Abstract
Catalases, heme or manganese, are efficient biocatalysts that split hydrogen peroxide into water and oxygen. We have cloned a manganese catalase from thermophilic bacterium, Geobacillus thermopakistaniensis, and expressed the corresponding gene in Escherichia coli. The gene product, CatGt, was synthesized in E. coli as inactive inclusion bodies. Solubilization and refolding of the inclusion bodies resulted in highly active CatGt with a specific activity of 18,521 μmol min−1 mg−1. The refolded protein exhibited apparent Km and kcat values of 260 mM and 10,360 s−1 subunit−1, respectively. It exhibited a half-life of 1 h at 100 °C. The unique features of CatGt are its high activity and thermostability. These features make it a valuable catalyst for industrial applications. To the best of our knowledge, CatGt is the most thermostable catalases characterized to date.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
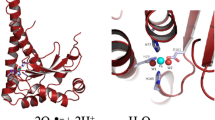
Catalases (EC1.11.1.6), more precisely hydroperoxidases, are a class of enzymes that specifically catalyze the decomposition of hydrogen peroxide to water and oxygen (Nicholls et al. 2001; Chelikani et al. 2004). Catalases are found in all the three domains of life and are involved in the cellular defense system to combat oxidative stress (Gonzalez-Flecha and Demple 1995; Fang 2004; Veal et al. 2007; Korshunov and Imlay 2010). On the basis of primary structure, they are classified into three groups. Two of them are comprised heme catalases while the third contains the non-heme catalases (Klotz et al.1997; Klotz and Loewen 2003). The most widespread and extensively characterized class consists of monofunctional heme-containing catalases. The second class is composed of bifunctional heme-containing catalase–peroxidases which exhibit catalase as well as peroxidase activity (Loewen et al. 2000). The active center in heme-catalases is a mononuclear iron porphyrin. The third class, non-heme catalases, utilizes manganese instead of iron, and is known as manganese catalase group (Chelikani et al. 2004; Loewen 2004; Nicholls 2012).
The major use of catalases is in food, medical, paper and textile industries for removing hydrogen peroxide from products or by-products (Rowe 1999; Akyilmaz and Kozgus 2009; Sharif et al. 2016). A minor use is in contact lens hygiene. Contact lenses are normally disinfected by hydrogen peroxide which is then decomposed by a catalase solution before using again (Cook and Worsley 1996). These applications require stable, particularly thermostable, catalases. The thermostable enzymes are usually obtained from thermophilic organisms, as these organisms are adapted to function optimally at high temperatures (Coolbear et al. 1992; Adams and Kelly 1998). In addition, the enzymes from these sources are more resistant to denaturing agents such as urea (Kristjansson1989; Owusu and Cowan 1989; Demirjian et al. 2001; Chohan and Rashid 2013; Gharib et al. 2016; Arif et al. 2018). Among catalases, heme-catalases are well characterized; however, they are generally thermolabile. In contrast, manganese catalases are more thermostable because of their characteristic four-helix bundle protein architecture (Chelikani et al. 2004; Whittaker 2011).
A protein data base search, provided by National Center for Biotechnology Information (https://www.ncbi.nlm.nih.gov/protein/?term=manganese-catalase), revealed nearly 28,000 protein sequences annotated as manganese catalases. However, a very few of them have been experimentally verified. There are only nine reports available on the characterization of bacterial and archaeal manganese catalases. These include Lactobacillus plantarum (Kono and Fridovich 1983; Barynin et al. 2001), Thermus thermophilus (Barynin and Grebenko 1986; Barynin et al. 1997; Antonyuk et al. 2000; Hidalgo et al. 2004), Thermoleophilum album (Allgood and Perry 1986), Thermus sp. YS 8–13 (Kagawa et al. 1999; Mizobata et al. 2000), Pyrobaculum calidifontis (Amo et al. 2002), Metallosphaera hakonensis (Ebara and Shigemori, 2008), Thermomicrobium roseum (Baginski and Sommerhalter 2017), Anabaena sp. PCC7120 (Bihani et al. 2016) and Geobacillus sp. WCH70 (Li et al. 2017). Purification of thermostable manganese catalases from their native organisms is difficult due to the low-level production. However, heterologous expression of these genes in a mesophilic host, such as Escherichia coli, provides large quantities of recombinant protein in less time and cost (Lee 1996; Sorensen and Mortensen 2005; Rosano and Ceccarelli 2014). Therefore, we attempted the production of recombinant manganese catalase from a thermophilic bacterium.
Geobacillus thermopakistaniensis is a thermophilic, aerobic, Gram-positive bacterium, which exhibits oxidase and catalase activities (Tayyab et al. 2011). The genome of the microorganism has been sequenced (Siddiqui et al. 2014) and available online with an accession number AYSF01000034. The genome sequence contains an open reading frame annotated as manganese catalase (accession # ESU72991). Here, we have described cloning, expression in Escherichia coli, and characterization of the thermostable manganese catalase, a product of ESU72991.
Materials and methods
Chemicals, bacterial strains and cultivation conditions
Analytical grade chemicals were used in this study and procured either from Fluka Chemical Corporation, Sigma-Aldrich Company or Thermo Fisher Scientific Inc. Restriction enzymes, DNA polymerase, DNA and protein standards, and DNA purification kits were obtained from Thermo Fisher Scientific Inc. Specific oligonucleotide primers were obtained from Macrogen, Inc. Cloning vector pTZ57R/T (Thermo Fisher Scientific Inc) and expression vector pET-21a(+) (Novagen–Merck Millipore) were used for cloning and expression of the target gene, respectively. Similarly, E. coli DH5α was used for plasmid preparation, and E. coli CodonPlus (DE3)-RIL (Stratagene, La Jolla, CA, USA) was employed for heterologous gene expression. G. thermopakistaniensis and E. coli were cultivated in LB medium (yeast extract 0.5%, tryptone1%, NaCl 0.5%) and cultivated at 60 and 37 °C, respectively.
Recombinant plasmid constructs
The gene encoding CatGt (accession # AYSF01000034) was amplified by polymerase chain reaction (PCR) using a set of sequence-specific forward (5′-CATATGGTATTTCAGCGGATCGACCGGCTGGCG) and reverse (5′-TCACAACCCTGCCGACCGCTGCAGCCGCTTGG) primers. An NdeI restriction site (bold underlined sequence) was incorporated in the forward primer. G. thermopakistaniensis genomic DNA was used as target for amplification of CatGt gene. The PCR-amplified gene product was inserted in pTZ57R/T cloning vector and the resulting plasmid was designated as pTZ-Cat. CatGt gene was excised from pTZ-Cat plasmid utilizing NdeI and HindIII restriction enzymes and ligated at the corresponding sites in pET-21a (+) expression vector. The resulting plasmid was named pET-Cat. The gene, in pET-Cat, was sequenced using T7 promoter and T7 terminator primers prior to expression of the gene in E. coli.
Synthesis of recombinant CatGt in E. coli
Transformation of E. coli BL21-CodonPlus (DE3)-RIL cells was carried out using pET-Cat recombinant plasmid. Isopropyl β-D-1-thiogalactopyranoside (IPTG) 0.2 mM was used to induce the gene expression. The cells were harvested by centrifugation at 6000×g, after 4 h of induction. The cell pellet, after resuspension in 50 mM Tris–Cl (pH 8.0), was lysed by sonication. After lysis, soluble and insoluble proteins were separated by centrifugation at 10,000×g. Analysis of proteins was done by 14% denaturing polyacrylamide gel electrophoresis (SDS-PAGE).
Solubilization and refolding of recombinant CatGt
Inclusion bodies, containing recombinant CatGt, were washed four–five times with 50 mM Tris–Cl (pH 8.0) containing 1% Triton X-100 and dissolved in solubilization buffer (8 M urea, 2 mM DTT, 50 mM glycine and 20 mM Tris–Cl pH 8.0) and heated at 100 °C for 10 min. The solubilized CatGt was dialyzed at 4 °C against refolding buffer (6 M urea in 20 mM Tris–Cl buffer, pH 8.0) for 4 h and gradually decreasing the concentration of urea to 5, 4, 3, 2, 1, 0.5 and 0 M by exchanging the refolding buffer after 3–4-h intervals. After complete removal of urea, the samples were centrifuged and supernatant was analyzed by SDS-PAGE and activity assays.
Gel filtration chromatography
Oligomeric nature of CatGt was determined by gel filtration chromatography using Superdex 200 10/300 GL column (GE Healthcare, NJ). The column was equilibrated with 50 mM Tris–Cl, pH 8.0, containing 150 mM NaCl. Protein sample (prepared in the same buffer) was loaded and eluted at a flow rate of 0.3 mL/min.
Catalase activity assays
Catalase activity was determined using a spectrophotometer (Shimadzu UV-1601) equipped with a thermoelectric cell. Routine activity assays were carried out at 60 °C using 20 mM hydrogen peroxide in 50 mM sodium phosphate buffer (pH 7.0). Substrate decomposition was measured at 240 nm (ε240 = 43.6 M−1 cm−1) (Beers and Sizer 1952; Hildebrandt and Roots 1975). One unit of catalase activity was defined as the amount of enzyme needed to convert 1 µmol of hydrogen peroxide to water and oxygen per min.
Peroxidase activity, using various substrates, was measured spectrophotometrically. Assays were performed at 60 °C unless mentioned otherwise. The reactions were carried out in 50 mM sodium phosphate buffer (pH 7.0) containing 3 mM hydrogen peroxide as electron acceptor and appropriate amount of electron donor including pyrogallol, o-dianisidine, glutathione, NADH and NADPH. The oxidation of pyrogallol, at a final concentration of 42 mM, was monitored by measuring the increase in absorbance at 430 nm. The oxidation o-dianisidine, at a final concentration of 0.3 mM, was monitored by measuring the decrease in absorbance at 460 nm. The oxidation of NADH or NADPH, at a final concentration of 0.2 mM, was monitored by measuring the decrease in absorbance at 340 nm. The oxidation of glutathione, at a final concentration of 2 mM, was monitored by coupling with glutathione reductase (1 U) in the presence of 0.2 mM NADPH and measuring the decrease in absorbance at 340 nm.
Temperature, pH and metal ions effect
To estimate the optimum temperature and pH, catalase activity of CatGt was measured at different temperatures (40–80 °C) without changing the pH, or altering the pH values (5.5–9.0) and keeping the temperature unchanged. A control experiment (without CatGt) was included at each temperature and pH, and the control values were subtracted from the values obtained with CatGt.
To study the effect of metal ions on enzyme activity, the chloride salts of Ni2+, Hg2+, Mn2+, Fe2+, Cu2+, Mg2+, Ca2+ and Co2+ were added at a fixed concentration of 100 μM. The activity assays were performed in 50 mM sodium phosphate buffer (pH 7.0) at 60 °C. The activity assays were carried out in triplicate. Control experiments without the addition of any metal ions and addition of EDTA were also employed.
Thermostability analysis
To determine thermostability, CatGt samples in 50 mM sodium phosphate buffer (pH 7.0) were incubated at different temperatures (60, 80 and 100 °C). Aliquots were taken out at various intervals and assayed for residual activity as described above. The activity of the protein sample kept on ice was taken as 100%.
Structural analysis by circular dichroism
Structural stability of CatGt was examined by circular dichroism (CD) spectroscopy utilizing Chirascan™-Plus CD Spectrometer (Applied Photophysics, UK). CatGt samples, in 30 mM Tris–Cl (pH 8.0), were incubated at 60 and 90 °C for 5 min. CD spectra were monitored in far-UV range (200–260 nm). The values of solvent spectra were subtracted from those of CatGt values.
Chemical denaturation analysis
CatGt, at a final concentration of 0.5 mg mL−1, was prepared in various concentrations of urea (0–8 M) or guanidine hydrochloride (0–6 M) and incubated at room temperature for 30 min and residual activity was examined as described above. Similarly, the structural stability of CatGt, in the presence of denaturants, was examined by circular dichroism spectroscopy as described above.
Kinetic analysis
Initial velocities of decomposition of hydrogen peroxide were measured in 50 mM phosphate buffer (pH 7.0) using various concentrations of hydrogen peroxide and fixed amount of CatGt. The apparent Km and Vmax were calculated using the Michaelis–Menten equation.
Analysis of metal content
The metal ion contents of the recombinant CatGt were analyzed by inductively coupled plasma-optical emission spectrometry (ICP-OES) using an Avio 500 ICP-OES spectrophotometer (PerkinElmer Inc). Calibration curves were obtained by the analysis of the PerkinElmer multi-element standard solutions. All solutions were prepared in deionized double-distilled water.
Results
Sequence analysis
CatGt gene (accession # ESU72991) comprises 900 nucleotides encoding a 33,556 Da polypeptide of 299 amino acids and a pI of 5.88. A comparison of the amino acid sequence of CatGt displayed the highest identity of 99.67% with an uncharacterized manganese catalase from Geobacillus stearothermophilus (accession # KFL16886). Among the characterized enzymes, the highest identity of 68.15% was found with a manganese catalase from thermophilic bacterium, T. roseum (Baginski and Sommerhalter 2017), followed by 61.56% to an archaeal catalase from P. calidifontis (Amo et al. 2002) and 58.75% to the counterpart from T. thermophilus (Barynin and Grebenko 1986; Barynin et al. 1997; Antonyuk et al. 2000). Manganese ions have been shown to coordinate with Glu36, Glu70, His73, Glu155 and His188 amino acids in the crystal structure of manganese catalase from T. thermophilus (Barynin et al. 2001). Furthermore, Lys158 is shown to be involved in catalytic proton transfer in this catalase. These amino acids are completely conserved in CatGt (Fig. 1). Surprisingly, CatGt exhibited only a 22% identity with a recently characterized manganese catalase from Geobacillus sp. WCH70 (Li et al. 2017). The genome sequence search of G. thermopakistaniensis revealed a similar protein with 92.59% identity, annotated as a hypothetical protein harboring a dimanganese centre, which could be a possible counterpart of manganese catalase from Geobacillus sp. WCH70.
Comparison of amino acid sequence of CatGt and characterized manganese catalases. Abbreviations at the left terminal are Tr (Thermomicrobium roseum; WP_012642239), Pc (Pyrobaculum calidifontis VA1; BAB97198), Tt (Thermus thermophilus; WP_014630234), Ts (Thermus sp. YS 8–13; BAA75627), Mh (Metallosphaera hakonensis; BAF81110), Ta (Thermoleophilum album; BAB87719), Lp (Lactobacillus plantarum; WP_103420789), As (Anabaena sp. PCC7120; WP_016949195), and Gs (Geobacillus sp. WCH70; ACS24898). Alignment was carried out using CLUSTAL W. Amino acids conserved in all the sequences are shown by asterisks. The structure of T. thermophilus catalase was used to determine the domain boundaries (Antonyuk et al. 2000). The amino acid residues involved in coordination with manganese atoms are shown with a grey background. The Lys158 (in T. thermophilus) or Glu174 (in L. plantarum) residues considered to be involved in catalytic proton transfer are encircled. The box shows calcium-binding site of L. plantarum catalase
Cloning and expression of CatGt
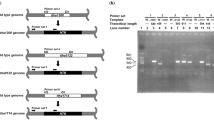
A 0.9-kb DNA fragment, matching exactly the size of CatGt gene, was amplified by PCR and inserted in pTZ-Cat. Double digestion of pTZ-Cat using NdeI and HindIII showed the release of a 0.9-kb DNA fragment confirming the presence of CatGt gene in the recombinant plasmid. Similarly, digestion of recombinant pET-Cat utilizing the same pair of restriction enzymes confirmed the presence of CatGt gene in the expression vector. DNA sequencing showed the absence of any unintended mutation in the cloned gene.
CatGt gene expression in E. coli BL21-CodonPlus (DE3)-RIL at 37 °C resulted in the synthesis of the recombinant protein mainly in insoluble and inactive form (Fig. 2). Various attempts were made by changing the expression conditions to get the recombinant protein in active and soluble form. Neither reduction of the cultivation temperature nor the inducing concentration of IPTG affected solubilization of recombinant CatGt. Furthermore, activation of host chaperonins, by giving a heat shock to the host culture at 45 °C, prior to induction with IPTG, and supply of Mn2+ in the growth medium were also proved unsuccessful for the production of soluble CatGt (data not shown).
Coomassie brilliant blue-stained SDS-PAGE demonstrating the production and refolding of recombinant CatGt. Lane M, molecular weight marker; lane 1, crude cellular extract of E. coli BL21-CodonPlus(DE3)-RIL harboring pET-21a(+); lane 2, crude cellular extract of E. coli BL21-CodonPlus(DE3)-RIL-harboring pET-Cat; lane 3, soluble fraction of E. coli BL21-CodonPlus(DE3)-RIL harboring pET-Cat; lane 4, insoluble fraction of E. coli BL21-CodonPlus(DE3)-RIL harboring pET-Cat; lane 5, refolded and active CatGt
Solublization of inclusion bodies of CatGt in 8 M urea and fractional dialysis for subsequent removal of urea resulted in attaining soluble and active protein. From 1-L culture, 2920 mg (wet weight) cells, and nearly 187 mg inclusion bodies were obtained. Denaturation and refolding of inclusion bodies yielded approximately 41 mg of the soluble and active CatGt. These results indicate that about 22% of the initially used insoluble protein was recovered in soluble and active form. SDS-PAGE analysis of the refolded CatGt showed a single protein band (Fig. 2).
When the refolded CatGt was passed through gel filtration column, the protein eluted at a retention volume of 10.2 mL that corresponded to a molecular weight of ~ 135 kDa, calculated from the standard curve. These results indicated that enzymatically active CatGt exists in tetrameric form in solution.
Temperature and pH effects
To find the effect of temperature on the activity of CatGt, we examined the activity at various temperatures without changing the pH. The activity gradually increased with an increase in temperature till 60 °C and thereafter it decreased (Fig. 3a). The optimum temperature for catalase activity of CatGt complemented the optimal growth temperature of G. thermopakistaniensis (60 °C).
Effect of temperature, pH and denaturants on CatGt. a Optimal temperature. CatGt activity was measured at different temperatures while keeping the pH unaltered. b Stability of CatGt against temperature. CatGt was incubated at 60 (circles), 80 (squares) and 90 °C (triangles) for different time intervals and remaining activity was measured under optimal conditions. c Optimal pH. CatGt activity was measured by varying the pH and keeping the temperature unchanged at 60 °C. The buffers used were 50 mM sodium phosphate, pH 5.5–7.5 (circles) and 50 mM Tris–Cl, pH 7.5–9.0 (squares). d Far-UV spectrum of CatGt at 60 °C (circles) and 90 °C (triangles). e Far-UV spectrum of CatGt without the addition of urea (triangles), with addition of 2 M urea (empty diamonds), 4 M urea (circles), 8 M urea (squares). f Far-UV spectrum of CatGt without the addition of guanidine hydrochloride (triangles), with addition of 2 M guanidine hydrochloride (squares), 4 M guanidine hydrochloride (empty diamonds), 6 M guanidine hydrochloride (circles). The error bars represent standard deviation (SD)
The effect of temperature was also examined by incubating CatGt at various temperatures for different intervals of time and the examining the residual activity at 60 °C. Even after 8 h of incubation at this temperature, no significant effect on enzyme activity was observed. CatGt exhibited half-lives of nearly 8 h at 80 °C and 1 h at 100 °C (Fig. 3b).
To find the effect of pH on activity of CatGt, catalase activity was observed at different pH while keeping the temperature unchanged. Highest activity was found at pH 7.0–7.5 in sodium phosphate buffer (Fig. 3c).
The stability of CatGt structure was examined by CD spectroscopy at 60 and 90 °C. CD spectra for both samples did not show any significant change, indicating that CatGt preserves the secondary structure even at 90 °C (Fig. 3d).
Addition of denaturants such as urea or guanidine hydrochloride to proteins usually results in loss of protein activity. However, some hyperthermophilic proteins are reported as an exception (Chohan and Rashid 2013; Gharib et al. 2016; Arif et al. 2018; Rasool et al. 2010). When we examined the effect of urea on CatGt, we found that enzyme activity was almost unaffected. When CatGt was incubated in the presence of 8 M urea for 30 min, 90% of the enzyme activity remained intact. These results indicated that although there was no significant decrease in catalase activity of the sample, the secondary structure was affected as reflected by the CD spectra (Fig. 3e).
In contrast to urea, incubation of CatGt in the presence of 2 M guanidine hydrochloride caused a 25% decrease in enzyme activity. The activity was almost completely lost when CatGt was incubated for 30 min in the presence of 6 M guanidine hydrochloride. The CD spectrum of this sample was also significantly changed. This agrees to the fact that guanidine hydrochloride is a more effective denaturant being a charged molecule, whereas urea being an uncharged molecule is devoid of ionic strength effects.
A comparison of structural changes (Fig. 3e, f) revealed that the spectral shift is higher in the CatGt sample incubated with 8 M urea compared to 2 M guanidine hydrochloride, in contrast to their catalase activities. Therefore, we examined the manganese content of both the samples assuming that guanidine hydrochloride affects not only secondary structure but also the binuclear manganese active site, resulting in loss of catalase activity. However, we could not come to any logical conclusion as manganese contents of both the samples were almost similar. Therefore, we speculate that the denaturation of CatGt is reversible in the presence of urea (as it is denaturant only), and irreversible in the presence of guanidine hydrochloride (as it is a denaturant as well as a salt, hence exhibits ionic strength). Similar results have been reported by Rajagopalan and co-workers (Rajagopalan et al. 1961) who have concluded that the enzymes which require metal ions as cofactors or which contain a metal as an integral part of the molecule have been found to be resistant to inhibition by urea. They have reported that urea causes denaturation which can be reversed upon dilution due to subsequent restoration of structural and catalytic properties. We speculate the same in case of CatGt because the protein treated with urea or guanidine hydrochloride was 100 times diluted in our reaction mixture. The observed spectral shift in the presence of urea or guanidine hydrochloride was of the undiluted sample.
Effect of metal ions
Incubation of CatGt with 1 mM EDTA or addition of 1 mM EDTA in the assay mixture considerably decreased the activity of enzyme. This indicated that enzymatic activity of CatGt might be activated by metal ions. When we examined the effect of various metal ions, a twofold enhancement in the CatGt activity was found in the presence of 100 μM Mn2+ (Fig. 4a). Therefore, the activity was examined in the presence of different concentrations of Mn2+, and there was more than tenfold increase in activity with 2 mM Mn2+ compared to the control containing no additional metal ions (Fig. 4b).
Effect of metal ions on CatGt activity. a Effect of various metal ions. CatGt activity was measured in the presence of various metal ions at a final concentration of 100 µM. b Effect of various concentrations of Mn2+ on CatGt activity. Catalase activity was determined using various concentrations of Mn2+. Activity assays were carried out under optimal conditions. The error bars represent standard deviation (SD)
When we examined the catalase activity of CatGt, refolded in the presence and absence of 2 mM Mn2+, a twofold enhancement was observed in the protein sample refolded with the addition of Mn2+. These activity measurements were performed without the addition of MnCl2 in the assay mixture. This indicated that most of the CatGt refolded without Mn2+ might be in an apo-form which could be converted to holo-form by Mn2+ supplementation.
When CatGt, refolded in the presence or absence of 2 mM Mn2+, was incubated with 2 mM Mn2+ at 60 °C for 1 h and assayed for catalase activity, there was a tenfold enhancement in the activity of both the samples as compared to their respective samples without incubation. These results indicate that higher temperature might be contributing to the incorporation of Mn2+ in active catalytic center which was confirmed by comparing the manganese contents of both the samples.
Metal content of CatGt
When we examined the metal content of the refolded CatGt, it was found that one subunit of CatGt, refolded in the absence of Mn2+, contained 0.45 ± 0.05 manganese atoms. However, CatGt refolded in the presence of Mn2+ contained 0.8 ± 0.04 atoms of manganese per subunit of CatGt. These results are consistent with catalase activity of both the samples as described above. We further examined the manganese contents of CatGt, refolded in the presence or absence of Mn2+, and later on incubated with 2 mM Mn2+ at 60 °C. The results showed 1.22 ± 0.02 manganese atoms per subunit of CatGt sample refolded in the absence of Mn2+ and 1.51 ± 0.04 atoms per subunit in CatGt sample refolded in the presence of Mn2+. The higher content of Mn2+ in the CatGt sample incubated at 60 °C may be attributed to flexible structure at higher temperature (60 °C), which may have facilitated the incorporation of Mn2+ in catalytic center. Similar results have also been previously reported by Mizobata and co-workers (Mizobata et al. 2000).
Effect of sodium azide
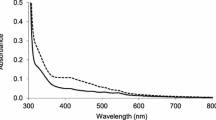
Sodium azide is a potent inhibitor of heme-catalases whereas manganese catalases are considerably resistant to it (Nicholls 1964). Inhibition by sodium azide is usually employed to differentiate between these catalases. We, therefore, compared the inhibition of CatGt and a heme-catalase from bovine liver (Sigma-Aldrich). Activity assays were conducted with the addition of 0–5 mM sodium azide. CatGt exhibited more than 90% activity at 1 mM sodium azide. In contrast, under same concentration of sodium azide, the activity of heme-catalase from bovine liver was completely inhibited. Even at 0.1 mM sodium azide there was a 50% inhibition of the heme-catalase (Fig. 5). Furthermore, the absorption spectra of CatGt and heme-catalase from bovine liver were compared. The heme-catalase, due to the heme prosthetic group, showed a Soret band at 407 nm (Fig. 6a), while no such band was found in CatGt spectra (Fig. 6b). These results indicated that the CatGt is a manganese catalase, not a heme-catalase.
Substrate specificity
When the substrate specificity of CatGt was analyzed for different substrates including hydrogen peroxide, pyrogallol, o-dianisidine, NADH, NADPH, and glutathione. Highest activity was observed against hydrogen peroxide (18,521 U mg−1) followed by pyrogallol (463 U mg−1). No significant peroxidase activity was observed with o-dianisidine, NADH, NADPH, or glutathione.
Kinetic analysis
Kinetic analysis of CatGt was carried out under optimal conditions (at 60 °C and pH 7.0). Initial velocities against various concentrations of hydrogen peroxide were plotted. The reactions followed the Michaelis–Menten kinetics (Fig. 7a). CatGt displayed apparent Km and Vmax values of 260 mM and 18,521 μmol min−1 mg−1, respectively. A kcat value of 10,360 s−1 subunit−1 was calculated from the data.
a Michaelis–Menten plot of initial reaction velocities against various concentrations of hydrogen peroxide to calculate the apparent Km and Vmax values. b Arrhenius plot (ln k versus reciprocal absolute temperature) of the reaction catalyzed by CatGt. c Eyring–Polanyi plot. ln (k/T) versus reciprocal absolute temperature of the reaction catalyzed by CatGt
Arrhenius plot was drawn to calculate the activation energy for the reaction catalyzed by CatGt (Fig. 7b). The activation energy was found 8 ± 0.5 kJ mol−1. Similarly, Eyring–Polanyi plot was used to calculate activation enthalpy and entropy of the reaction (Fig. 7c). These values were 15 ± 0.5 kJ mol−1 and − 158 ± 3 J mol−1 K−1, respectively.
Discussion
We have described here a catalase from G. thermopakistaniensis and produced substantial evidence that it is a manganese catalase because it contains manganese-binding residues, it is tenfold activated by Mn2+, Mn2+ are found in the protein, it is resistant to inhibition by sodium azide, and does not show a Soret band in the absorption spectrum. Three domains, including an N-terminal domain, a central four-helix bundle domain, and a C-terminal domain, have been identified in manganese catalases (Antonyuk et al. 2000; Barynin et al. 2001). The central domain contains the amino acids involved in coordination with manganese atoms, these amino acids are conserved in CatGt and all other characterized manganese catalases (Fig. 1). However, the amino acid residue considered to be involved in catalytic proton transfer is either Lys (Lys158) (Antonyuk et al. 2000) or Glu (Glu174) (Barynin et al. 2001). In CatGt it is Lys154 similar to the counterpart from Thermus sp. YS 8–13 (Kagawa et al. 1999) and in contrast to Glu161 and Glu174 in the counterparts from Geobacillus sp. WCH70 (Li et al. 2017) and L. plantarum, respectively (Barynin et al. 2001) (Fig. 1).
Previously characterized manganese catalases are either homotetrameric or homohexameric proteins. The molecular weight of one subunit of CatGt, estimated from SDS-PAGE, was ~ 33 kDa (Fig. 1) whereas the molecular weight determined by gel filtration chromatography was ~ 135 kDa. This indicated that CatGt is a homotetrameric protein. The manganese catalase from T. roseum which has the highest identity with CatGt is also a homotetrameric protein (Baginski and Sommerhalter 2017).
Previously reported crystal structures of manganese catalases indicated the presence of two manganese atoms per subunit of the protein (Antonyuk et al. 2000; Barynin et al. 2001). In this study, we could detect 1.22 ± 0.02 manganese atoms per subunit of CatGt sample refolded without addition of 2 mM Mn2+ and 1.51 ± 0.04 atoms per subunit in CatGt refolded in the presence of 2 mM Mn2+. These results are comparable with manganese contents of previously reported counterparts from Lactobacillus plantarum (1.12 ± 0.37) (Kono and Fridovich1983; Barynin et al. 2001), Thermoleophilum album (1.4 ± 0.4) (Allgood and Perry 1986), Thermus sp. YS 8–13 (1.2) (Kagawa et al. 1999), Pyrobaculum calidifontis (1.32 ± 0.03) (Amo et al. 2002), Thermomicrobium roseum (1.31 μg manganese per mg protein) (Baginski and Sommerhalter 2017), Anabaena sp. PCC7120 (1.8 ± 0.03) (Bihani et al. 2016) and Geobacillus sp. WCH70 (1.42 ± 0.08) (Li et al. 2017). It can, therefore, be assumed that CatGt also contains two manganese atoms per subunit.
A novel feature of CatGt is its high thermostability. The amino acid composition of a protein contributes significantly to the thermostability (Baldwin 2007; Pace 2009). When we analyzed the amino acid composition of CatGt, we found that the number of Ala residues (9.7%), the best α-helix former, was highest. The content of heat-sensitive amino acids such as Gln (3.3%), Asn (3.7%) and Cys (0.7%) was low. These residues are usually in low percentage in thermostable enzymes (Hensel 1993; Muir et al. 1995; Russell and Taylor 1995). When compared the thermostability of characterized catalases, we found CatGt more thermostable than its counterparts (Table 1). CatGt exhibited a half-life of 60 min at 100 °C while the nearest counterpart, in terms of thermostability, was a manganese catalase from the hypethermophilic archaeon P. calidifontis which showed a half-life of 50 min at this temperature (Amo et al. 2002). We, therefore, compared the amino acid contents and found that the enzyme from P. calidifontis contains 26 Ala residues compared to 29 in CatGt. Furthermore, the proportion of charged residues, Asp, Glu, Lys and Arg, is reported to be higher in thermostable enzymes (Arif et al. 2018). Among these residues, Glu tends to form multiple ion pairs and hydrogen bonds which makes the structure rigid and hence thermostable. High fraction (24.8%) of these residues was found in CatGt compared to 23.5% in the enzyme from P. calidifontis. The thermostability of a protein is correlated to the ratio of preferred (Glu and Lys) and avoided (Gln, Cys and His) amino acids (Farias and Bonato 2003). The percentage of Glu and Lys in CatGt is 12.8 while that of Gln, Cys and His is 6.3. A high ratio of charged amino acids contributes to stabilization of the exposed regions of a protein fold by forming additional electrostatic interactions (Bagautdinov and Yutani 2011). High thermostability of CatGt may partly be due to the lower number of thermolabile and higher number of charged amino acids. Among other factors, abundance of Ala and the tetrameric nature of CatGt may contribute to its thermostability.
In conclusion, CatGt was produced in inactive and insoluble form in E. coli, which was refolded into soluble and active form. The refolded protein is highly active and thermostable, probably the most thermostable catalase characterized to date. These features make it a promising candidate for use in textile and paper industries. Further studies are needed for its applications in industry for the removal of residual hydrogen peroxide in products or by-products.
References
Adams MW, Kelly RM (1998) Finding and using hyperthermophilic enzymes. Trends Biotechnol 16:329–332. https://doi.org/10.1016/S0167-7799(98)01193-7
Akyilmaz E, Kozgus O (2009) Determination of calcium in milk and water samples by using catalase enzyme electrode. Food Chem 115:347–351. https://doi.org/10.1016/j.foodchem.2008.11.075
Allgood GS, Perry JJ (1986) Characterization of a manganese containing catalase from the obligate thermophile Thermoleophileum album. J Bacteriol 168:563–567. https://doi.org/10.1128/jb.168.2.563-567.1986
Amo T, Atomi H, Imanaka T (2002) Unique presence of a manganese catalase in a hyperthermophilic archaeon, Pyrobaculum calidifontis VA1. J Bacteriol 184:3305–3312. https://doi.org/10.1128/JB.184.12.3305-3312
Antonyuk SV, Melik-Adamyan WR, Popov AN, Lamzin VS, Hempstead PD, Harrison PM, Artymiuk PJ, Barynin SM (2000) Three-dimensional structure of the enzyme dimanganese catalase from Thermus thermophilus at 1 A° resolution. Crystallogr Rep 45:105–116. https://doi.org/10.1134/1.171145
Arif M, Rashid N, Perveen S, Bashir Q, Akhtar M (2018) Extremely stable indole-3-glycerol-phosphate synthase from hyperthermophilic archaeon Pyrococcus furiosus. Extremophiles 23:69–77. https://doi.org/10.1007/s00792-018-1061-4
Bagautdinov B, Yutani K (2011) Structure of indole-3-glycerol phosphate synthase from Thermus thermophilus HB8: implications for thermal stability. Acta Crystallogr Sect D D67:1054–1064. https://doi.org/10.1107/S0907444911045264
Baginski R, Sommerhalter M (2017) A manganese catalase from Thermomicrobium roseum with peroxidase and catecholase activity. Extremophiles 21:201–210. https://doi.org/10.1007/s00792-016-0896-9
Baldwin RL (2007) Energetics of protein folding. J Mol Biol 371:283–301. https://doi.org/10.1016/j.jmb.2007.05.078
Barynin VV, Grebenko AI (1986) T-catalase in nonheme catalase of extremely thermophilic bacterium Thermus thermophilus HB-8. Dolk Akad Nauk USSR 286:461–464. https://doi.org/10.1128/JB.184.12.3305-3312
Barynin VV, Hempstead PD, Vagin AA, Antonyuk SV, Melik- Adamyan WR, Lamzin VS, Harrison PM, Artymiuk PJ (1997) The three-dimensional structure of the di-Mn catalase and the environment of the di-Mn sites in different redox states. J Ind Biochem 67:196. https://doi.org/10.1128/AEM.70.7.3839-3844
Barynin VV, Whittaker MM, Antonyuk SV, Lamzin VS, Harrison PM, Artymiuk PJ, Whittaker JM (2001) Crystal structure of manganese catalase from Lactobacillus plantarum. Structure 9:725–738. https://doi.org/10.1016/S0969-2126(01)00628-1
Beers RF, Sizer W (1952) A spectrophotometric method for measuring the breakdown of hydrogen peroxide by catalase. J Biol Chem 195:133–140. https://www.jbc.org/content/195/1/133.long. Accessed 11 June 2018
Bihani SC, Chakravarty D, Ballal A (2016) KatB, a cyanobacterial Mn-catalase with unique active site configuration: Implications for enzyme function. Free Radic Biol Med 93:118–129. https://doi.org/10.1016/j.freeradbiomed.2016.01.022
Chelikani P, Fita I, Chelikani P, Fita I, Loewen PC (2004) Diversity of structures and properties among catalases. Cell Mol Life Sci 61:192–208. https://doi.org/10.1007/s00018-003-3206-5
Chohan SM, Rashid N (2013) TK1656, a thermostable l-asparaginase from Thermococcus kodakaraensis, exhibiting highest ever reported enzyme activity. J Biosci Bioeng 116:438–443. https://doi.org/10.1016/j.jbiosc.2013.04.005
Cook JN, Worsley JL (1996) Compositions and method for destroying hydrogen peroxide on contact lens. US Patent No. 5,521,091. https://patents.google.com/patent/US5521091. Accessed 26 June 2018
Coolbear TR, Daniel M, Morgan HW (1992) The enzymes from extreme thermophiles: bacterial sources, thermostabilities and industrial relevance. Adv Biochem Eng Biotechnol 45:58–98. https://doi.org/10.1007/bfb0008756
Demirjian DC, Morís-Varas F, Cassidy CS (2001) Enzymes from extremophiles. Curr Opin Chem Biol. 5:144–151. https://doi.org/10.1016/S1367-5931(00)00183-6
Ebara S, Shigemori Y (2008) Alkali-tolerant high-activity catalase from a thermophilic bacterium and its overexpression in Escherichia coli. Protein Expr Purif 57:255–260. https://doi.org/10.1016/j.pep.2007.09.015
Fang FC (2004) Antimicrobial reactive oxygen and nitrogen species: concepts and controversies. Nat Rev Microbiol 2:820–832. https://doi.org/10.1038/nrmicro1004
Farias ST, Bonato MC (2003) Preferred amino acids and thermostability. Genet Mol Res 2:383–393. https://www.funpecrp.com.br/gmr/year2003/vol4-2/pdf/gmr0079.pdf. Accessed 22 Apr 2019
Gharib G, Rashid N, Bashir Q, Gardner QA, Muhammad Akhtar M, Imanaka T (2016) Pcal_1699, an extremely thermostable malate dehydrogenase from hyperthermophilic archaeon Pyrobaculum calidifontis. Extremophiles 20:57–67. https://doi.org/10.1007/s00792-015-0797-3
Gonzalez-FlechaFlecha B, Demple B (1995) Metabolic sources of hydrogen peroxide in aerobically growing Escherichia coli. J Biol Chem 270:13681–13687. https://doi.org/10.1074/jbc.270.23.13681
Hensel R (1993) Proteins of extreme thermophiles. In: Kates M, Kushner DJ, Matheson AT (eds) The biochemistry of the Archaea. Elsevier, Amsterdam, pp 209–221https://doi.org/10.1016/0307-4412(95)90682-7
Hidalgo A, Betancor L, Moreno R, Zafra O, Cava F, Fernández-Lafuente R, Guisán JM, Berenguer J (2004) Thermus thermophilus as a cell factory for the production of a thermophilic Mn-dependent catalase which fails to be synthesized in an active form in Escherichia coli. Appl Environ 70:3839–3844. https://doi.org/10.1128/AEM.70.7.3839-3844.2004
Hildebrandt AG, Roots I (1975) Reduced nicotinamide adenine dinucleotide phosphate (NADPH)-dependent formation and breakdown of hydrogen peroxide during mixed function oxidation reactions in liver microsomes. Arch Biochem Biophys 171:385–397. https://doi.org/10.1016/0003-9861(75)90047-8
Kagawa MN, Murakoshi Y, Nishikawa G, Matsumoto Y, Kurata T, Mizobata Y, Kawata Nagai J (1999) Purification and cloning of a thermostable manganese catalase from a thermophilic bacterium. Arch Biochem Biophys 362:346–355. https://doi.org/10.1006/abbi.1998.1041
Klotz MG, Klassen GR, Loewen PC (1997) Phylogenetic relationships among prokaryotic and eukaryotic catalases. Mol Biol Evol 14:951–958. https://doi.org/10.1093/oxfordjournals.molbev.a025838
Klotz MG, Loewen PC (2003) The molecular evolution of catalytic hydroperoxidases: evidence for multiple lateral transfer of genes between prokaryota and from bacteria into eukaryota. Mol Biol Evol 20:1098–1112. https://doi.org/10.1093/molbev/msg129
Kono Y, Fridovich I (1983) Isolation and characterization of the pseudocatalase of Lactobacillus plantarum. J Biol Chem 258:6015–6019
Korshunov S, Imlay JA (2010) Two sources of endogenous hydrogen peroxide in Escherichia coli. Mol Microbiol 75:1389–1401. https://doi.org/10.1111/j.1365-2958.2010.07059.x
Kristjansson JK (1989) Thermophilic organisms as sources of thermostable enzymes. Trends Biotechnol 7:349–353. https://doi.org/10.1016/0167-7799(89)90035-8
Lee SY (1996) High cell density culture of Escherichia coli. Trends Biotechnol 14:98–105. https://doi.org/10.1016/0167-7799(96)80930-9
Li HC, Yu Q, Wang H, Cao XY, Ma L, Li ZQ (2017) A new homo-hexamer Mn containing catalase from Geobacillus sp. WCH70. Catalysts 7:277–289. https://doi.org/10.3390/catal7090277
Loewen PC (2004) Diversity of structures and properties among catalases. Cell Mol Life Sci 61:192–208. https://doi.org/10.1007/s00018-003-3206-5
Loewen PC, Klotz MG, Hassett DJ (2000) Catalase—an “old” enzyme that continues to surprise us. ASM News 66:76–82
Mizobata T, Kagawa M, Murakoshi N, Kusaka E, Kameo K, Kawata Y, Nagai J (2000) Overproduction of Thermus sp. YS 8–13 manganese catalase in Escherichia coli; Production of soluble apoenzyme and in vitro formation of active holoenzyme. Eur J Biochem 267:4264–4271. https://doi.org/10.1046/j.1432-1033.2000.01474.x
Muir JM, Russell RJM, Hough DW, Danson MJ (1995) Citrate synthase from the hyperthermophilic archaeon Pyrococcus furiosus. Protein Eng 8:583–592. https://doi.org/10.1093/protein/8.6.583
Nicholls P (1964) The reaction of azide with catalase and their significance. Biochem J 90:331–343. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1202621/. Accessed 11 Aug 2018
Nicholls P (2012) Classical catalase: ancient and modern. Arch Biochem Biophys 525:95–101
Nicholls P, Fita I, Loewen PC (2001) Enzymology and structure of catalases. Adv Inorg Chem 51:51–106. https://doi.org/10.1016/S0898-8838(00)51001-0
Owusu RK, Cowan DA (1989) Correlation between microbial protein thermostability and resistance to denaturation in aqueous organic solvent two-phase systems. Enzyme Microb Technol 11:568–574. https://doi.org/10.1016/0141-0229(89)90084-7
Pace CN (2009) Energetics of protein hydrogen bonds. Nat Struct Mol Biol 16:681–682. https://doi.org/10.1038/nsmb0709-681
Rajagopalan KV, Fridovich I, Handler P (1961) Competitive inhibition of enzyme activity by urea. J Biol Chem 236:1059–1065. https://www.jbc.org/content/236/4/1059.long. Accessed 2 July 2019
Rasool N, Rashid N, Iftikhar S, Akhtar M (2010) N-terminal deletion of Tk1689, a subtilisin-like serine protease from Thermococcus kodakaraensis, copes with its cytotoxicity in Escherichia coli. J BiosciBioeng 110:381–385. https://doi.org/10.1016/j.jbiosc.2010.04.005
Rosano GL, Ceccarelli EA (2014) Recombinant protein expression in Escherichia coli: advances and challenges. Front Microbiol 5:172. https://doi.org/10.3389/fmicb.2014.00172
Rowe HD (1999) Biotechnology in the textile/clothing industry—a review. J Consumer Stud Home Econ 23:53–61. https://doi.org/10.1046/j.1365-2737.2000.00093.x
Russell RJM, Taylor GL (1995) Engineering thermostability: lessons from thermophilic proteins. Curr Opin Biotechnol 6:370–374. https://doi.org/10.1016/0958-1669(95)80064-6
Sharif A, Ashraf M, Javeed A, Anjum AA, Akhtar MF, Akhtar B, Saleem A (2016) Oxidative stress responses in Wistar rats on subacute exposure to pharmaceutical waste water. Environ Sci Pollut Res 23:24158–24165. https://doi.org/10.1007/s11356-016-7717-7
Siddiqui MA, Rashid N, Ayyampalayam S, Whitman WB (2014) Draft genome sequence of Geobacillus thermopakistaniensis strain MAS1. Genome Announc. https://doi.org/10.1128/genomeA.00559-14
Sørensen HP, Mortensen KK (2005) Soluble expression of recombinant proteins in the cytoplasm of Escherichia coli. Microb Cell Fact 4:1. https://doi.org/10.1186/1475-2859-4-1
Tayyab M, Rashid N, Akhtar M (2011) Isolation and identification of lipase producing thermophilic Geobacillus sp. SBS-4S: cloning and characterization of the lipase. J Biosci Bioeng 111:272–278. https://doi.org/10.1016/j.jbiosc.2010.11.015
Veal EA, Day AM, Morgan BA (2007) Hydrogen peroxide sensing and signaling. Mol Cell 26:1–14. https://doi.org/10.1016/j.molcel.2007.03.016
Whittaker JW (2011) Non-heme manganese catalase—the ‘other’ catalase. Arch Biochem Biophys 525:111–120. https://doi.org/10.1016/j.abb.2011.12.008
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by H. Atomi.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Shaeer, A., Aslam, M. & Rashid, N. A highly stable manganese catalase from Geobacillus thermopakistaniensis: molecular cloning and characterization. Extremophiles 23, 707–718 (2019). https://doi.org/10.1007/s00792-019-01124-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-019-01124-5