Abstract
L-proline catabolism is emerging as a key pathway that is critical to cellular metabolism, growth, survival, and death. Proline dehydrogenase (PRODH) enzyme, which catalyzes the first step of proline catabolism, has diverse functional roles in regulating many pathophysiological processes, including apoptosis, autophagy, cell senescence, and cancer metastasis. Notably, accumulated evidence demonstrated that PRODH plays complex role in many types of cancers. In this review, we briefly introduce the function of PRODH, then its expression in different types of cancer. We next discuss the regulation of PRODH in cancer, the downstream pathways of PRODH and the therapies that are under investigation. Finally, we propose novel insights for future perspectives on the modulation of PRODH.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cancer cells trigger metabolic reprogramming during their initiation and progression in response to the tumor microenvironment stimuli by directly or indirectly activating aberrant growth and survival signals (Pavlova and Thompson 2016; Agnihotri and Zadeh 2016; He et al. 2016; Chen et al. 2019; Yu et al. 2020; Y et al. 2019; He et al. 2019). The high rate of aerobic glycolysis and glutamine utilization are the two most significant features of cancer cell metabolic reprogramming (Byun et al. 2020; Bernfeld and Foster 2019; Lunt and Vander Heiden 2011; Chen et al. 2014). In addition to glutamine, other amino acids (e.g., serine, glycine, alanine, proline, etc.) are consumed in cancer cells for the generation of nucleotides, reactive oxygen, proteins, and oncometabolites (Muhammad et al. 2020). Based on dietary necessity, amino acids can be divided into essential amino acids (EAAs) and non-essential amino acids (NEAAs) (Choi and Coloff 2019). Several recent reports have uncovered the important role of NEAAs in the pathology of cancer (Coloff et al. 2016). Strategies that target NEAAs metabolism are still in clinical trials and therapy. For example, the allosteric inhibitor of glutaminase, compound BPTES (bis-2-(5-phenylacetamido-1,2,4-thiadiazol-2-yl) ethyl sulfide) that blocks glutamine utilization has been reported to play an anti-tumor role in a variety of cancers (Rajeshkumar et al. 2017; Yuneva et al. 2012; Le et al. 2012; Gross et al. 2014; Xiang et al. 2015). Other inhibitors of glutaminase, like CB-839 and compound 968, that block glutaminase activity inhibit the growth of transformed/cancer cells (Gross et al. 2014). Except for blocking glutamine utilization, limiting cellular cysteine could induce a unique cell death program known as ferroptosis and result in an anti-tumor effect(Mao et al. 2018). Inhibitors of the transporter xCT, like sorafenib and sulfasalazine, that block the transport of cystine, an oxidized dimer form of NEAA cysteine, have been already approved by the U.S. Food and Drug Administration (FDA) for tumor treatment (Koppula et al. 2020; Lei et al. 2020; Lo et al. 2008).
As one of the NEAAs, proline is the sole proteinogenic secondary amino acid that endows several functions not possible with other amino acids (Liu et al. 2020). Hence, proline is essential for collagen synthesis to support collagen physical stability (Shoulders and Raines 2009). Proline can also act as a ‘helix breaker’, as it contributes to the 3D structure of proteins by introducing a kink to disrupt the α-helix conformation (Cordes et al. 2002; Williams and Deber 1991; Burke et al. 2020). It is well established that proline plays a critical role in molecular recognition, protein stability, signaling transduction and cell redox reactions (Phang et al. 2010). Because proline has α-amino nitrogen within a pyrrolidine ring, proline has its own enzyme family that is distinct from most amino acids.
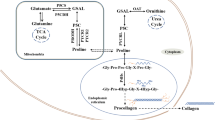
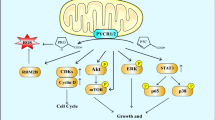
Proline dehydrogenase (PRODH), also known as proline oxidase (POX), is a mitochondrial inner-membrane protein that catalyzes proline to produce pyrroline-5-carboxylate (P5C) in the first step of proline catabolism. In this reaction, generated electrons transferred to the mitochondrial electron transport chain (ETC) for ATP or ROS generation, which finally mediates downstream signal pathways and biological processes (Fig. 1). For instance, PRODH generates ATP to promote tumor cell survival in some nutrient stress like hypoxia and glucose depletion (Liu and Phang 2012). In other cases, PRODH also induces apoptosis and autophagy in cancer cells through ROS generation, and functions as a tumor suppressor (Liu et al. 2006, 2009). As the complex and integral role of PRODH in cancer, PRODH has sparked great attention in the proline metabolism research field. Interestingly, recent reports suggested that PRODH may play a promote tumor or anti-tumor role depending on the environment and cell types.
Potential roles of PRODH in cancer cells through different mechanisms. PRODH has been demonstrated regulated by miR-23b*, AMPK, p53 and PPARγ. Inducible PRODH results in ATP/ROS generation via catalyzing L-proline to P5C. Then, the increasing ROS and ATP contribute to a broad range of cell actions that depend on microenvironment
A series of new discoveries have reported noval effects and mechanism of PRODH and proline catabolism in cancer. Hence, we will conclude the previous studies and related discoveries during the last few years. The principal focus of this review is the specific relationship between proline catabolism and cancer progression. We will discuss the regulation of PRODH in cancer, the downstream pathways of PRODH, and therapies that are under investigation. Finally, we will explore the possible potential mechanism of PRODH function in inhibiting cancer or promoting cancer in some circumstances.
Proline metabolism in cancer
PRODH is identified as one of the p53-induced genes related to apoptosis, which binds to mitochondrial inner membranes and catalyzes the oxidation of proline to P5C. P5C is an intermediate in the metabolic interconversions between the tricarboxylic acid (TCA) cycle and urea cycle (Adams 1970; Phang 2019). P5C converts to glutamic-γ-semialdehyde (GSA) spontaneously and carries out two dehydrogenation reactions for α-KG generation, a critical intermediate of the TCA cycle. In the other reaction pathway, P5C converts to ornithine catalyzed by ornithine aminotransferase and participates in the urea cycle (Phang 2019). The reverse conversion of P5C to proline is proline biosynthesis that contains three isoform P5C reductases (PYCRS), which also play important roles in cancer progression. Existing studies have identified the protumor role of PYCR1 in different cancers, including melanoma (De Ingeniis et al. 2012; Ye et al. 2018), renal cell carcinoma (Weijin et al. 2019), breast cancer (Loayza-Puch et al. 2016; Ding et al. 2017), hepatocellular carcinoma (Zhuang et al. 2019) and colorectal cancer (Yan et al. 2019; Burke et al. 2020). Several reviews conclude the unequivocal role of PYCR1 and proline biosynthesis in cancers based on different mechanisms (Burke et al. 2020; Phang 2019). However, PRODH seems to play complex roles which depend on cancer types and microenvironment. Hence, this review will concentrate on PRODH and proline catabolism in cancer.
PRODH is the enzyme in the first step of proline catabolism, which donates electrons to the electron transport chain for ROS or ATP production contributing to a series of biology reaction including signaling transduction, oxidation reaction, immune-inflammatory reaction, etc. Previous studies have established a series of observations focus on PRODH-mediate proline metabolism on several cancers (Phang 2019; Liu et al. 2008, 2005, 2010). Recent reports have discovered novel aspects of PRODH in different cancer types (Burke et al. 2020; Cappelletti et al. 2018; Fang et al. 2019); it is necessary to sum up the roles and mechanisms related to PRODH.
Regulatory mechanisms of PRODH
Transcriptional regulation of PRODH by transcriptional factors
To ensure appropriate functioning of PRODH in cancer cell regulation, the expression and activity of PRODH are subjected to a variety of pathways regulation. PRODH was first identified as a p53-induced gene-6 (PIG6) in a model for apoptosis(Polyak et al. 1997), and this discovery opened a new direction with a surge of reports related to the proline metabolism in subsequent studies. Since this discovery, the p53 response elements (REs) in the promoter and introns of PRODH were identified (Maxwell and Kochevar 2008; Raimondi et al. 2013; Liu et al. 2020). Interestingly, one of the p53 Res, located in PRODH introns, is efficiently transactivated by p53 members, p63 and p73 (Raimondi et al. 2013). The transcriptional activation of PRODH by p53 also has been demonstrated in other independent investigations (Nagano et al. 2016; Donald et al. 2001). Mutant p53 has been shown to reduce mRNA expression of PRODH in renal cancer (Maxwell and Rivera 2003). Similarly, mutant p53 reduced PRODH expression in colon cancer cell lines compared with wild-type p53 (Liu et al. 2020). However, p53 may not be the only factor that transactivates PRODH expression; other factors may also result in the variation of expression and activity of PRODH. The characteristics of PRODH finally showed in cancer cells mainly depended on the roles of predominant factors which may counteract the role of others. This transformation might account for discrepancies of PRODH in different cancer types.
PRODH expression can also be promoted by peroxisome proliferator activated receptor gamma (PPARγ), a ligand-dependent transcription factor that belongs to the nuclear hormone receptor superfamily (Pandhare et al. 2006). PPARγ can play a role in controlling the expression of networks of genes related to inflammation, lipid metabolism, adipogenesis, and metabolic homeostasis through forming obligate heterodimers with retinoid X receptor (RXR) and binding to specific response elements in the promoter regions of target genes (Ahmadian et al. 2013). PRODH promoter was found to be activated by PPARγ ligand troglitazone in colon cancer cells with both PPARγ-dependent and independent mechanisms (Pandhare et al. 2006). The oxidized low-density lipoprotein (oxLDL) is a potential factor that increased cancer risk(Tian et al. 2019). oxLDL could markedly increase PRODH expression based on one of its components, 7-ketocholesterol, through PPARγ in several cancer cells(Zabirnyk et al. 2010).
Posttranscriptional regulation and post-translational modification of PRODH
Apart from transcriptional regulation of PRODH as discussed above, PRODH expression can also be regulated at mRNA levels by microRNAs. MicroRNAs were first discovered in 1993 (Lee et al. 1993), and many studies have identified microRNAs roles in cancer biology for patients’ prognosis, disease classification, and clinical treatment trials (Rupaimoole and Slack 2017; Bertoli et al. 2015). MicroRNAs negatively regulate mRNA level through binding to complementary sequences in the 3′ untranslated region (UTR) of target mRNAs (Krol et al. 2010; Hayes et al. 2014). PRODH expression was found to be suppressed by miR-23b* in renal cancer (Liu et al. 2010). Moreover, oncogene Myc suppresses PRODH expression indirectly through miR-23b*-mediated pathway (Liu et al. 2012b). However, there are no more studies related to posttranscriptional regulation of PRODH in cancer. Considering the multi-functions of microRNAs in regulating cancer biology, further investigations about searching for potential microRNA target to PRODH is worthy.
To date, accumulating evidence demonstrate that PRODH is specifically regulated by several transcriptional factors like tp53 and PPARγ. There are few studies focus on PRODH related post-translational modification, including ubiquitination, phosphorylation, acetylation, methylation, etc. An epigenome-wide gene–age interaction analysis reveals that the elderly LUAD patients have better survival with lower methylation of PRODH (Chen et al. 2020). However, no more related studies and experiments reports the regulation of PRODH by post-translational modification. The underlying mechanisms of its enzyme activity and protein stability may be the key diver cause complex roles of PRODH on specific cancer. The post-translational regulation has been proved to contribute to tumor metabolism regulation, immunological tumor microenvironment (TME) modulation and cancer stem cell (CSC) stemness maintenance (Telerman and Amson 2009; Deng et al. 2020). Hence, the post-translational modification of PRODH, which mediates the “quantity” and “quality” of PRODH, needs further investigations.
PRODH expression and cancer
Since PRODH was first identified as a p53-induced gene-6 (PIG6) in 1997(Polyak et al. 1997), there have been several reports demonstrating its down expression in various cancers and multiple effects on cancer cell cycle arrest, cell senescence, apoptosis, and autophagy. Subsequently, some findings have reported that PRODH is up-regulated in some challenging circumstances and contributes to cancer progression through influencing tumor growth, EMT, metastasis, and T cell infiltration (Table1).
PRODH expression is markedly down-regulated compared to the corresponding normal tissues in renal and digestive tract tumors, including colorectal, rectum, stomach, and liver (Liu et al. 2009). Indeed, subsequent reports confirmed the ability of PRODH to trigger apoptosis through both intrinsic and extrinsic apoptotic pathway, and demonstrated that PRODH functions as a tumor suppressor in a variety of cancer cell type such as colorectal, renal, and tongue squamous cell cancer (Maxwell and Rivera 2003; Liu et al. 2005, 2009; Celińska-Janowicz et al. 2018). Several reports have proved this characteristic of PRODH independently (Hu et al. 2007; Liu et al. 2008; Toloczko-Iwaniuk et al. 2020). Intriguingly, PRODH activates apoptosis through multiple mechanisms, most of which are mediated by ROS production. By this mitochondrial (intrinsic) apoptotic pathway, cytochrome c is released from mitochondria into cytosol and results in the activation of caspase-9 and caspase-3. What is more, PRODH was shown to activate the extrinsic apoptotic pathway via increasing the expression of NFATc1, a transcription factor, and promoting its localization to the nucleus, which results in the induction of the death receptor TRAIL (extrinsic) and activation of caspase-8 (Liu et al. 2006). In addition to inducing apoptosis, PRODH also plays a role in arresting DLD-1 cells in G2 phase via upregulating the expression of GADDs, a gene that affects growth arrest and DNA damage (Liu et al. 2009). However, the potential mechanism of PRODH upregulating GADDs to block cell cycle was not carried out for further research.
Autophagy, a multistep lysosomal-mediated pathway that eliminates damaged organelles and invading pathogens to support nutrient recycling, is intimately linked to a cell's live/die decision(Amaravadi et al. 2019; Zhang et al. 2019). Follow-up experiments showed that PRODH was involved in autophagy and promoted cells survival when cells counteract nutrient deprivation or hypoxia. PRODH has been identified as an important regulator in oxLDL-mediated prosurvival autophagy through the generation of superoxide and subsequent up-regulation of beclin-1(Zabirnyk et al. 2010). PRODH also acts as an energy source for providing ATP under nutrient stress conditions. The expression of PRODH is up-regulated in various cancer cells including colon, breast, prostate, melanoma, lung, and ovarian under hypoxia tumor microenvironment which is mainly dependent on AMPK activation (Liu et al. 2012a). Similarly, glucose deprivation increased PRODH expression and promoted PRODH catalytic activity for maintenance of ATP levels that is AMPK dependent (Pandhare et al. 2009).
Although PRODH is down-expressed in renal and digestive tumor samples, PRODH is up-regulated in non-small cell lung cancer (NSCLC), pancreatic ductal adenocarcinoma (PDAC), prostate cancer (PCa), and some breast invasive carcinoma (BRCA) subtypes (Table 1). This up-regulation of PRODH in tumor tissues promotes cell growth, survival or metastasis through another mechanism. PRODH has been reported to induce IKKα activity through generation of ROS and results in the up-regulation of related inflammatory genes (Liu et al. 2020). PRODH has also been shown to promote NSCLC cell growth, migration and invasion (Liu et al. 2020). An epigenome-wide gene-age interaction analysis has revealed the reversed role of PRODH on survival in different age stage of lung cancer patients (Chen et al. 2020). Young patients with high methylation of PRODH was the best survival group, and low methylation of PRODH takes as an increased protective effect on LUAD patients survival with increased age (Chen et al. 2020). Interestingly, significant correlation was observed between the specific CpG probe and expression of PRODH in LUAD patients (r = -0.23, p = 3.38*10–5). This significant heterogeneity of PRODH methylation effect on different age stage may contribute to providing new evidence for researching the switch roles of PRODH in context and searching for new specific biomarkers and therapeutic strategies for improving prediction accuracy and treatment efficacy.
PRODH has also been shown to contribute to promoting breast cancer by supporting growth and metastasis of breast cancer cells (Fang et al. 2019). PRODH activity, dependent on P5C recycling via PYCR1, was shown to support spheroidal growth by sustaining ATP production. What is more, PRODH is significantly higher expressed in metastases compared to primary breast cancers in patients. Strikingly, the number of metastases was significantly reduced once PRODH activity was inhibited by L-THFA, whereas the weight of primary breast tumors remained unchanged. These data indicated that PRODH seems to be more specifically important in metastatic growth compared to primary growth (Elia et al. 2017). In accordance, a differential analysis mentioned PRODH as one of the significantly up-regulated genes in the multifocal/multicentric breast cancer ( MMBC) patients between invasive MMBC and unifocal breast cancer (UFBC). However, this study did not present any actual data for PRODH expression (Lang et al. 2018). These data indicate that PRODH activity seems to be important in supporting metastasis in this specific site or organ (Fang et al. 2019).
A role for PRODH in promoting prostate cancer has also been reported (Yan et al. 2018), which mainly focuses its role on inhibiting T cell infiltration. High levels of P5C, a metabolite converted from proline by PRODH, have shown to be released by prostate cancer cells and result in T cells signaling suppression by increasing ROS but decreasing cytokines and ATP production. Moreover, these aforementioned phenotypes were reversed with PRODH knockdown. Similarly, the up-regulation of PRODH increased tumor growth in animal model and decreased CD3+, CD4+ and CD8+ T cells infiltration. The expression of PRODH was up-regulated in human prostate cancer tissues compared with its corresponding non-neoplastic tissue. Further, the expression of PRODH was higher in the advanced tumors among the different stages of PCa. Collectively, this study provides a novel perspective of PRODH on impairing immune cell functions through promoting the release of P5C from prostate cancer and finally creating a microenvironment that improves cancer cell survival. In the other hand, this study provides a new standpoint of PRODH for tumor immunotherapy.
Given that PRODH is closely related to the pathological processes in multiple types of cancers by serving as an anti-tumor or protumor member, mechanistic insights into how PRODH converts its dual role on specific context and cancer types could be valuable for its translational implications in clinical settings, such as the development of precision medicine on gene-oriented treatment.
The role of PRODH and L-proline catabolism in cancer
The physiological activity of PRODH is mainly involved in regulating redox statues, inflammatory reaction, intercellular signaling and cell death fate. As aforementioned, PRODH is up-regulated in some challenging circumstances, such as hypoxia or glucose restriction, or some specific type of cancer; whereas, PRODH is down-regulated in several tumors and plays an anti-tumor role through different signaling pathway. In this part, we will summarize the above specific functions of PRODH and the related mechanism in cancer.
PRODH functions in tumor process through ROS-mediated mechanism
ROS is identified as a group of molecular oxygen in different patterns, which are formed by a series of reduction–oxidation reactions and the electron transport chain (Sabharwal and Schumacker 2014). H2O2 and O2− are the most key terms of ROS generated by the ETC and various enzymes, including NADPH oxidases, pyruvate dehydrogenase, acyl-CoA dehydrogenase, proline oxidase and et al. (Prasad et al. 2017; Srinivas et al. 2019). With the variation of ROS intracellular concentration stimulated by various stressors or metabolic enzymes, ROS seems to play in a beneficial or deleterious role by various mechanisms(Sies and Jones 2020). The low concentration of ROS is associated with some cellular responses like proliferation, migration and differentiation. The high concentration ROS exposure leads to inflammation, metastasis, growth arrest and cell death (Sies and Jones 2020). Because mitochondria are the major source of ROS (Sies and Jones 2020; Sabharwal and Schumacker 2014), PRODH, an enzyme that located in mitochondrial inner membrane, which donates an electron to the ETC for generating ROS via oxidizing proline to P5C, may contribute to triggering redox signaling under normal condition or the initiation of cancer under pathophysiological conditions.
As mentioned above, multiple experiments independently revealed that PRODH activates apoptosis through ROS generation both in intrinsic and extrinsic pathways in a variety of cancer types (Fig. 1) (Celińska-Janowicz et al. 2018; Liu et al. 2005, 2006; Maxwell and Rivera 2003). Previous works have demonstrated that PRODH activates apoptosis through both p53-dependent and p53-independent pathways (Rivera and Maxwell 2005; Maxwell and Davis 2000). Although PRODH was not up-regulated in DECV, a derivative cell line of ECV-304 cell that is resistant to p53-mediated apoptosis, apoptosis was induced in both cell lines for P5C production, which indicates that PRODH is capable of inducing apoptosis in a p53-independent pathway (Maxwell and Davis 2000). Interestingly, this study revealed the contribution of P5C but not ROS in inducing apoptosis. The potential role and mechanism of P5C for apoptosis still need further elucidation.
Other reports demonstrated the role of ROS for serving as an intracellular second messenger in signaling cascades and regulating gene expression through stimulating signal transduction and protein phosphorylation (Chio and Tuveson 2017). PRODH activates nuclear factor of activated T cells (NFAT), an indicator of activated calcineurin, through ROS production, and induces cytochrome c release from the mitochondria into the cytoplasm which finally results in apoptosis (Rivera and Maxwell 2005). Another study based on this finding has discovered that PRODH also stimulates the expression of tumor necrosis factor-related apoptosis inducing ligand (TRAIL) mediated by NFAT. Additionally, PRODH was involved in blocking MAPK signaling through reducing the phosphorylation of ERK, JNK and p38 (Liu et al. 2006). MnSOD, an antioxidant enzyme that defense against oxidative stress, reverses this reduction of MAPK signaling and inhibits PRODH-induced apoptosis (Liu et al. 2006). However, the detailed mechanism by which PRODH blocks the phosphorylation and the interaction of these pathways that mediates PRODH-induced apoptosis still need further investigation.
In addition to NFAT and TRAIL, COX-2/PGE2, EGFR and β-catenin signaling have shown to play important roles in PRODH-induced apoptosis. COX-2/PGE2 pathway contributes to metastatic spread, tumor development and maintenance (Greenhough et al. 2009; Echizen et al. 2018; Luo and Zhang 2017). PRODH suppresses COX-2/PGE2, EGFR and β-catenin/APC activities, and this suppression was reversed by MnSOD, indicating that ROS/superoxides generated by PRODH was involved in this process (Liu et al. 2008). Similarly, phosphorylation of EGFR and COX-2 was reduced with PRODH addition. Furthermore, this study suggested that celecoxib, a COX-2 inhibitor, could increase the expression of PRODH and induce apoptosis in oral squamous cell carcinoma (Toloczko-Iwaniuk et al. 2020).
Accumulated evidence suggest that ROS contributes to the induction and maintenance of cellular senescence through diverse pathways including mitochondrial DNA damage, signaling pathways and induction of microRNAs (Davalli et al. 2016). PRODH has also been identified as a senescence-specific gene induced by low dose of etoposide treatment (Nagano et al. 2016). The following study explored PRODH functions in senescence process; they revealed that PRODH promotes senescence and DNA damage via ROS production. And this promotion effect could be impaired by N-acetyl-L-cysteine (NAC), a potent ROS scavenger, which indicates that ROS may be involved in PRODH-mediated senescence. Moreover, etoposide-induced senescence and ROS production were suppressed by L-tetrahydro-2-furoic acid (L-THFA), an inhibitor for PRODH enzymic activity, which indicates that PRODH induces senescence through its enzymic activity (Nagano et al. 2017).
Autophagy (Nazio et al. 2019) is a process that promotes cell survival in response to multiple stimuli like viral infection, nutrient deprivation and genotoxic stress. ROS has been demonstrated to be involved in sustaining autophagy for its signal transduction role (Filomeni et al. 2015). Contrary to the anti-tumor roles described above, PRODH also serves as a survivor in some stress circumstance through ROS. For instance, PRODH is induced by AMPK pathway to initiate protective autophagy under hypoxia via generating ROS. However, PRODH switches its way from generating ROS to ATP under low glucose for cell energy to promote cell survival (Liu and Phang 2012). The potential mechanism of this switch needs more further investigation.
In addition, increased ROS with PRODH promotes phosphorylation of IKKα, which results in the up-regulation of several inflammatory genes in NSCLC cells (Liu et al. 2020). As previously described, ROS production by PRODH is intracellular and affects phosphorylation level of MAPK and COX-2. These findings indicate PRODH-mediated ROS may function as a signaling molecule that contributes to activating multiple critical elements of pathway. PRODH-mediated ROS generation whether inhibits or supports tumor progression depends on certain circumstance and cancer types; therefore, PRODH takes a complicated and flexible role in cancer.
PRODH functions in the tumor process through ATP-mediated mechanism
Proline catabolism via PRODH switches to support ATP production in several condition. Under nutrient stress conditions, especially with glucose deprivation, proline functions as a stress substrate accompanied with increased PRODH enzyme activity for ATP production to maintain cellular energy levels (Pandhare et al. 2009). Glucose depletion induced phosphorylation of AMPK, which indirectly induced PRODH activity through inhibiting mTOR (Pandhare et al. 2009). Consistently, PRODH activity was induced in a time- and dose-dependent tendency with a synthetic AMPK activator—5-aminoimidazole-4-carboxamide ribonucleoside (AICAR), addition (Pandhare et al. 2009). In addition to response to nutrient stress conditions, PRODH contributes to ATP production for supporting energy during spheroidal growth (Elia et al. 2017). Consistently, inhibition of complex III of ETC by antimycin A impaired spheroidal growth, which indicates the critical role of ATP in sustaining spheroidal growth (Elia et al. 2017).
However, additional related reports focus on PRODH-ATP axis functions and triggering mechanisms on cancer progression still need to be further studied.
PRODH regulates tumor process through pyrroline-5-carboxylate generation
P5C is conversed from proline in the first step of proline catabolism, and can be converted into glutamate catalyzed by the P5CDH enzyme (Yan et al. 2018). Glutamate-derived α-Ketoglutarate (α-KG) is a key intermediate of the TCA cycle. Hence, PRODH may take regulatory functions for tumor processes by contributing to P5C-glutamate-α-KG generation. In yeast and plants, P5C seems to cause ROS accumulation, stress response, and cell death (Zhu 2002; Zareba and Palka 2016; Borsani et al. 2005). However, P5C regulates these tumor processes directly or indirectly through multiple mechanisms. P5C plays a harmful role on T cells by inhibiting proliferation and function via inducing SHP1 expression in prostate cancer cells (Yan et al. 2018). Interestingly, P5C recycling to proline via PYCRs sustains PRODH activity during spheroidal growth, which contributes to energy production and metastasis formation in breast cancer cells (Elia et al. 2017). Some studies also found that PRODH generated α-KG by P5C conversion and increased α-KG led to HIF-1 signaling suppression (Verma 2006; Koivunen et al. 2007). Moreover, α-KG decreased the expression of HIF1α and Wnt/β-catenin target genes significantly through enhancing α-KG-mediated degradation of HIF1α in colon cancer (Wen et al. 2019). On the other hand, α-KG has been shown to directly bind to IKKβ and nuclear factor κB (NF-κB) signaling, which results in the increasing uptake of glucose and tumor cell survival by upregulating GLUT1. This finding reveals a critical role of α-KG-mediated signal pathway in brain tumor, and also provides a potential interpretation for the dual role of PRODH in some context (Wang et al. 2019). α-KG is also a cofactor of the KDM5, which can actively remove lysine trimethylation of H3K4me3. Hence, α-KG could enhance KDM5 activity for H3K4 demethylation. Given that α-KG plays an important role on histone modifications, the study that connects proline catabolism to histone modifications may be a novel area for investigation (Su et al. 2021).
Therapeutic strategies targeting PRODH in cancer
So far, various anticancer chemicals and molecules approach to activate or inhibit PRODH activity have been explored, including succinate, MnSOD et al.(Table 2). It is challenging to decide appropriate treatments targeting PRODH. Thus, summarizing the role and mechanism of PRODH and the potential therapeutic drug may provide alternatives to step out dilemma.
It is conceivable that PRODH could be indirectly regulated by compounds targeting upstream pathways. One study showed that etoposide activates p53 and promotes p53-mediated induction of PRODH, causing apoptosis and cell senescence (Maxwell and Davis 2000). Additionally, TZDs treatment markedly increased PRODH activity and protein through promoting the binding of PPARγ to the PRODH promoter, leading to PRODH-mediated ROS generation and apoptosis (Pandhare et al. 2006; Kim et al. 2007).
Moreover, compounds targeting PRODH activity directly have shown anticancer pharmacological properties. L-lactate is a reversible competitive inhibitor of PRODH in several bacterial (Kowaloff et al. 1977). L-THFA, a compound reported to inhibit PRODH activity in several studies, has also been shown to reduce spheroidal growth. Moreover, L-THFA treatment on mice significantly inhibits metastasis formation via impairing PRODH activity without any obvious adverse effects in normal cells and organ functions (Elia et al. 2017). Interestingly, inhibition of PRODH activity by L-THFA impairs cell migration and invasion formation in non-small lung cancer cells (Liu et al. 2020). L-THFA has also been reported to be a competitive inhibitor of PRODH in bacterial (Lee et al. 2003) and other mammalian cells. These data suggest that PRODH may function as a promising drug target in specific cancer types.
Other molecules targeting PRODH downstream signal have also been reported. N-acetyl cysteine, a ROS scavenger that is widely used as a pharmacological antioxidant (Ezeriņa et al. 2018), reduced ROS level generated by PRODH (Yan et al. 2018; Hancock et al. 2016), blocking apoptosis or reversing T cell cytokines secretion (Yan et al. 2018). MnSOD has also been found to inhibit PRODH-mediated apoptosis through reducing the release of cytochrome c (Liu et al. 2005).
As PRODH plays a complex and important role in tumor process, the detailed therapeutic schedules targeting PRODH are subjected to future investigation.
Conclusions and future perspectives
Recent discoveries have solidified the importance of PRODH and proline catabolism in cancer process. PRODH promotes apoptosis and tumor suppression in several cancer cells such as renal and colorectal cancer cells. The expression of PRODH is often down-regulated in these tumors, limiting PRODH-mediated apoptosis and anti-tumor roles. However, PRODH also functions as an oncogenic protein to support tumor cells survival, growth and metastasis in other contexts. The expression of PRODH are up-regulated in these cancer cells like PCa and PDAC. Hence, searching for clinical pharmacological compounds that target the proline pathway is meaningful for tumor-targeted therapy in different types of tumor.
Although pioneering studies of PRODH have advanced our understanding of its fundamental functions and roles in pathological process, studies of post-translational modifications of PRODH are still in blank area. Hence, further studies should focus on investigating post-translational modifications that controls PRODH expression and biological functions in pathophysiological conditions.
In summary, PRODH is involved in the pathogenesis of various cancers. The upstream regulators, downstream signal pathway, biological function, and potential targeted drugs of PRODH are required for prospective therapeutic approach in the future.
Abbreviations
- PRODH:
-
Proline dehydrogenase
- POX:
-
Proline oxidase
- EAAs:
-
Essential amino acids
- NEAAs:
-
Non-essential amino acids
- TNBC:
-
Triple-negative breast cancer
- P5C:
-
Pyrroline-5-carboxylate
- ETC:
-
Electron transport chain
- TCA:
-
Tricarboxylic acid cycle
- PYCRS:
-
P5C reductase
- PIG6:
-
P53-induced gene-6
- PPARγ:
-
Peroxisome proliferator activated receptor gamma
- TZDs:
-
Thiazolidinediones
- oxLDL:
-
Oxidized low-density lipoprotein
- TME:
-
Tumor microenvironment
- CSC:
-
Cancer stem cell
- EMT:
-
Epithelial–mesenchymal transition
- NSCLC:
-
Non-small cell lung cancer
- PDAC:
-
Pancreatic ductal adenocarcinoma
- PCa:
-
Prostate cancer
- BRCA:
-
Breast invasive carcinoma
- LUAD:
-
Lung adenocarcinoma
- MMBC:
-
Multifocal/multicentric breast cancer
- UFBC:
-
Unifocal breast cancer
- NFAT:
-
Nuclear factor of activated T cells
- TRAIL:
-
Tumor necrosis factor-related apoptosis inducing ligand
- NAC:
-
N-acetyl-L-cysteine
- L-THFA:
-
L-tetrahydro-2-furoic acid
- AICAR:
-
5-Aminoimidazole-4-carboxamide ribonucleoside
- NF-κB:
-
Nuclear factor κB
References
Adams E (1970) Metabolism of proline and of hydroxyproline. Int Rev Connect Tissue Res 5:1–91. https://doi.org/10.1016/b978-0-12-363705-5.50007-5
Agnihotri S, Zadeh G (2016) Metabolic reprogramming in glioblastoma: the influence of cancer metabolism on epigenetics and unanswered questions. Neuro Oncol 18(2):160–172. https://doi.org/10.1093/neuonc/nov125
Ahmadian M, Suh JM, Hah N, Liddle C, Atkins AR, Downes M, Evans RM (2013) PPARγ signaling and metabolism: the good, the bad and the future. Nat Med 19(5):557–566. https://doi.org/10.1038/nm.3159
Amaravadi RK, Kimmelman AC, Debnath J (2019) Targeting autophagy in cancer: recent advances and future directions. Cancer Discov 9(9):1167–1181. https://doi.org/10.1158/2159-8290.Cd-19-0292
Bernfeld E, Foster D (2019) Glutamine as an essential amino acid for KRas-driven cancer cells. Trends Endocrinol Metab 30(6):357–368. https://doi.org/10.1016/j.tem.2019.03.003
Bertoli G, Cava C, Castiglioni I (2015) MicroRNAs: new biomarkers for diagnosis, prognosis, therapy prediction and therapeutic tools for breast cancer. Theranostics 5(10):1122–1143. https://doi.org/10.7150/thno.11543
Borsani O, Zhu J, Verslues PE, Sunkar R, Zhu JK (2005) Endogenous siRNAs derived from a pair of natural cis-antisense transcripts regulate salt tolerance in Arabidopsis. Cell 123(7):1279–1291. https://doi.org/10.1016/j.cell.2005.11.035
Burke L, Guterman I, Palacios Gallego R, Britton RG, Burschowsky D, Tufarelli C, Rufini A (2020) The Janus-like role of proline metabolism in cancer. Cell Death Discov 6:104. https://doi.org/10.1038/s41420-020-00341-8
Byun J, Park M, Lee S, Yun J, Lee J, Kim J, Cho S, Jeon H, Lee I, Choi Y, Park K (2020) Inhibition of glutamine utilization synergizes with immune checkpoint inhibitor to promote antitumor immunity. Mol Cell 80(4):592-606.e598. https://doi.org/10.1016/j.molcel.2020.10.015
Cappelletti P, Tallarita E, Rabattoni V, Campomenosi P, Sacchi S, Pollegioni L (2018) Proline oxidase controls proline, glutamate, and glutamine cellular concentrations in a U87 glioblastoma cell line. PLoS ONE 13(4):e0196283. https://doi.org/10.1371/journal.pone.0196283
Celińska-Janowicz K, Zaręba I, Lazarek U, Teul J, Tomczyk M, Pałka J, Miltyk W (2018) Constituents of propolis: chrysin, caffeic acid, p-coumaric acid, and ferulic acid induce PRODH/POX-dependent apoptosis in human tongue squamous cell carcinoma cell (CAL-27). Front Pharmacol 9:336. https://doi.org/10.3389/fphar.2018.00336
Chen L, Shi Y, Liu S, Cao Y, Wang X, Tao Y (2014) PKM2: the thread linking energy metabolism reprogramming with epigenetics in cancer. Int J Mol Sci 15(7):11435–11445. https://doi.org/10.3390/ijms150711435
Chen L, Shi Y, Liu N, Wang Z, Yang R, Yan B, Liu X, Lai W, Liu Y, Xiao D, Zhou H, Cheng Y, Cao Y, Liu S, Xia Z, Tao Y (2019) DNA methylation modifier LSH inhibits p53 ubiquitination and transactivates p53 to promote lipid metabolism. Epigenetics Chromatin 12(1):59. https://doi.org/10.1186/s13072-019-0302-9
Chen C, Wei Y, Wei L, Chen J, Chen X, Dong X, He J, Lin L, Zhu Y, Huang H, You D, Lai L, Shen S, Duan W, Su L, Shafer A, Fleischer T, Bjaanæs MM, Karlsson A, Planck M, Wang R, Staaf J, Helland Å, Esteller M, Zhang R, Chen F, Christiani DC (2020) Epigenome-wide gene-age interaction analysis reveals reversed effects of PRODH DNA methylation on survival between young and elderly early-stage NSCLC patients. Aging (albany NY) 12(11):10642–10662. https://doi.org/10.18632/aging.103284
Chio IIC, Tuveson DA (2017) ROS in cancer: the burning question. Trends Mol Med 23(5):411–429. https://doi.org/10.1016/j.molmed.2017.03.004
Choi BH, Coloff JL (2019) The diverse functions of non-essential amino acids in cancer. Cancers (Basel). https://doi.org/10.3390/cancers11050675
Coloff JL, Murphy JP, Braun CR, Harris IS, Shelton LM, Kami K, Gygi SP, Selfors LM, Brugge JS (2016) Differential glutamate metabolism in proliferating and quiescent mammary epithelial cells. Cell Metab 23(5):867–880. https://doi.org/10.1016/j.cmet.2016.03.016
Cordes FS, Bright JN, Sansom MS (2002) Proline-induced distortions of transmembrane helices. J Mol Biol 323(5):951–960. https://doi.org/10.1016/s0022-2836(02)01006-9
Davalli P, Mitic T, Caporali A, Lauriola A, D’Arca D (2016) ROS, cell senescence, and novel molecular mechanisms in aging and age-related diseases. Oxid Med Cell Longev 2016:3565127. https://doi.org/10.1155/2016/3565127
De Ingeniis J, Ratnikov B, Richardson AD, Scott DA, Aza-Blanc P, De SK, Kazanov M, Pellecchia M, Ronai Z, Osterman AL, Smith JW (2012) Functional specialization in proline biosynthesis of melanoma. PLoS ONE 7(9):e45190. https://doi.org/10.1371/journal.pone.0045190
Deng L, Meng T, Chen L, Wei W, Wang P (2020) The role of ubiquitination in tumorigenesis and targeted drug discovery. Signal Transduct Target Ther 5(1):11. https://doi.org/10.1038/s41392-020-0107-0
Ding J, Kuo ML, Su L, Xue L, Luh F, Zhang H, Wang J, Lin TG, Zhang K, Chu P, Zheng S, Liu X, Yen Y (2017) Human mitochondrial pyrroline-5-carboxylate reductase 1 promotes invasiveness and impacts survival in breast cancers. Carcinogenesis 38(5):519–531. https://doi.org/10.1093/carcin/bgx022
Donald SP, Sun XY, Hu CA, Yu J, Mei JM, Valle D, Phang JM (2001) Proline oxidase, encoded by p53-induced gene-6, catalyzes the generation of proline-dependent reactive oxygen species. Cancer Res 61(5):1810–1815
Echizen K, Oshima H, Nakayama M, Oshima M (2018) The inflammatory microenvironment that promotes gastrointestinal cancer development and invasion. Adv Biol Regul 68:39–45. https://doi.org/10.1016/j.jbior.2018.02.001
Elia I, Broekaert D, Christen S, Boon R, Radaelli E, Orth MF, Verfaillie C, Grünewald TGP, Fendt SM (2017) Proline metabolism supports metastasis formation and could be inhibited to selectively target metastasizing cancer cells. Nat Commun 8:15267. https://doi.org/10.1038/ncomms15267
Ezeriņa D, Takano Y, Hanaoka K, Urano Y, Dick TP (2018) N-acetyl cysteine functions as a fast-acting antioxidant by triggering intracellular H(2)S and sulfane sulfur production. Cell Chem Biol 25(4):447-459.e444. https://doi.org/10.1016/j.chembiol.2018.01.011
Fang H, Du G, Wu Q, Liu R, Chen C, Feng J (2019) HDAC inhibitors induce proline dehydrogenase (POX) transcription and anti-apoptotic autophagy in triple negative breast cancer. Acta Biochim Biophys Sin (shanghai) 51(10):1064–1070. https://doi.org/10.1093/abbs/gmz097
Filomeni G, De Zio D, Cecconi F (2015) Oxidative stress and autophagy: the clash between damage and metabolic needs. Cell Death Differ 22(3):377–388. https://doi.org/10.1038/cdd.2014.150
Greenhough A, Smartt HJM, Moore AE, Roberts HR, Williams AC, Paraskeva C, Kaidi A (2009) The COX-2/PGE 2 pathway: key roles in the hallmarks of cancer and adaptation to the tumour microenvironment. Carcinogenesis 30(3):377–386. https://doi.org/10.1093/carcin/bgp014
Gross MI, Demo SD, Dennison JB, Chen L, Chernov-Rogan T, Goyal B, Janes JR, Laidig GJ, Lewis ER, Li J, Mackinnon AL, Parlati F, Rodriguez ML, Shwonek PJ, Sjogren EB, Stanton TF, Wang T, Yang J, Zhao F, Bennett MK (2014) Antitumor activity of the glutaminase inhibitor CB-839 in triple-negative breast cancer. Mol Cancer Ther 13(4):890–901. https://doi.org/10.1158/1535-7163.Mct-13-0870
Hancock CN, Liu W, Alvord WG, Phang JM (2016) Co-regulation of mitochondrial respiration by proline dehydrogenase/oxidase and succinate. Amino Acids 48(3):859–872. https://doi.org/10.1007/s00726-015-2134-7
Hayes J, Peruzzi PP, Lawler S (2014) MicroRNAs in cancer: biomarkers, functions and therapy. Trends Mol Med 20(8):460–469. https://doi.org/10.1016/j.molmed.2014.06.005
He X, Yan B, Liu S, Jia J, Lai W, Xin X, Tang CE, Luo D, Tan T, Jiang Y, Shi Y, Liu Y, Xiao D, Chen L, Liu S, Mao C, Yin G, Cheng Y, Fan J, Cao Y, Muegge K, Tao Y (2016) Chromatin remodeling factor LSH drives cancer progression by suppressing the activity of fumarate hydratase. Cancer Res 76(19):5743–5755. https://doi.org/10.1158/0008-5472.Can-16-0268
He Y, Gao M, Tang H, Cao Y, Liu S, Tao Y (2019) Metabolic Intermediates in Tumorigenesis and Progression. Int J Biol Sci 15(6):1187–1199. https://doi.org/10.7150/ijbs.33496
Hu CA, Donald SP, Yu J, Lin WW, Liu Z, Steel G, Obie C, Valle D, Phang JM (2007) Overexpression of proline oxidase induces proline-dependent and mitochondria-mediated apoptosis. Mol Cell Biochem 295(1–2):85–92. https://doi.org/10.1007/s11010-006-9276-6
Kim KY, Ahn JH, Cheon HG (2007) Apoptotic action of peroxisome proliferator-activated receptor-gamma activation in human non small-cell lung cancer is mediated via proline oxidase-induced reactive oxygen species formation. Mol Pharmacol 72(3):674–685. https://doi.org/10.1124/mol.107.035584
Koivunen P, Hirsilä M, Remes AM, Hassinen IE, Kivirikko KI, Myllyharju J (2007) Inhibition of hypoxia-inducible factor (HIF) hydroxylases by citric acid cycle intermediates: possible links between cell metabolism and stabilization of HIF. J Biol Chem 282(7):4524–4532. https://doi.org/10.1074/jbc.M610415200
Koppula P, Zhuang L, Gan B (2020) Cystine transporter SLC7A11/xCT in cancer: ferroptosis, nutrient dependency, and cancer therapy. Protein Cell. https://doi.org/10.1007/s13238-020-00789-5
Kowaloff EM, Phang JM, Granger AS, Downing SJ (1977) Regulation of proline oxidase activity by lactate. Proc Natl Acad Sci U S A 74(12):5368–5371. https://doi.org/10.1073/pnas.74.12.5368
Krol J, Loedige I, Filipowicz W (2010) The widespread regulation of microRNA biogenesis, function and decay. Nat Rev Genet 11(9):597–610. https://doi.org/10.1038/nrg2843
Lang Z, Wu Y, Pan X, Qu G, Zhang T (2018) Study of differential gene expression between invasive multifocal/ multicentric and unifocal breast cancer. J Buon 23(1):134–142
Le A, Lane AN, Hamaker M, Bose S, Gouw A, Barbi J, Tsukamoto T, Rojas CJ, Slusher BS, Zhang H, Zimmerman LJ, Liebler DC, Slebos RJ, Lorkiewicz PK, Higashi RM, Fan TW, Dang CV (2012) Glucose-independent glutamine metabolism via TCA cycling for proliferation and survival in B cells. Cell Metab 15(1):110–121. https://doi.org/10.1016/j.cmet.2011.12.009
Lee RC, Feinbaum RL, Ambros V (1993) The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75(5):843–854. https://doi.org/10.1016/0092-8674(93)90529-y
Lee YH, Nadaraia S, Gu D, Becker DF, Tanner JJ (2003) Structure of the proline dehydrogenase domain of the multifunctional PutA flavoprotein. Nat Struct Biol 10(2):109–114. https://doi.org/10.1038/nsb885
Lei G, Zhang Y, Koppula P, Liu X, Zhang J, Lin SH, Ajani JA, Xiao Q, Liao Z, Wang H, Gan B (2020) The role of ferroptosis in ionizing radiation-induced cell death and tumor suppression. Cell Res 30(2):146–162. https://doi.org/10.1038/s41422-019-0263-3
Liu W, Phang JM (2012) Proline dehydrogenase (oxidase), a mitochondrial tumor suppressor, and autophagy under the hypoxia microenvironment. Autophagy 8(9):1407–1409. https://doi.org/10.4161/auto.21152
Liu Y, Borchert GL, Donald SP, Surazynski A, Hu CA, Weydert CJ, Oberley LW, Phang JM (2005) MnSOD inhibits proline oxidase-induced apoptosis in colorectal cancer cells. Carcinogenesis 26(8):1335–1342. https://doi.org/10.1093/carcin/bgi083
Liu Y, Borchert GL, Surazynski A, Hu CA, Phang JM (2006) Proline oxidase activates both intrinsic and extrinsic pathways for apoptosis: the role of ROS/superoxides. NFAT and MEK/ERK Signaling Oncogene 25(41):5640–5647. https://doi.org/10.1038/sj.onc.1209564
Liu Y, Borchert GL, Surazynski A, Phang JM (2008) Proline oxidase, a p53-induced gene, targets COX-2/PGE2 signaling to induce apoptosis and inhibit tumor growth in colorectal cancers. Oncogene 27(53):6729–6737. https://doi.org/10.1038/onc.2008.322
Liu Y, Borchert GL, Donald SP, Diwan BA, Anver M, Phang JM (2009) Proline oxidase functions as a mitochondrial tumor suppressor in human cancers. Cancer Res 69(16):6414–6422. https://doi.org/10.1158/0008-5472.Can-09-1223
Liu W, Zabirnyk O, Wang H, Shiao YH, Nickerson ML, Khalil S, Anderson LM, Perantoni AO, Phang JM (2010) miR-23b targets proline oxidase, a novel tumor suppressor protein in renal cancer. Oncogene 29(35):4914–4924. https://doi.org/10.1038/onc.2010.237
Liu W, Glunde K, Bhujwalla ZM, Raman V, Sharma A, Phang JM (2012a) Proline oxidase promotes tumor cell survival in hypoxic tumor microenvironments. Cancer Res 72(14):3677–3686. https://doi.org/10.1158/0008-5472.Can-12-0080
Liu W, Le A, Hancock C, Lane AN, Dang CV, Fan TW, Phang JM (2012b) Reprogramming of proline and glutamine metabolism contributes to the proliferative and metabolic responses regulated by oncogenic transcription factor c-MYC. Proc Natl Acad Sci U S A 109(23):8983–8988. https://doi.org/10.1073/pnas.1203244109
Liu Y, Mao C, Wang M, Liu N, Ouyang L, Liu S, Tang H, Cao Y, Liu S, Wang X, Xiao D, Chen C, Shi Y, Yan Q, Tao Y (2020) Cancer progression is mediated by proline catabolism in non-small cell lung cancer. Oncogene 39(11):2358–2376. https://doi.org/10.1038/s41388-019-1151-5
Lo M, Wang YZ, Gout PW (2008) The x(c)- cystine/glutamate antiporter: a potential target for therapy of cancer and other diseases. J Cell Physiol 215(3):593–602. https://doi.org/10.1002/jcp.21366
Loayza-Puch F, Rooijers K, Buil LC, Zijlstra J, Oude Vrielink JF, Lopes R, Ugalde AP, van Breugel P, Hofland I, Wesseling J, van Tellingen O, Bex A, Agami R (2016) Tumour-specific proline vulnerability uncovered by differential ribosome codon reading. Nature 530(7591):490–494. https://doi.org/10.1038/nature16982
Lunt S, Vander Heiden M (2011) Aerobic glycolysis: meeting the metabolic requirements of cell proliferation. Annu Rev Cell Dev Biol 27:441–464. https://doi.org/10.1146/annurev-cellbio-092910-154237
Luo C, Zhang H (2017) The role of proinflammatory pathways in the pathogenesis of colitis-associated colorectal cancer. Mediators Inflamm 2017:5126048. https://doi.org/10.1155/2017/5126048
Mao C, Wang X, Liu Y, Wang M, Yan B, Jiang Y, Shi Y, Shen Y, Liu X, Lai W, Yang R, Xiao D, Cheng Y, Liu S, Zhou H, Cao Y, Yu W, Muegge K, Yu H, Tao Y (2018) A G3BP1-interacting lncRNA promotes ferroptosis and apoptosis in cancer via nuclear sequestration of p53. Cancer Res 78(13):3484–3496. https://doi.org/10.1158/0008-5472.Can-17-3454
Maxwell SA, Davis GE (2000) Differential gene expression in p53-mediated apoptosis-resistant vs. apoptosis-sensitive tumor cell lines. Proc Natl Acad Sci U S A. 97(24):13009–13014. https://doi.org/10.1073/pnas.230445997
Maxwell SA, Kochevar GJ (2008) Identification of a p53-response element in the promoter of the proline oxidase gene. Biochem Biophys Res Commun 369(2):308–313. https://doi.org/10.1016/j.bbrc.2008.01.171
Maxwell SA, Rivera A (2003) Proline oxidase induces apoptosis in tumor cells, and its expression is frequently absent or reduced in renal carcinomas. J Biol Chem 278(11):9784–9789. https://doi.org/10.1074/jbc.M210012200
Muhammad N, Lee HM, Kim J (2020) Oncology therapeutics targeting the metabolism of amino acids. Cells. https://doi.org/10.3390/cells9081904
Nagano T, Nakano M, Nakashima A, Onishi K, Yamao S, Enari M, Kikkawa U, Kamada S (2016) Identification of cellular senescence-specific genes by comparative transcriptomics. Sci Rep 6:31758. https://doi.org/10.1038/srep31758
Nagano T, Nakashima A, Onishi K, Kawai K, Awai Y, Kinugasa M, Iwasaki T, Kikkawa U, Kamada S (2017) Proline dehydrogenase promotes senescence through the generation of reactive oxygen species. J Cell Sci 130(8):1413–1420. https://doi.org/10.1242/jcs.196469
Nazio F, Bordi M, Cianfanelli V, Locatelli F, Cecconi F (2019) Autophagy and cancer stem cells: molecular mechanisms and therapeutic applications. Cell Death Differ 26(4):690–702. https://doi.org/10.1038/s41418-019-0292-y
Olivares O, Mayers JR, Gouirand V, Torrence ME, Gicquel T, Borge L, Lac S, Roques J, Lavaut M-N, Berthezène P, Rubis M, Secq V, Garcia S, Moutardier V, Lombardo D, Iovanna JL, Tomasini R, Guillaumond F, Vander Heiden MG, Vasseur S (2017) Collagen-derived proline promotes pancreatic ductal adenocarcinoma cell survival under nutrient limited conditions. Nat Commun 8(1):16031. https://doi.org/10.1038/ncomms16031
Pandhare J, Cooper SK, Phang JM (2006) Proline oxidase, a proapoptotic gene, is induced by troglitazone: evidence for both peroxisome proliferator-activated receptor gamma-dependent and -independent mechanisms. J Biol Chem 281(4):2044–2052. https://doi.org/10.1074/jbc.M507867200
Pandhare J, Donald SP, Cooper SK, Phang JM (2009) Regulation and function of proline oxidase under nutrient stress. J Cell Biochem 107(4):759–768. https://doi.org/10.1002/jcb.22174
Pavlova NN, Thompson CB (2016) The emerging hallmarks of cancer metabolism. Cell Metab 23(1):27–47. https://doi.org/10.1016/j.cmet.2015.12.006
Phang JM (2019) Proline Metabolism in Cell Regulation and Cancer Biology: Recent Advances and Hypotheses. Antioxid Redox Signal 30(4):635–649. https://doi.org/10.1089/ars.2017.7350
Phang JM, Liu W, Zabirnyk O (2010) Proline metabolism and microenvironmental stress. Annu Rev Nutr 30:441–463. https://doi.org/10.1146/annurev.nutr.012809.104638
Polyak K, Xia Y, Zweier JL, Kinzler KW, Vogelstein B (1997) A model for p53-induced apoptosis. Nature 389(6648):300–305. https://doi.org/10.1038/38525
Prasad S, Gupta SC, Tyagi AK (2017) Reactive oxygen species (ROS) and cancer: role of antioxidative nutraceuticals. Cancer Lett 387:95–105. https://doi.org/10.1016/j.canlet.2016.03.042
Raimondi I, Ciribilli Y, Monti P, Bisio A, Pollegioni L, Fronza G, Inga A, Campomenosi P (2013) P53 family members modulate the expression of PRODH, but not PRODH2, via intronic p53 response elements. PLoS ONE 8(7):e69152. https://doi.org/10.1371/journal.pone.0069152
Rajeshkumar NV, Yabuuchi S, Pai SG, De Oliveira E, Kamphorst JJ, Rabinowitz JD, Tejero H, Al-Shahrour F, Hidalgo M, Maitra A, Dang CV (2017) Treatment of pancreatic cancer patient-derived xenograft panel with metabolic inhibitors reveals efficacy of phenformin. Clin Cancer Res 23(18):5639–5647. https://doi.org/10.1158/1078-0432.Ccr-17-1115
Rivera A, Maxwell SA (2005) The p53-induced gene-6 (proline oxidase) mediates apoptosis through a calcineurin-dependent pathway. J Biol Chem 280(32):29346–29354. https://doi.org/10.1074/jbc.M504852200
Rupaimoole R, Slack FJ (2017) MicroRNA therapeutics: towards a new era for the management of cancer and other diseases. Nat Rev Drug Discov 16(3):203–222. https://doi.org/10.1038/nrd.2016.246
Sabharwal SS, Schumacker PT (2014) Mitochondrial ROS in cancer: initiators, amplifiers or an Achilles’ heel? Nat Rev Cancer 14(11):709–721. https://doi.org/10.1038/nrc3803
Shoulders MD, Raines RT (2009) Collagen structure and stability. Annu Rev Biochem 78:929–958. https://doi.org/10.1146/annurev.biochem.77.032207.120833
Sies H, Jones DP (2020) Reactive oxygen species (ROS) as pleiotropic physiological signalling agents. Nat Rev Mol Cell Biol 21(7):363–383. https://doi.org/10.1038/s41580-020-0230-3
Srinivas US, Tan BWQ, Vellayappan BA, Jeyasekharan AD (2019) ROS and the DNA damage response in cancer. Redox Biol 25:101084. https://doi.org/10.1016/j.redox.2018.101084
Su H, Liang Z, Weng S, Sun C, Huang J, Zhang T, Wang X, Wu S, Zhang Z, Zhang Y, Gong Q, Xu Y (2021) miR-9–5p regulates immunometabolic and epigenetic pathways in β-glucan-trained immunity via IDH3α. JCI Insight. https://doi.org/10.1172/jci.insight.144260
Telerman A, Amson R (2009) The molecular programme of tumour reversion: the steps beyond malignant transformation. Nat Rev Cancer 9(3):206–216. https://doi.org/10.1038/nrc2589
Tian Y, Yang B, Qiu W, Hao Y, Zhang Z, Yang B, Li N, Cheng S, Lin Z, Rui Y, Cheung O, Yang W, Wu W, Cheung Y, Lai P, Luo J, Sung J, Chen R, Wang H, Cheng A, Yang P (2019) ER-residential Nogo-B accelerates NAFLD-associated HCC mediated by metabolic reprogramming of oxLDL lipophagy. Nat Commun 10(1):3391. https://doi.org/10.1038/s41467-019-11274-x
Toloczko-Iwaniuk N, Dziemianczyk-Pakiela D, Celinska-Janowicz K, Zareba I, Klupczynska A, Kokot ZJ, Nowaszewska BK, Reszec J, Borys J, Miltyk W (2020) Proline-dependent induction of apoptosis in oral squamous cell carcinoma (OSCC)-the effect of celecoxib. Cancers (Basel) 12(1):136. https://doi.org/10.3390/cancers12010136
Verma A (2006) Oxygen-sensing in tumors. Curr Opin Clin Nutr Metab Care 9(4):366–378. https://doi.org/10.1097/01.mco.0000232895.28674.79
Wang X, Liu R, Qu X, Yu H, Chu H, Zhang Y, Zhu W, Wu X, Gao H, Tao B, Li W, Liang J, Li G, Yang W (2019) α-ketoglutarate-activated NF-κB signaling promotes compensatory glucose uptake and brain tumor development. Mol Cell 76(1):148-162.e147. https://doi.org/10.1016/j.molcel.2019.07.007
Wang CY, Chiao CC, Phan NN, Li CY, Sun ZD, Jiang JZ, Hung JH, Chen YL, Yen MC, Weng TY, Chen WC, Hsu HP, Lai MD (2020) Gene signatures and potential therapeutic targets of amino acid metabolism in estrogen receptor-positive breast cancer. Am J Cancer Res 10(1):95–113
Weijin F, Zhibin X, Shengfeng Z, Xiaoli Y, Qijian D, Jiayi L, Qiumei L, Yilong C, Hua M, Deyun L, Jiwen C (2019) The clinical significance of PYCR1 expression in renal cell carcinoma. Medicine (baltimore) 98(28):e16384. https://doi.org/10.1097/md.0000000000016384
Wen YA, Xiong X, Scott T, Li AT, Wang C, Weiss HL, Tan L, Bradford E, Fan TWM, Chandel NS, Barrett TA, Gao T (2019) The mitochondrial retrograde signaling regulates Wnt signaling to promote tumorigenesis in colon cancer. Cell Death Differ 26(10):1955–1969. https://doi.org/10.1038/s41418-018-0265-6
Williams KA, Deber CM (1991) Proline residues in transmembrane helices: structural or dynamic role? Biochemistry 30(37):8919–8923. https://doi.org/10.1021/bi00101a001
Xiang Y, Stine ZE, Xia J, Lu Y, O’Connor RS, Altman BJ, Hsieh AL, Gouw AM, Thomas AG, Gao P, Sun L, Song L, Yan B, Slusher BS, Zhuo J, Ooi LL, Lee CG, Mancuso A, McCallion AS, Le A, Milone MC, Rayport S, Felsher DW, Dang CV (2015) Targeted inhibition of tumor-specific glutaminase diminishes cell-autonomous tumorigenesis. J Clin Invest 125(6):2293–2306. https://doi.org/10.1172/jci75836
Y H, M G, H T, Y C, S L, Y T, (2019) Metabolic intermediates in tumorigenesis and progression. Int J Biol Sci 15(6):1187–1199
Yan Y, Chang L, Tian H, Wang L, Zhang Y, Yang T, Li G, Hu W, Shah K, Chen G, Guo Y (2018) 1-Pyrroline-5-carboxylate released by prostate Cancer cell inhibit T cell proliferation and function by targeting SHP1/cytochrome c oxidoreductase/ROS Axis. J Immunother Cancer 6(1):148. https://doi.org/10.1186/s40425-018-0466-z
Yan K, Xu X, Wu T, Li J, Cao G, Li Y, Ji Z (2019) Knockdown of PYCR1 inhibits proliferation, drug resistance and EMT in colorectal cancer cells by regulating STAT3-Mediated p38 MAPK and NF-κB signalling pathway. Biochem Biophys Res Commun 520(2):486–491. https://doi.org/10.1016/j.bbrc.2019.10.059
Ye Y, Wu Y, Wang J (2018) Pyrroline-5-carboxylate reductase 1 promotes cell proliferation via inhibiting apoptosis in human malignant melanoma. Cancer Manag Res 10:6399–6407. https://doi.org/10.2147/cmar.S166711
Yu T, Dong T, Eyvani H, Fang Y, Wang X, Zhang X, Lu X (2020) Metabolic interventions: a new insight into the cancer immunotherapy. Arch Biochem Biophys. https://doi.org/10.1016/j.abb.2020.108659
Yuneva MO, Fan TW, Allen TD, Higashi RM, Ferraris DV, Tsukamoto T, Matés JM, Alonso FJ, Wang C, Seo Y, Chen X, Bishop JM (2012) The metabolic profile of tumors depends on both the responsible genetic lesion and tissue type. Cell Metab 15(2):157–170. https://doi.org/10.1016/j.cmet.2011.12.015
Zabirnyk O, Liu W, Khalil S, Sharma A, Phang JM (2010) Oxidized low-density lipoproteins upregulate proline oxidase to initiate ROS-dependent autophagy. Carcinogenesis 31(3):446–454. https://doi.org/10.1093/carcin/bgp299
Zareba I, Palka J (2016) Prolidase-proline dehydrogenase/proline oxidase-collagen biosynthesis axis as a potential interface of apoptosis/autophagy. BioFactors 42(4):341–348. https://doi.org/10.1002/biof.1283
Zareba I, Surazynski A, Chrusciel M, Miltyk W, Doroszko M, Rahman N, Palka J (2017) Functional consequences of intracellular proline levels manipulation affecting PRODH/POX-dependent pro-apoptotic pathways in a novel in vitro cell culture model. Cell Physiol Biochem 43(2):670–684. https://doi.org/10.1159/000480653
Zhang M, White TA, Schuermann JP, Baban BA, Becker DF, Tanner JJ (2004) Structures of the Escherichia coli PutA proline dehydrogenase domain in complex with competitive inhibitors. Biochemistry 43(39):12539–12548. https://doi.org/10.1021/bi048737e
Zhang Y, Zhang L, Gao J, Wen L (2019) Pro-death or pro-survival: contrasting paradigms on nanomaterial-induced autophagy and exploitations for cancer therapy. Acc Chem Res 52(11):3164–3176. https://doi.org/10.1021/acs.accounts.9b00397
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273. https://doi.org/10.1146/annurev.arplant.53.091401.143329
Zhuang J, Song Y, Ye Y, He S, Ma X, Zhang M, Ni J, Wang J, Xia W (2019) PYCR1 interference inhibits cell growth and survival via c-Jun N-terminal kinase/insulin receptor substrate 1 (JNK/IRS1) pathway in hepatocellular cancer. J Transl Med 17(1):343. https://doi.org/10.1186/s12967-019-2091-0
Acknowledgements
We apologize to those whose work has not been cited.
Funding
This work is supported by National Natural Science Foundation of China [82002916 (CM), 81874139 and 82073097 [SL], 81872285[YS], 82072594 [YT], 82073136 [DX]], Natural Science Foundation of Hunan Province [2020JJ5790 (CM)], China Postdoctoral Science Foundation [2019M652804 (CM)] and Postdoctoral Foundation of Central South University [220372 (CM)], Shenzhen Science and Technology Program (KQTD20170810160226082 [YT]), Shenzhen Municipal Government of China (JCYJ20180507184647104 [YT]), and the Hunan Provincial Key Area R&D Program (2019SK2253[YT]).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
None.
Additional information
Handling editor: J. M. Phang.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Liu, Y., Mao, C., Liu, S. et al. Proline dehydrogenase in cancer: apoptosis, autophagy, nutrient dependency and cancer therapy. Amino Acids 53, 1891–1902 (2021). https://doi.org/10.1007/s00726-021-03032-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-021-03032-5