study can likely be explained by the time scale of the simulations or the absence of a bound G-protein. Data from the literature suggest that GPCR activation may rather take place on a millisecond than on a microsecond time scale [62]. Moreover, a full activation is at least for some GPCRs thought to require the stabilizing effect of a bound G-protein [46], which was not present in our simulations.
Changes in the wild-type histamine H1 receptor structure upon histamine binding. Structural overlays of the doxepin-bound crystal structure (white) and the final structures of the 2 μs wild-type MD simulations with bound histamine (cyan and orange). a Structural overlay for the whole receptor from two different perspectives. b Structural overlay of helix H7 demonstrating the downward rotation of the Tyr4687.53 side chain
Conclusions
In this work, we used MD simulations to investigate histamine binding to the H1 receptor. The following section is intended to set our observations in context with results from previous research. A comprehensive compilation of all residues that have been reported in the GPCRdb [51] to influence histamine binding is provided in Table 2.
In agreement with site-directed mutagenesis studies performed by Ohta et al. [12], we saw that the residues Asp1073.32, Thr1945.43, and Asn1985.46 formed extensive contacts with histamine (Fig. 6). The saltbridge between Asp1073.32 and the histamine amino group was particularly relevant for a stable binding mode because simulations of the D107A mutant showed much more flexibility for histamine and even a complete dissociation in one of two runs (Fig. 3, 4, Online Resources 3, 4). This result fits to an experimental investigation of D107A, D107N, and D107E mutations leading all to a complete abolishment of histamine binding [12]. Further studies have shown that the well-conserved Asp3.32 residue in transmembrane helix 3 is of general importance for the binding of both agonists and antagonists to the whole group of biogenic amine receptors [53, 63, 64]. For the histamine H1 receptor, this particular residue is known to be involved not only in interactions with histamine but also in the binding of doxepin from the crystal structure by Shimamura et al. [10], and several other antagonists such as KW-4679, acrivastine, or cetirizine [53, 65].
Besides residues Asp1073.32, Thr1945.43, and Asn1985.46, a computational study, which was performed on a homology model before the H1 receptor crystal structure became available, suggested a role of Tyr1083.33, Ser1113.36, and Lys1915.40 for histamine binding [13]. In our simulations, we observed that especially the first two of these residues formed a high number of contacts with the ligand (Fig. 6). For Lys1915.40, there exist also data from experimental studies indicating that a mutation to alanine leads to a moderate loss in histamine binding affinity [52, 54]. This is in line with the observation that this residue also contributed to binding in our MD trajectories, albeit only to a minor extent. Further residues, which were contacted in our MDs and for which experimental evidence is available, are the two phenylalanines Phe4326.52 and Phe4356.55 [52, 58]. Especially Phe4326.52 seems to be of vital importance since a mutation of this residue was shown to abolish histamine binding [52]. Phe4356.55, and additionally Tyr4316.51, were also proposed as interacting residues in a later docking investigation by Panula et al. [14]. Our simulations revealed a high number of interactions with both of them.
Taken together, it can be concluded that the binding mode from our unbiased MD study is in rather good agreement with the results from previous investigations. We feel that the insights from our work may be useful for a better understanding of ligand binding at the H1 receptor. In the future, this knowledge could be valuable for drug design given that the H1 receptor plays an important role in the context of allergic reactions.
References
Akdis CA, Simons FER (2006) Histamine receptors are hot in immunopharmacology. Eur J Pharmacol 533(1–3):69–76
Hill SJ (1990) Distribution, properties, and functional characteristics of three classes of histamine receptor. Pharmacol Rev 42(1):45–83
Akdis CA, Jutel M, Akdis M (2008) Regulatory effects of histamine and histamine receptor expression in human allergic immune responses. In: Chemical immunology and allergy, KARGER, pp 67–82. https://doi.org/10.1159/000154858
Canonica GW, Blaiss M (2011) Antihistaminic, anti-inflammatory, and antiallergic properties of the nonsedating second-generation antihistamine desloratadine: a review of the evidence. World Allergy Organ J 4 (2):46–52. https://doi.org/10.1097/wox.0b013e3182093e19
Hill SJ, Ganellin CR, Timmerman H, Schwartz JC, Shankley NP, Young JM, Schunack W, Levi R, Haas HL (1997) International union of pharmacology. XIII. Classification of histamine receptors. Phamacol Rev 49(3):253–278
Simons FER (2004) Advances in H1-antihistamines. N Engl J Med 351 (21):2203–2217. https://doi.org/10.1056/nejmra033121
Berridge MJ (1993) Inositol trisphosphate and calcium signalling. Nature 361(6410):315–325. https://doi.org/10.1038/361315a0
Monczor F, Fernandez N (2016) Current knowledge and perspectives on histamine H1 and H2 receptor pharmacology: functional selectivity, receptor crosstalk, and repositioning of classic histaminergic ligands. Mol Pharmacol 90(5):640–648. https://doi.org/10.1124/mol.116.105981
Simons FER, Simons KJ (2011) Histamine and H1-antihistamines: celebrating a century of progress. J Allergy Clin Immunol 128(6):1139–1150, e4. https://doi.org/10.1016/j.jaci.2011.09.005
Shimamura T, Shiroishi M, Weyand S, Tsujimoto H, Winter G, Katritch V, Abagyan R, Cherezov V, Liu W, Han GW, Kobayashi T, Stevens RC, Iwata S (2011) Structure of the human histamine H1 receptor complex with doxepin. Nature 475(7354):65–70. https://doi.org/10.1038/nature10236
Ballesteros JA, Weinstein H (1995) Integrated methods for the construction of three-dimensional models and computational probing of structure-function relations in G protein-coupled receptors. In: Methods in neurosciences. Elsevier, pp 366–428. https://doi.org/10.1016/s1043-9471(05)80049-7
Ohta K, Hayashi H, Mizuguchi H, Kagamiyama H, Fujimoto K, Fukui H (1994) Site-directed mutagenesis of the histamine H1 receptor: roles of aspartic acid 107, asparagine 198 and threonine 194. Biochem Biophys Res Commun 203(2):1096–1101
Jongejan A, Bruysters M, Ballesteros JA, Haaksma E, Bakker RA, Pardo L, Leurs R (2005) Linking agonist binding to histamine H1 receptor activation. Nat Chem Biol 1:98–103. https://doi.org/10.1038/nchembio714
Panula P, Chazot PL, Cowart M, Gutzmer R, Leurs R, Liu WLS, Stark H, Thurmond RL, Haas HL (2015) International union of basic and clinical pharmacology. XCVIII. Histamine receptors. Pharmacol Rev 67(3):601–655. https://doi.org/10.1124/pr.114.010249
Kim S, Thiessen PA, Bolton EE, Chen J, Fu G, Gindulyte A, Han L, He J, He S, Shoemaker BA, Wang J, Yu B, Zhang J, Bryant SH (2015) PubChem substance and compound databases. Nucleic Acids Res 44(D1):D1202–D1213. https://doi.org/10.1093/nar/gkv951
Paiva TB, Tominaga M, Paiva ACM (1970) Ionization of histamine, N-acetylhistamine, and their iodinated derivatives. J Med Chem 13(4):689–692. https://doi.org/10.1021/jm00298a025
Hanwell MD, Curtis DE, Lonie DC, Vandermeersch T, Zurek E, Hutchison GR (2012) Avogadro: an advanced semantic chemical editor, visualization, and analysis platform. J Cheminformatics 4(1):17. https://doi.org/10.1186/1758-2946-4-17
Wang J, Wolf RM, Caldwell JW, Kollman PA, Case DA (2004) Development and testing of a general amber force field. J Comput Chem 25(9):1157–1174. https://doi.org/10.1002/jcc.20035
Vanquelef E, Simon S, Marquant G, Garcia E, Klimerak G, Delepine JC, Cieplak P, Dupradeau FY (2011) R.E.D. server: a web service for deriving RESP and ESP charges and building force field libraries for new molecules and molecular fragments. Nucleic Acids Res 39(suppl):W511–W517. https://doi.org/10.1093/nar/gkr288
Granovsky AA (2010) Firefly version 7.1. http://classic.chem.msu.su/gran/firefly/index.html
Schmidt MW, Baldridge KK, Boatz JA, Elbert ST, Gordon MS, Jensen JH, Koseki S, Matsunaga N, Nguyen KA, Su S, Windus TL, Dupuis M, Montgomery JA (1993) General atomic and molecular electronic structure system. J Comput Chem 14(11):1347–1363. https://doi.org/10.1002/jcc.540141112
Case DA, Cerutti D, Cheatham IIIT, Darden T, Duke R, Giese T, Gohlke H, Goetz A, Greene D, Homeyer N, Izadi S, Kovalenko A, Lee T, LeGrand S, Li P, Lin C, Liu J, Luchko T, Luo R, Mermelstein D, Merz K, Monard G, Nguyen H, Omelyan I, Onufriev A, Pan F, Qi R, Roe DA, Roitberg A, Sagui C, Simmerling C, Botello-Smith W, Swails J, Walker R, Wang J, Wolf R, Wu X, Xiao L, York D, Kollman P (2017) AMBER 2017. University of California, San Francisco
Mobley DL, Chodera JD, Dill KA (2006) On the use of orientational restraints and symmetry corrections in alchemical free energy calculations. J Chem Phys 125(8):084902. https://doi.org/10.1063/1.2221683
Fiser A, Do RKG, Sali A (2000) Modeling of loops in protein structures. Protein Sci 9(9):1753–1773. https://doi.org/10.1110/ps.9.9.1753
Fiser A, Sali A (2003) ModLoop: automated modeling of loops in protein structures. Bioinformatics 19 (18):2500–2501. https://doi.org/10.1093/bioinformatics/btg362
Tripos International, St Louis, MO, USA (2006) Sybyl 7.3
Hornak V, Abel R, Okur A, Strockbine B, Roitberg A, Simmerling C (2006) Comparison of multiple Amber force fields and development of improved protein backbone parameters. Proteins 65(3):712–725. https://doi.org/10.1002/prot.21123
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79(2):926–935. https://doi.org/10.1063/1.445869
Lomize MA, Pogozheva ID, Joo H, Mosberg HI, Lomize AL (2011) OPM database and PPM web server: resources for positioning of proteins in membranes. Nucleic Acids Res 40(D1):D370–D376. https://doi.org/10.1093/nar/gkr703
Siu SWI, Vȧcha R, Jungwirth P, Böckmann RA (2008) Biomolecular simulations of membranes: physical properties from different force fields. J Chem Phys 128(12):125103. https://doi.org/10.1063/1.2897760
Toukan K, Rahman A (1985) Molecular-dynamics study of atomic motions in water. Phys Rev B 31 (5):2643–2648. https://doi.org/10.1103/physrevb.31.2643
Wolf MG, Hoefling M, Aponte-Santamaría C, Grubmüller H, Groenhof G (2010) g_membed: efficient insertion of a membrane protein into an equilibrated lipid bilayer with minimal perturbation. J Comput Chem 31 (11):2169–2174. https://doi.org/10.1002/jcc.21507
Berendsen H, van der Spoel D, van Drunen R (1995) GROMACS: a message-passing parallel molecular dynamics implementation. Comput Phys Commun 91(1-3):43–56. https://doi.org/10.1016/0010-4655(95)00042-e
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera–a visualization system for exploratory research and analysis. J Comput Chem 25(13):1605–1612. https://doi.org/10.1002/jcc.20084
Berendsen HJC, Postma JPM, van Gunsteren WF, DiNola A, Haak JR (1984) Molecular dynamics with coupling to an external bath. J Chem Phys 81(8):3684–3690. https://doi.org/10.1063/1.448118
Hess B, Bekker H, Berendsen HJC, Fraaije JGEM (1997) LINCS: a linear constraint solver for molecular simulations. J Comput Chem 18(12):1463–1472
Roe DR, Cheatham TE (2013) PTRAJ and CPPTRAJ: software for processing and analysis of molecular dynamics trajectory data. J Chem Theory Comput 9:3084–3095. https://doi.org/10.1021/ct400341p
Söldner C A, Horn AHC, Sticht H (2018) Interaction of glycolipids with the macrophage surface receptor Mincle – a systematic molecular dynamics study. Sci Rep 8:1. https://doi.org/10.1038/s41598-018-23624-8
Williams T, Kelley C, et al. (2018) Gnuplot 4.6: an interactive plotting program, 2012. http://www.gnuplot.info
Saleh N, Ibrahim P, Saladino G, Gervasio FL, Clark T (2017) An efficient metadynamics-based protocol to model the binding affinity and the transition state ensemble of G-protein-coupled receptor ligands. J Chem Inf Model 57(5):1210–1217. https://doi.org/10.1021/acs.jcim.6b00772
Saleh N, Kleinau G, Heyder N, Clark T, Hildebrand PW, Scheerer P (2018a) Binding, thermodynamics, and selectivity of a non-peptide antagonist to the Melanocortin-4 receptor. Front Pharmacol, 9. https://doi.org/10.3389/fphar.2018.00560
Saleh N, Hucke O, Kramer G, Schmidt E, Montel F, Lipinski R, Ferger B, Clark T, Hildebrand PW, Tautermann CS (2018b) Multiple binding sites contribute to the mechanism of mixed agonistic and positive allosteric modulators of the cannabinoid CB1 receptor. Angewandte Chemie Int Edn 57(10):2580–2585. https://doi.org/10.1002/anie.201708764
Bonomi M, Branduardi D, Bussi G, Camilloni C, Provasi D, Raiteri P, Donadio D, Marinelli F, Pietrucci F, Broglia RA, Parrinello M (2009) PLUMED: a portable plugin for free-energy calculations with molecular dynamics. Comput Phys Commun 180(10):1961–1972. https://doi.org/10.1016/j.cpc.2009.05.011
Saleh N, Saladino G, Gervasio FL, Haensele E, Banting L, Whitley DC, de Oliveira-Santos JS, Bureau R, Clark T (2016) A three-site mechanism for agonist/antagonist selective binding to vasopressin receptors. Angewandte Chemie Int Edn 55(28):8008–8012. https://doi.org/10.1002/anie.201602729
Dror RO, Pan AC, Arlow DH, Borhani DW, Maragakis P, Shan Y, Xu H, Shaw DE (2011) Pathway and mechanism of drug binding to G-protein-coupled receptors. Proc Natl Acad Sci 108(32):13118–13123. https://doi.org/10.1073/pnas.1104614108
Clark T (2017) G-protein coupled receptors: answers from simulations. Beilstein J Org Chem 13:1071–1078. https://doi.org/10.3762/bjoc.13.106
Marino KA, Filizola M (2017) Investigating small-molecule ligand binding to G protein-coupled receptors with biased or unbiased molecular dynamics simulations. In: Methods in molecular biology. Springer, New York, pp 351–364 https://doi.org/10.1007/978-1-4939-7465-8_17
Peeters M, van Westen G, Li Q, IJzerman A (2011) Importance of the extracellular loops in G protein-coupled receptors for ligand recognition and receptor activation. Trends Pharmacol Sci 32(1):35–42. https://doi.org/10.1016/j.tips.2010.10.001
Strasser A, Wittmann HJ, Seifert R (2017) Binding kinetics and pathways of ligands to GPCRs. Trends Pharmacol Sci 38(8):717–732. https://doi.org/10.1016/j.tips.2017.05.005
Wheatley M, Wootten D, Conner M, Simms J, Kendrick R, Logan R, Poyner D, Barwell J (2012) Lifting the lid on GPCRs: the role of extracellular loops. Br J Pharmacol 165(6):1688–1703. https://doi.org/10.1111/j.1476-5381.2011.01629.x
Pȧndy-Szekeres G, Munk C, Tsonkov TM, Mordalski S, Harpsøe K, Hauser AS, Bojarski AJ, Gloriam DE (2017) GPCRdb in 2018: adding GPCR structure models and ligands. Nucleic Acids Res 46 (D1):D440–D446. https://doi.org/10.1093/nar/gkx1109
Bruysters M, Pertz HH, Teunissen A, Bakker RA, Gillard M, Chatelain P, Schunack W, Timmerman H, Leurs R (2004) Mutational analysis of the histamine H1-receptor binding pocket of histaprodifens. Eur J Pharmacol 487(1–3):55–63. https://doi.org/10.1016/j.ejphar.2004.01.028
Nonaka H, Otaki S, Ohshima E, Kono M, Kase H, Ohta K, Fukui H, Ichimura M (1998) Unique binding pocket for KW-4679 in the histamine H1 receptor. Eur J Pharmacol 345(1):111–117. https://doi.org/10.1016/s0014-2999(97)01620-8
Gillard M (2002) Binding characteristics of cetirizine and levocetirizine to human H1 histamine receptors: Contribution of Lys191 and Thr194. Mol Pharmacol 61(2):391–399. https://doi.org/10.1124/mol.61.2.391
Bakker RA (2004) 8R-lisuride is a potent stereospecific histamine H1-receptor partial agonist. Molec Pharmacol 65(3):538–549. https://doi.org/10.1124/mol.65.3.538
Leurs R, Smit M, Tensen C, Terlaak A, Timmerman H (1994) Site-directed mutagenesis of the histamine H1-receptor reveals a selective interaction of asparagine207 with subclasses of H1-receptor agonists. Biochem Biophys Res Commun 201(1):295–301. https://doi.org/10.1006/bbrc.1994.1701
Moguilevsky N, Varsalona F, Guillaume JP, Noyer M, Gillard M, Daliers J, Hėnichart J P, Bollen A (1995) Pharmacological and functional characterisation of the wild—type and site—directed mutants of the human H1 histamine receptor stably expressed in CHO cells. J Recept Signal Transd 15(1-4):91–102. https://doi.org/10.3109/10799899509045210
Cordova-Sintjago TC, Fang L, Bruysters M, Leurs R, Booth RG (2012) Molecular determinants of ligand binding at the human histamine H1 receptor: site-directed mutagenesis results analyzed with ligand docking and molecular dynamics studies at H1 homology and crystal structure models. J Chem Pharm Res 4(6):2937–2951
Kobilka BK (2007) G protein coupled receptor structure and activation. Biochimica et Biophysica Acta (BBA) - Biomembranes 1768(4):794–807. https://doi.org/10.1016/j.bbamem.2006.10.021
Kimata N, Reeves PJ, Smith SO (2015) Uncovering the triggers for GPCR activation using solid-state NMR spectroscopy. J Magn Reson 253:111–118. https://doi.org/10.1016/j.jmr.2014.12.014
Venkatakrishnan AJ, Deupi X, Lebon G, Heydenreich FM, Flock T, Miljus T, Balaji S, Bouvier M, Veprintsev DB, Tate CG, Schertler GFX, Babu MM (2016) Diverse activation pathways in class a GPCRs converge near the G-protein-coupling region. Nature 536(7617):484–487. https://doi.org/10.1038/nature19107
Vilardaga JP, Bünemann M, Krasel C, Castro M, Lohse MJ (2003) Measurement of the millisecond activation switch of G protein-coupled receptors in living cells. Nat Biotechnol 21(7):807–812. https://doi.org/10.1038/nbt838
Schwartz TW, Rosenkilde MM (1996) Is there a lock for all agonist keys in 7TM receptors? Trends Pharmacol Sci 17:6. https://doi.org/10.1016/0165-6147(96)10017-1
Strader CD, Fong TM, Graziano MP, Tota MR (1995) The family of G-protein coupled receptors. FASEB J 9(9):745– 754
Wieland K, Laak AMT, Smit MJ, Kühne R, Timmerman H, Leurs R (1999) Mutational analysis of the antagonist-binding site of the histamine H1 receptor. J Biol Chem 274(42):29994–30000. https://doi.org/10.1074/jbc.274.42.29994
Acknowledgements
The authors gratefully acknowledge the computer resources and support provided by the Erlangen Regional Computing Center (RRZE) and the Leibniz Rechenzentrum, Munich. C.A.S. would like to thank Jonas Kaindl from the Computer-Chemie-Centrum (CCC) of the FAU Erlangen-Nürnberg for fruitful discussions and valuable advice. This paper is dedicated to Prof. Tim Clark, an eminent computational chemist, on the occasion of his 70th birthday.
Funding
The study was funded by Deutsche Forschungsgemeinschaft (DFG) in the graduate school Graduiertenkolleg GRK1910. In addition, the work was supported by a grant of computer time on SuperMUC at the Leibniz Rechenzentrum, Munich (project pr74su).
Author information
Authors and Affiliations
Contributions
H.S. and C.A.S. conceived the study. A.H.C.H. and C.A.S. carried out the parametrization of histamine. C.A.S. performed the simulations and subsequent analyses. All authors interpreted the results and contributed to the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
This paper belongs to the Topical Collection Tim Clark 70th Birthday Festschrift
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Söldner, C.A., Horn, A.H.C. & Sticht, H. Binding of histamine to the H1 receptor—a molecular dynamics study. J Mol Model 24, 346 (2018). https://doi.org/10.1007/s00894-018-3873-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-018-3873-7