Abstract
The binding of [Ru(H2O)2(bpydip)]2+/3+ (bpydip = N,N′-bis[1-(2-pyridinyl)ethylidene]-1,3-propanediamine) to human serum albumin was investigated by spectroscopic techniques. The hydrophobic interaction between ruthenium(II) complex and HSA was spontaneous with the binding constants calculated to be 1.27 × 103 dm3 mol−1 (298 K), 7.74 × 103 dm3 mol−1 (303 K), and 9.46 × 103 dm3 mol−1 (308 K). The binding of ruthenium(III) complex to HSA induced the hybrid quenching mechanism with the binding constants estimated to be 1.25 × 109 dm3 mol−1 (298 K), 1.31 × 109 dm3 mol−1 (303 K), and 3.57 × 109 dm3 mol−1 (308 K). Thermodynamic parameters (ΔH = 76 kJ mol−1; ΔS = 515 J mol−1 K−1) indicated that hydrophobic forces played a major role in the binding of ruthenium(III) complex to HSA. In addition, the number of binding sites (n) for ruthenium(II) complex was approximately 1, whereas ruthenium(III) complex recorded 2. CD data indicated that the secondary structure of HSA is not changed in the presence of complexes.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
With advances in the development of new metallodrugs with antitumor functions, the platinum drugs, which are used in treatment today, are gradually being replaced by other ones which are more efficient and less toxic [1–6]. Among these new drugs, those with ruthenium as a central metal are the most promising, thanks to their biocompatibility, rapid elimination by the body, and lower toxicity [7–10]. Some compounds based on ruthenium show a promising impact on curing cancer. For example, cis-[Ru(DMSO)4Cl2] and cis-[Ru(NH3)4Cl2] are active against the P388 leukemia [11, 12], whereas binuclear ruthenium(II) complexes exhibit activity against cervical cancer and gastric cancer tumour cell lines [13]. Two ruthenium based drugs NAMI-A and KP1019 are already in the advanced stages of approval for medical use [14–16].
The mechanism by which ruthenium drugs act in tumor treatment was investigated and originally described by Kelman et al. [17], and is known as the action by reduction mechanism. Recently, the redox chemistry of ruthenium has been studied as an aspect of the mechanism of action of metallodrugs [18–22].
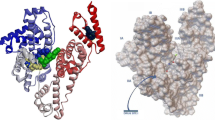
To reach the zone of action, drugs need to be bound to carrier systems, traveling through the body without any undesired reactions on the way. Many drug delivery systems have been developed with this characteristic, like liposomes, surfactant mediums, and modified proteins, for example [23–26]. However, in the mammalian body, there are serum albumin proteins, which have the ability to distribute all of these substances throughout the body [27, 28]. In the human body, human serum albumin (HSA) performs this role. With 585 amino acid residues, separated into 3 different subdomains (I, II, and III), each of which in turn is separated into another two (A and B), HSA differs from the other forms of mammalian albumin thanks to the presence of only one tryptophan residue (Trp214), allocated in subdomain IIA [29, 30]. This specific residue is the main factor responsible for the intrinsic fluorescence of this protein, allied with the lower fluorescent intensity of tyrosine residues spread throughout the structure.
According to Sudlow et al. [31, 32], two main binding sites to endogenous and exogenous substances can be found in the HSA structure; these are known as Sudlow’s site I (subdomain IIA and a small part of subdomain IIB) and Sudlow’s site II (subdomain IIIA). Subsequent studies revealed that most drugs have a preference for binding to site I, thanks to its malleability, hydrophobic framework, and diversity of amino acids [33]. Thus, fluorescence quencher studies using drugs with a known binding site (site markers) compared with new ones can provide a better understanding of the location of this new species in the albumin framework and, consequently, understand its body distribution behavior until the drug action zone [34, 35].
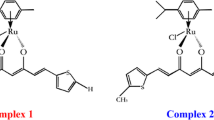
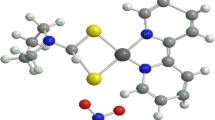
In this work, we used fluorescence studies and knowledge of the HSA structure to determine the binding behavior of a new ruthenium–Schiff base complex with protein and influence of the metal centre oxidation state on this interaction. The complex chosen was Ru(H2O)2(bpydip)]Cl2 (bpydip = N,N′-bis[1-(2-pyridinyl)ethylidene]-1,3-propanediamine), illustrated in Fig. 1. The binding Schiff base metal complexes and serum albumin has been the subject of extensive research in recent years because of its implications in numerous physiological functions [36–42]. Furthermore, it has been shown that linkage of Schiff base metal complexes to serum albumin increases the antioxidant capacity of the protein [43–45].
Results and discussion
Fluorescence quenching studies
HSA contain tryptophan, tyrosine, and phenylalanine residues which contribute to their intrinsic fluorescence [46]. When the excitation wavelength is 295 nm, tyrosine and phenylalanine residues are not excited, and only tryptophan emission is seen [47]. However, the fluorescence of HSA may be reduced or quenched as a result of molecular rearrangements, energy transfer, ground-state complex formation, or collisional quenching [48].
Two types of mechanisms for fluorescence quenching have been recognized [48]: (a) dynamic quenching, where the fluorophore groups, during their excited state lifetimes, are able to collide with quenching molecules; (b) static quenching, where there is a ground-state complex formation between the fluorophore and the quencher. The dynamic and static mechanisms can be distinguished by their different dependences on temperature and viscosity. Since higher temperatures favor larger diffusion coefficients, the dynamic quenching constants are expected to increase with increasing temperature. On the other hand, increased temperature is most likely to result in the decreased stability of the ground-state complex, and thus smaller values for the static quenching constants [48].
The dependence of the fluorescence intensity on quencher concentration can be determined by use of Stern–Volmer equation:
where F 0 and F is the fluorescence intensity in the absence and presence of ruthenium complex, respectively. K sv is the Stern–Volmer quenching constant, and [Q] is the concentration of ruthenium complex [49]. Thus, changes in intrinsic fluorescence of HSA as a function of the quencher concentration can be used to measure accessibility of small molecules in the protein [47, 50]
As shown in Fig. 2, the fluorescence spectrum of HSA in the absence of ruthenium(II) complex has emission band at 345 nm, when excited at 295 nm (curve a), and the fluorescence intensity of HSA decreases gradually with the increasing concentration of ruthenium(II) complex. In the inset of Fig. 2, Stern–Volmer plots for interaction of ruthenium(II) complex with HSA at 298, 303, and 308 K are shown. In three temperatures the plots are linear, which indicate that only one type of quenching occurs (dynamic or static quenching) [48]. The values of Stern–Volmer constant decrease lightly with increase in the temperature (Table 1), which suggest that quenching is due to the formation of a nonfluorescent complex between the HSA and ruthenium(II) derivative, i.e. static quenching [48, 51]. For ruthenium complexes, the most likely mechanism of protein quenching is due to the formation of a ground-state charge-transfer complex, where the metal center is connected by a tryptophan residue [52, 53].
Therefore, the experimental data were reanalyzed using modified Stern–Volmer equation:
where ΔF = F 0 − F, f a is the fraction of accessible fluorophores, and K a is the quenching constant of the accessible fraction [48].
The modified Stern–Volmer plots are shown in Fig. 3. In all temperatures the plots are linear and yield 1/(f a K a) as their slope. 1/f a can be obtained from the intercept of the plots of F 0/ΔF vs. 1/[Q]. As seen in Table 1, the quenching constant (K sv and K a) from each method are in good agreement. The differences in quenching constants determined by Eqs. (1) and (2) reflect the accessibility of ruthenium(II) complex in the protein. In this case, the fraction of the initial fluorescence which is accessible to the ruthenium(II) complex (f a) is nearly 90 %. For ruthenium(III) complex, Fig. 4, the fluorescence intensity of HSA also decreases gradually with the increasing concentration of complex.
Following a similar approach to the of ruthenium(II) complex, Stern–Volmer plots for interaction of ruthenium(III) complex with HSA at 298, 303, and 308 K can be obtained from expression of Eq. (1). The Stern–Volmer plots for quenching by Ru(III) complex show clear upward curvature (inset Fig. 4). The positive deviations from linearity indicates that both static and dynamic quenching occur for the same fluorophore. According to Lakowicz [48], the fluorescence decay can be interpreted in terms of a sphere of action static quenching model (or hybrid quenching mechanism). This quenching mechanism can be observed when the quencher species brings a sphere of action (e.g., a solvation shell) that can primarily approach the fluorophore to form the ground-state complex, which is characteristic in static quench. This nonfluorescent complex (or dark complex) can be undone and the fluorophore decays by collisional mechanism [48].
Thus, in order to obtain the values of quenching constant for interaction of ruthenium(III) complex with HSA, a new mathematical approach is necessary [54, 55]. Equation (3) reflects the Stern–Volmer equation with the quench sphere of action consideration, where w is the ratio of HSA molecules quenched by the collisional mechanism, (1 − w) is the contribution of protein molecules that are immediately quenched by the complex sphere of action, and K app is apparent quenching constant.
The modified Stern–Volmer plots are shown in Fig. 5. Modified Stern–Volmer quenching constants for ruthenium(III) complex are listed in Table 2.
The differences in fluorescence quenching measured in the ruthenium(II) and ruthenium(III) systems probably are consequences of the fluctuations in the solvation shell surrounding the complex after the redox process and the effects of charge on the quencher. Ruthenium(III) is a hard acid and it is less polarizable than ruthenium(II), and is less richly covalent [56]. The change in the oxidation state of the metallic center causes a rearrangement of the Schiff base ligand, steric repulsion between pyridine rings on the Schiff base and structural changes in the solvent shell [57–59]. The electronic, structural rearrangement, and steric effects facilitate the diffusion of ruthenium(III) complex by protein structure and, consequently, leads to a strong quenching of tryptophan fluorescence.
Binding constants and thermodynamic parameters also are affected by environment surrounding of ruthenium complex (solvation shell) and charge of metallic center, as can be seen below.
Binding constants and binding sites
In order to completely understand the ruthenium complex-albumin interaction, we determined the apparent binding constant (K b) and the number of binding sites (n). These parameters indicate the affinity between ruthenium complex and HSA and are important for developing new medicinal products [60, 61]. To obtain the binding parameters, we made use of the equation:
As shown in Table 3, the K b value for ruthenium(II) complex varies around 103 dm3 mol−1, whereas ruthenium(III) complex record 109 dm3 mol−1. In addition, the values of n for ruthenium(II) complex are approximately 1, whereas ruthenium(III) complex record 2. This difference can be attributed to the preference of amino acids residues, which are better electron donor than water, to bind and stabilize ruthenium(III) complex. The strongest interaction between ruthenium(III) complex with HSA is consistent with the likely electronic, structural rearrangement, and steric effects that accompany the change in the oxidation state of the metallic center.
Thermodynamic studies
Different types of intermolecular forces contribute to stabilize the interaction of complexes with biomolecules, such as hydrogen bonds, electrostatic forces, and hydrophobic interactions [62]. In this work, we investigated the nature of the interactions between ruthenium(II/III) complex and HSA based on the van’t Hoff equation [47, 62], where ΔH is the enthalpy change, ΔS is the entropy change and R is the gas constant (8.314 J mol−1 K−1).
To calculate the free energy (ΔG), the following equation was used:
As shown in Table 3, the interaction of ruthenium(II) complex with HSA is a spontaneous process in which the Gibbs free energy change is negative, and is accompanied by positive enthalpy and entropy changes. The changes in the values of the Gibbs energy, enthalpy, and entropy that accompany the formation of the ruthenium(III)—HSA adduct can be explained on the grounds that as amino acids are stronger σ-donors than water, it increases the electron density at the ruthenium and hence facilitates the formation of adduct. According to Ross and Subramanian [62], positive values for ΔH and ΔS suggest that the insertion process of the compound on the HSA is determined by hydrophobic interactions [46].
The adduct formation leads to minimal reorganization of the proteic microenvironment, and exposes hydrophobic moieties of HSA to the solvent, maximizing the hydrophobic effect and the stabilization of the ruthenium—HSA adducts [63].
Circular dichroism
The CD technique was used to elucidate possible changes of the secondary structure of the protein promoted by binding of the ruthenium complex to HSA [64]. In Fig. 6, we display the CD spectrum of HSA that shows two negative bands at 208 and 220 nm related to the α-helical secondary structure [52]. The CD spectra of HSA in the presence of a five times higher ruthenium complex concentration showed no significant difference between the bands.
We estimate the α-helical residues according to the following Eq. (7) where θ is determined by Eq. (8) and θ obs is the emission observed in millidegrees, n the number of amino acids in the HSA structure, l the path length, and Cp the molar concentration of protein [65].
For both complex interactions, we observed a reduction of the α-helix ratio from 53.6 % in free HSA to 53.4 % in the HSA-ruthenium complex (molar ratio 1:0.5), 49.4 % (at 1:2.5), and 43.0 % (at 1:5).
As there is no significant modification between the ratio values, we suggest that an association between HSA and ruthenium complexes does not significantly alter the secondary structure of the protein; this is quite similar to that observed in the nickel Schiff base complex [66].
Conclusion
We studied the interaction of the same ruthenium–Schiff base complex with HSA in two different oxidation states (RuII and RuIII). Both complexes revealed a good interaction with the protein, acting as quencher to the intrinsic albumin fluorescence. The RuII complex quenches the protein fluorescence by a static mechanism, while the RuIII complex reveals a hybrid quenching mechanism (static + dynamic). The complexes bind to HSA with different intensities. The K b value for the RuII complex is in the order of 103 dm3 mol−1, while for RuIII, the order is about 109 dm3 mol−1. Positive values of ΔG reveal a spontaneous interaction of both complexes of protein, and the values of ΔS and ΔH allow us to infer that both interact with proteins by electrostatic interactions. Moreover, neither of the complexes changes the conformation of the protein, as shown by CD spectra. Thus, this work shows that modification of the oxidation state of the center metal in ruthenium complexes can influence the way this complex interacts with HSA and must be considered in the development of new metallodrugs, which will bind to this protein for transportation throughout the body.
Experimental
All reagents used in this work were of analytical grade and used without further purification. Human serum albumin (HSA, ≥96 % lyophilized powder, A1653), RuCl3·nH2O, 2-acetylpyridine, and 1,3-diaminopropane were purchased from Sigma-Aldrich. All compounds were dissolved in Tris–HCl buffer solution (0.05 mol dm−3 Tris, 0.10 mol dm−3 NaCl, pH 7.4). The pH values were checked with a suitably standardized pH meter. HSA solutions were prepared based on a molecular mass of 66,500 g mol−1. The doubly purified water used in all experiments was from a Millipore® system.
Synthesis and characterization of the ruthenium Schiff base complex, [RuCl2(bpydip)], were reported by de Souza et al. [67]. IR and 1H NMR spectra were found to be identical with the ones described in the literature [67]. When [RuCl2(bpydip)] was dissolved in water, it underwent an aquation reaction, where two chloride ligands were replaced by two water molecules, generating a product with the formula [Ru(H2O)2(bpydip)]Cl2. 1H NMR and UV–Vis spectra were found to be identical with the ones described in the literature [67]. [RuII(H2O)2(bpydip)]2+ was converted to a [RuIII(H2O)2(bpydip)]3+ upon electrooxidation at 0.45 V vs NHE. UV–Vis spectra were found to be identical with the ones described in the literature [67]. The electrolytic oxidation of [RuII(H2O)2(bpydip)]2+ was carried out using a PARC potentiostat, model 173. A platinum disc working electrode, a platinum wire auxiliary electrode and Ag/AgNO3 reference electrode were employed for the measurements.
Fluorescence analysis
We performed all of the steady decay fluorescence analysis using a spectrofluorimeter model Cary Eclipse by Varian. The emission and excitation slit was fixed at 5 for all analysis. Preliminary tests allowed determination of the HSA concentration in 1.4 × 10−5 mol dm−3, and 0.7–8.6 × 10−5 mol dm−3 for ruthenium complex titration. The titration process and analysis was performed in a 1.00 quartz cell, with the additions of a complex solution and a 3 min wait between the addition and obtaining the spectrum. We performed all of the analysis at temperatures of 298, 303, and 308 K, at an excitation wavelength of 295 nm, for both complex forms. The spectra range was from 300 to 540 nm and emission maximum was established at 340 nm. Using these values and the specific equations, we obtained the K app, K Sv, w, K b, and n values. To obtain the ΔH, ΔS, and ΔG values was used the Vant’ Hoff equation method.
The fluorescence data were adjusted for inner filter effect using Eq. (9), where F cor and F obs are fluorescence intensity corrected and observed, respectively, and A ex and A em are the absorption and emission wavelength, respectively [48].
Circular dichroism
We performed the analyses in a J-810 Spectropolarimeter, in the absence and presence of ruthenium complex and recorded in the range 200–260 nm min−1. In all analyses, we used 0.1 cm quartz cells of and titration was made in molar ratios of 1:0, 1:0.5, 1:2.5, and 1:5, HSA:ruthenium complex.
References
Tiekink ER, Gielen M (eds) (2005) Metallotherapeutic drugs and metal-based diagnostic agents—the use of metals in medicine. Wiley, Hoboken
Kostova I (2014) Recent Pat Anticancer Drug Discov 1:1
Banti CN, Kyros L, Geromichalos GD, Kourkoumelis N, Kubicki M, Hadjikakou SK (2014) Eur J Med Chem 77:388
Cutillas N, Yellol GS, Haro C, Vicente CK, Rodrigues V, Ruiz J (2013) Coord Chem Rev 257:2784
Novio F, Simmchen J, Vázquez-Mera N, Amorin-Ferré L, Ruiz-Molina D (2013) Coord Chem Rev 257:2839
Maldonado CR, Salassa L, Gomez-Blanco N, Mareque-Rivas JC (2013) Coord Chem Rev 257:2668
Leijen S, Burgers SA, Baas P, Pluim D, Tibben M, van Werkhoven E, Alessio E, Sava G, Beijnen JH, Schellens JHM (2015) Invest New Drugs 33:201
Clarke MJ (2003) Coord Chem Rev 236:209
Barry NPE, Sadler PJ (2012) Chem Soc Rev 41:3264
Joshi T, Pierroz V, Mari C, Gemperle L, Ferrari S, Gasser G (2014) Angew Chem Int Ed 53:2960
Coluccia M, Sava G, Loseto F, Nassi A, Boccarelli A, Giordano D, Alessio E, Mestroni G (1993) Eur J Cancer 29A:1873
Bergamo A, Gaiddon C, Schellens JH, Beijnen JH, Sava G (2012) J Inorg Biochem 106:90
Zang Y, Lai L, Cai P, Cheng GZ, Xu XM, Liu Y (2015) New J Chem 39:5805
Alessio E, Mestroni G, Bergamo A, Sava G (2004) Curr Top Med Chem 4:1525
Cocchietto M, Zajac J, Brabec V, Mestroni G, Sava G (2015) Dalton Trans 44:1905
Blunden BM, Stenzel MH (2015) J Chem Technol Biotechnol 90:1177
Kelman AD, Clarke MJ, Edmonds SD, Peresie HJ (1977) J Clin Hematol Oncol 7:274
Reisner E, Arion VB, Silva MFCG, Lichtenecker R, Eichinger A, Keppler BK, Kukushkin VY, Pombeiro AJL (2004) Inorg Chem 43:7083
Webb MI, Chard RA, Al-Jobory YM, Jones MR, Wong EWY, Walsby CJ (2012) Inorg Chem 51:954
Graf N, Lippard SJ (2012) Adv Drug Deliv Rev 64:993
Lima AP, Pereira FC, Almeida MAP, Mello FMS, Pires WC, Pinto TM, Delella FK, Felisbino SL, Moreno V, Batista AA, Lacerda EPS (2014) PLoS ONE. doi:10.1371/journal.pone.0105865
Adigun RA, Martincigh B, Nyamori VO, Omondi B, Masimirembwa C, Simoyi RH (2014) Dalton Trans 43:12943
Ye H, Karim AA, Loh XJ (2014) Mat Sci Eng C-Biomim 45:609
Liu J, Huang Y, Kumar A, Tan A, Jin S, Mozhi A, Liang XJ (2014) Biotechnol Adv 32:693
Choonara BF, Choonara YE, Kumar P, Bijukumar D, Toit LC, Pillay V (2014) Biotechnol Adv 32:1269
Estevão BM, Pellosi DS, Freitas CF, Vanzin D, Franciscato DS, Caetano W, Hioka N (2014) J Photochem Photobiol, A 287:30
Peters JT, Stewart AJ (2013) Biochim Biophys Acta 1830:5351
Sleep D, Cameron J, Evans LR (2013) Biochim Biophys Acta 1830:5526
He XM, Carter DC (1992) Nature 358:209
Carter D, Ho JX (1994) Adv Protein Chem 45:153
Sudlow G, Birkett DJ, Wade DN (1975) Mol Pharmacol 11:824
Sudlow G, Birkett DJ, Wade DN (1976) Mol Pharmacol 12:1052
Yamasaki K, Chuang VTG, Maruyama T, Otagiri M (2013) Biochim Biophys Acta 1830:5435
Anand U, Mukherjee S (2013) Biochim Biophys Acta 1830:5394
Dömötör O, Hartinger CG, Bytzek AK, Kiss T, Keppler BK, Enyedy EA (2013) J Biol Inorg Chem 18:9
Li P, Niu MJ, Hong M, Cheng S, Dou JM (2014) J Inorg Biochem 137:101
Fani N, Bordbar AK, Ghayeb Y (2013) J Lumin 141:166
Chen Z, Zhang J, Liu C (2013) Biometals 26:827
Yang L, Wang B, Tan J, Zhu L (2013) Mini-Rev Med Chem 13:1487
Ali I, Haque A, Saleem K, Hsieh MF (2013) Bioorg Med Chem 21:3808
Noureen A, Saleem S, Fatima T, Siddiqi H, Mirza B (2013) Pak J Pharm Sci 26:113
Moorthy NS, Hari N, Cerqueira NS, Ramos MJ, Fernandes PA (2012) Med Chem Res 21:133
Kostova I, Saso L (2013) Curr Med Chem 20:4609
Ray A, Seth BK, Pal U, Basu S (2012) Spectrochim Acta A Mol Biomol Spectrosc 92:164
Wang RM, Mao JJ, Song JF, Huo CX, He YF (2007) Chin Chem Lett 18:1416
Peters T (1996) All About Albumin: Biochemistry. Academic Press, San Diego, USA, Genetics and Medicinal Applications
Naveenraj S, Anandan S (2013) J Photochem Photobiol, C 14:53
Lakowicz JR (2006) Principles of Fluorescence Spectroscopy, 3rd edn. Springer Science, New York
Santos RLSR, Sanches RNF, Silva DD (2015) J Coord Chem 68:3209
Giralt E, Peczuh MW, Salvatella X (2011) Protein Surface Recognition: Approaches for Drug Discovery. John Wiley & Sons Ltd, Chichester
Sun J, Huang Y, Zheng C, Zhou Y, Liu Y, Liu J (2015) Biol Trace Elem Res 163:266
Trynda-Lemiesz L, Karaczyn A, Keppler BK, Kozlowski H (2000) J Inorg Biochem 78:341
Trynda-Lemiesz L, Kozlowski H, Katsaros N (2000) Metal Based Drugs 7:293
Kadadevarmath JS, Malimath GH, Melavanki RM, Patil NR (2014) Spectrochim Acta A Mol Biomol Spectrosc 117:630
Geethanjali HS, Nagaraja D, Melavanki RM (2015) J Mol Liq 209:669
Seddon EA, Seddon KR (1984) The Chemistry of Ruthenium. Elsevier, New York
Vigato PA, Tamburini S (2004) Coord Chem Rev 248:1717
Hui JW-S, Wong W-T (1996) Coord Chem Rev 172:389
Richmond MG (1995) Coord Chem Rev 141:63
Fanali G, di Masi A, Trezza V, Marino M, Fasano M, Ascenzi P (2012) Mol Aspects Med 33:209
Bal W, Sokolowska M, Kurowska E, Faller P (2013) Biochem Biophys Acta 1830:5444
Ross P, Subramanian S (1981) Biochemistry 20:3091
Raicu V, Popescu A (2008) Integrated Molecular and Cellular Biophysics. Springer Science, New York
Fasman GD (1996) Circular Dichroism and the Conformational Analysis of Biomolecules. Springer Science, New York
Daneshgar P, Moosavi-Movahedi A, Norouzi P, Ganjali M, Madadkar-Obhani A, Saboury A (2009) Int J Biol Macromol 45:129
Ray A, Seth B, Pal U, Basu S (2012) Spectrochim Acta A Mol Biomol Spectrosc 92:164
Souza VR, Nunes GS, Rocha CR, Toma HE (2002) Inorg Chim Acta 348:50
Acknowledgments
We are grateful to CNPq for financial support.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Franciscato, D.S., de Souza, V.R. Influence of the oxidation state of the metal center on the interaction of ruthenium complex with HSA. Monatsh Chem 147, 1315–1321 (2016). https://doi.org/10.1007/s00706-016-1659-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00706-016-1659-0