Abstract
During the 2019 winter wheat season, symptoms of severe chlorosis and stunting were observed on wheat in the irrigation production areas of South Africa. RNAtag-seq data were generated for seven samples from KwaZulu-Natal province and one from Limpopo. Analysis of assembled contigs indicated the presence of a putatively novel member of the genus Tenuivirus, tentatively named "wheat yellows virus" (WhYV). The genome is made up of four segments, which are 8952, 3451, 2338, and 2045 nucleotides in length and code for a total of seven ORFs. Phylogenies of each segment (nucleotide) and the polymerase gene (amino acid), as well as amino acid sequence comparisons of each gene product, showed that WhYV is most closely related to rice stripe virus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Wheat (Triticum aestivum L.) is one of the most important crops in the global production of cereal for both human and animal consumption. In South Africa, wheat is second only to maize as the country's most important grain crop [1]. The production of South African wheat under irrigation conditions regularly comes under significant disease pressure, such as Fusarium head blight [2], various rust fungi [3], and the recently described wheat stripe mosaic virus [4].
The genus Tenuivirus (family Phenuiviridae) comprises several viruses of poaceous cereal crops [5]. Tenuiviruses are vectored by delphacid planthopper vectors in a circulative-propagative manner with high virus-vector specificity [6]. There are currently nine accepted species [7]: Rice stripe tenuivirus, Echinochloa hoja blanca tenuivirus, European wheat striate mosaic tenuivirus, Iranian wheat stripe tenuivirus, Maize stripe tenuivirus, Melon tenuivirus, Rice grassy stunt tenuivirus, Rice hoja blanca tenuivirus, and Urochloa hoja blanca tenuivirus. Tenuiviruses has long been suspected to be present in wheat in Brazil and Europe, and recent high-throughput sequencing efforts have confirmed the presence of these viruses through the genomic characterization of Brazilian wheat spike virus [8] and European wheat striate mosaic virus [9]. There are no previous records of tenuiviruses infecting wheat in South Africa.
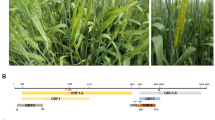
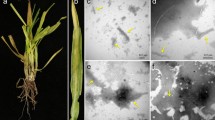
During 2019, winter wheat in the irrigation areas of KwaZulu-Natal (KZN) and Limpopo provinces showed symptoms of stunting and leaf chlorosis (Supplementary Fig. S1). Total RNA from one sample from Limpopo (sample W23) and seven samples from KZN (samples W24-W30) was extracted from leaves, using a modified CTAB protocol [10]. An RNAtag-seq library was prepared according to Shishkin et al. [11] and sequenced using an Illumina HiSeq 2500 sequencer (Illumina, San Diego, CA, USA) with 125-bp paired-end reads at the Agricultural Research Council Biotechnology Platform, Pretoria, South Africa. Datasets were demultiplexed using Je [12]. Trimming for quality and adapter content was performed using CLC Genomics Workbench 9 (QIAGEN Bioinformatics, Aarhus, Denmark) with parameters set as follows: minimum read length of 20 nt (nucleotides), quality limit of 0.05, and adapter trimming with Illumina universal (5’-AGATCGGAAGAG-3’) and RNAtag-seq (5’-TACACGACGCTCTTC CGATCTNNNNNNNNT-3’) adapters. The numbers of reads associated with each dataset are listed in Supplementary Table S1. All raw trimmed datasets are available as a National Center for Biotechnology Information (NCBI) sequence read archive with the accession number PRJNA816710. De novo assembly of contigs was performed using metaSPAdes 3.14.0 [13], and plant-virus-associated contigs were identified using BLASTn [14] with the viral refseq database.
All samples were associated with a putatively novel member of the genus Tenuivirus with four genome segments. Given the leaf chlorosis observed in the sampled plants, the virus has been tentatively named "wheat yellows virus" (WhYV). In addition to WhYV, nearly complete sequences for all three genome segments of brome mosaic virus were assembled for samples W25 and W26 (data not shown). The average lengths for each genome segment of WhYV were 8,952, 3,451, 2,338, and 2,045 nt, respectively, with a total average genome length of 16,786 nt. The specific length, coverage, and NCBI GenBank number of the genome segments from each sample are listed in Supplementary Table S1. The gene organization of each segment was determined using NCBI ORF Finder [14], which together with the putative functions of each gene, is shown in Figure 1. The genome organization is typical of the genus Tenuivirus [15], particularly those tenuiviruses with four genome segments. Initial BLASTn analysis indicated that WhYV shared the greatest sequence similarity with rice stripe virus (RSV) in all genome segments. Nucleotide sequence identity values were determined using the “pairwise comparison” function in CLC Genomics Workbench 21 (QIAGEN Bioinformatics, Aarhus, Denmark). The identity values for comparison of WhYV and RSV were 72%, 65.2%, 59.5%, and 62.7% for RNA 1, 2, 3, and 4, respectively. BLASTp analysis of each putative gene product of WhYV confirmed that the virus is most closely related to RSV. Average amino acid identity (AAI) values shared between cognate gene products of WhYV and RSV were determined using the AAI calculator in the enveomics collection [16] and are shown in Fig. 1. The current species demarcation thresholds are < 85% amino acid sequence identity shared between any cognate gene products and < 60% nt identity between cognate intergenic regions [8]. For WhYV, the amino acid sequence identity values between the corresponding gene products are all below 85%, with the exception of NScv4 on RNA4, for which the value was just above the threshold, at 85.9%. The nucleotide sequences of the intergenic regions of WhYV RNA 2, 3, and 4 are 47%, 50%, and 38% identical to those of the cognate regions of RSV, and these values are below the threshold of 60%.
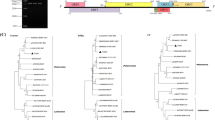
(A) Genome segment lengths (in nucleotides [nt]), genes, gene locations, putative gene product sizes (in amino acids [aa]), product functions, and average amino acid sequence identity (AAI) shared between wheat yellows virus (WhYV) and the cognate gene products of rice stripe virus. a JQ927432 DaL08, b JQ927427 HuZ10, c AJ620313 DLi1, d DQ299173 TXQ3-04; e AF221830 BS; f JQ927417 KunM08. The WhYV genome derived from sample W26 is shown here (ON156482-ON156485). MW, molecular weight; pI, isoelectric point; AAI, average amino acid identity. (B) Maps of the genome segments of WhYV. Each genome or genome segment is shown in the positive sense, and each open reading frame (ORF) is represented by a grey block with the direction of each ORF indicated. The putative functions of the gene products expressed by each ORF are shown. RdRp, RNA-dependent RNA polymerase; NS2, movement protein; NSvc2, glycoprotein; NS3, non-structural protein; CP, nucleocapsid; SP, disease-specific protein; NSvc4, NSvc4 protein
The nucleotide sequences of each RNA segment and the amino acid sequence of the polymerase gene were aligned with the cognate sequences of other tenuiviruses using Clustal Omega [17], and these alignments were then used for to maximum-likelihood phylogenetic analysis, using the best-fit substitution model in MEGA X [18], which was the general time-reversible model [19] for segment 1 and the Kimura 3-parameter model [20] for the remaining three segments. The Le Gascuel substitution model [21] with empirical base frequencies was used to infer the L-gene phylogeny. All phylogenies were run with gamma distribution to account for among-site rate variation, as well as bootstrapping with 1000 replicates, using the same model parameters to determine branch support. The L-gene phylogeny is shown in Fig. 2, and the nucleotide-sequence-based phylogenies of each gene segment are shown in Supplementary Figure S2.
Maximum-likelihood phylogeny based on an amino acid sequence alignment of the L-gene product (RNA-dependent RNA polymerase) of wheat yellows virus (sequences from this study are indicated by black circles) and relevant members of the genus Tenuivirus. The phylogeny represents the tree with the highest log likelihood and was generated in MEGA X using the Le Gascuel substitution model with empirical base frequencies, a proportion of invariant sites, and gamma distribution to account for among-site rate variation (n = 5). Bootstrapping was applied (1000 replicates), and the percentage of trees in which the associated taxa clustered together is shown next to the branches. Bootstrap percentages lower than 50 are not shown
To confirm the presence of WhYV in each sample, an RT-PCR assay was designed to target the coat protein (CP) gene, using the following oligonucleotide primers. WhYV-CP-F, 5’-TCA CAC ACT AGA GCA AAG ATC ACA-3’; WhYV-CP-R: 5’-AGT ATT CTG GCT ATG ATG CTG CAA-3’. Reactions were performed in two steps, using M-MuLV reverse transcriptase and OneTaq® 2X Master Mix (New England Biolabs, Ipswich, MA, USA). Bands of the expected size (950 bp) were generated for each sample, and amplicons from W24, W25, and W26 were used for unidirectional (forward) Sanger sequencing (Inqaba Biotechnical Industries, Pretoria South Africa). The resulting sequences were 100% identical to the cognate sequences of their respective contigs.
The sequences of the terminal nucleotides of WhYV genome segments were confirmed for both the viral (vRNA) and complementary RNA (vcRNA), using a 3’ RACE System for Rapid Amplification of cDNA Ends (Life Technologies, Carlsbad, CA, USA), according to the manufacturer's specifications. The following gene-specific primers were used: segment 1, vRNA (5’-ATC AAG TTG CAT GGA GTG GTC AAG-3’) and vcRNA (5’-TCA CAA AGC TCA GAG ACA CAG AGC-3’); segment 2, vRNA (5’-TGG ATG TAA GGG TAG TAC TGG ATT G-3’) and vcRNA (5’-CAA CAT TGT GTG ACA AGG GAA TCT C-3’); segment 3, vRNA (5’-GCC AGT TCC TCT GAC ATA TCT CAT-3’) and vcRNA (5’-CTC CCA CTA ATA GGT GGC TTA TCT G-3’), segment 4, vRNA (5’-GAG CCT TGA CAG AGT AGT CCT GT-3’) and vcRNA (5’-TCT CTG CCT TGG CCA TCT TTA CGA-3’). Total RNA from sample W26 was polyadenylated using E. coli poly(A) polymerase (New England Biolabs, Ipswich, MA, USA) prior to performing RACE. The RACE amplicons were sequenced by the Sanger method. The complete sequence of each genome segment confirmed the presence of inverted complementarity in the terminal nucleotides of each segment. This is typical of all segmented negative-stranded RNA viruses [15] and leads to the formation of panhandle structures that act as promoters for replication and transcription [5].
Complementarity was observed in the terminal 18 nucleotides of segments 1, 3, and 4 and the terminal 17 nucleotides of segment 2. The lengths of the complementary sequences is comparable with those of RSV [22], and the terminal sequences are completely conserved between RSV and WhYV in segments 2, 3, and 4 and have only two mismatches in segment 1. A mismatched cytosine residue was consistently observed in all segments at the 11th position, which has also been reported for RSV [22]. The terminal 10 nucleotides are completely conserved among the 5’ ends of all segments and the 3’ends of segments 2, 3, and 4, with the sequence ACACAAAGUC. Segment 1 has the sequence ACACAUAGUC at the 5’ end, with a single nucleotide mismatch at position 6 (underlined). This mismatch has also been reported in RSV genomes has been suggested to be involved in replicative or transcriptional control [22].
Phylogenetic analysis indicated that WhYV clusters with other grass-infecting tenuiviruses, and most closely with RSV when comparing the polymerase genes and all four genome segments. This is corroborated by AAI values for each gene product, as well as the fact that RSV and WhYV each have four genome segments and sequence conservation in the terminal nucleotides. Despite this clear relationship between the two viruses, WhYV is distinct from RSV, based on current species demarcation thresholds, as well as the fact that WhYV infects a host other than rice. The high levels of sequence similarity among the eight viruses sequenced in this study hint at a recent introduction of the virus into the country, although many more populations need to be characterized to confirm this. There is also very little data available regarding the distribution and diversity of delphacid leafhoppers in South Africa, and therefore, no connections can be made regarding a putative vector, although this will form part of ongoing research.
Availability of data and material
The data that support the findings of this study are openly available in the NCBI public databases.
References
Nhemachena CR, Kirsten J (2017) A historical assessment of sources and uses of wheat varietal innovations in South Africa. S Afr J Sci 113:2016–2018
Minnaar-Ontong A, Herselman L, Kriel W, Leslie JF (2017) Morphological characterization and trichothecene genotype analysis of a Fusarium Head Blight population in South Africa. Eur J Plant Pathol 148:261–269
Pretorius ZA, Prins R, Wessels E, Bender CM, Visser B, Boshoff WHP (2020) Accomplishments in wheat rust research in South Africa. S Afr J Sci 116:7688
Terefe TG, Visser B, Botha W, Kozana A, Roberts R, Thompson GD, Prinsloo G, Read DA (2021) Detection and molecular characterization of Wheat stripe mosaic virus on wheat in South Africa. Crop Prot 143:105464
Kormelink R, Verchot J, Tao X, Desbiez C (2021) The Bunyavirales: the plant-infecting counterparts. Viruses 13:842
Falk BW, Tsai JH (1998) Biology and molecular biology of viruses in the genus Tenuivirus. Annu Rev Phytopathol 36:139–163
Gaafar YZA, Rabenstein F, Zia A, Gaafar AZA, Ziebell H (2021) Molecular characterisation of a new tenuivirus from Festuca sp. Virus Res 304:198509
Pereira FS, Stempkowski LA, Fajardo TVM, Júnior AN, Lau D, Mar TB, do Nascimento SC, Bogo A, Casa RT, da Silva FN (2022) A novel tenuivirus infecting wheat in Brazil. Arch Virol 167:989–993
Sõmera M, Kvarnheden A, Desbiez C, Blystad D, Sooväli P, Kundu JK, Gantsovski M, Nygren J, Lecoq H, Verdin E, Spetz C, Tamisier L, Truve E, Massart S (2020) Sixty years after the first description: Genome sequence and biological characterization of European wheat striate mosaic virus infecting cereal crops. Phytopathology 110:68–79
White EJ, Venter M, Hiten NF, Burger JT (2008) Modified cetyltrimethylammonium bromide method improves robustness and versatility: the benchmark for plant RNA extraction. Biotechnol J 3:1424–1428
Shishkin AA, Giannoukos G, Kucukural A, Ciulla D, Busby M, Surka C, Chen J (2015) Simultaneous generation of many RNA- seq libraries in a single reaction. Nat Methods 12:323–325
Girardot C, Scholtalbers J, Sauer S, Su SY, Furlong EEM (2016) Je, a versatile suite to handle multiplexed NGS libraries with unique molecular identifiers. BMC Bioinform 17(1):419
Nurk S, Meleshko D, Korobeynikov A, Pevzner PA (2017) metaSPAdes: a new versatile metagenomic assembler. Genome Res 27(5):824–834
Wheeler DL, Church DM, Federhen S, Lash AE, Madden TL, Pontius JU, Schuler GD, Schriml LM, Sequeira E, Tatusova TA, Wagner L (2003) Database resources of the National Center for Biotechnology. Nucleic Acids Res 31:28–33
Kormelink R, Garcia ML, Goodin M, Sasaya T, Haenni A (2011) Negative-strand RNA viruses: the plant-infecting counterparts. Virus Res 162:184–202
Rodriguez-R LM, Konstantinidis KT (2016) The enveomics collection: a toolbox for specialized analyses of microbial genomes and metagenomes. PeerJ Prepr 4:e1900v1
Madeira F, Park YM, Lee J, Buso N, Gur T, Madhusoodanan N, Basutkar P, Tivey ARN, Potter SC, Finn RD, Lopez R (2019) The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res 47:W636–W641
Stecher G, Tamura K, Kumar S (2020) Molecular Evolutionary Genetics Analysis (MEGA) for macOS. Mol Biol Evol 37:1237–1239
Tavaré S (1986) Some probabilistic and statistical problems in the analysis of DNA sequences. Lect Math Life Sci 17:57–86
Tamura K (1992) Estimation of the number of nucleotide substitutions when there are strong transition-transversion and G+C content biases. Mol Biol Evol 9:678–687
Le SQ, Gascuel O (2008) An improved general amino acid replacement matrix. Mol Biol Evol 25:1307–1320
Takahashi M, Toriyama S, Kikuchi Y, Hayakawa T, Ishihama A (1990) Complementarity between the 5’- and 3’-terminal sequences of rice stripe virus RNAs. J Gen Virol 71:2817–2821
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflicts of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Handling Editor: Ralf Georg Dietzgen.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Read, D.A., Roberts, R., Prinsloo, G. et al. Genomic and phylogenetic characterization of wheat yellows virus, a novel tenuivirus infecting wheat in South Africa. Arch Virol 168, 3 (2023). https://doi.org/10.1007/s00705-022-05649-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00705-022-05649-7