Abstract
Newcastle disease viruses (NDV) represent a major threat to poultry production worldwide. Recently in Egypt NDV circulated extensively, even in vaccinated farms. In the present study samples were collected from sixteen vaccinated broiler farms in animals exhibiting the typical gross lesions of NDV. Virus isolation and pathogenicity studies for positive samples were carried out in accordance to reference procedures and phylogenetic analysis was carried out based on partial sequences of the Fusion gene. Furthermore, in vivo investigation of the ability of heterologous antibody, induced by commercially available lentogenic strain-based vaccines, to efficiently reduce viral shedding was examined. Results revealed that all the sixteen farms were positive for the presence of NDV. Out of these fifteen were confirmed to due to velogenic viruses, based on a main death time (MDT) ≤ 48 hours and partial sequencing of the F gene that showed the presence of a polybasic amino acid motif. However, three patterns in the cleavage site of these velogenic viruses were identified in the present study. Phylogenetic analysis revealed that all fifteen isolates were clustered with class II genotype VIIb while the remaining isolate (B81) was class II genotype II. Results of the in vivo study revealed that adequate heterologous antibody levels, induced by the proposed vaccination program, sufficiently protected birds from morbidity and mortality. However, virus shedding was quantitatively affected in relation to the time of challenge after vaccination. Altogether, with an absence of vaccines able to induce homologous antibody to the presently circulating viruses, higher antibody levels, which depend on efficient and timely implementation of the vaccination program, are considered as highly important in relation to the reduction of virus shedding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Newcastle disease (ND) is a contagious viral disease that commonly affects most avian species, inducing a high morbidity and mortality up to 100% [5]. ND virus (NDV) is an enveloped, single-stranded, negative-sense RNA virus classified within the family Paramyxoviridae with isolates designated as avian paramyxovirus-1 (APMV-1) [30]. NDV is composed of six genes and their encoding six structural proteins: nucleoprotein (NP), phosphoprotein (P), matrix (M), fusion (F), hemagglutinin-neuraminidase (HN), and the RNA polymerase (L) [26]. The sequence of the F protein’s cleavage site represents the primary molecular determinant of NDV virulence [15]. However, NDV strains with identical cleavage site sequences have shown significant differences in their virulence [12], which highlights the ultimate importance of characterizing the pathogenicity of NDV strains via internationally recognized tests such as mean death time (MDT), intracerebral pathogenicity index (ICPI), and intravenous pathogenicity index (IVPI) [6]. Isolates of NDV are categorized into 4 pathotypes: a- Lentogenic or avirulent forms; b- Mesogenic isolates: of moderate virulence but not fatal; c- Velogenic viscerotropic isolates of high virulence causing fatal enteric disease. d- Velogenic neurotropic isolates of high virulence causing high mortalities and signs of nervous system infection [17]. Genotypically, APMV-1 is classified into two clades designated as class I and class II. The majority of APMV-1 strains, including both virulent and non-pathogenic strains, belong to class II within which are classified the viruses of economic importance to chickens, while class I isolates mainly circulate in wild waterfowl [2]. The class II genotypes I and II include mainly avirulent strains and NDV vaccine strains [32] while virulent strains belong to class II genotypes III to IX and XI to XVI [9]. Virulent viruses belong to genotypes V, VI and VII are highly mobile, spread globally and are responsible for the majority of recent outbreaks in poultry and wild birds worldwide [14].
In Egypt, NDV was identified for the first time in 1947 [10]. Since then, devastating NDV outbreaks are frequently reported in Egypt. Based on genotypic characterization both class II genotype II and VI have been frequently identified in Egypt [20, 35, 36]. However, class II genotype VII is purported to be the cause of the current outbreaks in Egypt [21, 40].
Despite the routine intensive vaccination programs implemented in Egypt and worldwide genotype VII NDV still represents a major threat to poultry production [13, 22, 42]. Several factors could be responsible for this failure in vaccination such as the persistence of immunosuppressive diseases [24], improper vaccination strategies, low cross-protective-immunity between the vaccines used and field strains [50], continuous evolution of the virus, the role of wild birds in the spread of the infection [34], inadequate application and storage of vaccines [23] in addition to, differing genetic resistance to NDV among various breeds of chickens [25].
It is widely accepted that a homologous vaccine could provide increased protection against infection with genotype VII NDV, with a significant reduction in virus shedding compared to what is seen with the LaSota vaccine [19, 50]. However, even with a homologous inactivated vaccine a higher level of specific antibodies may be required to reduce virus shedding significantly. On the other hand, heterologous vaccination can induce significantly higher levels of antibodies after an optimum period of time that, theoretically, could prevent NDV transmission [33]. Altogether, the effective level of antibodies required to reduce NDV shedding depend on: 1- using a genotype matched vaccine (to the field strain); 2- allowing sufficient time for the vaccinated birds to develop the optimum immune response; and 3- the ultimate level of antibodies produced.
Thus, the present study was performed to genotypically and pathologically characterize NDV viruses isolated from vaccinated broilers farms, in addition, evaluate the efficiency of the commonly used (in the Egyptian market) NDV vaccines with a new vaccination strategy.
Material and methods
Ethics statement
The present study was conducted in strict accordance with the recommendations of the Ministry of Agriculture and Land Reclamation and Ministry of Environment, Egypt for experimental animal infection. The protocol of the study was reviewed and approved by the committee of laboratory biosafety for the reference laboratory for quality control on poultry production (RLQP). All virus isolation and animal experiments were performed in the virology unit and biological experiments and research unit at RLQP, under strict biosafety level 3 control facilities.
Sample collection and direct detection of NDV using RT-PCR
Tracheal swabs were collected from sixteen vaccinated broiler farms (30-45 days old) with suspected Newcastle disease (ND), located in nine separate governorates (Table 1). Swabs were collected from twenty birds per farm and pooled, as one sample for each farm, in buffered saline solution with antibiotic (10,000 U/ml penicillin, 10 mg/ml streptomycin, 0.25 mg/ml gentamicin and 5,000 IU/ml nystatin), adjusted to pH 7.0-7.4 [38]. Collected samples were directly screened for the presence of NDV by RT-PCR. Briefly, RNA was extracted from the pools of tracheal swabs using the QIAamp Viral RNA Mini Kit (Qiagen) according to the manufacturer’s instructions. Samples were amplified using the QuantiTect probe RT-PCR kit (Qiagen), primers and a thermal profile, as previously described by Wise, et al. [49]. Furthermore, the extracted RNA was screened for presence of AI-H5, AI-H7, AI-H9 and IB using primers and a thermal profile, as previously described by Slomka, et al. [8, 43, 45] respectively.
Virus isolation and pathogenicity assessment
The clarified supernatant from the swab pools (100 µl/egg) was inoculated into the allantoic cavity of 10 day-old embryonated specific pathogen free (SPF) eggs (SPF farm, Koam Osheim, El-Fayoum, Egypt). The inoculated eggs were incubated at 37 °C and candled daily for 5 days. Allantoic fluid was collected from dead embryos after the first 24 hours and examined for hemagglutination (HA) and hemagglutination inhibition (HI) activity using four HA units, according to OIE guidelines [38]. The egg infective dose 50 (EID50) was calculated, according to Reed and Muench [41].
The level of pathogenicity was determined by MDT in accordance with the reference procedure [38]. Briefly, ten-fold serial dilutions of each of the 16 isolates was inoculated intra-allantoically into 5 ten-day old SPF eggs, before being incubated at 37 οC and monitored daily for dead embryos. Eggs with dead embryos were discarded after the first 24 hours. The ICPI was carried out for selected isolates: B7, B270, F460 and F388 in 10 one-day old SPF chicks and calculated according to OIE [38] instructions.
Genetic and phylogenetic analysis
Viral RNA was extracted from the allantoic fluid using the QIAamp viral RNA mini kit (Qiagen, Germany). The first quarter of the coding region of the F gene was amplified using a OneStep RT-PCR kit (Qiagen), primers and a thermal profile as previously described by Mase, et al. [29]. The PCR products (766 bp) generated from the sixteen isolates was subjected to electrophoresis on a 2 % agarose gel and the specific band cut precisely and purified using a QIA-quick gel extraction kit (Qiagen- USA), according to the manufacturer’s instructions. The purified PCR products were used as templates for sequencing on an Applied Biosystems 3130 genetic analyzer (Hitachi, Japan) using a Big dye Terminator V3.1 cycle sequencing kit (Perkin-Elmer, Foster city, CA, USA) according to the manufacturer’s instructions. Nucleotide and amino acid sequences were aligned using BioEdit software v7.0 [16]. Representative strains for all NDV genotypes and NDV subgenotype VII were retrieved from the GenBank database (accession numbers are shown in Figs. 1 and 2 and sequence details are available in Supplementary Table 1) and aligned with the sequences of the isolates from the present study. A phylogenetic tree was constructed using the Maximum likelihood method based on the Kimura 2-parameter model with 1000 bootstrap replicates, using MEGA version 4 software [47]. The trees were subsequently viewed and edited using FigTree v1.4.2 software (http://tree.bio.ed.ac.uk/software/figtree/).
The gene sequences generated in this study (n = 16) were submitted to GenBank and assigned accession numbers: KM288609 to KM288622 and KP316015, KP316016.
Vaccine protection study
Sixty-five SPF chicks were purchased from Koum Oshiem SPF farm, Fayoum, Egypt and housed in BSL3 isolators in the animal experimental facilities of RLQP, Egypt, with continuous access to food and water. All chicks were serologically tested prior to vaccination using hemagglutination inhibition (HI) test for antibodies against NDV. Chicks were randomly divided into 3 vaccinated groups of 15 birds each and 2 control groups of 10 birds each. Groups 1, 2 and 3 were vaccinated intranasally (IN) with 100 ul of CLONE 30 per bird (live lentogenic strain- LaSota) at 1 and 11 days and subcutaneously (S/C) with 0.3 ml of killed NDV vaccine per bird (inactivated lentogenic strain - Ulster strain) at one day old. The first and second groups were challenged with 100 µl of inoculum containing 106 median egg infectious doses (EID50) of B7 strain (KM288609) at 21 and 30 days respectively while the 3rd group was kept unchallenged and received 100 µl of buffered saline solution. The 4th group was challenged with 106 EID50 of B7 without previous vaccination.
Birds were observed daily for 14 days post challenge. Trachea, lung, proventriculus and duodenum samples from all dead birds and 3 chickens from the vaccinated groups were collected for histopathology. Oropharyngeal and cloacal swabs were collected in buffered saline solution with antibiotic for quantification of virus shedding. Briefly, oropharyngeal and cloacal swabs were collected at 2, 7 and 14 days post challenge and the viral RNA extracted using a QIAamp viral RNA mini kit (QIAGEN, Germany) according to the manufacturer’s instruction. The specific primers, probe and PCR conditions used were as previously described by Wise, et al. [49]. Quantification of virus shedding was performed by absolute quantification using a standard curve for a known titred strain.
Statistical analysis
Statistically significant differences in the level of viral shedding between different groups were evaluated using the Student’s t-test in Microsoft Excel.
Results
NDV detection, confirmation and pathogenicity evaluation
The sixteen samples, each representing one farm, were positive for NDV and negative for IB, AIV-H5, H7 and H9 by real-time PCR. These results were confirmed by HA and HI results (Table 1). Furthermore, the results of the pathogenicity characterization of the sixteen isolates using MDT and ICPI indicated that fifteen of the tested isolates showed a MDTs ≤ 48 hrs. The ICPI was calculated for the 4 following strains, B7, B270, F460 and F388, and they were characterized as velogenic strains based on their respective ICPIs of 1.73, 1.72, 1.71 and 1.66 (Table 1). Only one strain “B81” showed a MDT > 108 hrs (Table 1).
Genetic and phylogenetic analysis
Based on the results of the partial F gene sequencing, the cleavage site amino acid sequence of the sixteen isolates was consistent with the pathotyping results identified by MDT and ICPI. However, three separate patterns of cleavage site sequence were identified in the velogenic isolates (Table 1).
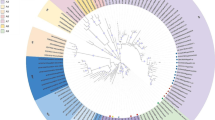
Phylogenetic analysis results revealed that fifteen isolates were assigned to class II genotype VII while isolate B81 was assigned to class II genotype II (Fig. 1). Further, the sequences of the fifteen velogenic isolates were investigated based on: the criteria for identification of NDV subgenotypes proposed by Diel, et al. [12]; and the genetic characterization of virulent viruses circulating in west and central Africa investigated by Snoeck, et al. [46]. The results revealed that all fifteen velogenic viruses in the present study belong to genotype VIIb (Fig. 2).
Protection study
A vaccination program which includes vaccination at one day of age with inactivated and live vaccines followed by live vaccination at 10 days old was investigated. Geometric mean HI (log2) titers exhibited following this program were 7.26, 7.86 and 7.6 at 7, 14 and 21 days post vaccination (dpv) respectively. To determine whether these titers could protect and efficiently reduce viral shedding and, in addition, to evaluate whether the timing of challenge could effect vaccine efficiency, we inoculated vaccinated chickens with 100 µl of inoculum containing 106 EID50 of B7 virus at 21 or 30 dpv.
In non-vaccinated challenged groups, all chickens developed severe depression, respiratory and nervous signs that began at 2 days post challenge (dpc). By 5 dpc mortality had reached 100% with a high level of viral shedding (Table 2). Neither clinical signs nor mortalities were observed in the two vaccinated groups. However, higher virus shedding was observed after challenge at 21 dpv compared to 30 dpv (at 2 and 7 dpc); however, at 14 dpc no virus shedding was observed in both groups (Table 3).
The histopathological examination of trachea, lung, proventriculus and small intestine samples revealed severe inflammatory and necrotic reactions in response to infection without prior vaccination (Fig. 3). On the other hand mild or no reactions were recorded in the vaccinated but challenged groups (Fig. 4). No pronounced differences were observed between the birds challenged at 21 and 30 dpv.
The histopathological examination of trachea, lung, proventriculus and small intestine samples revealed severe inflammatory and necrotic reaction in response to infection without prior vaccination. Briefly, (A) Trachea showed sloughed epithelium with haemorrhages (x20). (B) Air capillaries were filled with cellular exudates (x20). (C) Proventriculus showed haemorrhages in the mucosal epithelium and severe lymphocytic inflammation (x10). (D) Severe sloughed necrotic intestinal villi (x20)
The histopathological examination of trachea, lung, proventriculus and small intestine samples revealed mild or non-existent reactions within the vaccinated-challenged groups. Briefly, (A) The group of vaccinated-challenged birds showed normal histological structure within the trachea, (B) lungs, (C) proventriculus and (D) intestine (all figures are x20, Haemotoxylin and Eosin, H&E staining)
Discussion
ND represents one of the most important avian viral diseases and has a devastating impact on poultry production worldwide. Velogenic NDV is widely spread in many areas affecting intensive poultry production globally [32], including Egypt, with recurrent outbreaks being frequently reported [36].
NDVs with an ICPI ≥0.7, multiple basic amino acids (at least three arginine (R) or lysine (K)) within the cleavage site of the fusion protein [38] and an MDT of less than 60 hours [37] are designated as velogenic viruses. Based on these criteria, 15 out of the 16 samples tested in the present study were velogenic viruses. The MDT was 36 to 48 hours for all fifteen isolates and the ICPI for the four tested isolates was between 1.66 and 1.73. Furthermore, partial sequencing of the F gene showed the presence of polybasic amino acid motifs 112RRQKRF117 in all fifteen isolates except B7 and B109. This cleavage site pattern has previously been reported in European genotype VII isolates [1] as well as isolates in China [27]. The B7 sequence 112RKQKRF117 motif was also previously identified in highly virulent NDV isolates [44].
In the context of our phylogenetic analysis, it is worth noting that globally three main panzootics have been identified [4, 28]. The first started in the mid 1920s continuing until the 1950s and caused by NDV genotypes II, III, and IV. This seemed to be restricted to the South East of Asia where the outbreaks began. In the late 1960s and 1970s genotypes V and VI emerged and caused the second pandemic. The third pandemic, caused by VIb emerged during the 1980s. Since then, genotype VII has spread worldwide and is thought to be responsible for outbreaks reported in the Far East, Middle East, Africa and Europe [18, 28, 32] as well as South America [39], constituting the fourth panzootic [7]. In the present study, phylogenetic analysis revealed that all the virulent viruses belonged to genotype VIIb, according to the unified nomenclature and classification system of NDV proposed by Diel, et al. [12]. VIIb along with VIId are the predominant genotypes and still continue to spread in the world; however, other new sub-genotypes (VIIi and VIIh) have emerged and seem to have replaced older subgroups in countries like Pakistan and Israel [34].
In addition to direct airborne transmission, NDV can be transmitted via a fecal-oral route and through feed and water contaminated with infected respiratory secretions [3]. It is worth noting that all APMV1 viruses belong to a single serotype and vaccination with any NDV vaccine should protect against morbidity and mortality from any virulent NDV challenge; however this does not equally inhibit virus shedding [31]. If we bear in mind that mass-vaccination can protect only about 60% of the flock with the remaining animals having insufficient immunity [11] limiting NDV shedding represents a major concern for the control and prevention of this disease. Considering these factors, in the present study we examined whether heterologous vaccines could provide immunity that prohibits virus shedding, and whether this depends on antibodies titers and the timing between vaccination and challenge.
In the present study isolate B7, which had been confirmed to be velogenic with the highest ICPI among the tested isolates, was selected for in vivo challenge. Challenge of birds with 106 EID50 (4th group) caused the deaths of all birds at 5 dpc with very high viral shedding (Table 2) and typical respiratory and nervous system manifestations. Using the proposed vaccination program significantly reduced the viral shedding at 2 dpc with no clinical signs, when compared to the positive control group. Furthermore no viral shedding could be detected at 14 dpc. These findings indicate that heterologous vaccination not only prevent mortality and morbidity but also could reduce viral shedding to a very low level. This could be attributed to the very high levels of antibody induced by the proposed vaccination program as well as the efficiency of vaccines applied under experimental conditions (flock immunity) [48]. However, it is worth noting that under field conditions multiple factors could contribute in a reduction in the effectiveness of vaccination, which highlights the importance of antibody specificity [33]. Also, it is worth noting that challenging at 30 dpv significantly reduced viral shedding, when compared to challenging at 21 dpv. This finding supported by the Miller, et al. [33] conclusion that highlighted the importance of having enough time between vaccination and challenge to develop a proper immune response to prevent transmission. Further work is required to select vaccine strains that confer high antibody titers and specificity to the currently circulating field strains in Egypt. Until this is achieved vaccines inducing high antibody titres and a vaccination process providing higher flock (herd) immunity is strongly recommended to control ND in Egypt.
References
Abolnik C, Horner R, Bisschop S, Parker M, Romito M, Viljoen G (2004) A phylogenetic study of South African Newcastle disease virus strains isolated between 1990 and 2002 suggests epidemiological origins in the Far East. Arch Virol 149:603–619
Aldous E, Mynn J, Banks J, Alexander D (2003) A molecular epidemiological study of avian paramyxovirus type 1 (Newcastle disease virus) isolates by phylogenetic analysis of a partial nucleotide sequence of the fusion protein gene. Avian Pathol 32:237–255
Alexander D (1988) Newcastle disease: methods of spread. In: Alexander DJ (ed) Newcastle disease. Kluwer Academic Publishers, Boston, pp 256–272
Alexander D (1988) Newcastle disease: historical aspect. In: Alexander DJ (ed) Newcastle disease. Kluwer Academic Publishers, Boston, pp 1–10
Alexander D, Campbell G, Manvell R, Collins M, Parsons G, McNulty M (1992) Characterisation of an antigenically unusual virus responsible for two outbreaks of Newcastle disease in the Republic of Ireland in 1990. Vet Rec 130:65–68
Alexander D (2009) Newcastle, disease: OIE terrestrial manual. In: Manual of diagnostic tests and vaccines for terrestrial animals, chapter 2, p 3
Berhanu A, Ideris A, Omar AR, Bejo MH (2010) Molecular characterization of partial fusion gene and C-terminus extension length of haemagglutinin-neuraminidase gene of recently isolated Newcastle disease virus isolates in Malaysia. Virol J 7:1
Callison S, Jackwood M, Hilt D (2001) Molecular characterization of infectious bronchitis virus isolates foreign to the United States and comparison with United States isolates. Avian Dis 492–499
Courtney SC, Susta L, Gomez D, Hines NL, Pedersen JC, Brown CC, Miller PJ, Afonso CL (2013) Highly divergent virulent isolates of Newcastle disease virus from the Dominican Republic are members of a new genotype that may have evolved unnoticed for over 2 decades. J Clin Microbiol 51:508–517
Daubney R, Mansy W (1948) The occurrence of Newcastle disease in Egypt. J Comp Pathol Therap 58:189–200
Degefa T, Dadi L, Yami A, Nassir M (2004) Technical and economic evaluation of different methods of Newcastle disease vaccine administration. J Vet Med Ser A 51:365–369
Diel DG, da Silva LH, Liu H, Wang Z, Miller PJ, Afonso CL (2012) Genetic diversity of avian paramyxovirus type 1: proposal for a unified nomenclature and classification system of Newcastle disease virus genotypes. Infect Genetics Evol 12:1770–1779
Diel DG, Susta L, Garcia SC, Killian ML, Brown CC, Miller PJ, Afonso CL (2012) Complete genome and clinicopathological characterization of a virulent Newcastle disease virus isolate from South America. J Clin Microbiol 50:378–387
Dimitrov KM, Ramey AM, Qiu X, Bahl J, Afonso CL (2016) Temporal, geographic, and host distribution of avian paramyxovirus 1 (Newcastle disease virus). Infect Genetics Evol 39:22–34
Glickman RL, Syddall RJ, Iorio RM, Sheehan JP, Bratt MA (1988) Quantitative basic residue requirements in the cleavage-activation site of the fusion glycoprotein as a determinant of virulence for Newcastle disease virus. J Virol 62:354–356
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. In: Nucleic acids symposium series, pp 95–98
Hanson R, Brandly C (1955) Identification of vaccine strains of Newcastle disease virus. Science 122:156–157
Herczeg J, Wehmann E, Bragg R, Dias PT, Hadjiev G, Werner O, Lomniczi B (1999) Two novel genetic groups (VIIb and VIII) responsible for recent Newcastle disease outbreaks in Southern Africa, one (VIIb) of which reached Southern Europe. Arch Virol 144:2087–2099
Hu Z, Hu S, Meng C, Wang X, Zhu J, Liu X (2011) Generation of a genotype VII Newcastle disease virus vaccine candidate with high yield in embryonated chicken eggs. Avian Dis 55:391–397
Hussein H, El-Sanousi A, Yousif A (2005) Sequence analysis of fusion and matrix protein genes of the velogenic viscerotropic New castle disease virus Egyptian strain SR/76. Int J Virol 1:38
Hussein H, Emara M, Rohaim M (2014) Molecular Characterization of Newcastle disease virus genotype VIID in avian influenza H5N1 infected broiler flock in Egypt. Int J Virol 10:46–54
Jeon W-J, Lee E-K, Lee Y-J, Jeong O-M, Kim Y-J, Kwon J-H, Choi K-S (2008) Protective efficacy of commercial inactivated Newcastle disease virus vaccines in chickens against a recent Korean epizootic strain. J Vet Sci 9:295–300
Kapczynski DR, Afonso CL, Miller PJ (2013) Immune responses of poultry to Newcastle disease virus. Dev Comp Immunol 41:447–453
Kattenbelt JA, Stevens MP, Selleck PW, Gould AR (2010) Analysis of Newcastle disease virus quasispecies and factors affecting the emergence of virulent virus. Arch Virol 155:1607–1615
King D (1996) Influence of chicken breed on pathogenicity evaluation of velogenic neurotropic Newcastle disease virus isolates from cormorants and turkeys. Avian Dis 40(1):210–217
Lamb RA, Kolakofsky D (1996) Paramyxoviridae: the viruses and their replication. In: Fields BN, Knipe DM, Howley PM (eds) Virology. Lipincott-Raven, Philadelphia, pp 1177–1203
Liu X, Wan H, Ni X, Wu Y, Liu W (2003) Pathotypical and genotypical characterization of strains of Newcastle disease virus isolated from outbreaks in chicken and goose flocks in some regions of China during 1985–2001. Arch Virol 148:1387–1403
Lomniczi B, Wehmann E, Herczeg J, Ballagi-Pordany A, Kaleta E, Werner O, Meulemans G, Jorgensen P, Mante A, Gielkens A (1998) Newcastle disease outbreaks in recent years in western Europe were caused by an old (VI) and a novel genotype (VII). Arch Virol 143:49–64
Mase M, Imai K, Sanada Y, Sanada N, Yuasa N, Imada T, Tsukamoto K, Yamaguchi S (2002) Phylogenetic analysis of Newcastle disease virus genotypes isolated in Japan. J Clin Microbiol 40:3826–3830
Mayo M (2002) A summary of taxonomic changes recently approved by ICTV. Arch Virol 147:1655–1656
Miller PJ, Estevez C, Yu Q, Suarez DL, King DJ (2009) Comparison of viral shedding following vaccination with inactivated and live Newcastle disease vaccines formulated with wild-type and recombinant viruses. Avian Dis 53:39–49
Miller PJ, Decanini EL, Afonso CL (2010) Newcastle disease: evolution of genotypes and the related diagnostic challenges. Infect Genetics Evol 10:26–35
Miller PJ, Afonso CL, El Attrache J, Dorsey KM, Courtney SC, Guo Z, Kapczynski DR (2013) Effects of Newcastle disease virus vaccine antibodies on the shedding and transmission of challenge viruses. Dev Comp Immunol 41:505–513
Miller PJ, Haddas R, Simanov L, Lublin A, Rehmani SF, Wajid A, Bibi T, Khan TA, Yaqub T, Setiyaningsih S (2015) Identification of new sub-genotypes of virulent Newcastle disease virus with potential panzootic features. Infect Genetics Evol 29:216–229
Mohamed MH, Kumar S, Paldurai A, Megahed MM, Ghanem IA, Lebdah MA, Samal SK (2009) Complete genome sequence of a virulent Newcastle disease virus isolated from an outbreak in chickens in Egypt. Virus Genes 39:234–237
Mohamed MH, Kumar S, Paldurai A, Samal SK (2011) Sequence analysis of fusion protein gene of Newcastle disease virus isolated from outbreaks in Egypt during 2006. Virol J 8:1
OIE (2004) Newcastle disease (chapter 2.1.15) from OIE manual of diagnostic tests and vaccines for terrestrial animals. http://www.cfsph.iastate.edu/HPAI/resources/Additional%20Resources/OIEManual2_1_15.pdf
OIE (2012) Newcastle disease, chap 2.3.14. In: OIE terrestrial manual 2012: manual of diagnostic tests and vaccines for terrestrial animals (World Organisation for Animal Health, Paris), pp 576–589. http://www.oie.int/fileadmin/Home/eng/Health_standards/tahm/2.03.14_NEWCASTLE_DIS.pdf
Perozo F, Marcano R, Afonso CL (2012) Biological and phylogenetic characterization of a genotype VII Newcastle disease virus from Venezuela: efficacy of field vaccination. J Clin Microbiol 50:1204–1208
Radwan M, Darwish S, El-Sabagh I, El-Sanousi A, Shalaby M (2013) Isolation and molecular characterization of Newcastle disease virus genotypes II and VIId in Egypt between 2011 and 2012. Virus Genes 47:311–316
Reed LJ, Muench H (1938) A simple method of estimating fifty per cent endpoints. Am J Epidemiol 27:493–497
Rui Z, Juan P, Jingliang S, Jixun Z, Xiaoting W, Shouping Z, Xiaojiao L, Guozhong Z (2010) Phylogenetic characterization of Newcastle disease virus isolated in the mainland of China during 2001–2009. Vet Microbiol 141:246–257
Shabat MB, Meir R, Haddas R, Lapin E, Shkoda I, Raibstein I, Perk S, Davidson I (2010) Development of a real-time TaqMan RT-PCR assay for the detection of H9N2 avian influenza viruses. J Virol Methods 168:72–77
Shabbir MZ, Zohari S, Yaqub T, Nazir J, Shabbir MAB, Mukhtar N, Shafee M, Sajid M, Anees M, Abbas M (2013) Genetic diversity of Newcastle disease virus in Pakistan: a countrywide perspective. Virol J 10:1
Slomka M, Pavlidis T, Banks J, Shell W, McNally A, Essen S, Brown I (2007) Validated H5 Eurasian real-time reverse transcriptase-polymerase chain reaction and its application in H5N1 outbreaks in 2005–2006. Avian Dis 51:373–377
Snoeck CJ, Owoade AA, Couacy-Hymann E, Alkali BR, Okwen MP, Adeyanju AT, Komoyo GF, Nakouné E, Le Faou A, Muller CP (2013) High genetic diversity of Newcastle disease virus in poultry in West and Central Africa: cocirculation of genotype XIV and newly defined genotypes XVII and XVIII. J Clin Microbiol 51:2250–2260
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Van Boven M, Bouma A, Fabri TH, Katsma E, Hartog L, Koch G (2008) Herd immunity to Newcastle disease virus in poultry by vaccination. Avian Pathol 37:1–5
Wise MG, Suarez DL, Seal BS, Pedersen JC, Senne DA, King DJ, Kapczynski DR, Spackman E (2004) Development of a real-time reverse-transcription PCR for detection of Newcastle disease virus RNA in clinical samples. J Clin Microbiol 42:329–338
Yi J, Liu C, Chen B, Wu S (2011) Molecular characterization of a virulent genotype VIId strain of Newcastle disease virus from farmed chickens in Shanghai. Avian Dis 55:279–284
Acknowledgements
The authors would like to thank collogues in Virology and Biotechnology Unit, Reference Laboratory for Veterinary Quality Control on Poultry Production for their technical help. They would also like to thank the broiler farms’ owners for their cooperation and patience during sample collection.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Funding
A. Samy is currently a postdoctoral fellow in Anses France, funded by the Science & Technology Development Fund in Egypt (STDF) and Institut Français d’Egypte (IFE), Project ID 18551.
Conflict of interest
All authors declare that they have no conflict of interest.
Ethical approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed.
This article does not contain any studies with human participants performed by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Saad, A.M., Samy, A., Soliman, M.A. et al. Genotypic and pathogenic characterization of genotype VII Newcastle disease viruses isolated from commercial farms in Egypt and evaluation of heterologous antibody responses. Arch Virol 162, 1985–1994 (2017). https://doi.org/10.1007/s00705-017-3336-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-017-3336-y