Abstract
Oral squamous cell carcinoma (OSCC) is the most common malignant tumor in the head and neck region. The aim of this study was to identify the key molecules and to elucidate the molecular mechanisms of OSCC carcinogenesis through a microarray analysis of RNA extracted from normal epithelium, dysplasia, and squamous cell carcinoma components. Out of molecules that showed changes in gene expression in the microarray analysis, we focused on Sulfite oxidase (SUOX), which correlated significantly with carcinogenic process and exhibited a stepwise decrease in expression. The expression of SUOX was evaluated in detail at the protein level using samples from 58 patients with cancer of the tongue, and correlating clinicopathological factors were also comprehensively examined. SUOX expression declined significantly from normal epithelium to dysplasia to squamous cell carcinoma components in line with carcinogenic process. With regard to squamous cell carcinoma, SUOX expression was significantly lower when T classification was high. Our findings indicated that SUOX is negatively associated with the progression and proliferation of tongue cancer, and suggest that SUOX may be a key molecule in tongue tumors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oral squamous cell carcinoma (OSCC) is the most common malignant tumor in the head and neck region [1], and represents over 90% of oral cancers [2]. More than half of all oral cancers occur in the tongue, floor of the mouth, and gingiva, with areas of mechanical stimulation being the most frequently affected [3,4,5]. OSCCs have the same genetic profile beyond the sites including the tongue, floor of the mouth, and gingiva. Globally, the 5-year survival rate for oral cancer is 55–60%, while tumor diameter, presence or absence of lymph node metastasis, and presence or absence of distant metastasis are regarded to be the most important prognostic factors [6]. Additionally, perineural invasion [7, 8], vascular invasion [8], and bone infiltration [9] are also cited as prognostic factors and it is important that they each be evaluated histopathologically. On the other hand, as regards recurrence, oral epithelial dysplasia (OED) may progress to OSCC, and OED located in the proximity of OSCC is reportedly associated with an increased risk of local recurrence and progression to OSCC [10, 11]. The 5-year survival in such cases has been reported to be as low as 30% [12]. Since OED is closely involved in the development of OSCC, it is important to examine the mechanism of OED occurrence using a molecular pathological approach, and to identify the key molecules involved in carcinogenesis.
Approximately 12% of all OED progress to OSCC [13]. Tumor protein p53 (TP53) [14, 15], cyclin-dependent kinase inhibitor 2A (CDKN2A) [15], phosphatase and tensin homolog deleted from chromosome 10 (PTEN) [15], Harvey rat sarcoma (HRAS) [15], phosphatidylinositol-4, and 5-bisphosphate 3-kinase catalytic subunit alpha (PIK3CA) [15] are all reported to be important genes linked to oncogenesis potentially; however, cases not dependent on these known factors are also known to exist, so it is important to search for other factors.
In this study, we extracted RNA from normal epithelium, dysplasia, and squamous cell carcinoma components and performed a microarray analysis to identify the key molecules involved in OSCC carcinogenesis and elucidate the molecular mechanism of OSCC pathogenesis. Of the potential key molecules identified in the assay, we focused on SUOX. Until now, there have been no reports on the role that SUOX plays in OSCC. In this study, SUOX expression was further evaluated in detail at the protein level, and its relationship with other clinicopathological features was examined.
Materials and methods
Microarray analysis
For the extraction of RNA, formalin-fixed paraffin-embedded (FFPE) blocks from 3 OSCC cases were used. The FFPE blocks were sectioned to a thickness of 7 μm using a Leica RM2245 microtome (Leica Microsystems K.K., Tokyo, Japan), with an RNase-free water-treated blade. For each case, one section was stained with hematoxylin eosin stain (H.E.) while three sections were used for extracting RNA. The normal epithelium, dysplasia, and squamous cell carcinoma components were marked using the previously mentioned H.E-stained specimen, superimposed on the unstained specimen, and each component was collected by scraping with a scalpel treated with RNase-free water. RNA from each component was isolated, linearly amplified, hybridized to the Affymetrix GeneChip Human X3P Array (Affymetrix, Santa Clara, CA, USA) and labeled in accordance with the manufacturer’s instructions for the Arcturus Paradise PLUS Reagent System (Life Technologies, Grand Island, NY, USA) and GeneChip 3′ IVT Express Reagent kit (Affimetrix). The extracted RNA was measured using Nano drop ® ND 1000 (Thermo Fisher Scientific, Waltham, MA, USA). Affymetrix array CEL files were processed by the RNA algorithm [16] to obtain probe set-level gene expression data, using the Expression Console software (Affymetrix).
Patients
We selected patients with primary tongue cancer who had not undergone preoperative treatments such as chemotherapy and/or radiotherapy at Kurume University Hospital between 2010 and 2015. Fifty-eight patients filled this condition. Of these 58 patients, 30 were male and 28 were female. The mean age was 63.4 ± 15.3 (range 28–90). The mean tumor size was 18.3 ± 9.32 mm (range 2–45 mm). All cases accompanied dysplastic components adjacent to carcinoma components. Twenty-six patients were classified T1, 31 were T2 and 1 was T3. Regarding histological grading, 47 were well-differentiated, 10 were moderately-differentiated, and 1 was poorly-differentiated. Thirteen cases had lymphatic vessel invasion, while four cases had vascular invasion (Table 1).
The excised tissues were fixed using 10% buffered formalin, sectioned at 4-µm-thickness, followed by HE staining. Histopathological evaluations were performed by three pathologists (K.N., J.A. and H.Y.). Pathological diagnosis was performed according to the WHO classification of Head and Neck Tumors 4th Edition [6].
This study was approved by the ethics committee of Kurume University (#330).
Immunohistochemistry
We performed immunohistochemistry (IHC) on paraffin-embedded sections using the SUOX antibody (ab88346, dilution 1: 300, Abcam plc., Cambridge, UK). IHC was performed using the Ventana Benchmark (Ventana, Tucson, AZ). All IHC analyses were evaluated by two experienced observers who were unaware of the patients’ clinical conditions. We considered only nuclear expression of SUOX as positive. We used the Allred score system [17] in the staining evaluation to calculate a total score (TS) from a population score (PS) and an intensity score (IS). Allred score is usually used in breast carcinoma, and is in various carcinomas including OSCC [18, 19]. This system is easy to learn and highly reproducible [20]. Briefly, a PS was assigned representing the estimated proportion of positive staining cells (0 = none; 1 = < 1/100; 2 = 1/100 to < 1/10; 3 = 1/10 to < 1/3; 4 = 1/3 to < 2/3; 5 = > 2/3). Average estimated intensity of staining in positive cells was assigned an IS (0 = none; 1 = weak; 2 = intermediate; 3 = strong). PS and IS were added to obtain a total score that ranged from 0 to 8. We compared the expression of SUOX in each component and examined the correlation between the SUOX staining intensity of the carcinoma component and clinicopathological factors, such as tumor size, differentiation (well / moderately to poor), T classification, lymphatic vessel invasion, and vascular invasion.
Statistical analysis
The inter-rater reliability coefficient for TS was calculated for two doctors (K.N and M.N). The inter-rater reliability coefficient was κ = 0.8 (excellent). Additionally, the scores for (K.N) were statistically analyzed and the results were compared with those for the other pathologist (M.N). It was confirmed that there was no significant difference between the resulting scores. Differences in expression intensity for normal epithelium vs dysplasia components as well as for dysplasia vs squamous cell carcinoma components were evaluated by T-test, with p < 0.05 considered significant. Correlation between TS and clinicopathological features (tumor size: mm, T classification, histological grade, lymphatic vessel invasion, vascular invasion) in the carcinoma component was evaluated using a bivariate logistic regression model (p < 0.05). The statistical software used was JMP® Pro 13 (SAS Institute Inc., Cary, NC, USA).
Results
Microarray analysis
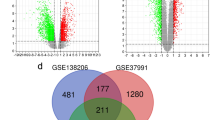
Table 2 shows the clinicopathological features and RNA yield for each component of 3 cases. The expression of 61,298 molecules was confirmed by microarray analysis, and a heat map was prepared (Fig. 1). Among these molecules, we focused on 22 that showed significant differences between normal epithelium vs dysplasia component and between dysplasia component vs squamous cell carcinoma component. Carcinogenesis was associated with decreased expression in four molecules and increased expression in 18 molecules (Table 3). Of the 22 molecules, SUOX showed the most significant differences in expression between each group. Exact p-values of SUOX for normal epithelium vs dysplasia component, normal epithelium vs carcinoma and dysplasia component and carcinoma were p = 0.023, 0.018 and 0.014, respectively. Moreover SUOX expression decreased gradually in the order of normal epithelium to dysplasia component to squamous cell carcinoma component (Table 3). These changes were the highest in SUOX and statistically significant. Therefore, we selected SUOX for further examination.
Immunohistochemical analysis
Immunohistochemistry was performed using samples from the aforementioned 58 tongue cancer patients. In normal sections, SUOX was mainly expressed in the nuclei of intermediate and basal layer squamous cells, but there was little expression on the surface layer.
On the other hand, in the dysplastic epithelia, expression was not readily seen in the basal layer, but was mainly observed in cells from the intermediate to surface layer, while in the carcinoma component nucleic expression was almost completely absent (Fig. 2a). Further, the intensity of SUOX expression by Allred score was normal epithelium = 7.2 ± 0.107 (range 6–8), dysplasia = 5.27 ± 0.153 (range 3–7), squamous cell carcinoma = 3.17 ± 0.274 (range 0–6), and there was a significant difference (p < 0.01) between each component (Fig. 2b).
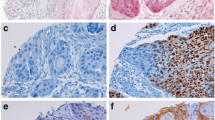
A Immunohistochemical findings of SUOX in normal epithelium, dysplasia and carcinoma components. Expression of SUOX was observed in the nuclei from the basal layer to the intermediate layer of the normal squamous epithelium (a, d). In the dysplasia component, expression was observed mainly in the nucleus of atypical cells from the intermediate layer to the surface layer, but was not readily observed in nuclei of the basal layer (b, e). Conversely, almost no expression was observed in the carcinoma component (c, f). (Hematoxylin eosin a–c ×200, SUOX d–f ×200) Scale bar 50 μm. b Comparison of total score (TS) of SUOX expression among normal epithelium (N), dysplasia (D) and squamous cell carcinoma (Ca) components. TS of SUOX of each component was significantly and gradually decreased as carcinogenesis progressed (*p < 0.01). TS of N, D and Ca was 7.29 ± 0.107, 5.27 ± 0.153 and 3.17 ± 0.274, respectively
As for carcinoma, nucleic expression was scatteringly observed in T1 squamous carcinoma but rarely in ≥ T2 squamous carcinoma (Fig. 3a). TS in the carcinoma component was 3.84 ± 0.39 in T1, and 2.62 ± 0.35 ≥ in ≥ T2 squamous carcinoma, and this decrease in SUOX expression from T1 to ≥ T2 was significant (p = 0.025) (Fig. 3b).
A Immunohistochemical findings of SUOX in T1 and ≥ T2 oral squamous cell carcinoma. Tumor cells with nuclear SUOX expression are scatteringly observed in a case of T1 squamous cell carcinoma (a, c), while a case of ≥ T2 squamous cell carcinoma shows almost no SUOX expression (b, d). (Hematoxylin eosin: a, b ×200, SUOX: c, d ×200) Scale bar indicates 50 μm. b Comparison of total score (TS) of SUOX expression between T1 and ≥ T2 oral squamous cell carcinoma cases. TS average was 3.84 ± 0.39 in T1 squamous cell carcinoma, and 2.62 ± 0.35 in ≥ T2 squamous cell carcinoma. This difference in SUOX expression by T classification was significant (p = 0.025)
In addition, when clinicopathological features and TS were examined in the carcinoma component, a significant negative correlation was found between SUOX expression and tumor diameter. However, TS in carcinoma component showed no significant correlation with any other factors, including degree of differentiation or vascular invasion.
Discussion
Many studies have examined the relationship between various genes and carcinogenesis in OSCC. However, most of these reports have focused either on expression occurring exclusively in the carcinoma [21,22,23,24,25] or on comparisons between non-cancerous and cancerous components [26,27,28]; few have investigated for factors that showed significant gradual differences among normal, dysplastic, and carcinoma components [29]. For this reason, we extracted and comprehensively examined RNA from normal, dysplastic, and carcinoma components using microarray analysis. Based on our results, we isolated SUOX, a factor that exhibited a significant and gradual decrease in expression from normal epithelium to dysplasia component to squamous cell carcinoma component, and evaluated its clinicopathological significance.
SUOX is a metallo-enzyme present in the mitochondria of all eukaryotes that utilizes, as coenzymes, molybdenum and heme, [30]. Via cytochrome c, it transfers electrons, produced through the oxidation of sulfurous acid into sulfuric acid, to the electron transport for use in ATP synthesis through oxidative phosphorylation [31,32,33]. This is the final step in the metabolism of sulfur-containing compounds, and sulfite is excreted [34]. In general, SUOX deficiency is the most common disease associated with SUOX [30, 35, 36] and there are few studies of SUOX in malignant tumors. In particular, though there have been no reports examining the expression of SUOX in OSCC, it has been reported in recent years that carcinogenesis of liver tumors is accompanied by diminished SUOX expression [37]. Jin GZ et al. demonstrated that the expression of SUOX decreased in a stepwise fashion along with the carcinogenic process and, furthermore, the expression of SUOX decreased along as tumor size increased [37]. Although SUOX in OSCC shows similar kinetics in liver tumor, it is difficult to compare the role of SUOX between OSCC and liver tumor as the mechanisms of carcinogenesis and the processes of tumor progression are different. Out of 22 genes, 17 were reported as cancer-related genes [19, 37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53]. However, the number of reports on each gene is limited. Extensive investigation on these genes has not been conducted so far.
Many immunohistochemical studies on OSCC have been conducted and many biomarkers have been reported to be associated with various clinicopathological factors, including prognosis. Most of the biomarkers demonstrated that higher expressions were associated with aggressive clinicopathological factors. These types of biomarkers have been listed as follows: epithelial cell adhesion molecule, CD44s, cyclooxygenase-2, autophagy-related 16-like 1, glucose-regulated protein 78, cysteine-rich 61, Aurora B, urokinase-type plasminogen activator receptor [54,55,56,57,58,59,60,61]. On the other hand, the expressions of SUOX decreased with tumor progression and showed an inverse association with tumor size. These types of biomarkers on OSCC were relatively limited. Deleted in liver cancer, keratin 13 and matriptase-2 were reported as these types of biomarkers [29, 62, 63].
In this study, we demonstrated that the reduction of SUOX expression was significantly associated with the stage but not with the tumor differentiation. Most biomarkers described before were reported to be related with several worse clinicopathological factors, including the stage and the tumor differentiation. A few markers, such as cysteine-rich 61 and Aurora B, were associated with only the stage and not with the tumor differentiation as well as SUOX [59, 60]. On the other hand, the expression of urokinase-type plasminogen activator receptor showed significant correlations with the tumor differentiation but not the tumor size. The reasons for these differences to the findings of our study are still unclear. Patient background, race, smoking and/or alcohol intake and cohort size might influence these differences.
The mechanisms by which down-expression of SUOX affected the carcinogenesis remain poorly understood. The following possibilities were considered. First, SUOX is an enzyme present in all normal eukaryotic mitochondria and is involved in ATP synthesis through oxidative phosphorylation. On the other hand, in cancer cells oxidative phosphorylation is suppressed and ATP is produced by the glycolytic pathway [64, 65]. Second, other known or unknown molecules that are altered with carcinogenesis in OSCC may regulate SUOX expression. Indeed, we found that the expression level of some molecules was correlated with SUOX alteration in our microarray analysis; however, the exact mechanisms are still unclear as we did not confirm the correlation between them by IHC.
It is possible that the difference in metabolic pathways between these normal cells and cancer cells may be involved in the reduction of SUOX expression in cancer cells.
The fact that SUOX acts to suppress carcinogenesis in tongue cancer suggests that SUOX may be a key molecule useful in clarifying patient condition and diagnosis, or as a potential target for treatment. There are many aspects of the role of SUOX in cancer that are yet unclear. Although not a detailed examination, our present microarray study identified multiple molecules either positively or inversely correlated with SUOX. Detailed in vitro and in vivo studies, involving factors related to SUOX, and not limited to OSCC, but targeting multiple carcinomas, are necessary to further clarify the specific functions of SUOX in relation to cancer.
References
Neville BW, Day TA (2002) Oral cancer and precancerous lesions. CA Cancer J Clin 52:195–215
Warnakulasuriya S (2009) Causes of oral cancer–an appraisal of controversies. Br Dent J 207:471–475
Larsen SR, Johansen J, Sorensen JA, Krogdahl A (2009) The prognostic significance of histological features in oral squamous cell carcinoma. J Oral Pathol Med 38:657–662
Marocchio LS, Lima J, Sperandio FF, Correa L, de Sousa SO (2010) Oral squamous cell carcinoma: an analysis of 1,564 cases showing advances in early detection. J Oral Sci 52:267–273
Troeltzsch M, Knosel T, Eichinger C, Probst F, Troeltzsch M, Woodlock T, Mast G, Ehrenfeld M, Otto S (2014) Clinicopathologic features of oral squamous cell carcinoma: do they vary in different age groups? J Oral Maxillofac Surg 72:1291–1300
El-Naggar AK, Chan JKC, Grandis JR, Takata T, Slootweg PJ (2016) WHO classification of head and neck tumours 4th Edition. WHO Press World Health Organization, Geneva, pp 109–118
Woolgar JA (2006) Histopathological prognosticators in oral and oropharyngeal squamous cell carcinoma. Oral Oncol 42:229–239
Woolgar JA, Triantafyllou A (2011) Squamous cell carcinoma and precursor lesions: clinical pathology. Periodontol 2000 57:51–72
Ebrahimi A, Murali R, Gao K, Elliott MS, Clark JR (2011) The prognostic and staging implications of bone invasion in oral squamous cell carcinoma. Cancer 117:4460–4467
Li Y, Bai S, Carroll W, Dayan D, Dort JC, Heller K, Jour G, Lau H, Penner C, Prystowsky M, Rosenthal E, Schlecht NF, Smith RV, Urken M, Vered M, Wang B, Wenig B, Negassa A, Brandwein-Gensler M (2013) Validation of the risk model: high-risk classification and tumor pattern of invasion predict outcome for patients with low-stage oral cavity squamous cell carcinoma. Head Neck Pathol 7:211–223
Weijers M, Snow GB, Bezemer PD, van der Wal JE, van der Waal I (2002) The clinical relevance of epithelial dysplasia in the surgical margins of tongue and floor of mouth squamous cell carcinoma: an analysis of 37 patients. J Oral Pathol Med 31:11–15
Safi AF, Kauke M, Grandoch A, Nickenig HJ, Zoller JE, Kreppel M (2017) Analysis of clinicopathological risk factors for locoregional recurrence of oral squamous cell carcinoma—retrospective analysis of 517 patients. J Craniomaxillofac Surg 45(10): 1749–1753
Mehanna HM, Rattay T, Smith J, McConkey CC (2009) Treatment and follow-up of oral dysplasia - a systematic review and meta-analysis. Head Neck 31:1600–1609
Gaykalova DA, Mambo E, Choudhary A, Houghton J, Buddavarapu K, Sanford T, Darden W, Adai A, Hadd A, Latham G, Danilova LV, Bishop J, Li RJ, Westra WH, Hennessey P, Koch WM, Ochs MF, Califano JA, Sun W (2014) Novel insight into mutational landscape of head and neck squamous cell carcinoma. PLoS One 9:e93102
Stransky N, Egloff AM, Tward AD, Kostic AD, Cibulskis K, Sivachenko A, Kryukov GV, Lawrence MS, Sougnez C, McKenna A, Shefler E, Ramos AH, Stojanov P, Carter SL, Voet D, Cortes ML, Auclair D, Berger MF, Saksena G, Guiducci C, Onofrio RC, Parkin M, Romkes M, Weissfeld JL, Seethala RR, Wang L, Rangel-Escareno C, Fernandez-Lopez JC, Hidalgo-Miranda A, Melendez-Zajgla J, Winckler W, Ardlie K, Gabriel SB, Meyerson M, Lander ES, Getz G, Golub TR, Garraway LA, Grandis JR (2011) The mutational landscape of head and neck squamous cell carcinoma. Science 333:1157–1160
Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, Speed TP (2003) Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4:249–264
Allred DC, Harvey JM, Berardo M, Clark GM (1998) Prognostic and predictive factors in breast cancer by immunohistochemical analysis. Mod Pathol 11:155–168
Sasahira T, Ueda N, Yamamoto K, Kurihara M, Matsushima S, Bhawal UK, Kirita T, Kuniyasu H (2014) Prox1 and FOXC2 act as regulators of lymphangiogenesis and angiogenesis in oral squamous cell carcinoma. PLoS One 9:e92534
Takada K, Toyokawa G, Okamoto T, Akamine T, Takamori S, Katsura M, Fujishita T, Shoji F, Oda Y, Maehara Y (2016) An immunohistochemical analysis of PD-L1 protein expression in surgically resected small cell lung cancer using different antibodies and criteria. Anticancer Res 36:3409–3412
Harvey JM, Clark GM, Osborne CK, Allred DC (1999) Estrogen receptor status by immunohistochemistry is superior to the ligand-binding assay for predicting response to adjuvant endocrine therapy in breast cancer. J Clin Oncol 17:1474–1481
Suzuki F, Oridate N, Homma A, Nakamaru Y, Nagahashi T, Yagi K, Yamaguchi S, Furuta Y, Fukuda S (2005) S100A2 expression as a predictive marker for late cervical metastasis in stage I and II invasive squamous cell carcinoma of the oral cavity. Oncol Rep 14:1493–1498
Yanagawa T, Omura K, Harada H, Nakaso K, Iwasa S, Koyama Y, Onizawa K, Yusa H, Yoshida H (2004) Heme oxygenase-1 expression predicts cervical lymph node metastasis of tongue squamous cell carcinomas. Oral Oncol 40:21–27
Fillies T, Werkmeister R, van Diest PJ, Brandt B, Joos U, Buerger H (2005) HIF1-alpha overexpression indicates a good prognosis in early stage squamous cell carcinomas of the oral floor. BMC Cancer 5:84
de Carvalho-Neto PB, dos Santos M, de Carvalho MB, Mercante AM, dos Santos VP, Severino P, Tajara EH, Louro ID, da Silva-Conforti AM (2013) FAS/FASL expression profile as a prognostic marker in squamous cell carcinoma of the oral cavity. PLoS One 8:e69024
Bova RJ, Quinn DI, Nankervis JS, Cole IE, Sheridan BF, Jensen MJ, Morgan GJ, Hughes CJ, Sutherland RL (1999) Cyclin D1 and p16INK4A expression predict reduced survival in carcinoma of the anterior tongue. Clin Cancer Res 5:2810–2819
Tsai ST, Jin YT, Tsai WC, Wang ST, Lin YC, Chang MT, Wu LW (2005) S100A2, a potential marker for early recurrence in early-stage oral cancer. Oral Oncol 41:349–357
Liu LK, Jiang XY, Zhou XX, Wang DM, Song XL, Jiang HB (2010) Upregulation of vimentin and aberrant expression of E-cadherin/beta-catenin complex in oral squamous cell carcinomas: correlation with the clinicopathological features and patient outcome. Mod Pathol 23:213–224
Pande P, Soni S, Kaur J, Agarwal S, Mathur M, Shukla NK, Ralhan R (2002) Prognostic factors in betel and tobacco related oral cancer. Oral Oncol 38:491–499
Tripathi SC, Kaur J, Matta A, Gao X, Sun B, Chauhan SS, Thakar A, Shukla NK, Duggal R, Choudhary AR, DattaGupta S, Sharma MC, Ralhan R, Siu KW (2012) Loss of DLC1 is an independent prognostic factor in patients with oral squamous cell carcinoma. Mod Pathol 25:14–25
Kisker C, Schindelin H, Pacheco A, Wehbi WA, Garrett RM, Rajagopalan KV, Enemark JH, Rees DC (1997) Molecular basis of sulfite oxidase deficiency from the structure of sulfite oxidase. Cell 91:973–983
D’Errico G, Di Salle A, La Cara F, Rossi M, Cannio R (2006) Identification and characterization of a novel bacterial sulfite oxidase with no heme binding domain from Deinococcus radiodurans. J Bacteriol 188:694–701
Tan WH, Eichler FS, Hoda S, Lee MS, Baris H, Hanley CA, Grant PE, Krishnamoorthy KS, Shih VE (2005) Isolated sulfite oxidase deficiency: a case report with a novel mutation and review of the literature. Pediatrics 116:757–766
Cohen HJ, Betcher-Lange S, Kessler DL, Rajagopalan KV (1972) Hepatic sulfite oxidase. Congruency in mitochondria of prosthetic groups and activity. J Biol Chem 247:7759–7766
Feng C, Wilson HL, Hurley JK, Hazzard JT, Tollin G, Rajagopalan KV, Enemark JH (2003) Essential role of conserved arginine 160 in intramolecular electron transfer in human sulfite oxidase. Biochemistry 42:12235–12242
Johnson JL (2003) Prenatal diagnosis of molybdenum cofactor deficiency and isolated sulfite oxidase deficiency. Prenat Diagn 23:6–8
Dublin AB, Hald JK, Wootton-Gorges SL (2002) Isolated sulfite oxidase deficiency: MR imaging features. Am J Neuroradiol 23:484–485
Jin GZ, Yu WL, Dong H, Zhou WP, Gu YJ, Yu H, Yu H, Lu XY, Xian ZH, Liu YK, Cong WM, Wu MC (2013) SUOX is a promising diagnostic and prognostic biomarker for hepatocellular carcinoma. J Hepatol 59:510–517
Muys BR, Lorenzi JC, Zanette DL, Lima e Bueno Rde B, de Araujo LF, Dinarte-Santos AR, Alves CP, Ramao A, de Molfetta GA, Vidal DO, Silva WA Jr (2016) Placenta-enriched LincRNAs MIR503HG and LINC00629 decrease migration and invasion potential of JEG-3 cell line. PLoS One 11:e0151560
Yue H, Cai Y, Song Y, Meng L, Chen X, Wang M, Bian Z, Wang R (2017) Elevated TARP promotes proliferation and metastasis of salivary adenoid cystic carcinoma. Oral Surg Oral Med Oral Pathol Oral Radiol 123:468–476
Rotunno M, Hu N, Su H, Wang C, Goldstein AM, Bergen AW, Consonni D, Pesatori AC, Bertazzi PA, Wacholder S, Shih J, Caporaso NE, Taylor PR, Landi MT (2011) A gene expression signature from peripheral whole blood for stage I lung adenocarcinoma. Cancer Prev Res (Phila) 4:1599–1608
Porpaczy E, Bilban M, Heinze G, Gruber M, Vanura K, Schwarzinger I, Stilgenbauer S, Streubel B, Fonatsch C, Jaeger U (2009) Gene expression signature of chronic lymphocytic leukaemia with Trisomy 12. Eur J Clin Investig 39:568–575
Das ND, Park JH, Jung KH, Lee HT, Park KS, Choi MR, Chai YG (2011) Sodium arsenite dependent protein expression analysis on human embryonic carcinoma (NCCIT) cell line. Toxicol Lett 207:149–158
Fan S, Liu B, Sun L, Lv XB, Lin Z, Chen W, Chen W, Tang Q, Wang Y, Su Y, Jin S, Zhang D, Zhong J, Li Y, Wen B, Zhang Z, Yang P, Zhou B, Liang Q, Yu X, Zhu Y, Hu P, Chu J, Huang W, Feng Y, Peng H, Huang Q, Song E, Li J (2015) Mitochondrial fission determines cisplatin sensitivity in tongue squamous cell carcinoma through the BRCA1-miR-593-5p-MFF axis. Oncotarget 6:14885–14904
Lin CC, Hsu YC, Li YH, Kuo YY, Hou HA, Lan KH, Chen TC, Tzeng YS, Kuo YY, Kao CJ, Chuang PH, Tseng MH, Chiu YC, Chou WC, Tien HF (2017) Higher HOPX expression is associated with distinct clinical and biological features and predicts poor prognosis in de novo acute myeloid leukemia. Haematologica 102:1044–1053
Del Valle PR, Milani C, Brentani MM, Katayama ML, de Lyra EC, Carraro DM, Brentani H, Puga R, Lima LA, Rozenchan PB, Nunes Bdos S, Goes JC, Azevedo Koike Folgueira M A (2014) Transcriptional profile of fibroblasts obtained from the primary site, lymph node and bone marrow of breast cancer patients. Genet Mol Biol 37: 480–489
Nagy A, Pongor LS, Szabo A, Santarpia M, Gyorffy B (2017) KRAS driven expression signature has prognostic power superior to mutation status in non-small cell lung cancer. Int J Cancer 140:930–937
Subramanian S, West RB, Marinelli RJ, Nielsen TO, Rubin BP, Goldblum JR, Patel RM, Zhu S, Montgomery K, Ng TL, Corless CL, Heinrich MC, van de Rijn M (2005) The gene expression profile of extraskeletal myxoid chondrosarcoma. J Pathol 206:433–444
Porpaczy E, Tauber S, Bilban M, Kostner G, Gruber M, Eder S, Heintel D, Le T, Fleiss K, Skrabs C, Shehata M, Jager U, Vanura K (2013) Lipoprotein lipase in chronic lymphocytic leukaemia - strong biomarker with lack of functional significance. Leuk Res 37:631–636
Kuang J, Li QY, Fan F, Shen NJ, Zhan YJ, Tang ZH, Yu WL (2017) Overexpression of the X-linked ribosomal protein S4 predicts poor prognosis in patients with intrahepatic cholangiocarcinoma. Oncol Lett 14:41–46
Shen L, Du X, Ma H, Mei S (2017) miR-1193 suppresses the proliferation and invasion of human T-cell leukemia cells through directly targeting the transmembrane 9 superfamily 3 (TM9SF3). Oncol Res 25:1643–1651
Zhao L, Zhao Y, He Y, Mao Y (2017) miR-19b promotes breast cancer metastasis through targeting MYLIP and its related cell adhesion molecules. Oncotarget 8:64330–64343
Bruggemann M, Gromes A, Poss M, Schmidt D, Klumper N, Tolkach Y, Dietrich D, Kristiansen G, Muller SC, Ellinger J (2017) Systematic analysis of the expression of the mitochondrial ATP synthase (Complex V) subunits in clear cell renal cell carcinoma. Transl Oncol 10:661–668
Zilberg C, Lee MW, Yu B, Ashford B, Kraitsek S, Ranson M, Shannon K, Cowley M, Iyer NG, Palme CE, Ch’ng S, Low T, O’Toole H, Clark S, Gupta JR R (2017) Analysis of clinically relevant somatic mutations in high-risk head and neck cutaneous squamous cell carcinoma. Mod Pathol. https://doi.org/10.1038/modpathol.2017.128
Sen S, Carnelio S (2016) Expression of epithelial cell adhesion molecule (EpCAM) in oral squamous cell carcinoma. Histopathology 68:897–904
Hema K, Rao K, Devi HU, Priya N, Smitha T, Sheethal H (2014) Immunohistochemical study of CD44s expression in oral squamous cell carcinoma-its correlation with prognostic parameters. J Oral Maxillofac Pathol 18:162–168
Byatnal AA, Byatnal A, Sen S, Guddattu V, Solomon MC (2015) Cyclooxygenase-2—an imperative prognostic biomarker in oral squamous cell carcinoma—an immunohistochemical study. Pathol Oncol Res 21:1123–1131
Tang JY, Hsi E, Huang YC, Hsu NC, Yang WC, Chang HW, Chai CY, Chu PY (2015) Overexpression of autophagy-related 16-like 1 in patients with oral squamous cell carcinoma. Pathol Oncol Res 21:301–305
Xia F, Xu JC, Zhang P, Zhang YY, Zhang QW, Chao ZH, Wang F (2014) Glucose-regulated protein 78 and heparanase expression in oral squamous cell carcinoma: correlations and prognostic significance. World J Surg Oncol 12:121
Kok SH, Chang HH, Tsai JY, Hung HC, Lin CY, Chiang CP, Liu CM, Kuo MY (2010) Expression of Cyr61 (CCN1) in human oral squamous cell carcinoma: An independent marker for poor prognosis. Head Neck 32:1665–1673
Pannone G, Hindi SA, Santoro A, Sanguedolce F, Rubini C, Cincione RI, De Maria S, Tortorella S, Rocchetti R, Cagiano S, Pedicillo C, Serpico R, Lo Muzio L, Bufo P (2011) Aurora B expression as a prognostic indicator and possible therapeutic target in oral squamous cell carcinoma. Int J Immunopathol Pharmacol 24:79–88
Bacchiocchi R, Rubini C, Pierpaoli E, Borghetti G, Procacci P, Nocini PF, Santarelli A, Rocchetti R, Ciavarella D, Lo Muzio L, Fazioli F (2008) Prognostic value analysis of urokinase-type plasminogen activator receptor in oral squamous cell carcinoma: an immunohistochemical study. BMC Cancer 8:220
Kitamura R, Toyoshima T, Tanaka H, Kawano S, Kiyosue T, Matsubara R, Goto Y, Hirano M, Oobu K, Nakamura S (2012) Association of cytokeratin 17 expression with differentiation in oral squamous cell carcinoma. J Cancer Res Clin Oncol 138:1299–1310
Cheng MF, Lin LH, Huang MS, Lee HS, Ji DD, Lin CS, Hsia KT (2017) Downexpression of matriptase-2 correlates with tumor progression and clinical prognosis in oral squamous-cell carcinoma. Appl Immunohistochem Mol Morphol 25:481–488
Gatenby RA, Gillies RJ (2004) Why do cancers have high aerobic glycolysis? Nat Rev Cancer 4:891–899
Kim JW, Dang C V (2006) Cancer’s molecular sweet tooth and the Warburg effect. Cancer Res 66:8927–8930
Acknowledgements
We thank Mr. Hiroto Fukushima and Ms. Kumiko Tsubone for their assistance in our experiments.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors have declared no conflicts of interest.
Rights and permissions
About this article
Cite this article
Nakamura, K., Akiba, J., Ogasawara, S. et al. SUOX is negatively associated with multistep carcinogenesis and proliferation in oral squamous cell carcinoma. Med Mol Morphol 51, 102–110 (2018). https://doi.org/10.1007/s00795-017-0177-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00795-017-0177-4