Abstract
The complete genome sequence of a new potato virus M (PVM) isolate (PVM-YN), collected from potato (Solanum tuberosum) in Yunnan, China, was determined. It was 8,530 nucleotides (nt) in length, excluding the poly(A) tail at the 3′ end, and shared 71.4–72.0% nucleotide sequence identity with available PVM isolates in the NCBI database. The coat proteins (CP) of PVM-YN shared 79.0–97.4% amino acid sequence identity with that of other isolates. It is the first report of the complete genomic sequence of a new PVM isolate infecting S. tuberosum in China.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The species Potato virus M is a member of the genus Carlavirus in the family Betaflexiviridae. The virions of potato virus M (PVM) are slightly flexuous filaments of 610–700 nm in length and 12–15 nm in diameter, containing a linear, positive sense (+), single-stranded RNA (ssRNA) genome of about 8.5 kb in length [1]. The genomic RNA of PVM contains a cap-like structure at the 5′-UTR (untranslated region) and a poly(A) tail at the 3′-UTR [2, 3]. PVM has six open reading frames (ORFs) [4]. ORF1 encodes a polyprotein responsible for RNA replication. The overlapping ORFs 2, 3, and 4 encode three putative proteins of 25, 12, and 7 kDa, respectively, recognized as the triple gene block (TGB) responsible for cell-to-cell movement. ORFs 5 and 6 encode the coat protein (CP, 34 kDa) and a cysteine-rich protein (11 kDa), respectively. The 11-kDa protein encoded by ORF6 is a nucleic acid-binding protein that can bind single- or double-stranded RNA and DNA [4, 5]. Recently, the function of this protein was demonstrated to be suppression of host antiviral gene silencing [6]. PVM is economically important and a common virus of potato (Solanum tuberosum) with a worldwide distribution. It was first isolated and identified in the United States from S. tuberosum in 1923 [7]. Since then, the virus has been found in all potato producing countries in the world, alone or in mixed infection with potato virus S (PVS) [8]. Some isolates of PVM induce only mild symptoms in potato and can reduce tuber yield by 10–18%, whereas other isolates cause severe symptoms and can result in tuber yield losses of 40 to 75% [9]. This virus is transmitted non-persistently by several aphid species and also by mechanical inoculation with sap from young leaves [10].
In 2015, several potato plants with typical symptoms, including mosaic, mottle, crinkling and mild abaxial rolling of leaves, as well as stunting of shoots, were observed in China’s Yunnan province. The result of ELISA experiments (PVM ELISA kit, Agdia, Elkhart, IN) demonstrated that these potato plants were infected by PVM. In this study, the complete genomic sequence of this PVM isolate was determined. This is the first report on the complete genomic sequence of a new isolate of PVM (PVM-YN) infecting potato in China, which is a clearly distinct isolate from other previously identified PVM isolates.
Material and methods
Total RNA was extracted from virus-infected tissue culture potato seedlings using TriPure total RNA Isolation Reagent (Invitrogen, Roche, CA, USA). First-strand cDNA was synthesized with M-MLV reverse transcriptase (Promega, Madison, WI, USA), according to the manufacturer’s instructions. For amplification of the DNA fragments, primers were designed based on the complete sequences of other PVM isolates (Supplementary Table 1). The 5′ terminal end sequence of the 5′-UTR of the genomic sequence was determined with the 5′ rapid amplification of cDNA ends (RACE) method, using the SMARTer® RACE cDNA amplification kit (Clontech, USA) following the manufacturer’s instructions and using a gene specific primer. The amplified fragments (PCR products) were purified using an agarose gel DNA purification kit (TaKaRa Biotechnology Dalian Co, Ltd, China). All the cDNA and RACE fragments were cloned into the pMD18-T vector (TaKaRa, Dalian, China) according to the manufacturer’s instructions. These plasmid clones were used to transform competent Escherichia coli DH5 alpha cells. Plasmid DNAs were sequenced by the dideoxynucleotide chain termination method using an automatic DNA sequencer (Model 377XL Perkin-Elmer Applied Biosystem Co., Foster City, CA).The complete sequence of PVM was assembled and analyzed with DNAMAN version 5.0 (Lynnon Biosoft, QC, Canada). Phylogenetic trees were constructed using the neighbor-joining method with 1,000 bootstrap replications in MEGA 6.0 [11]. PVM sequences used for comparison were obtained from the GenBank database (Supplementary Table 2).
Sequence properties
The complete genome sequence of PVM-YN was 8,530 nucleotides (nts) long, excluding the 3′ poly(A) tail (KY364848). PVM-YN shares 71.4 to 72% identity, at the nucleotide level, with the full-length sequences of other PVM isolates available in the NCBI database. The genome contains six ORFs as found in all carlaviruses. The 5′-UTR consists of 83 nts. ORF1 begins at the AUG start codon at nt 84 and terminates at the UAA stop codon at nt 5,978. It encodes a 222 kDa polypeptide of 1,963 amino acid (aa) residues, consisting of the methyl transferase (aa 43–354), the carlavirus endopeptidase (aa 990–1077), the viral helicase (aa 1162–1421), and the RNA-dependent RNA polymerase domains (aa 1546–1955), which share 86.9–91.6%, 69.3–73.8%, 80.3–82.2%, and 92.7–95.1% of aa sequence identity, respectively, with the corresponding domains of other PVM isolates. Overall, this polypeptide shares 75.9–77.9% amino acid sequence identity (Supplementary Table 2) with the polypeptide of other PVM isolates. This polypeptide is the most divergent, with local sequence identity levels of 53.8–56.5% from aa 355 to 989.
ORFs 2–4 include the triple gene block (TGB) proteins. ORF2 begins at nt 6,016 and ends at nt 6,705 and encodes a 25-kDa protein with a viral helicase 1 (aa 24–221) and a RecD (aa 184–229) domain. This protein shares 78.60–82.10% amino acid identity with other PVM isolates. There was an intergene region between ORF1 and ORF2 from nts 5,979–6,015. ORF3 begins at nt 6,683 and ends at nt 7,012 and encodes a 12-kDa protein; the level of sequence identity ranged from 69.72 to 71.56%. ORF4 starts at nt 7,009 and ends at nt 7,206 and codes for a 7-kDa protein that showed the lowest level of identity among the TGB protein sequences (about 55.38–58.6%). ORF5 begins at nt 7,222 and ends at nt 8,136 and encodes the 34-kDa viral CP, sharing 79.0–97.4% amino acid sequence identity with the PVM isolates available in the GenBank database. Lastly, ORF6 begins at nt 8,133 and ends at nt 8,459, overlapping with the CP and encoding a cysteine-rich nucleic acid-binding protein (NABP) of 11 kDa; this protein had 71.1–83.3% amino acid identity with that of other isolates.
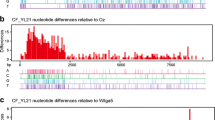
To further understand the molecular relationships between PVM-YN and other PVM isolates, phylogenetic trees were constructed using MEGA (version 6.0). Phylogenetic analysis of the complete PVM genome sequences available in GenBank indicated that PYM-YN clustered in the outer branch of other PVM isolates belonging to PVM ordinary strains (PVM-O), due to the lack of ORF1 in PVM divergent strains (PVM-D) [12] (Fig. 1A). The phylogenetic tree based on the amino acid sequences of the CP gene of PVM isolates grouped these PVM isolates into three main clusters: PVM-O, PVM-D, and a third cluster including PVM-YN as well as PVM isolates from India (Fig. 1B). Different isolates from the same country were grouped into different clusters. This result suggests that there are no geographical correlations between or within these clusters as previously described [13]. Two other isolates collected from tomato and ginseng fruit in China clustered with PVM-O [8].
Phylogenetic tree of PVM isolates. Sequences were aligned using Clustal W. The tree was constructed by the neighbor-joining algorithm (MEGA 6 package) based on: (A) the nucleotide sequence alignment of the available complete PVM genome sequences, using potato virus S (PVS) isolate as an out-group, and the amino acid sequences of PVM coat proteins (B). The data set was exposed to 1,000 bootstrap replicates. Bootstrap values higher than 60 are indicated at the corresponding branch. The bar represent 0.1 exchanged per 100 nucleotides
Criteria for species delineation in the genus Carlavirus include: serological specificity, natural host range, and size and sequence identity of the CP gene [1]. Sequence analysis revealed that members of distinct species of carlaviruses share less than 72% nucleotide sequence identity (or 80% amino acid sequence identity) in their entire CP or polymerase genes. In this study, the PVM-YN isolate shared more than 83.6% and 75.9–77.4% aa sequence identity in the CP and replicase genes, respectively, with other PVM isolates. This study is the first report of a complete PVM genome sequence isolated from potato in the Yunnan province of China.
References
Adams MJ, Antoniw JF (2004) The new plant virus family Flexiviridae and assessment of molecular criteria for species demarcation. Arch Virol 149:1045–1060
Flarken S, Ungewickell V, Mengel W, Maiss E (2008) Construction of an infectious full-length cDNA clone of potato virus M. Arch Virol 153:1385–1389
Ge BB, He Z, Jiang DM, Zhang ZX, Liu GI, Wang HQ (2012) Characterization and complete nucleotide sequence of potato virus M isolated in China. Acta Virol 56:261–263
Zavriev SK, Kanyuka KV, Levay KE (1991) The genome organization of potato virus M RNA. J Gen Virol 72:9–14
Gramstat A, Courtpozanis A, Rohde W (1990) The 12-kDa protein of potato virus M displays properties of a nucleic acid binding regulatory protein. FEBS Lett 276:34–38
Senshu H, Yamaji Y, Minato N, Shiraishi T, Maejima K (2011) A dual strategy for the suppression of host antiviral silencing: two distinct suppressors for viral replication and viral movement encoded by potato virus M. J Virol 85:10269–10278
Schultz ES, Folsom D (1923) Transmission, variation, and control of certain degeneration diseases of Irish potatoes. J Agric Res 25:43–147
Cox BA, Jones RAC (2010) Genetic variability in the coat protein gene of Potato virus S isolates and distinguishing its biologically distinct strains. Arch Virol 155:1163–1169
Brunt AA (2001) Potato virus M (PVM; Genus Carlavirus). In: Loebenstein G, Berger PH, Brunt AA, Lawson RH (eds) Virus and virus-like diseases of potatoes and production of seed potatoes. Kluwer, Dordrecht, pp 101–108
Kerlan C (2008) Potato viruses. In: Mahy BWJ, Van Regenmortel MHV (eds) Desk encyclopedia of plant and fungal virology. San Diego, California, pp 458–470
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol Biol Evol. 30:2725–2729
Fatemeh T, Behrooz J, Mohammad Z, Majid S, Hamid R, Mohsen M (2014) Genetic structure and molecular variability of potato virus M populations. Arch Virol 159:2081–2090
Xu H, D’Aubin J, Nie J (2010) Genomic variability in Potato virus M and the development of RT-PCR and RFLP procedures for the detection of this virus in seed potatoes. Virol J 7:25 http://www.virologyj.com/content/7/1/25
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Funding
This study was partially supported by grants from the Yunnan Provincial Modern Agriculture for Potatoes Industrialization System (2017KJTX003), Top Scientists Input Program of Yunnan (2013HA028), and Yuling Scholar Program (Zhongkai Zhang).
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflicts of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
705_2017_3380_MOESM1_ESM.docx
Supplementary Table 1 Primers used in RT-PCR for amplification of genome fragments. Supplementary Table 2 Percent nucleotide (nt) and amino acid (aa) sequence identities of available complete PVM genomes compared to PVM-YN (DOCX 29 kb)
Rights and permissions
About this article
Cite this article
Su, X., Wu, K., Zhang, L.Z. et al. Complete genome sequence of a new isolate of potato virus M in Yunnan, China. Arch Virol 162, 2485–2488 (2017). https://doi.org/10.1007/s00705-017-3380-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-017-3380-7