Abstract
Amino acids are important metabolites for tissue metabolism, growth, maintenance, and repair, which are basic life necessities. Therefore, summarizing analytical methods for amino acid determination in organisms is important. In the past decades, analytical methods for amino acids have developed rapidly but have not been fully explored. Thus, this article provides reference to analytical methods for amino acids in organisms for food and human research. Present amino acid analysis methods include thin-layer chromatography, high-performance liquid chromatography, liquid chromatography–mass spectrometer, gas chromatography–mass spectrometry, capillary electrophoresis, nuclear magnetic resonance, and amino acid analyzer analysis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Amino acids (AAs) are the most important chemical structures in organisms and generally categorized into nonessential and essential AAs (Marouzi et al. 2017; Choi et al. 2007). The quantity and quality of AAs are required for analysis in the fields of medicine, food, feed, agriculture, and chemistry. Moreover, essential AAs constitute approximately 20–37% of the protein requirement of a human adult, and some AAs are potential biomarkers of diseases.
Apart from participating in protein biosynthesis (Johnson et al. 2014), AAs also serve as precursors for many hormones, neurotransmitters (Tian et al. 2018), and other specialized metabolites (Broer and Broer 2017; Hildebrandt et al. 2015). For example, glu can be used as an acidic AA in metabolism and as an excitatory neurotransmitter of information. Dietary AA patterns with high levels of gly, cys, arg, and try may be associated with reduced risk of cardiovascular events (Mirmiran et al. 2017). Thus, some AAs in food should be explored, especially those from animals and plants (Mondanelli et al. 2019; Berrazaga et al. 2019; Tosti et al. 2018; Young and Pellett 1987). In addition, AA catabolism can influence plant growth and development, such as intracellular pH control and metabolic energy generation. Furthermore, analytical methods for AAs research enable the evaluation of plant quality because AAs provide nutrition to humans and can be extracted for medicinal components. D-AAs are considered unnatural AAs (Gao et al. 2015) but are recognized as naturally occurring physiologically active substances and biomarkers in mammals (Miyoshi et al. 2012). Some D-AAs occur in food under high temperatures (Hayase et al. 1975) and alkali treatment (Friedman et al. 1984). Furthermore, D-Asp and D-Asn are found in the peptidoglycans of some bacteria (Veiga et al. 2006). D-AAs have many applications; for example, they can be used as sweeteners. The functions of D-AAs have attracted attention, but sensitive and high-throughput analytical methods for analyzing D-AAs remains inadequate (Muller et al. 2014). To date, electrochemical sensors are generally used to detect D-AAs. Given that most AAs are small aliphatic molecules incapable of fluorescence or UV absorption, analyzing AAs is difficult (Ou et al. 2013). To better analyze AAs, pre-column or post-column derivation of AAs is performed for detection, then the AAs are detected by HPLC, LC–MS, or GC–MS (Furst et al. 1990; Fierabracci et al. 1991; Gogichaeva and Alterman 2012). Advanced techniques for quantifying AAs include CE, NMR, and amino acid analysis.
Other articles discuss several analytical methods for AAs without introducing derivatizations. The current article comprehensively compares current analytical methods and discusses AA applications in food and human research. Induced derivatizations are used to supply information for drug discovery, disease detection, and food nutrition exploration.
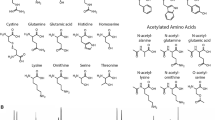
The structure diagram of the methods and analytical techniques mentioned in this review and their properties are summarized in Fig. 1.
Derivatizations
Given that most AAs lack natural UV or fluorescence absorption functional groups, chemical derivation has become an effective way for increasing the sensitivity of AA detection (Sharma et al. 2014; Pretorius et al. 2018; Sakaguchi et al. 2015; Stocchi et al. 1992; Sherwood 2000; Hess 2012; Fonseca et al. 2018; de Puit et al. 2014; Toue et al. 2014; Rebane et al. 2012; Oldekop et al. 2017a, b; Yang et al. 2017; Miyoshi et al. 2014). The structure for most common derivation reagents is displayed in Fig. 2. Information about common chemical derivation reagents and derivation conditions is summarized in Table 1. Of these reagents, only with 6-aminoquinolyl-N-hydroxysuccinimidyl carbamate (AQC), fluorenylmethyl chloroformate (FMOC-Cl), and phenylisothiocyanate (PITC) can react with primary and secondary AAs simultaneously. However, AQC and FMOC-Cl hydrolysis products sometimes interfere with detection, and the derivatives of PITC are unstable. Moreover, PITC must be removed from a sample for the prevention of column contamination. o-Phthalaldehyde (OPA) does not react with secondary AAs and its derivatives are sometimes unstable. Therefore, derivation reagents should be carefully selected. It must react with AAs directly as soon as possible and stable for a long time.
Amino acid analysis
Thin-layer chromatography
TLC is widely used in the separation and identification of AAs, peptides, lipids, and alkaloids. It is simple, convenient, and cost effective (Yousefinejad et al. 2015; Lu and Olesik 2013). Reversed-phase chromatographic analysis is usually performed on AAs, such as isoleu, leu, val, ala, gly, and orn (Nguyen et al. 2016). Separating racemic mixtures is necessary due to different pharmacological activities. In a previous paper, aromatic AAs were analyzed by using Spi(τ-dec) as a chiral selector for high-performance thin-layer chromatography (Remelli et al. 2014).
High-performance liquid chromatography
HPLC can be used for the qualitative and quantitative analysis of AAs and exhibits high efficiency, high sensitivity, a wide range of applications, and other advantages. In recent years, numerous HPLC methods capable of FL/UV detection have been developed for the analysis of AAs. The concrete detection conditions of the methods are summarized in Table 2.
The analysis of free AAs in biological samples requires the removal of proteins by hydrolysis before derivation and detection. For example, porcine gelatin and bovine gelatins are heated with NaOH to hydrolyze before derivation by OPA (Rezazadeh et al. 2015). Furthermore, pulsed electromembrane extraction is an interesting way for extracting AA derivatives. Finally, Asn and Gln were analyzed within 20 min at 330 nm. Small volumes of plasma samples from children and individuals with critical illnesses are useful in testing. Only 50 μL of plasma volume is required for the simultaneous determination of 33 kinds of derivatized AAs (Wang et al. 2013). A total of 23 kinds of AA PTH derivatives were constructed with HPLC–DAD; this method provides information on the AA contents of peptides (Tasakis and Touraki 2018). Chloroprocaine, especially its inactive metabolite 4-amino-2-chlorobenzoic acid (ACBA), can be quantified through HPLC/MS (De Gregori et al. 2018). A simple HPLC method was developed to directly determine gly in immunoglobulins (Rounova et al. 2018). Plant–animal linkages, such as insect–plant interactions, can be explored by analyzing AAs. First, samples are hydrolyzed into AAs, then plant and insect samples are reconstituted. AA derivatives can be separated by using HPLC–PDA within 36 min (Dhillon et al. 2014). This method is highly sensitive and reproducible and is essential to the analysis of AAs. Core–shell particle columns are potential tools for clarifying the biological activities of AAs (Song et al. 2013). LC–FLD method for quantitative determination neuroactive AAs in rat brain is essential to several neurological diseases (Fonseca et al. 2018). Furthermore, a method for simultaneously detecting 17 kinds of AAs through HPLC is currently available (Nagasaki et al. 2017). Dr. Daniel Armstrong’s group suggested that mammalian brains have unreported D-AAs (Gao et al. 2015; Weatherly et al. 2017). They also analyzed D-AAs in mammals by HPLC with FMOC derivation.
AAs in plants play different roles. For example, gly can promote plant photosynthesis. Similar to biological samples, plant samples need to be hydrolyzed and derivatized. Most studies showed that glu is the most abundant AA in plant proteins; however, in manketti seed kernel flour, the most abundant AA is arg (Gwatidzo et al. 2013). AAs are the important components of chamomile flowers, and 14 kinds of AAs were analyzed with HPLC (Ma et al. 2015). The effect of rhizobial strains on the accumulation of AAs in nodules can be analyzed by HPLC with UV (Bertrand et al. 2016). Furthermore, 19 kinds of AAs are usually employed in HPLC–CAD for the detection of underivatized quantization (Furota et al. 2018).
Diethyl ethoxymethylenemalonate (DEEMM) derivation is followed by UHPLC separation and was used to quantify 21 kinds of AAs in beer (Redruello et al. 2017) with high resolution, accuracy, and sensitivity. Seventeen kinds of AAs in different feeds were analyzed with UPLC (Szkudzińska et al. 2017). HPLC–FL was used for the quantification of free AAs in rice and this method has good linearity, repeatability, and reproducibility (Liyanaarachchi et al. 2018). In addition, 20 kinds of AAs were examined at 338 nm and 266 nm by HPLC (Lamp et al. 2018), and 26 kinds of AAs were extracted at 340/450 and 266/305 nm with UPLC (Manninen et al. 2018).
Liquid chromatography–mass spectrometer
With the development of mass spectrometry and separation methods, LC–MS has become an essential analytical tool for separating AAs (Tsai et al. 2016). Here, LC–MS/MS and UPLC–MS/MS are listed. Concrete detection conditions of the methods are summarized in Table 3.
There are some examples of AAs analyzed by LC–MS. The plasma contains a variety of AAs, and each kind of AA has its own effect. For example, met is involved in the formation of hemoglobin, tissue, and serum and promotes the function of the spleen, pancreas, and lymph. By analyzing underivatized AAs by LC–MS/MS, the means to eliminate the variation can be discovered; a method for all clinically relevant AAs is presently available (Le et al. 2014). HCY concentrations in human sera can indicate some diseases, and HCY concentrations can be quantified by LC–MS/MS bioanalytical method (Ghassabian et al. 2014). LC–MS/MS serve as a useful tool for diabetes because LC–MS/MS directly determines branched chain amino acid (BCAAs) and aromatic AAs in human sera (Yang et al. 2013). Furthermore, plasma AA concentrations in patients with major depressive disorder can be analyzed by LC–MS/MS (Woo et al. 2015). Twenty-four kinds of AAs in human plasma were simultaneously quantified for studying the effects of renal function in de novo kidney (Klepacki et al. 2016). Nakano et al. (2017) simultaneously analyzed 18 kinds of D-AAs without derivation process and applied the method to vinegar for the validation which successfully quantified D-AAs in samples. Multiple AA enantiomers were simultaneously determined in human serum (Han et al. 2018). Moreover, D-Ser in human plasma (Xie et al. 2014) or mouse brains (Kinoshita et al. 2013) can be determined by LC–MS/MS.
Twenty kinds of plant extract AAs with derivation were analyzed by LC–MS/MS based on MRM (Ziegler and Abel 2014). The AAs in natural waters were measured with SPE by LC–MS/MS (How et al. 2014). Guerrasio et al. (2014) developed a novel hydrophilic interaction liquid chromatography combined with electrospray tandem mass spectrometry (HILIC–MS/MS) analytical method for the quantitation of 17 kinds of AAs, and they use a Pichia pastoris cell extract grown on uniformly 13C-labeled glu as an internal standard. Free AAs of Polish and Slovak honeys were characterized by using LC–MS/MS without derivation (Kowalski et al. 2017).
UPLC is another new technology that uses small particles as a stationary phase to achieve ultra-high resolution, sensitivity, and analysis speed. MS can analyze mass-to-charge ratios. Furthermore, UPLC is one of the most optimal entrances of MS. The combination of UPLC and MS significantly improves the reproducibility, the reliability, and the accuracy of qualitative analysis. Many AAs are analyzed by UPLC–MS.
Twenty kinds of AAs and their tracer(s) in human plasma and skeletal muscle can be quantified, and the LC–MS/MS method may be applied to other matrices (Borno and van Hall 2014). The derivation procedure is capable of measuring low enrichment levels, and this procedure is important for human plasma (Oosterink et al. 2014). Simultaneously determining 20 kinds of AAs in plasma at different collecting time points can be achieve by UPLC–MS/MS (Xia et al. 2016b). UPLC–ESI–MS/MS can completely separate pairs of nine kinds of AAs and propose a differential analysis of D/L-amino using light and heavy l-PGA-OSu (Mochizuki et al. 2014). A HILIC column can simultaneously quantify 18 kinds of free AAs in the urine, and this method involves simple samples without any derivation (Joyce et al. 2016). AA enantiomers are usually distinguished by RP–UHPLC–Q–TOF–MS method. D-AAs in the different regions of rat brains can be quantified by UPLC–MS/MS (Li et al. 2017). Gao et al. (2015) identified and quantified D-AAs using chemical derivation coupled with nanoliquid chromatography, and this method may open up a window for studying the organic composition of individual micrometeorites.
In the analysis of AAs from food, 22 kinds of DEEMM-derived AAs in 11 herbs and 4 honeys with LC–ESI–MS/MS in positive and negative ion ESI modes were found; sample dilution was used for the evaluation of matrix effect (Oldekop et al. 2017b). A rapid, reliable, and high-throughput method for simultaneously measuring AAs, polyamines, and dipeptides in complex biological samples is currently available (Ubhi et al. 2013).
Gas chromatography–mass spectrometry
GC–MS is a widely employed technique for doping test, clinical disease diagnosis, and pharmacokinetics study. GC–MS has a universal detector with high efficiency, simplicity, high sensitivity, and high quantitative accuracy. The concrete detection conditions of the methods are summarized in Table 4.
GC–MS can be used to identify not only the methylated AAs but also the human plasma AAs (Reddy et al. 2016). Lopes et al. (2015) validated GC–MS method for the measurement of six kinds of AAs in canine serum samples and assessed the stability of AAs after sample storage. In TBDMS-derived AAs, 24 novel fragment ions were analyzed by GC–MS/MS; additionally, the precision of 13C-MFA in Escherichia coli central carbon metabolism could be improved by introducing the MID data of novel fragment ions (Okahashi et al. 2016). A simple AA extraction method by MAD was developed and 16 kinds of AAs were simultaneously quantified through GC–MS (de Paiva et al. 2013). Furthermore, a rapid method for precisely determining AAs in whole blood is currently available (Kawana et al. 2010). Free and combined AAs in cinobufacini injection were measured with GC–MS (Wu et al. 2015). Twenty kinds of MCF-derived AA enantiomers in serum and urine were separated by comprehensive two-dimensional gas chromatography–time-of-flight mass spectrometry (Waldhier et al. 2011). Trp was used as a chiral probe molecule and applied GC was used in the determination of the enantiomeric excess of AAs in solutions that do not have chromophores (Fujihara and Maeda 2017).
With regard to food, GC–MS could simultaneously analyze nine kinds of AAs in mixed starch waste (Liu et al. 2016). This method achieved good linearity and low limit of detection and quantification. The GC–MS can serve as a system for the separation and detection of AAs in potatoes (Uri et al. 2014) and for the quantification of AAs in plant tissues (Vancompernolle et al. 2016). Li et al. (2013) developed a new derivation and microextraction technique for the quantification of AAs in tobacco by GC–MS. Rubino et al. (2014) analyzed the soil AAs by gas chromatography–combustion–isotope ratio mass spectrometry (GC–C–IRMS).
Capillary electrophoresis
CE is a new type of separation technology that uses capillary as separation channel and is driven by high-voltage DC electric field. CE is fast and has high resolution and good repeatability and is widely used in the analysis of AAs, peptides, and proteins. Concrete detection conditions of the methods are summarized in Table 5.
High-speed capillary electrophoresis (HSCE) which can separate 10 kinds of AAs was applied to analyze the composition of AAs in laver, where five kinds of AAs could be completely separated and quantified (Wang et al. 2015). Schiavone et al. (2015) and colleagues employed an electrokinetically pumped, nanospray sheath-flow CE–ESI–MS interface which can illustrate 20 kinds of AA separation in 7 min. Their work led to the separation of leu and isoleu by the structural isomers and reduced the peak trailing and overlapping. A robust, highly selective, and highly sensitive CE–MS method for the direct analysis of phe and tyr in DBS was described. In this method, the CE run time was less than 3 min and exhibited good linearity and lower detection limit (Jeong et al. 2013). A new method which could avoid using chiral selector was presented (Prior et al. 2016). In this method, FLEC was used as a chiral AA-deriving agent and ammonium perfluorooctanoate as volatile pseudo-stationary phase for the separation of the formed diastereomers. They optimized the CE–MS for the analysis of chiral AAs in CSF and indicated that this method has good linearity, acceptable peak area, and electrophoretic mobility and repeatability. Acket et al. (2018) compared CE-HRMS without derivation with classical GC–MS for 13C labeling analysis of AAs form flaxseed. CE as a low-cost method was used for the quantitation of BCAAs in two commercial sport nutritional supplements where good recovery and precision were obtained. The results indicated the analysis of BCAAs in human bioliquid in supplementing the protein to ensure BCAA demand. A method of CE with indirect UV for the separation and determination of nine kinds of AAs (including Asp, Glu, Ser, Thr, Pro, Ile,Trp, Lys, and Met) (Qiu et al. 2017) and this method were applied to determine AAs in honey from different nectar plants and origins (Zhou and Shi 2013).
CE was also used to separate chiral AAs (Yuan et al. 2011). D-ser, D- and L-asp, and L-glu were measured with CE-LIF method by Jakó T (Jako et al. 2014). Furthermore, CE-LIF also measured d-Ala (Ota et al. 2014). Eight kinds of chiral AAs are completely separated with in-capillary derivation (Moldovan et al. 2016). A simple, rapid, and robust method for D-Orn and D-Ser in human plasma based on CE-LIF was developed (Lorenzo et al. 2013). Prof. Soga (Hirayama et al. 2019) described CE–MS to analyze AAs in detail.
Nuclear magnetic resonance
NMR is a useful tool in studying the composition and structure of various organic and inorganic substances. In addition, NMR does not require complex sample preparation (Munz et al. 2016; Yuan et al. 2017). The main drawback of the method is its limited sensitivity. The AA (Gly, Ala, Glu, Leu, Ser, etc.) composition of spider dragline silk was determined by 1H NMR (Shi et al. 2013). This method is used to quantify the changes of AA (Ile, Leu, Val, Ala, Met, etc.) concentration occurring in Bogue fish during storage. The result indicates that the greatest concentration change was ala and gly which is a key role in determining the individual taste of different fish species (Ciampa et al. 2012). NMR also provides a reliable method to determine AAs in Lycii Fructus (Hsieh et al. 2018).
Amino acid analyzer
The AA analyzer is used to analyze the content of protein hydrolysate and free AAs by the post-column derivation of three-ketone column by cation exchange chromatography. Plasma AAs in female rats were measured by automatic AA analyzer (Okame et al. 2015). Zhao et al. (2014) used automatic AA analyzer to analyze the concentrations of free AAs in the lungs. They investigated the change of AA concentrations (Try, Gly, Orn, Pro, Phe) in plasma free AAs and the change in AA concentrations (Tau, Glu, Gly, Lys, and Orn) in TFAAs and concluded that plasma free AA profiles may reflect the status of cancer tissues. In addition, 17 kinds of AAs in tobacco leaves were eluted on an ion-exchange column (Zeng et al. 2015). Reacting with ninhydrin, the derivatives of AAs were detected by ultraviolet detection. The AAs of beef jerky were analyzed with an AA analyzer for the evaluation of the quality traits of beef jerky (Shikha Ojha et al. 2018). Thirty-eight kinds of free AAs in human plasma were detected with a automated pre-column derivatization AA analyzer (Hirayama et al. 2019).
Electrochemical sensor
Electrochemical (bio-) sensor is a fast, simple, and reliable tool for simultaneously resolving and determining D-AAs (Martin et al. 2015; Wang et al. 2016; Zor et al. 2013). D-Thr and L-Thr were distinguished by novel potential-type electrochemical chiral biosensing system, and the distinguished and quantitative determination of Tyr enantiomers was achieved (Guo et al. 2017). An electrochemical sensor based on 2,2,6,6-tetramethylpiperidine-1-oxyl cellulose nanocrystals and a L-cys-modified Au electrode can be used for the detection and discrimination of phe, leu, and Val enantiomers (Bi et al. 2016). A biosensor based on 3,4,9, 10-perylene tetracarboxylic acid-functionalized multiwalled carbon nanotubes and D-AA oxidase showed high sensitivity and selectivity for the chiral recognition of D-Ala (Xia et al. 2016a). Furthermore, organic electrochemical transistors with gate electrodes modified with molecularly imprinted polymer films dramatically improved the sensitivity of chiral recognition biosensors for D/L-Trp and D/L-Tyr (Zhang et al. 2018). Electrochemical (bio-) sensors can be applied to monitor D/L-Trp or other D/L AAs (Wang et al. 2016; Zor et al. 2013).
Outlook
This current review generalizes the analytical methods of AAs in recent years. Furthermore, we found that the HPLC, LC–MS, and GC–MS are the commonly used analytical methods. Compared with HPLC, LC–MS and GC–MS are more sensitive and more effective; however, the HPLC is more cost-effective. Moreover, TLC, CE, NMR, and AA analyzer can also be used for the analysis AAs. Moreover, there are specific detection methods for D-AAs that provide great convenience for AA analysis. Presently, using an AA analyzer may be the most convenient method, but the necessary equipment is expensive and has many limits. According to literature, LC–MS is the most popular method. However, the problem for the analysis of cost and time has not been dissolved.
There is room for improved methodology for AA analysis, such as simplification of sample preparation process and optimization analysis method (including increasing sensitivity, etc.). With the development of modern science and technology, more sensitive and accurate methods of analyzing AAs are expected. Thus, these methods promote biological metabolism and synthesis of polypeptide drugs.
Abbreviations
- AA:
-
Amino acid
- ACBA:
-
4-Amino-2-chlorobenzoic acid
- AQC:
-
6-Aminoquinolyl-N-hydroxysuccinimidyl carbamate
- BCAA:
-
Branched chain amino acid
- DEEMM:
-
Diethyl ethoxymethylenemalonate
- FMOC-Cl:
-
Fluorenylmethyl chloroformate
- HILIC:
-
Hydrophilic interaction liquid chromatography combined
- HSCE:
-
High-speed capillary electrophoresis
- OPA:
-
o-Phthalaldehyde
- PITC:
-
Phenylisothiocyanate
References
Acket S, Degournay A, Gosset M, Merlier F, Troncoso-Ponce MA, Thomasset B (2018) Analysis of (13)C labeling amino acids by capillary electrophoresis—high resolution mass spectrometry in developing flaxseed. Anal Biochem 547:14–18. https://doi.org/10.1016/j.ab.2018.02.009
Berrazaga I, Micard V, Gueugneau M, Walrand S (2019) The role of the anabolic properties of plant- versus animal-based protein sources in supporting muscle mass maintenance: a critical review. Nutrients. https://doi.org/10.3390/nu11081825
Bertrand A, Bipfubusa M, Dhont C, Chalifour FP, Drouin P, Beauchamp CJ (2016) Rhizobial strains exert a major effect on the amino acid composition of alfalfa nodules under NaCl stress. Plant Physiol Biochem 108:344–352. https://doi.org/10.1016/j.plaphy.2016.08.002
Bi Q, Dong S, Sun Y, Lu X, Zhao L (2016) An electrochemical sensor based on cellulose nanocrystal for the enantioselective discrimination of chiral amino acids. Anal Biochem 508:50–57. https://doi.org/10.1016/j.ab.2016.05.022
Borno A, van Hall G (2014) Quantitative amino acid profiling and stable isotopically labeled amino acid tracer enrichment used for in vivo human systemic and tissue kinetics measurements. J Chromatogr B Anal Technol Biomed Life Sci 951–952:69–77. https://doi.org/10.1016/j.jchromb.2014.01.019
Broer S, Broer A (2017) Amino acid homeostasis and signalling in mammalian cells and organisms. Biochem J 474(12):1935–1963. https://doi.org/10.1042/BCJ20160822
Callahan MP, Martin MG, Burton AS, Glavin DP, Dworkin JP (2014) Amino acid analysis in micrograms of meteorite sample by nanoliquid chromatography-high-resolution mass spectrometry. J Chromatogr A 1332:30–34. https://doi.org/10.1016/j.chroma.2014.01.032
Choi KM, Yoon H-H, Seo Y-K, Song K-Y, Kwon S-Y, Lee H-S, Park YS, Kim Y-J, Park J-K (2007) Effect of essential and nonessential amino acid compositions on the in vitro behavior of human mesenchymal stem cells. Korean J Chem Eng 24(6):1058–1063
Ciampa A, Picone G, Laghi L, Nikzad H, Capozzi F (2012) Changes in the amino acid composition of Bogue (Boops boops) fish during storage at different temperatures by 1H-NMR spectroscopy. Nutrients 4(6):542–553. https://doi.org/10.3390/nu4060542
De Gregori S, De Gregori M, Bloise N, Bugada D, Molinaro M, Filisetti C, Allegri M, Schatman ME, Cobianchi L (2018) In vitro and in vivo quantification of chloroprocaine release from an implantable device in a piglet postoperative pain model. J Pain Res 11:2837–2846. https://doi.org/10.2147/JPR.S180163
de Paiva MJ, Menezes HC, Christo PP, Resende RR, Cardeal Zde L (2013) An alternative derivatization method for the analysis of amino acids in cerebrospinal fluid by gas chromatography-mass spectrometry. J Chromatogr B Anal Technol Biomed Life Sci 931:97–102. https://doi.org/10.1016/j.jchromb.2013.05.014
de Puit M, Ismail M, Xu X (2014) LCMS analysis of fingerprints, the amino acid profile of 20 donors. J Forensic Sci 59(2):364–370. https://doi.org/10.1111/1556-4029.12327
Dhillon MK, Kumar S, Gujar GT (2014) A common HPLC-PDA method for amino acid analysis in insects and plants. Indian J Exp Biol 52(1):73–79
Fierabracci V, Masiello P, Novelli M, Bergamini E (1991) Application of amino acid analysis by high-performance liquid chromatography with phenyl isothiocyanate derivatization to the rapid determination of free amino acids in biological samples. J Chromatogr 570(2):285–291. https://doi.org/10.1016/0378-4347(91)80531-g
Fonseca BM, Cristovao AC, Alves G (2018) An easy-to-use liquid chromatography method with fluorescence detection for the simultaneous determination of five neuroactive amino acids in different regions of rat brain. J Pharmacol Toxicol Methods 91:72–79. https://doi.org/10.1016/j.vascn.2018.02.002
Friedman M, Gumbmann MR, Masters PM (1984) Protein-alkali reactions: chemistry, toxicology, and nutritional consequences. Adv Exp Med Biol 177:367–412. https://doi.org/10.1007/978-1-4684-4790-3_18
Fujihara A, Maeda N (2017) Quantitative chiral analysis of amino acids in solution using enantiomer-selective photo dissociation of cold gas-phase tryptophan via chiral recognition. Anal Chim Acta 979:31–35. https://doi.org/10.1016/j.aca.2017.04.027
Furota S, Ogawa NO, Takano Y, Yoshimura T, Ohkouchi N (2018) Quantitative analysis of underivatized amino acids in the sub- to several-nanomolar range by ion-pair HPLC using a corona-charged aerosol detector (HPLC-CAD). J Chromatogr B Anal Technol Biomed Life Sci 1095:191–197. https://doi.org/10.1016/j.jchromb.2018.07.033
Furst P, Pollack L, Graser TA, Godel H, Stehle P (1990) Appraisal of four pre-column derivatization methods for the high-performance liquid chromatographic determination of free amino acids in biological materials. J Chromatogr 499:557–569. https://doi.org/10.1016/s0021-9673(00)97000-6
Gao X, Ma Q, Zhu H (2015) Distribution, industrial applications, and enzymatic synthesis of D-amino acids. Appl Microbiol Biotechnol 99(8):3341–3349. https://doi.org/10.1007/s00253-015-6507-3
Ghassabian S, Rethwan NS, Griffiths L, Smith MT (2014) Fully validated LC-MS/MS method for quantification of homocysteine concentrations in samples of human serum: a new approach. J Chromatogr B Anal Technol Biomed Life Sci 972:14–21. https://doi.org/10.1016/j.jchromb.2014.09.032
Gogichaeva NV, Alterman MA (2012) Amino acid analysis by means of MALDI TOF mass spectrometry or MALDI TOF/TOF tandem mass spectrometry. Methods Mol Biol (Clifton, NJ) 828:121–135. https://doi.org/10.1007/978-1-61779-445-2_12
Guerrasio R, Haberhauer-Troyer C, Mattanovich D, Koellensperger G, Hann S (2014) Metabolic profiling of amino acids in cellular samples via zwitterionic sub-2 mum particle size HILIC-MS/MS and a uniformly 13C labeled internal standard. Anal Bioanal Chem 406(3):915–922. https://doi.org/10.1007/s00216-013-7456-2
Guo Y, Yao R, Wang Z, Zhang Y, Cui M, Zhao Q, Wang H (2017) Novel potential type electrochemical chiral recognition biosensor for amino acid. J Solid State Electrochem 22(1):41–49. https://doi.org/10.1007/s10008-017-3719-8
Gwatidzo L, Botha BM, McCrindle RI (2013) Determination of amino acid contents of manketti seeds (Schinziophyton rautanenii) by pre-column derivatisation with 6-aminoquinolyl-N-hydroxysuccinimidyl carbamate and RP-HPLC. Food Chem 141(3):2163–2169. https://doi.org/10.1016/j.foodchem.2013.04.101
Han M, Xie M, Han J, Yuan D, Yang T, Xie Y (2018) Development and validation of a rapid, selective, and sensitive LC-MS/MS method for simultaneous determination of D- and L-amino acids in human serum: application to the study of hepatocellular carcinoma. Anal Bioanal Chem 410(10):2517–2531. https://doi.org/10.1007/s00216-018-0883-3
Hayase F, Kato H, Fujimaki M (1975) Racemization of amino acid residues in proteins and poly (L-amino acids) during roasting. J Agric Food Chem 23(3):491–494. https://doi.org/10.1021/jf60199a055
Hess S (2012) A universal HPLC-MS method to determine the stereochemistry of common and unusual amino acids. Methods Mol Biol (Clifton, NJ) 828:63–75. https://doi.org/10.1007/978-1-61779-445-2_7
Hildebrandt TM, Nunes Nesi A, Araujo WL, Braun HP (2015) Amino acid catabolism in plants. Mol Plant 8(11):1563–1579. https://doi.org/10.1016/j.molp.2015.09.005
Hirayama A, Ikeda S, Sato A, Soga T (2019) Amino acid analysis by capillary electrophoresis-mass spectrometry. Methods Mol Biol (Clifton, NJ) 2030:307–313. https://doi.org/10.1007/978-1-4939-9639-1_23
How ZT, Busetti F, Linge KL, Kristiana I, Joll CA, Charrois JW (2014) Analysis of free amino acids in natural waters by liquid chromatography-tandem mass spectrometry. J Chromatogr A 1370:135–146. https://doi.org/10.1016/j.chroma.2014.10.040
Hsieh LY, Chan HH, Kuo PC, Hung HY, Li YC, Kuo CL, Peng Y, Zhao ZZ, Kuo DH, Sun IW, Wu TS (2018) A feasible and practical (1)H NMR analytical method for the quality control and quantification of bioactive principles in Lycii Fructus. J Food Drug Anal 26(3):1105–1112. https://doi.org/10.1016/j.jfda.2018.01.001
Jako T, Szabo E, Tabi T, Zachar G, Csillag A, Szoko E (2014) Chiral analysis of amino acid neurotransmitters and neuromodulators in mouse brain by CE-LIF. Electrophoresis 35(19):2870–2876. https://doi.org/10.1002/elps.201400224
Jeong JS, Kim SK, Park SR (2013) Amino acid analysis of dried blood spots for diagnosis of phenylketonuria using capillary electrophoresis-mass spectrometry equipped with a sheathless electrospray ionization interface. Anal Bioanal Chem 405(25):8063–8072. https://doi.org/10.1007/s00216-013-6999-6
Johnson MA, Vidoni S, Durigon R, Pearce SF, Rorbach J, He J, Brea-Calvo G, Minczuk M, Reyes A, Holt IJ, Spinazzola A (2014) Amino acid starvation has opposite effects on mitochondrial and cytosolic protein synthesis. PLoS ONE 9(4):e93597. https://doi.org/10.1371/journal.pone.0093597
Joyce R, Kuziene V, Zou X, Wang X, Pullen F, Loo RL (2016) Development and validation of an ultra-performance liquid chromatography quadrupole time of flight mass spectrometry method for rapid quantification of free amino acids in human urine. Amino Acids 48(1):219–234. https://doi.org/10.1007/s00726-015-2076-0
Kawana S, Nakagawa K, Hasegawa Y, Yamaguchi S (2010) Simple and rapid analytical method for detection of amino acids in blood using blood spot on filter paper, fast-GC/MS and isotope dilution technique. J Chromatogr B Anal Technol Biomed Life Sci 878(30):3113–3118. https://doi.org/10.1016/j.jchromb.2010.09.015
Kinoshita K, Jingu S, Yamaguchi J (2013) A surrogate analyte method to determine D-serine in mouse brain using liquid chromatography-tandem mass spectrometry. Anal Biochem 432(2):124–130. https://doi.org/10.1016/j.ab.2012.09.035
Klepacki J, Klawitter J, Klawitter J, Karimpour-Fard A, Thurman J, Ingle G, Patel D, Christians U (2016) Amino acids in a targeted versus a non-targeted metabolomics LC-MS/MS assay. Are the results consistent? Clin Biochem 49(13–14):955–961. https://doi.org/10.1016/j.clinbiochem.2016.06.002
Kowalski S, Kopuncova M, Ciesarova Z, Kukurova K (2017) Free amino acids profile of Polish and Slovak honeys based on LC-MS/MS method without the prior derivatisation. J Food Sci Technol 54(11):3716–3723. https://doi.org/10.1007/s13197-017-2838-7
Lamp A, Kaltschmitt M, Ludtke O (2018) Improved HPLC-method for estimation and correction of amino acid losses during hydrolysis of unknown samples. Anal Biochem 543:140–145. https://doi.org/10.1016/j.ab.2017.12.009
Le A, Ng A, Kwan T, Cusmano-Ozog K, Cowan TM (2014) A rapid, sensitive method for quantitative analysis of underivatized amino acids by liquid chromatography-tandem mass spectrometry (LC-MS/MS). J Chromatogr B Anal Technol Biomed Life Sci 944:166–174. https://doi.org/10.1016/j.jchromb.2013.11.017
Li G, Wu D, Xie W, Sha Y, Lin H, Liu B (2013) Analysis of amino acids in tobacco by derivatization and dispersive liquid-liquid microextraction based on solidification of floating organic droplet method. J Chromatogr A 1296:243–247. https://doi.org/10.1016/j.chroma.2013.03.076
Li Z, Xing Y, Guo X, Cui Y (2017) Development of an UPLC-MS/MS method for simultaneous quantitation of 11 d-amino acids in different regions of rat brain: Application to a study on the associations of d-amino acid concentration changes and Alzheimer's disease. J Chromatogr B Anal Technol Biomed Life Sci 1058:40–46. https://doi.org/10.1016/j.jchromb.2017.05.011
Liu M, Zhang X, Tan T (2016) The effect of amino acids on lipid production and nutrient removal by Rhodotorula glutinis cultivation in starch wastewater. Bioresour Technol 218:712–717. https://doi.org/10.1016/j.biortech.2016.07.027
Liyanaarachchi GVV, Mahanama KRR, Somasiri H, Punyasiri PAN (2018) Validation of a reversed-phase high-performance liquid chromatographic method for the determination of free amino acids in rice using l-theanine as the internal standard. Food Chem 240:196–203. https://doi.org/10.1016/j.foodchem.2017.07.038
Lopes R, Grutzner N, Berghoff N, Lidbury JA, Suchodolski JS, Steiner JM (2015) Analytic validation of a gas chromatography-mass spectrometry method for quantification of six amino acids in canine serum samples. Am J Vet Res 76(12):1014–1021. https://doi.org/10.2460/ajvr.76.12.1014
Lorenzo MP, Villasenor A, Ramamoorthy A, Garcia A (2013) Optimization and validation of a capillary electrophoresis laser-induced fluorescence method for amino acids determination in human plasma: application to bipolar disorder study. Electrophoresis 34(11):1701–1709. https://doi.org/10.1002/elps.201200632
Lu T, Olesik SV (2013) Electrospun polyvinyl alcohol ultra-thin layer chromatography of amino acids. J Chromatogr B Anal Technol Biomed Life Sci 912:98–104. https://doi.org/10.1016/j.jchromb.2012.10.037
Ma X, Zhao D, Li X, Meng L (2015) Chromatographic method for determination of the free amino acid content of chamomile flowers. Pharmacogn Mag 11(41):176–179. https://doi.org/10.4103/0973-1296.149735
Manninen H, Rotola-Pukkila M, Aisala H, Hopia A, Laaksonen T (2018) Free amino acids and 5'-nucleotides in Finnish forest mushrooms. Food Chem 247:23–28. https://doi.org/10.1016/j.foodchem.2017.12.014
Marouzi S, Sharifi Rad A, Beigoli S, Teimoori Baghaee P, Assaran Darban R, Chamani J (2017) Study on effect of lomefloxacin on human holo-transferrin in the presence of essential and nonessential amino acids: spectroscopic and molecular modeling approaches. Int J Biol Macromol 97:688–699. https://doi.org/10.1016/j.ijbiomac.2017.01.047
Martin A, Batalla P, Hernandez-Ferrer J, Martinez MT, Escarpa A (2015) Graphene oxide nanoribbon-based sensors for the simultaneous bio-electrochemical enantiomeric resolution and analysis of amino acid biomarkers. Biosens Bioelectron 68:163–167. https://doi.org/10.1016/j.bios.2014.12.030
Mirmiran P, Bahadoran Z, Ghasemi A, Azizi F (2017) Contribution of dietary amino acids composition to incidence of cardiovascular outcomes: A prospective population-based study. Nutr Metab Cardiovasc Dis 27(7):633–641. https://doi.org/10.1016/j.numecd.2017.05.003
Miyoshi Y, Koga R, Oyama T, Han H, Ueno K, Masuyama K, Itoh Y, Hamase K (2012) HPLC analysis of naturally occurring free D-amino acids in mammals. J Pharm Biomed Anal 69:42–49. https://doi.org/10.1016/j.jpba.2012.01.041
Miyoshi Y, Nagano M, Ishigo S, Ito Y, Hashiguchi K, Hishida N, Mita M, Lindner W, Hamase K (2014) Chiral amino acid analysis of Japanese traditional Kurozu and the developmental changes during earthenware jar fermentation processes. J Chromatogr, B: Anal Technol Biomed Life Sci 966:187–192. https://doi.org/10.1016/j.jchromb.2014.01.034
Mochizuki T, Todoroki K, Inoue K, Min JZ, Toyo'oka T (2014) Isotopic variants of light and heavy L-pyroglutamic acid succinimidyl esters as the derivatization reagents for DL-amino acid chiral metabolomics identification by liquid chromatography and electrospray ionization mass spectrometry. Anal Chim Acta 811:51–59. https://doi.org/10.1016/j.aca.2013.12.016
Moldovan RC, Bodoki E, Kacso T, Servais AC, Crommen J, Oprean R, Fillet M (2016) A micellar electrokinetic chromatography-mass spectrometry approach using in-capillary diastereomeric derivatization for fully automatized chiral analysis of amino acids. J Chromatogr A 1467:400–408. https://doi.org/10.1016/j.chroma.2016.08.035
Mondanelli G, Iacono A, Carvalho A, Orabona C, Volpi C, Pallotta MT, Matino D, Esposito S, Grohmann U (2019) Amino acid metabolism as drug target in autoimmune diseases. Autoimmun Rev 18(4):334–348. https://doi.org/10.1016/j.autrev.2019.02.004
Muller C, Fonseca JR, Rock TM, Krauss-Etschmann S, Schmitt-Kopplin P (2014) Enantioseparation and selective detection of D-amino acids by ultra-high-performance liquid chromatography/mass spectrometry in analysis of complex biological samples. J Chromatogr A 1324:109–114. https://doi.org/10.1016/j.chroma.2013.11.026
Munz E, Jakob PM, Borisjuk L (2016) The potential of nuclear magnetic resonance to track lipids in planta. Biochimie 130:97–108. https://doi.org/10.1016/j.biochi.2016.07.014
Nagasaki T, Koito T, Nemoto S, Ushio H, Inoue K (2017) Simultaneous analysis of free amino acids and taurine-related compounds in deep-sea mussel tissues using reversed-phase HPLC. Fish Sci 84(1):1–8
Nakano Y, Konya Y, Taniguchi M, Fukusaki E (2017) Development of a liquid chromatography-tandem mass spectrometry method for quantitative analysis of trace d-amino acids. J Biosci Bioeng 123(1):134–138. https://doi.org/10.1016/j.jbiosc.2016.07.008
Nguyen M, Fried B, Sherma J (2016) Effects of Echinostoma caproni miracidia dose on the amino acid contents of Biomphalaria glabrata as determined by high-performance thin-layer chromatography. Acta Parasitol 61(1):108–112. https://doi.org/10.1515/ap-2016-0014
Okahashi N, Kawana S, Iida J, Shimizu H, Matsuda F (2016) GC-MS/MS survey of collision-induced dissociation of tert-butyldimethylsilyl-derivatized amino acids and its application to (13)C-metabolic flux analysis of Escherichia coli central metabolism. Anal Bioanal Chem 408(22):6133–6140. https://doi.org/10.1007/s00216-016-9724-4
Okame R, Nakahara K, Murakami N (2015) Plasma amino acid profiles at various reproductive stages in female rats. J Vet Med Sci 77(7):815–821. https://doi.org/10.1292/jvms.15-0095
Oldekop M-L, Herodes K, Rebane R (2017a) Comparison of amino acid derivatization reagents for liquid chromatography atmospheric pressure chemical ionization mass spectrometric analysis of seven amino acids in tea extract. Int J Mass Spectrom 421:189–195. https://doi.org/10.1016/j.ijms.2017.07.004
Oldekop ML, Rebane R, Herodes K (2017b) Dependence of matrix effect on ionization polarity during LC-ESI-MS analysis of derivatized amino acids in some natural samples. Eur J Mass Spectrom (Chichester) 23(5):245–253. https://doi.org/10.1177/1469066717711026
Oosterink JE, Buijs N, van Goudoever JB, Schierbeek H (2014) A novel method for simultaneous measurement of concentration and enrichment of NO synthesis-specific amino acids in human plasma using stable isotopes and LC/MS ion trap analysis. J Chromatogr B Anal Technol Biomed Life Sci 958:10–15. https://doi.org/10.1016/j.jchromb.2014.03.005
Ota N, Rubakhin SS, Sweedler JV (2014) D-Alanine in the islets of Langerhans of rat pancreas. Biochem Biophys Res Commun 447(2):328–333. https://doi.org/10.1016/j.bbrc.2014.03.153
Ou G, Feng X, Du W, Liu X, Liu BF (2013) Recent advances in microchip electrophoresis for amino acid analysis. Anal Bioanal Chem 405(25):7907–7918. https://doi.org/10.1007/s00216-013-6830-4
Pretorius CJ, McWhinney BC, Sipinkoski B, Wilce A, Cox D, McWhinney A, Ungerer JPJ (2018) Rapid amino acid quantitation with pre-column derivatization; ultra-performance reverse phase liquid chromatography and single quadrupole mass spectrometry. Clin Chim Acta 478:132–139. https://doi.org/10.1016/j.cca.2017.12.027
Prior A, Moldovan RC, Crommen J, Servais AC, Fillet M, de Jong GJ, Somsen GW (2016) Enantioselective capillary electrophoresis-mass spectrometry of amino acids in cerebrospinal fluid using a chiral derivatizing agent and volatile surfactant. Anal Chim Acta 940:150–158. https://doi.org/10.1016/j.aca.2016.08.040
Qiu J, Wang J, Xu Z, Liu H, Ren J (2017) Quantitation of underivatized branched-chain amino acids in sport nutritional supplements by capillary electrophoresis with direct or indirect UV absorbance detection. PLoS ONE 12(6):e0179892. https://doi.org/10.1371/journal.pone.0179892
Rebane R, Oldekop ML, Herodes K (2012) Comparison of amino acid derivatization reagents for LC-ESI-MS analysis. Introducing a novel phosphazene-based derivatization reagent. J Chromatogr, B: Anal Technol Biomed Life Sci 904:99–106. https://doi.org/10.1016/j.jchromb.2012.07.029
Reddy BS, Chary VN, Pavankumar P, Prabhakar S (2016) Characterization of N-methylated amino acids by GC-MS after ethyl chloroformate derivatization. J Mass Spectrom 51(8):638–650. https://doi.org/10.1002/jms.3788
Redruello B, Ladero V, Del Rio B, Fernandez M, Martin MC, Alvarez MA (2017) A UHPLC method for the simultaneous analysis of biogenic amines, amino acids and ammonium ions in beer. Food Chem 217:117–124. https://doi.org/10.1016/j.foodchem.2016.08.040
Remelli M, Faccini S, Conato C (2014) Chiral ligand-exchange resolution of underivatized amino acids on a dynamically modified stationary phase for RP-HPTLC. Chirality 26(6):313–318. https://doi.org/10.1002/chir.22324
Rezazadeh M, Yamini Y, Seidi S, Aghaei A (2015) Pulsed electromembrane extraction for analysis of derivatized amino acids: a powerful technique for determination of animal source of gelatin samples. Talanta 136:190–197. https://doi.org/10.1016/j.talanta.2015.01.007
Rounova O, Demin P, Korotkov M, Malkova V, Ustinnikova O (2018) Development of a hydrophilic interaction high-performance liquid chromatography method for the determination of glycine in formulations of therapeutic immunoglobulins. Anal Bioanal Chem 410(26):6935–6942. https://doi.org/10.1007/s00216-018-1297-y
Rubino M, Milin S, D'Onofrio A, Signoret P, Hatte C, Balesdent J (2014) Measurement of delta13C values of soil amino acids by GC-C-IRMS using trimethylsilylation: a critical assessment. Isot Environ Health Stud 50(4):516–530. https://doi.org/10.1080/10256016.2014.959444
Sakaguchi Y, Kinumi T, Yamazaki T, Takatsu A (2015) A novel amino acid analysis method using derivatization of multiple functional groups followed by liquid chromatography/tandem mass spectrometry. Analyst 140(6):1965–1973. https://doi.org/10.1039/c4an01672f
Schiavone NM, Sarver SA, Sun L, Wojcik R, Dovichi NJ (2015) High speed capillary zone electrophoresis-mass spectrometry via an electrokinetically pumped sheath flow interface for rapid analysis of amino acids and a protein digest. J Chromatogr B Anal Technol Biomed Life Sci 991:53–58. https://doi.org/10.1016/j.jchromb.2015.04.001
Sharma G, Attri SV, Behra B, Bhisikar S, Kumar P, Tageja M, Sharda S, Singhi P, Singhi S (2014) Analysis of 26 amino acids in human plasma by HPLC using AQC as derivatizing agent and its application in metabolic laboratory. Amino Acids 46(5):1253–1263. https://doi.org/10.1007/s00726-014-1682-6
Sherwood RA (2000) Amino acid measurement in body fluids using PITC derivatives. Methods Mol Biol (Clifton, NJ) 159:169–175. https://doi.org/10.1385/1-59259-047-0:169
Shi X, Holland GP, Yarger JL (2013) Amino acid analysis of spider dragline silk using (1)H NMR. Anal Biochem 440(2):150–157. https://doi.org/10.1016/j.ab.2013.05.006
Shikha Ojha K, Granato D, Rajuria G, Barba FJ, Kerry JP, Tiwari BK (2018) Application of chemometrics to assess the influence of ultrasound frequency, Lactobacillus sakei culture and drying on beef jerky manufacture: Impact on amino acid profile, organic acids, texture and colour. Food Chem 239:544–550. https://doi.org/10.1016/j.foodchem.2017.06.124
Song Y, Funatsu T, Tsunoda M (2013) Amino acid analysis using core-shell particle column. J Chromatogr B Anal Technol Biomed Life Sci 927:214–217. https://doi.org/10.1016/j.jchromb.2012.09.005
Stocchi V, Palma F, Piccoli G, Biagiarelli B, Magnani M, Masat L, Cucchiarini L (1992) Analysis of amino acids as DABS-derivatives with a sensitivity to the femtomole level using RP-HPLC narrow-bore columns. Amino Acids 3(3):303–309. https://doi.org/10.1007/bf00806005
Szkudzińska K, Smutniak I, Rubaj J, Korol W, Bielecka G (2017) Method validation for determination of amino acids in feed by UPLC. Accred Qual Assur 22(5):247–252. https://doi.org/10.1007/s00769-017-1281-9
Tasakis RN, Touraki M (2018) Identification of bacteriocins secreted by the probiotic Lactococcus lactis following microwave-assisted acid hydrolysis (MAAH), amino acid content analysis, and bioinformatics. Anal Bioanal Chem 410(4):1299–1310. https://doi.org/10.1007/s00216-017-0770-3
Tian K, Wang YX, Li LX, Liu YQ (2018) Neuronal death/apoptosis induced by intracellular zinc deficiency associated with changes in amino-acid neurotransmitters and glutamate receptor subtypes. J Inorg Biochem 179:54–59. https://doi.org/10.1016/j.jinorgbio.2017.11.014
Tosti V, Bertozzi B, Fontana L (2018) Health benefits of the Mediterranean diet: metabolic and molecular mechanisms. J Gerontol A Biol Sci Med Sci 73(3):318–326. https://doi.org/10.1093/gerona/glx227
Toue S, Sugiura Y, Kubo A, Ohmura M, Karakawa S, Mizukoshi T, Yoneda J, Miyano H, Noguchi Y, Kobayashi T, Kabe Y, Suematsu M (2014) Microscopic imaging mass spectrometry assisted by on-tissue chemical derivatization for visualizing multiple amino acids in human colon cancer xenografts. Proteomics 14(7–8):810–819. https://doi.org/10.1002/pmic.201300041
Tsai TH, Wang M, Ressom HW (2016) Preprocessing and analysis of LC-MS-based proteomic data. Methods in molecular biology (Clifton, NJ) 1362:63–76. https://doi.org/10.1007/978-1-4939-3106-4_3
Ubhi BK, Davenport PW, Welch M, Riley J, Griffin JL, Connor SC (2013) Analysis of chloroformate-derivatised amino acids, dipeptides and polyamines by LC-MS/MS. J Chromatogr, B: Anal Technol Biomed Life Sci 934:79–88. https://doi.org/10.1016/j.jchromb.2013.06.026
Uri C, Juhász Z, Polgár Z, Bánfalvi Z (2014) A GC–MS-based metabolomics study on the tubers of commercial potato cultivars upon storage. Food Chem 159:287–292. https://doi.org/10.1016/j.foodchem.2014.03.010
Vancompernolle B, Croes K, Angenon G (2016) Optimization of a gas chromatography-mass spectrometry method with methyl chloroformate derivatization for quantification of amino acids in plant tissue. J Chromatogr B Anal Technol Biomed Life Sci 1017–1018:241–249. https://doi.org/10.1016/j.jchromb.2016.02.020
Veiga P, Piquet S, Maisons A, Furlan S, Courtin P, Chapot-Chartier MP, Kulakauskas S (2006) Identification of an essential gene responsible for D-Asp incorporation in the Lactococcus lactis peptidoglycan crossbridge. Mol Microbiol 62(6):1713–1724. https://doi.org/10.1111/j.1365-2958.2006.05474.x
Waldhier MC, Almstetter MF, Nurnberger N, Gruber MA, Dettmer K, Oefner PJ (2011) Improved enantiomer resolution and quantification of free D-amino acids in serum and urine by comprehensive two-dimensional gas chromatography-time-of-flight mass spectrometry. J Chromatogr A 1218(28):4537–4544. https://doi.org/10.1016/j.chroma.2011.05.039
Wang H, McNeil YR, Yeo TW, Anstey NM (2013) Simultaneous determination of multiple amino acids in plasma in critical illness by high performance liquid chromatography with ultraviolet and fluorescence detection. J Chromatogr B Anal Technol Biomed Life Sci 940:53–58. https://doi.org/10.1016/j.jchromb.2013.09.016
Wang W, Ma L, Yao F, Lin X, Xu K (2015) High-speed separation and detection of amino acids in laver using a short capillary electrophoresis system. Electrophoresis 36(2):335–340. https://doi.org/10.1002/elps.201400246
Wang F, Gong W, Wang L, Chen Z (2016) Selective recognition of D-tryptophan from d/l-tryptophan mixtures in the presence of Cu(II) by electropolymerized L-lysine film. Anal Biochem 492:30–33. https://doi.org/10.1016/j.ab.2015.09.002
Weatherly CA, Du S, Parpia C, Santos PT, Hartman AL, Armstrong DW (2017) d-Amino acid levels in perfused mouse brain tissue and blood: a comparative study. ACS Chem Neurosci 8(6):1251–1261. https://doi.org/10.1021/acschemneuro.6b00398
Woo HI, Chun MR, Yang JS, Lim SW, Kim MJ, Kim SW, Myung WJ, Kim DK, Lee SY (2015) Plasma amino acid profiling in major depressive disorder treated with selective serotonin reuptake inhibitors. CNS Neurosci Ther 21(5):417–424. https://doi.org/10.1111/cns.12372
Wu X, Si N, Bo G, Hu H, Yang J, Bian B, Zhao HY, Wang H (2015) Characterization and quantitative amino acids analysis of analgesic peptides in cinobufacini injection by size exclusion chromatography, matrix-assisted laser desorption/ionization time of flight mass spectrometry and gas chromatography mass spectrometry. Biomed Chromatogr 29(1):138–147. https://doi.org/10.1002/bmc.3250
Xia Q, Huang Y, Lin X, Zhu S, Fu Y (2016a) Highly sensitive d-alanine electrochemical biosensor based on functionalized multi-walled carbon nanotubes and d-amino acid oxidase. Biochem Eng J 113:1–6. https://doi.org/10.1016/j.bej.2016.05.003
Xia T, Gao S, Shu C, Wen Y, Yun Y, Tao X, Chen W, Zhang F (2016b) Analysis of amino acids in human blood using UHPLC-MS/MS: potential interferences of storage time and vacutainer tube in pre-analytical procedure. Clin Biochem 49(18):1372–1378. https://doi.org/10.1016/j.clinbiochem.2016.09.018
Xie Y, Alexander GM, Schwartzman RJ, Singh N, Torjman MC, Goldberg ME, Wainer IW, Moaddel R (2014) Development and validation of a sensitive LC-MS/MS method for the determination of D-serine in human plasma. J Pharm Biomed Anal 89:1–5. https://doi.org/10.1016/j.jpba.2013.10.028
Yang R, Dong J, Guo H, Li H, Wang S, Zhao H, Zhou W, Yu S, Wang M, Chen W (2013) Rapid and precise measurement of serum branched-chain and aromatic amino acids by isotope dilution liquid chromatography tandem mass spectrometry. PLoS ONE 8(12):e81144. https://doi.org/10.1371/journal.pone.0081144
Yang Y, Fan TW, Lane AN, Higashi RM (2017) Chloroformate derivatization for tracing the fate of amino acids in cells and tissues by multiple stable isotope resolved metabolomics (mSIRM). Anal Chim Acta 976:63–73. https://doi.org/10.1016/j.aca.2017.04.014
Young VR, Pellett PL (1987) Protein intake and requirements with reference to diet and health. Am J Clin Nutr 45(5 Suppl):1323–1343. https://doi.org/10.1093/ajcn/45.5.1323
Yousefinejad S, Honarasa F, Saeed N (2015) Quantitative structure-retardation factor relationship of protein amino acids in different solvent mixtures for normal-phase thin-layer chromatography. J Sep Sci 38(10):1771–1776. https://doi.org/10.1002/jssc.201401427
Yuan B, Ding Y, Kamal GM, Shao L, Zhou Z, Jiang B, Sun P, Zhang X, Liu M (2017) Reconstructing diffusion ordered NMR spectroscopy by simultaneous inversion of Laplace transform. J Magn Reson 278:1–7. https://doi.org/10.1016/j.jmr.2017.03.004
Yuan B, Wu H, Sanders T, McCullum C, Zheng Y, Tchounwou PB, Liu YM (2011) Chiral capillary electrophoresis-mass spectrometry of 3,4-dihydroxyphenylalanine: evidence for its enantioselective metabolism in PC-12 nerve cells. Anal Biochem 416(2):191–195. https://doi.org/10.1016/j.ab.2011.05.025
Zeng Y, Cai W, Shao X (2015) Quantitative analysis of 17 amino acids in tobacco leaves using an amino acid analyzer and chemometric resolution. J Sep Sci 38(12):2053–2058. https://doi.org/10.1002/jssc.201500090
Zhang L, Wang G, Xiong C, Zheng L, He J, Ding Y, Lu H, Zhang G, Cho K, Qiu L (2018) Chirality detection of amino acid enantiomers by organic electrochemical transistor. Biosens Bioelectron 105:121–128. https://doi.org/10.1016/j.bios.2018.01.035
Zhao Q, Cao Y, Wang Y, Hu C, Hu A, Ruan L, Bo Q, Liu Q, Chen W, Tao F, Ren M, Ge Y, Chen A, Li L (2014) Plasma and tissue free amino acid profiles and their concentration correlation in patients with lung cancer. Asia Pac J Clin Nutr 23(3):429–436. https://doi.org/10.6133/apjcn.2014.23.3.13
Zhou X, Shi Y (2013) Determination of amino acids in honey by capillary electrophoresis with indirect ultraviolet detection. Se Pu 31(7):661–666
Ziegler J, Abel S (2014) Analysis of amino acids by HPLC/electrospray negative ion tandem mass spectrometry using 9-fluorenylmethoxycarbonyl chloride (Fmoc-Cl) derivatization. Amino Acids 46(12):2799–2808. https://doi.org/10.1007/s00726-014-1837-5
Zor E, Hatay Patir I, Bingol H, Ersoz M (2013) An electrochemical biosensor based on human serum albumin/graphene oxide/3-aminopropyltriethoxysilane modified ITO electrode for the enantioselective discrimination of D- and L-tryptophan. Biosens Bioelectron 42:321–325. https://doi.org/10.1016/j.bios.2012.10.068
Acknowledgements
The work was funded by the program of Shanghai Committee of Science and Technology in China (Grant number 17401902300).
Author information
Authors and Affiliations
Contributions
N Zhang and B Wang had the idea for the article, CC Zhong and CP Zou performed the literature search and data analysis, and WH Xu drafted the work. All authors read and approved the review.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Informed consent
Obtaining informed consent for this type of study is not required.
Research involving human participants and/or animals
This article reviews published studies and does not require either the approval of animal use or human consent.
Additional information
Handling editor: J. D. Wade.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Xu, W., Zhong, C., Zou, C. et al. Analytical methods for amino acid determination in organisms. Amino Acids 52, 1071–1088 (2020). https://doi.org/10.1007/s00726-020-02884-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-020-02884-7