Abstract
Here, we report the genome sequence of a feral pigeon alphaherpesvirus (columbid herpesvirus type 1, CoHV-1), strain HLJ, and compare it with other avian alphaherpesviruses. The CoHV-1 strain HLJ genome is 204,237 bp in length and encodes approximately 130 putative protein-coding genes. Phylogenetically, CoHV-1 complete genome resides in a monophyletic group with the falconid herpesvirus type 1 (FaHV-1) genome, distant from other alphaherpesviruses. Interestingly, the evolutionary analysis of partial genes of CoHV-1 isolated from different organisms and areas (currently accessible on GenBank) indicates that the CoHV-1 HLJ strain isolated from pigeon (Columba livia) is closely related to the strains isolated from peregrine falcon (Falco peregrinus) in Poland and owl (Bubo virginianus) in USA. These results may suggest possible transmission of the virus between different organisms and different geographic areas.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Columbid herpesvirus type 1 (CoHV-1), classified in the Mardivirus genus, subfamily Alphaherpesvirinae and family Herpesviridae, can lead to high mortality rates in young pigeons, with clinical signs in infected pigeons presenting as depression, anorexia or conjunctivitis [1]. Specific lesions can also occur as well as upper respiratory tract inflammation and ulceration as well as hepatic and splenic necrosis [2], and even neurological signs [3, 4]. The observed clinical signs in pigeon are called Smadel’s disease [1].
In May 2013, the CoHV-1 HLJ strain was isolated from an unvaccinated feral pigeon in the KangDa Livestock and Poultry Disease Consulting Service Centre in Heilongjiang Province, China [5]. The virus was propagated on primary chicken embryo fibroblasts, before the nucleocapsid DNA was purified using the sodium dodecyl sulfate-proteinase K and phenol/chloroform protocol [6]. A pair of primers targeting the CoHV-1 glycoprotein B gene (accession numbers: KU051711) was designed to identify CoHV-1 genome in the DNA we had purified. Furthermore, a total of 95 pairs of primers (Supplementary table 1) were designed according to other alphaherpesviruses (accession numbers: KJ668231; JN542536; JX844666; NC_002577; AF291866; JQ647509 and AY372243) to amplify the whole CoHV-1 genome. Subsequently, CoHV-1 genome sequences were assembled using the Seqman program (DNASTAR, Madison, WI) and mapped manually. The CoHV-1 HLJ strain genome sequence has been deposited in the GenBank database and has the accession number KX589235.
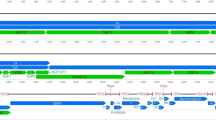
The CoHV-1 genome was 204,237 bp in length, with an overall G/C base composition of 61.5%, and it exhibited class E structural characteristics (Supplementary figure 1), similar to Falconid herpesvirus 1 (FaHV-1), Gallid herpesvirus 2 (GaHV-2), Gallid herpesvirus 3 (GaHV-3) and Meleagrid herpesvirus 1 (MeHV-1), but distinct from Gallid herpesvirus 1 (GaHV-1), Anatid herpesvirus 1 (AnHV-1) and Psittacid herpesvirus (PsHV-1), which contain class D genomes. The E class of herpesvirus genomes contain a UL and US, which are flanked by large inverted repeat regions. The CoHV-1 genome was annotated to contain 130 potential coding genes based on the criteria outlined in the methods (Supplementary table 2). The genomic organization was similar to the FaHV-1 genome and the distributions of specific regions were as follows: the TRL region extended from nt 1 to 6998 (67.4% G/C); the UL region extended from nt 6999 to 143548 (60.7% G/C); the IRL region extended from nt 143549 to 150546 (67.4% G/C); the IRS region extended from nt 150547 to 172445 (64% G/C); the US region extended from nt 172446 to 182338 (51.5% G/C) and finally the TRS region extended from nt 182339 to 204237 (64% G/C). The CoHV-1 genome had the largest genome of any avian alphaherpesvirus sequenced to date.
To examine possible incongruence between gene regions and gene region combinations, phylogenetic history was hypothesized using partitioned Bayesian inference (BI) to generate a tree to explore the phylogenetic and molecular evolution of 8 representative genomes, based on UL, IRL+IRS, US and UL+IRL+IRS+US. The trees in this study were constructed using MrBayes v.3.1 [7]. Phylogenetically, the dendrogram positioned CoHV-1 and FaHV-1 in a monophyletic group (Figure 1). Although multiple alignment of the nucleotide sequences of the FaHV-1 genome (130 gene sequences) along with CoHV-1 genome shared up to 99.4% identity, 81 amino acid sequences encoding by these genes were inconsistent, including LORF3, envelope glycoprotein C (gC), ICP4, LORF5, ICP22, US3, envelope glycoprotein D (gD), envelope glycoprotein I (gI) and envelope glycoprotein E (gE) etc (Supplementary table 3). ICP4 is closely related to the reactivation of latent virus [8]. In addition, glycoproteins play a vital role in virus attachment to and penetration of host cells and are directly related to host range diversity [9]. These mutations in the CoHV-1 genome sequence may affect the reactivation capacity of latent virus, virus attachment and spreading in host cells. These mutations may even be related to host range diversity.
Phylogenetic dendrograms analyzing the CoHV-1 HLJ strain and 8 other alphaherpesviruses. A. The phylogenetic tree was generated from a multiple alignment of UL. B. The phylogenetic tree was generated from a multiple alignment of IRL+IRS. C. The tree was generated from a multiple alignment of US. D. The phylogenetic tree was generated from a multiple alignment of whole genome sequences (UL+IRL+IRS+US). The GenBank accession numbers of the sequences used are as follows: HHV-1 (NC_001806), AnHV-1 (JQ647509), GaHV-1 (JN542536), PsHV-1 (AY372243), GaHV-2 (JX844666), GaHV-3 (NC_002577), MeHV-1 (AF291866) and FaHV-1 (KJ688231). The red circle represents the CoHV-1 HLJ strain (colour figure online)
Phylogenetic trees constructed using partial CoHV-1 DNA-dependent DNA polymerase gene sequences (available on GenBank) and the corresponding regions of the CoHV-1 HLJ strain genome as well as two alphaherpesviruses suggested that the CoHV-1 HLJ strain isolated from pigeon (Columba livia) is closely related to strains isolated from peregrine falcon (Falco peregrinus) in Poland and owl (Bubo virginianus) in the USA (Figure 2). These results suggest possible transmission of the virus between different host organisms and between different geographic areas.
Phylogenetic dendrograms comparing partial genes from CoHV-1 sequences accessible on GenBank and the CoHV-1 HLJ strain. A. The tree was generated from a multiple alignment of 20 partial genes (eighteen CoHV-1, one FaHV-1 and one HHV-1) and the CoHV-1 HLJ strain. The GenBank accession numbers of the sequences used were as follows: JX892992, JX892993, JX892994, JX892995, JX892996, JX892997, JX892998, JX892999, JX893000, JX893001, JX893002, JX893003, JX893004, JX893005, JX893006, JX893007, JX893008, JX893009, FaHV-1 (KJ668231) and HHV-1 (NC_001806), respectively. B. The tree was generated from a multiple alignment of 11 partial genes (nine CoHV-1, one FaHV-1 and one HHV-1) and the CoHV-1 HLJ strain. The GenBank accession numbers of the sequences used were as follows: EF522952, EF522953, EF522954, EF522955, EF522956, EF522957, EF522958, EF522959, EF522960, FaHV-1 (KJ668231) and HHV-1 (NC_001806). The red square represents the CoHV-1 HLJ strain (colour figure online)
References
Smadel JE, Jackson EB, Harman JW (1945) A new virus disease of pigeons: recovery of the virus. J Exp Med 81:385–398
Gailbreath KL, Oaks JL (2008) Herpesviral inclusion body disease in owls and falcons is caused by the pigeon herpesvirus (columbid herpesvirus 1). J Wildl Dis 44:427–433
Pinkerton ME, Wellehan JJ, Johnson AJ, Childress AL, Fitzgerald SD, Kinsel MJ (2008) Columbid herpesvirus-1 in two Cooper’s hawks (Accipiter cooperii) with fatal inclusion body disease. J Wildl Dis 44:622–628
Wozniakowski GJ, Samorek-Salamonowicz E, Szymanski P, Wencel P, Houszka M (2013) Phylogenetic analysis of Columbid herpesvirus-1 in rock pigeons, birds of prey and non-raptorial birds in Poland. BMC Vet Res 9:52
Zhao PP, Ma J, Guo Y, Tian L, Guo GY, Zhang KX, Xing MW (2015) Isolation and characterization of a herpesvirus from feral pigeons in China. Vet J 206:417–419
Goldenberger D, Perschil I, Ritzler M, Altwegg M (1995) A simple “universal” DNA extraction procedure using SDS and proteinase K is compatible with direct PCR amplification. PCR Methods Appl 4:368–370
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Huang W, Xie P, Xu M, Li P, Zao G (2011) The influence of stress factors on the reactivation of latent herpes simplex virus type 1 in infected mice. Cell Biochem Biophys 61:115–122
Wang J, Hoper D, Beer M, Osterrieder N (2011) Complete genome sequence of virulent duck enteritis virus (DEV) strain 2085 and comparison with genome sequences of virulent and attenuated DEV strains. Virus Res 160:316–325
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no financial or personal relationships with other people or organizations that could inappropriately influence or bias the content of this paper.
Ethical approval
All procedures used in this study were approved by the Institution Animal Care and Use Committee of Northeast Forestry University (UT-17; 25 July 2012).
Funding
This study was provided by the National Natural Science Foundation of China (Grant No. 31672619), the Fundamental Research Funds for the Central Universities (Grant No. 2572016EAJ5) and the Natural Science Foundation of Heilongjiang Province (Grant No. C2015061).
Electronic supplementary material
Below is the link to the electronic supplementary material.
705_2017_3329_MOESM1_ESM.tif
Supplementary Figure 1: Genetic organization of the CoHV-1 genome. This map shows the class E genome structure of CoHV-1 and indicates the location, direction and sizes of annotated ORFs, numbered based on the CoHV-1 locus tag and labeled according to homology with the characterized HHV-1 genome as well as other avian herpesviruses. The genomic genes were numbered from 5’ to 3’ accordant with the direction of the black arrows in the map. (TIFF 6115 kb)
Rights and permissions
About this article
Cite this article
Guo, Y., Li, S., Sun, X. et al. Complete genome sequence and evolution analysis of a columbid herpesvirus type 1 from feral pigeon in China. Arch Virol 162, 2131–2133 (2017). https://doi.org/10.1007/s00705-017-3329-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-017-3329-x