Abstract
The complete genome sequence of a novel mycovirus, Phoma matteucciicola RNA virus 1 (PmRV1), derived from Phoma matteucciicola strain LG-01, was sequenced and analyzed. The complete cDNA sequence of PmRV1 is 3432 bp in length with a GC content of 57.17%. The genome of PmRV1 contains two putative open reading frames (ORFs): ORF1 and ORF2. ORF1 encodes a hypothetical protein with significant similarity to a protein encoded by Periconia macrospinosa ambiguivirus 1 (PmAV1). ORF2 encodes a protein of 491 amino acids with a conserved RNA-dependent RNA polymerase (RdRp) domain. Additionally, the triad within domain III has an asparagine (GDN) instead of the nearly universally conserved aspartic acid (GDD). RdRp phylogeny showed that PmRV1 grouped together with PmAV1 as a sister branch of a new member of the recently proposed family of mycotombus-like viruses. This is first report of the complete sequence of a novel mycovirus, PmRV1, infecting Phoma matteucciicola strain LG-01, the causal agent of leaf blight of Curcuma wenyujin.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Mycoviruses are widespread and replicate in diverse filamentous fungi, yeasts and oomycetes [1,2,3]. Previous studies have shown that the genomes of mycoviruses are predominately double-stranded RNA (dsRNA), positive-sense single-stranded RNA (+ssRNA), or negative-sense single-stranded RNA (-ssRNA) [4]. However, mycoviruses with single-stranded DNA (ssDNA) genomes have been reported, such as Sclerotinia sclerotiorum hypovirulence-associated DNA virus 1 (SsHADV-1) and Fusarium graminearum gemytripvirus 1 (FgGMTV1) [5,6,7]. The +ssRNA mycoviruses are currently classified into nine families, including Alphaflexiviridae, Barnaviridae, Botourmiaviridae, Gammaflexiviridae, Hypoviridae, Endornaviridae, Narnaviridae, Deltaflexiviridae, and the recently proposed family "Mycotombusviridae" [8,9,10]. Members of the proposed family "Mycotombusviridae" possess non-segmented +ssRNA genomes, 2.6- 4.6 kb in length, that usually contain two noncontiguous open reading frames (ORFs), with the second ORF encoding an RNA-dependent RNA polymerase [11, 12].
Phoma matteucciicola is a filamentous fungus that causes leaf blight disease in Curcuma wenyujin, causing huge economic losses [13]. So far, only two mycoviruses, Phoma matteucciicola partitivirus 1 (PmPV1) and Phoma matteucciicola ourmia-like virus 1 (PmOLV1), have been reported in P. matteucciicola [14, 15]. In this study, a novel (+) ssRNA virus was isolated from P. matteucciicola strain LG-01 and identified. Based on its genome organization and phylogeny, the virus was named "Phoma matteucciicola RNA virus" (PmRV1), and it is proposed to be a new unassigned mycotombus-like virus.
Provenance of the virus material
The host of PmRV1, Phoma matteucciicola strain LG-01, was isolated from C. wenyujin showing symptoms of leaf blight disease Hainan, China, and identified as P. matteucciicola based on molecular phylogeny. LG-01 was cultured on potato dextrose agar (PDA) medium covered with a cellophane membrane at 28 °C for 10 days for dsRNA extraction. The dsRNAs of LG-01 were extracted from approximately 3.0 g of frozen mycelia according to the protocol described by Morris and Dodds [16]. Subsequently, the crude virus was treated with RNase-free S1 nuclease and DNase I (TaKaRa, Dalian, China), to remove contaminating nucleic acids. A cDNA library of purified PmRV1 dsRNA was then generated using tagged random dN6 primers (5'-CGATCGATCATGATGCAATGCNNNNNN-3') and Moloney murine leukaemia virus reverse transcriptase. To obtain the full-length PmRV1 cDNA, internal gaps between the initial sequences were obtained by RT-PCR using specific primers. The terminal sequences of PmRV1 dsRNA were obtained using rapid amplification of ligase-mediated rapid amplification of cDNA ends (RLM-RACE). All purified RT-PCR products were cloned into the pMD19-T vector (TaKaRa, Dalian, China) for Sanger sequencing. For each PCR product, at least three independent clones were sequenced in both directions by Sangon Biotech (Shanghai) Co., Ltd. The complete nucleotide sequence of PmRV1 was deposited in the GenBank database, with accession number MT590656. The amino acid (aa) sequence of the putative RNA-dependent RNA polymerase (RdRp) sequence of PmRV1 was aligned with other related viral RdRp aa sequences using the ClustalX2 program. A phylogenetic tree was created by the maximum-likelihood (ML) method using Molecular Evolutionary Genetic Analysis software version 7.0 (MEGA 7.0) with 1,000 bootstrap replicates.
Sequence properties
The full-length sequence of PmRV1 is 3432 nucleotides (nt) in length with two noncontiguous ORFs, separated by 115 nt and with 5' and 3' untranslated regions (UTRs) of 333 nt and 423 nt in length, respectively (Fig. 1a). Using the Mfold program (https://unafold.rna.albany.edu), the sequences at the PmRV1 5' and 3' UTR of PmRV1 were predicted to form two potential stem-loop structures, with initial ∆G values of -11.00 kcal/mol and -10.70 kcal/mol, respectively (Fig. 1b).
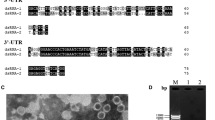
a Schematic representation of the genomic organization of PmRV1. The open reading frames (ORFs) and the untranslated regions (UTRs) are showed as open bars and single lines, respectively. The gray bar indicates the RdRp domain. The nucleotide positions of the initiation and termination codons are indicated under the ORFs. The molecular weights of the predicted proteins and the length of the non-coding sequence are shown above the ORFs. b Predicted secondary structures of the 5' (left) and 3' (right) termini of PmRV1. c Amino acid sequence alignment of PmRV1 RdRps motifs with those of the related mycotombus-like viruses. Horizontal lines above the alignment show the five motifs. Shaded areas show identical amino acid residues. Asterisks, colons, and dots indicate identical amino acid residues, conservative variations, and semi-conservative variations, respectively. Conserved abnormal GDN triplets that differ from the standard (+) ssRNA amino acid sequence are highlighted in red
ORF1, starting at nt 334 and ending at nt 1419, encodes a 361-aa polypeptide with a predicted molecular mass of 39.6 kDa (Fig. 1a). A database search with BLASTp showed that the aa sequence of PmRV1 shares 67% sequence identity with a similar hypothetical protein of Periconia macrospinosa ambiguivirus 1 (PmAV1).
ORF2, starting at nt 1534 and ending at nt 3009, encodes a 491-aa protein with a predicted molecular mass of 54.8 kDa. Database searches showed that ORF2 was most closely related to the RdRps of PmAV1 (69.7% identity) and soybean leaf-associated ssRNA virus 2 (62.1% identity). In addition, a search of the conserved domain database and multiple protein sequence alignment indicated that ORF2 encodes a protein containing a conserved viral RdRp domain with five conserved motifs (I–V) in the subfamily RdRP_3 (pfam00998) (Fig. 1c).
To analyze the relationship between PmRV1 and other mycoviruses, we constructed a phylogenetic tree using the RdRp regions of PmRV1 and other related RNA viruses. The results showed that PmRV1 and nine other unclassified ssRNA mycoviruses form a well-supported clade that is separate from the insect and plant viruses (Fig. 2). Moreover, PmRV1 is closely related to nine other unclassified mycoviruses in sequence and genome organization, which clearly differ from members of the family Tombusviridae. All 10 of these viruses contain two noncontiguous ORFs, with ORF2 containing the RdRp domain. Additionally, the PmRV1 RdRp has a GDN triad in motif III versus the GDD motif found in +ssRNA viruses (Fig. 1c). It has been reported that modification of the GDD to GDN has a detrimental effect on enzymatic activity in +ssRNA viruses [17]. Based on our results, we concluded that PmRV1 is a new member of the recently proposed family "Mycotombusviridae".
References
Ghabrial SA, Castón JR, Jiang D, Nibert ML, Suzuki N (2015) 50-plus years of fungal viruses. Virology 356:356–368
Rosseto P, Costa AT, Polonio JC, da Silva AA, Pamphile JA, Azevedo JL (2016) Investigation of mycoviruses in endophytic and phytopathogenic strains of Colletotrichum from different hosts. Genet Mol Res 15:15017651
Xie J, Jiang D (2014) New insights into mycoviruses and exploration for the biological control of crop fungal diseases. Annu Rev Phytopathol 52:45–68
Donaire L, Pagán I, Ayllón MA (2016) Characterization of Botrytis cinerea negative-stranded RNA virus 1, a new mycovirus related to plant viruses, and a reconstruction of host pattern evolution in negative-sense ssRNA viruses. Virology 499:212–218
Yu X, Li B, Fu Y, Jiang D, Ghabrial SA, Li G, Peng Y, Xie J, Cheng J, Huang J, Yi X (2010) A geminivirus-related DNA mycovirus that confers hypovirulence to a plant pathogenic fungus. Proc Natl Acad Sci USA 107:8387–8392
Yu X, Li B, Fu Y, Xie J, Cheng J, Ghabrial SA, Li G, Yi X, Jiang D (2013) Extracellular transmission of a DNA mycovirus and its use as a natural fungicide. Proc Natl Acad Sci USA 110:1452–1457
Li P, Wang S, Zhang L, Qiu D, Zhou X, Guo L (2020) A tripartite ssDNA mycovirus from a plant pathogenic fungus is infectious as cloned DNA and purified virions. Sci Adv 6:eaay9634
Li Q, Huang W, Hai D, Wang Y, Xie J, Wang M (2020) The complete genome sequence of a novel hypovirus infecting Bipolaris oryzae. Arch Virol 165:1027–1031
Muhammad H, Xie J, Wu S, Shahzeen M, Dan Z, Abdoulaye AH, Wang Q, Cheng J, Fu Y, Jiang D (2018) A novel deltaflexivirus that infects the plant fungal pathogen, Sclerotinia sclerotiorum, can be transmitted among host vegetative incompatiblestrains. Viruses 10:295
Li K, Zheng D, Cheng J, Chen T, Xie J (2015) Characterization of a novel Sclerotinia sclerotiorum RNA virus as the prototype of a new proposed family within the order Tymovirales. Virus Res 219:92–99
Gilbert KB, Holcomb EE, Allscheid RL, Carrington JC (2019) Hiding in plain sight: New virus genomes discovered via a systematic analysis of fungal public transcriptomes. PLoS ONE 14:e0219207
Ai Y, Zhong J, Chen C, Zhu H, Gao B (2016) A novel single-stranded RNA virus isolated from the rice-pathogenic fungus Magnaporthe oryzae with similarity to members of the family Tombusviridae. Arch Virol 161:725–729
Zheng F, Ma R, Xu G, Zheng F, Ding X, Xie C (2018) Leaf blight on Curcuma wenyujin caused by Phoma matteucciicola in China. Plant Dis 102:2042
Zheng F, Xu G, Zhou J, Xie C, Cui H, Miao W, Kang Z, Zheng L (2019) Complete genomic sequence and organization of a novel mycovirus from Phoma matteuccicola strain LG915. Arch Virol 164:2209–2213
Zhou J, Wang Y, Liang X, Xie C, Liu W, Miao W, Kang Z, Zheng L (2020) Molecular characterization of a novel Ourmia-Like virus infecting Phoma matteucciicola. Viruses 12:231
Morris TJ, Dodds JA (1979) Isolation and analysis of double-Stranded RNA from virus-infected plant and fungal tissue. Phytopathology 69:854–858
Vázquez AL, Alonso JM, Parra F (2000) Mutation analysis of the GDD sequence motif of a calicivirus RNAdependent RNA polymerase. J Virol 74:3888–3891
Funding
This study was financially supported by Hainan Province Key R&D Project (ZDYF2019143), the Scientific Research Foundation for Advanced Talents, Hainan University (no. KYQD(ZR)1873), Hainan Major Research Found of Science and Technology (no. ZDKJ201817), and the National Natural Science Foundation of China (no. 31701732).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with animals or human participants performed by any of the authors.
Additional information
Handling Editor: Ioly Kotta-Loizou.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhou, J., Hu, X., Liang, X. et al. Complete genome sequence of a novel mycovirus from Phoma matteucciicola. Arch Virol 166, 317–320 (2021). https://doi.org/10.1007/s00705-020-04865-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-020-04865-3