Abstract
Genetic factors play a crucial role for the pathophysiology of treatment-resistant depression (TRD). It has been established that Catechol-O-methyltransferase (COMT) and cyclic amp-response element-binding protein (CREB) are associated with antidepressant response. The aim of this study was to explore the association between single nucleotide polymorphisms (SNPs) in COMT and CREB1 genes and TRD in a Chinese population. We recruited 181 patients with major depressive disorder (MDD) and 80 healthy controls, including 81 TRD patients. Depressive symptoms were assessed with the Hamilton Depression Rating Scale-17 (HDRS). Genotyping was performed using mass spectrometry. Genetic analyses were conducted by PLINK Software. The distribution of COMT SNP rs4818 allele and genotypes were significantly different between TRD and controls. Statistical differences in allele frequencies were observed between TRD and non-TRD groups, including rs11904814 and rs6740584 in CREB1 gene, rs4680 and rs4818 in COMT gene. There were differences in the distribution of HDRS total scores among different phenotypes of CREB1 rs11904814, CREB1 rs6740584, COMT rs4680 and rs4818. Gene–gene interaction effect of COMT–CREB1 (rs4680 × rs6740584) revealed significant epistasis in TRD. There findings indicate that COMT and CREB1 polymorphisms influence the risk of TRD and affect the severity of depressive symptoms of MDD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Major depressive disorder (MDD) is a common, recurrent and disabling psychiatric disorder with a 12-month prevalence of approximately 6% and a lifetime risk of nearly 20% (Malhi and Mann 2018). It has been reported that MDD will be a major cause of global disease burden by 2030 (Briley and Lépine 2011). 1/3 MDD patients showed treatment-refractory after recerving multiple pharmacological treatments despite significant improvements in depressive symptoms under the treatment of antidepressants (Carvalho et al. 2014). Patients with treatment-resistant depression (TRD) have continuous psychiatric and somatic symptoms that deteriorate their social functioning abilities and quality of life (Lex et al. 2019; Fava 2003; Chen et al. 2019). The prognosis of TRD was terrible and the recurrence rate was higher than that of non-TRD, which greatly increased the burden on social functioning and economic impact (Demyttenaere and Van Duppen 2019; Johnston et al. 2019).
The etiology of TRD is complex and multifactorial. Genetic factor has a significant impact on the clinical outcomes of antidepressant treatment. Increasing researches focused on the association between gene variants and the susceptibility of TRD. Studies indicated that the polymorphisms in genes related to metabolic enzymes responsible for antidepressants transport and metabolic clearance play an important role in the progress of TRD (Halaris et al. 2021). Genome-wide association studies (GWAS) of TRD found that the genes related to cytoskeleton regulation and neural plasticity may be associated with TRD (Fabbri et al. 2018, 2019; O’Dushlaine et al. 2014). The transcription factor cAMP-responsive element-binding protein 1 (CREB1) is a member of the leucine zipper family and encoded by CREB1 that located in 2q33.3 (Xing et al. 1996). CREB1 is a transcription factor implicated in synaptic and neuronal plasticity base on mediated the BDNF pathway (Rafa-Zabłocka et al. 2018). Meta-analysis suggested that CREB1 as potentially crucial mediator for effects of antidepressant (Xiao et al. 2018). The literature provides some evidence of CREB1 gene implicated in the antidepressant response. Research found that the increasing of CREB levels in rodent models cause the antidepressant-like effect and the mice knocked-out of the CREB1 gene led to a antidepressants-resistant phenotype (Rafa-Zabłocka et al. 2018; Perlis et al. 2008). In addition, CREB was upregulate and neurogenesis was increased in the hippocampus after antidepressant treatment (Gundersen et al. 2013). A preliminary study detected that the haplotypes within CREB1 involved in the TRD (Serretti et al. 2011). The study by Raffaella Calati and colleagues on treatment resistance and CREB1 SNPs in MDD patients found that the variants of CREB1 were probably implicated in the treatment resistance (Calati et al. 2013). Although some studies failed to detected the relationship between CREB1 SNPs and the status of treatment-resistance, most of these studies supported the polymorphisms of CREB1 influence the remission of depressive symptoms (Serretti et al. 2011; Calati et al. 2013; Calabrò et al. 2018).
Based on the finding of “the European Group for the Study of Resistant Depression” (GSRD), the COMT gene responsible for the catecholamine inactivation also as a promising candidate genes in treatment-resistance (Fabbri et al. 2013; Kocabas et al. 2010; Schosser et al. 2012). Meta-analysis indicated that the polymorphism of COMT associated with the treatment response in MDD patients (Tang et al. 2020). COMT rs4680 (val158met) has been intensively investigated result in the decrease of enzyme activity and protein level (Calabrò et al. 2018). The role of COMT rs4680 and other COMT polymorphisms in antidepressant treatment response have been investigated, however, with inconsistent results. In addition, most findings on the genes associated with TRD have focused on Caucasians. Only one study has suggested that the COMT gene was associated with sex differences of fluoxetine therapeutic response in a Chinese population (Tsai 2009).
To our knowledge, there are no studies investigating the relationship between TRD and genes CREB1 and COMT in a Chinese Han population. As mentioned above, we hypothesized that the SNPs of COMT and CREB1 genes likely play a critical role in the pathophysiology and development of TRD. Thus, the aims of the preliminary study were: (1) to investigate the relationship between genes COMT and CREB1 SNPs and TRD; (2) to explore the association between the SNPs of the genes COMT and CREB1 and the severity of symptoms of TRD; (3) to examine the interaction effects of the genes COMT and CREB1 as they related to the risk for TRD.
Methods
Subjects
A total of 181 MDD patients (male/female: 63/118, average age: 44.22 ± 11.44 years, average duration: 25.20 ± 32.00 months) were recruited from the outpatient clinic of Tianjin Anding Hospital. The inclusion criteria for this study were as follows: (1) Han Chinese; (2) 18–65-years old; (3) patients met the DSM-IV diagnostic criteria of MDD, confirmed by two experienced psychiatrists through the Structured Clinical Interview for DSM-IV (SCID). The previous usage of drugs information was collected according to reports of close relatives and clinical medical records. All subjects were in good physical health. Subjects with a history of substance dependence or major medical illnesses (e.g., neurological disorder, cerebrovascular disease) were excluded.
MDD patients were divided into TRD group (male/female 34/47, age 45.99 ± 12.668 years) and Non-TRD group (male/female 29/71, age 42.79 ± 10.189 years). Patients diagnosed with TRD were those who did not respond to at least two different classes of antidepressants of adequate dose for at least 6 weeks during the current depressive episode based on the medical records. The choice of treatment medication was based on the previous treatment experience, efficacy, tolerance, long-term treatment plan, age, gender and economic status. The competent psychiatrist was select the appropriate drugs for the patient after comprehensive consideration these factors. The dosage of medication is determined by the psychiatrist according to the patient's condition and adverse reactions. In addition, the symptomatic treatment were applied for the comorbidity disorder including sleep disorders, anxiety disorders etc.
A total of 80 healthy controls (male/female 25/55, age 40.39 ± 11.589 years) were also recruited and were gender- and age-matched with the patients. All healthy controls were Han Chinese from the local community of Tianjin, screened by psychiatrists based on SCID diagnostic assessment to exclude subjects with any psychiatric disorders. Furthermore, none of the healthy controls had a family history of mental diseases.
Informed consent was obtained from all participants prior to the start of this research. This study was approved by the institutional review board and the ethic committee of Tianjin Anding Hospital in accordance with the Declaration of Helsinki. (No. 2019‐18).
Clinical assessments
The Hamilton Depression Rating Scale (HDRS) was established in 1960 (Hamilton 1960). The scale is still commonly used for the evaluation of depressive symptoms in current psychiatric research. In this study, the depressive symptoms of MDD patient were assessed by four psychiatrists using the HDRS (17-item). Before the study, all psychiatrists participated in an HDRS training session. Analysis of repeated assessments showed that the inter-observer correlation coefficient of the HDRS total score was more than 0.8.
Genotyping
After overnight fasting, blood samples were collected from peripheral vessels to 5-ml EDTA vacuum tubes. Genomic DNA samples were isolated from the whole blood samples using the high-salt method (Carpi et al. 2011). Genotyping was performed by primer extension of multiplex products with detection by matrix-assisted laser desorption time-of-flight mass spectrometry (Wolk et al. 2018). All genotyping was conducted by technicians who were blind to the clinical status of the participants. 10% of the samples were repeated for the genotyping assay, and the concordant rate was 100%.
Based on the prior knowledge about COMT and CREB1 involved in the pathogenesis of response to antidepressants, 6 SNPs of the two candidate genes were selected using a candidate gene approach, namely rs11904814 (CREB1), rs2253206 (CREB1), rs6740584 (CREB1), rs4633 (COMT), rs4680 (COMT) and rs4818 (COMT) (Table 1). Minor allele frequencies (MAFs) of Han Chinese were taken from dbSNP (http://www.ncbi.nlm.nih.gov/projects/SNP/).
Statistical analysis
In this study, counting data were shown as percentages, and descriptive data was shown as mean ± standard deviation (x ± s). Comparisons of the differences in demographic and clinical variables between MDD patients and healthy controls, TRD and non-TRD samples, t tests were used as continuous variables, and chi-squared tests or Fisher’s exact test were used as categorical variables.
The Hardy–Weinberg equilibrium (HWE) test was performed to estimate the representativeness of samples before the correlation analyses (Table 1). Comparisons of the distribution of alleles and genotypes of candidate genes SNPs between MDD, TRD, non-TRD and controls groups were performed using PLINK. Statistical analyses for specific phenotypes were carried out by SPSS software (version 25.0). The logistic regression model was used to test the association between candidate SNPs and depressive phenotypes. The odds ratios (ORs) with 95% confidence intervals (CIs) were presented.
Gene–gene interaction analyses (epistasis) were carried out by PLINK among candidate gene SNPs in all patients. Statistical significance was evaluated using two-tailed design-based tests and the P value was conservatively set at the 0.008 level of significance. Bonferroni corrections were used for multiple tests to reduce type I error.
Results
General information
As shown in Table 2, there were no significant differences in the demographic variables between MDD and controls, TRD and non-TRD groups. In addition, no statistical difference was found in the duration of illness reported in TRD and non-TRD groups.
Association analysis of the SNPs within candidate genes between TRD and non-TRD groups and controls
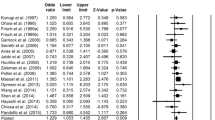
According to the stratified analysis of treatment response, the differences in allele and genotype distribution of the COMT SNP rs4818 between TRD and control groups were significant and the C allele showing higher frequency in TRD individuals than controls (allele C, P = 0.003, OR 2.477, 95% CI 1.350–4.543; P = 0.002) (Table 3). COMT SNP rs4680 was significantly different in the genotype frequencies (P < 0.001) between TRD and control groups (Table 3). In light of the low frequency of AA genotypes in non-TRD and control groups, we used the dominant model analysis (GG/GA + AA), demonstrating that the GG homozygote of COMT SNP rs4680 was more frequent in the TRD group than in healthy controls. There were no significant differences in SNPs of COMT and CREB1 genes between the non-TRD and the control groups (Fig. 1).
Association analysis of the SNPs of candidate genes between TRD and non-TRD groups
Based on the association analysis of the candidate gene SNPs and TRD, significant differences in allele and genotype distribution between the TRD and non-TRD groups were observed (Table 4). Significant differences were observed in rs11904814 of CREB1 gene (allele T, OR 2.096, 95% CI 1.312–3.350, P = 0.001) as well as rs4680 and rs4818 of COMT gene (allele G, OR 1.586, 95% CI 1.030–2.443, P = 0.036; allele C, OR 1.983, 95% CI 1.132–3.474, P = 0.016, respectively). The genotypes distribution of COMT rs4680 and rs4818 between the TRD and non-TRD groups were significant different (P < 0.001; P < 0.001, respectively). Compared to the non-TRD group, the allelic association of CREB1 rs6740584 was significant and more frequent C allele in the TRD group (allele C, OR 1.648, 95% CI 1.045–2.599, P = 0.031). After the Bonferroni corrections, most of the above significant variables passed the corrections, except for CREB1 rs6740584. For CREB1 rs2253206 and COMT rs4633, no significant association between TRD and SNPs was observed.
Comparing the severity of depressive symptoms in different genotypes of candidate gene SNPs
By comparing the two subgroups, it was noted that mean scores of HDRS in the TRD group were significantly higher than in the non-TRD group (29.40 ± 5.021 vs. 22.41 ± 3.626).
TRD subjects were classified according to their genotyping data, and total HDRS scores were compared between groups. HDRS scores differences among genotype of candidate gene SNPs in TRD patients can be seen in Table 5. Patients with GG genotype of rs11904814 of CREB1 demonstrated higher HDRS scores (29.1 ± 4.98) than GT and TT genotypes (P = 0.003). In addition, HDRS scores differed between the genotypes rs6740584 of CREB1 with patients with TT genotype demonstrating higher HDRS scores (28.5 ± 5.85) than CC genotype patients (P = 0.04). HDRS scores for the GA genotype of rs4680 of COMT were higher in MDD patients (27.66 ± 27.6) than AA and GG genotype patients (P < 0.001). Finally, TRD patients with the GC genotype of rs4818 of COMT displayed higher HDRS scores (27.26 ± 27.2) than CC and GG genotypes patients (P = 0.016). The association was not significant between genotype of CREB1 rs6740584 and COMT rs4818 and HDRS scores after Bonferroni corrections.
In non-TRD patients, there were no significant differences in HDRS total scores among different genotypes of candidate gene SNPs groups.
Gene–gene interaction analysis
The results of gene × gene interaction analysis showed that the COMT (rs4680) and CREB1 (rs6740584) genes have significant epistasis in TRD patients compare to non-TRD patients (P < 0.001). There was an interaction effect of COMT and CREB1 upon treatment-resistant.
Discussion
In this study, we explored the association between SNPs of candidate genes COMT and CREB1 in TRD to find genetic markers in the development of TRD in a Han Chinese population. This study first found that SNPs of COMT and CREB1 genes may be associated with TRD but not non-TRD. The main findings of this study were that: (1) CREB1 rs11904814 may increase the risk of TRD in MDD patients; (2) COMT rs4818 and rs4680 may confer the susceptibility to TRD; (3) TRD patients with GG genotype of rs11904818 (CREB1) and GA genotype of rs4680 (COMT) presented more severity depressive symptoms; (4) the interaction of COMT and CREB1 showed significant epistasis in TRD.
In the present study, we found that CREB1 rs11904814 was associated with TRD. The SNP rs11904814 located in intron 3 of CREB1 (Utge et al. 2010). Few study focus on the CREB1 rs11904814 for MDD patients. There were two studies found that the interaction between rs11904814 and the environment increased the susceptibility to MDD in Chinese Han populations (Wang et al. 2015; Ma et al. 2017). Another study had noted the association of CREB1 SNP rs11904814 with depression in men (Utge et al. 2010). To the best of our knowledge, no association analysis between CREB1 SNP rs11904814 and TRD has been reported. We observed that subjects carrying the T allele of CREB1 rs11904814 represented a lower risk to develop into TRD.
Mechanism of many antidepressants action was activated the NE or 5HT receptors and then coupled to adenylate cyclase/cAMP/PKA pathway or coupled to phosphatidylinositol and triggered the calcium influx. These signal pathways can induce CREB phosphorylation and modulated by set of target genes activated by CREB (Blendy 2006). Rats with overexpression of CREB in the hippocampal dentate gyrus showed significantly reduced immobility times in the FST.34 The expression of CREB1 mRNA and CREB was elevated in the hippocampus after antidepressant therapy in MDD patients (Xiao et al. 2018). CREB and CREB-dependent gene expression plays a critical role in response to signal transduction cascades stimulated by hormones, inflammatory factors and synaptic activity (Blendy 2006). Patients with TRD have the imbalance of inflammatory activity (Strawbridge et al. 2019). CREB1 encoded by CREB1 has been identified as a downstream target of the Mitogen-activated protein kinase (MAPK) pathway. MAPK1 and CREB1 have implicated in inflammatory processes and associated with the symptom remission after antidepressant treatment (Calati et al. 2013). The brain-derived neurotrophic factor (BDNF)—the neurotrophic tyrosine kinase receptor 2 (TrkB) pathway modulated the phosphorylation of CREB, and CREB mediated the up-regulation of BDNF that is closely related to therapeutic response (Chen et al. 2001). Previous study indicated that Polymorphisms of CREB1 affected response to paroxetine by altering the expression of transcription factor in BDNF signaling pathway (Murphy et al. 2013). In addition, CREB maintained the levels of BDNF in the hippocampus and prefrontal cortex (PFC) after antidepressant treatment (Rafa-Zabłocka et al. 2018). The CREB also has been shown to play a critical role in the neuronal plasticity and to alleviated the decrease gray matter volume in the hippocampus of TRD patients (Marchese et al. 2017).
In this study, we observed that COMT rs4818 and rs4680 increased the risk of TRD. The relationship between COMT SNPs and the antidepressants response has been widely studied. Evidence indicated that COMT SNP rs4680 is related to the efficacy of paroxetine in the early stage of depression, and that patients with GG genotype had a worse effect of paroxetine treatment than those with AA or AG genotypes (Benedetti et al. 2009). A similar result was found in response to fluvoxamine, with patients with GG homozygotes of COMT experiencing a poorer reaction than those with AA or AG genotypes (Benedetti et al. 2010). We also observed that the frequencies of GG genetype in TRD group was more frequent than non-TRD group. However, there were several studies finding no association between the COMT SNP rs4680 and reaction to antidepressants (Taranu et al. 2017; Brunoni et al. 2020; Chiesa et al. 2014). The opposites view have been suggested that patients with GG homozygotes were associated with better response to MECT treatment (Lin et al. 2015). Some possible explanations for this discrepancy could include differences in ethnicity, sample size, method of treatment for depression and some methodology differences (Li et al. 2014).
COMT encoded by the COMT gene is an enzyme implicated in monoamines degradation. As we know, dopamine regulated the process of motivation and emotion, reward, attention and psychomotor function that was impaired in MDD (Antypa et al. 2013). Study indicated that the DBS therapy on superolateral-branch of the medial forebrain bundle (MFB) increased the expression of dopamine D2 receptors in the hippocampus and present antidepressant responses in rats (Dandekar et al. 2017). In addition, dopamine agonists have been used as adjunctive drugs to improve the symptoms of TRD (Chen et al. 2013; Hori et al. 2013). COMT regulates the activity of noradrenaline and adrenaline which are essential for the action of antidepressants (Marchese et al. 2017). The polymorphisms of COMT lead to a change in dopamine concentration through changing enzymatic activity. It has been shown that the increase of stress-induced CRH mRNA depends on the noradrenergic pathway (Kovács 2013). The over-activation of the HPA axis also plays a critical role in TRD (Rosenblat et al. 2015). The increase in cortisol levels is associated with the severity of depressive symptoms, worse clinical response and prognosis (Li et al. 2021).
Our results suggested that the SNPs of CREB1 and COMT may affect the severity of depressive symptoms in patients with TRD. It was found that the CREB1 gene was associated with the severity and remission of depressive symptoms (Calati et al. 2013; Juhasz et al. 2011). Several studies have also suggested that the A allele of rs4680 correlated with lower depressive symptoms in Asians (Seib et al. 2016; Avinun et al. 2020). In contrast, another study found that COMT SNPs were not associated with the severity of depressive symptoms in a population of Chinese Han participants (Chao et al. 2019). These conflicting findings may be due to differences experimental design. In the current study, more severe depressive symptoms were observed in patients presenting with the specific genotype of CREB1 rs11904814 and COMT SNPs rs4680, these SNPs also influenced the susceptibility of TRD.
In present study, pairwise gene–gene interaction analyses (epistasis) were carried out by PLINK among candidate gene SNPs in all patients. Our results suggest an interaction of CREB1 and COMT polymorphisms on TRD susceptibility despite there were no allelic differences SNPs between TRD and healthy controls in the two SNPs. Some evidence indicated that epistatic interactions can be observed even if a single SNP individually leads to negative findings (Fabbri et al. 2013). CREB might be involved in the tissue-specific expression of COMT P1 promoter in rats (Tenhunen 1996). However, it is unclear that the specific effect of CREB1/COMT signaling pathway on TRD. The current results should be interpreted with caution and their interactions on development of TRD should be further examined.
The limitations of this research should be considered. (1) patients were treated with different types of antidepressants with different mechanisms of action. The effect of studied polymorphisms on a specific drug was difficult to interpreted and the effects of specific antidepressants should be explored in future. (2) the sample size relatively small and lack of independent replication, further studies with larger sample size should be conducted to understand the role of COMT and CREB1 in TRD susceptibility. (3) The interaction between COMT and CREB1 should be further investigated to elucidate the potential mechanism of TRD.
Conclusions
In conclusion, the SNPs of CREB1 and COMT increased the susceptibility of TRD and influenced the severity of depressive symptoms of TRD patients in a Chinese Han population. Based on our findings, we argue that our data support the idea that COMT and CREB1 genes have a genetic interaction that may increase the risk for developing TRD. Replicative studies of this work need a larger sample size to verify the relationship we observed above. In addition, we believe our data provides preliminary evidence to support a possible interaction effect between COMT and CREB1 genes in patients with TRD and points toward a potential mechanism of epistasis.
References
Antypa N, Drago A, Serretti A (2013) The role of COMT gene variants in depression: Bridging neuropsychological, behavioral and clinical phenotypes. Neurosci Biobehav Rev. 37(8):1597–610. https://doi.org/10.1016/j.neubiorev.2013.06.006
Avinun R, Nevo A, Radtke SR, Brigidi BD, Hariri AR (2020) Divergence of an association between depressive symptoms and a dopamine polygenic score in Caucasians and Asians. Eur Arch Psychiatry Clin Neurosci. 270:229–35. https://doi.org/10.1007/s00406-019-01040-x
Benedetti F, Colombo C, Pirovano A, Marino E, Smeraldi E (2009) The catechol-O-methyltransferase Val (108/158) Met polymorphism affects antidepressant response to paroxetine in a naturalistic setting. Psychopharmacology 203:155–160. https://doi.org/10.1007/s00213-008-1381-7
Benedetti F, Dallaspezia S, Colombo C, Lorenzi C, Pirovano A, Smeraldi E (2010) Effect of catechol-O-methyltransferase Val (108/158) Met polymorphism on antidepressant efficacy of fluvoxamine. Eur Psychiatry 25:476–478. https://doi.org/10.1016/j.eurpsy.2009.12.007
Blendy JA (2006) The role of CREB in depression and antidepressant treatment. Biol Psychiat 59:1144–1150. https://doi.org/10.1016/j.biopsych.2005.11.003
Brunoni AR, Carracedo A, Amigo OM, Pellicer AL, Talib L, Carvalho AF et al (2020) Association of BDNF, HTR2A, TPH1, SLC6A4, and COMT polymorphisms with tDCS and escitalopram efficacy: ancillary analysis of a double-blind, placebo-controlled trial. Braz J Psychiatr. 42:128–35. https://doi.org/10.1590/1516-4446-2019-0620
Calabrò M, Mandelli L, Crisafulli C, Lee SJ, Jun TY, Wang SM et al (2018) Neuroplasticity, neurotransmission and brain-related genes in major depression and bipolar disorder: focus on treatment outcomes in an asiatic sample. Adv Ther 35(10):1656–70. https://doi.org/10.1007/s12325-018-0781-2
Calati R, Crisafulli C, Balestri M, Serretti A, Spina E, Calabrò M et al (2013) Evaluation of the role of MAPK1 and CREB1 polymorphisms on treatment resistance, response and remission in mood disorder patients. Prog Neuropsychopharmacol Biol Psychiatry 44:271–278. https://doi.org/10.1016/j.pnpbp.2013.03.005
Carpi FM, Di Pietro F, Vincenzetti S, Mignini F, Napolioni V (2011) Human dna extraction methods: patents and applications. Recent Patents DNA Gene Seq. https://doi.org/10.2174/187221511794839264
Carvalho AF, Berk M, Hyphantis TN, McIntyre RS (2014) The integrative management of treatment-resistant depression: a comprehensive review and perspectives. Psychother Psychosom 83:70–88. https://doi.org/10.1159/000357500
Chao JK, Yang MC, Chen CS, Wang IC, Kao WT, Shi MD (2019) A gender-specific COMT haplotype contributes to risk modulation rather than disease severity of major depressive disorder in a Chinese population. J Affect Disord 246:376–386. https://doi.org/10.1016/j.jad.2018.12.088
Chen TY, Tzeng NS (2013) Aripiprazole: a dopamine modulator that mimics methylphenidate in producing faster antidepressant effects. Med Hypotheses 81:183–185. https://doi.org/10.1016/j.mehy.2013.05.009
Chen AC, Shirayama Y, Shin KH, Neve RL, Duman RS (2001) Expression of the cAMP response element binding protein (CREB) in hippocampus produces an antidepressant effect. Biol Psychiatry. 49(9):753–62. https://doi.org/10.1016/s0006-3223(00)01114-8
Chen MH, Lin WC, Wu HJ, Cheng CM, Li CT, Hong CJ et al (2019) Antisuicidal effect, BDNF Val66Met polymorphism, and low-dose ketamine infusion: Reanalysis of adjunctive ketamine study of Taiwanese patients with treatment-resistant depression (AKSTP-TRD). J Affect Disord 251:162–169. https://doi.org/10.1016/j.jad.2019.03.075
Chiesa A, Lia L, Alberti S, Lee SJ, Han C, Patkar AA et al (2014) Lack of influence of rs4680 (COMT) and rs6276 (DRD2) on diagnosis and clinical outcomes in patients with major depression. Int J Psychiatry Clin Pract. 18:97–102. https://doi.org/10.3109/13651501.2014.894073
Dandekar MP, Luse D, Hoffmann C, Cotton P, Peery T, Ruiz C et al (2017) Increased dopamine receptor expression and anti-depressant response following deep brain stimulation of the medial forebrain bundle. J Affect Disord. 217:80–88. https://doi.org/10.1016/j.jad.2017.03.074
Demyttenaere K, Van Duppen Z (2019) The impact of (the concept of) treatment-resistant depression: an opinion review. Int J Neuropsychopharmacol 22:85–92. https://doi.org/10.1093/ijnp/pyy052
Fabbri C, Porcelli S, Serretti A (2013) Genetics of Treatment-resistant Depression. Treatment-resistant Depression. Wiley, Oxford, pp 43–90. https://doi.org/10.1093/ijnp/pyy024
Fabbri C, Corponi F, Albani D, Raimondi I, Forloni G, Schruers K et al (2018) Pleiotropic genes in psychiatry: Calcium channels and the stress-related FKBP5 gene in antidepressant resistance. Prog Neuropsychopharmacol Biol Psychiatry 81:203–210. https://doi.org/10.1016/j.pnpbp.2017.10.005
Fabbri C, Kasper S, Kautzky A, Bartova L, Dold M, Zohar J et al (2019) Genome-wide association study of treatment-resistance in depression and meta-analysis of three independent samples. Br J Psychiatry 214:36–41. https://doi.org/10.1192/bjp.2018.256
Fava M (2003) Diagnosis and definition of treatment-resistant depression. Biol Psychiat 53:649–659. https://doi.org/10.1016/S0006-3223(03)00231-2
Gundersen BB, Briand LA, Onksen JL, Lelay J, Kaestner KH, Blendy JA (2013) Increased hippocampal neurogenesis and accelerated response to antidepressants in mice with specific deletion of CREB in the hippocampus: role of cAMP response-element modulator τ. J Neurosci. 33(34):13673–85. https://doi.org/10.1523/JNEUROSCI.1669-13.2013
Halaris A, Sohl E, Whitham EA (2021) Treatment-resistant depression revisited: a glimmer of hope. J Pers Med 11(2):155. https://doi.org/10.3390/jpm11020155
Hamilton M (1960) A RATING SCALE FOR DEPRESSION. J Neurol Neurosurg Psychiatry 23:56–62. https://doi.org/10.1136/jnnp.23.1.56
Hori H, Kunugi H (2013) Dopamine agonist-responsive depression: dopamine and depression. Psychogeriatrics 13:189–195. https://doi.org/10.1111/psyg.12014
Johnston KM, Powell LC, Anderson IM, Szabo S, Cline S (2019) The burden of treatment-resistant depression: a systematic review of the economic and quality of life literature. J Affect Disord 242:195–210. https://doi.org/10.1016/j.jad.2018.06.045
Juhasz G, Dunham JS, McKie S, Thomas E, Downey D, Chase D et al (2011) The CREB1-BDNF-NTRK2 pathway in depression: multiple gene-cognition-environment interactions. Biol Psychiatry 69:762–71. https://doi.org/10.1016/j.biopsych.2010.11.019
Kocabas NA, Faghel C, Barreto M, Kasper S, Linotte S, Mendlewicz J et al (2010) The impact of catechol-O-methyltransferase SNPs and haplotypes on treatment response phenotypes in major depressive disorder: a case–control association study. Int Clin Psychopharmacol 25:218–227. https://doi.org/10.1097/YIC.0b013e328338b884
Kovács KJ (2013) CRH: The link between hormonal-, metabolic- and behavioral responses to stress. J Chem Neuroanat 54:25–33. https://doi.org/10.1016/j.jchemneu.2013.05.003
Lépine JP, Briley M (2011) The increasing burden of depression. Neuropsychiatric Dis Treat 7:3. https://doi.org/10.2147/NDT.S19617
Lex H, Ginsburg Y, Sitzmann AF, Grayhack C, Maixner DF, Mickey BJ (2019) Quality of life across domains among individuals with treatment-resistant depression. J Affect Disord 243:401–407. https://doi.org/10.1016/j.jad.2018.09.062
Li M, Luo X, Rietschel M, Lewis CM, Mattheisen M, Müller-Myhsok B et al (2014) Allelic differences between Europeans and Chinese for CREB1 SNPs and their implications in gene expression regulation, hippocampal structure and function, and bipolar disorder susceptibility. Mol Psychiatry 19:452–461. https://doi.org/10.1038/mp.2013.37
Li Z, Ruan M, Chen J, Fang Y (2021) Major depressive disorder: advances in neuroscience research and translational applications. Neurosci Bull 37:863–880
Lin Z, He H, Zhang C, Wang Z, Jiang M, Li Q et al (2015) Influence of Val108/158Met COMT gene polymorphism on the efficacy of modified electroconvulsive therapy in patients with treatment resistant depression. Cell Biochem Biophys 71:1387–1393. https://doi.org/10.1007/s12013-014-0361-2
Ma J, Wang L, Yang Y, Qiao Z, Fang D, Qiu X et al (2017) GNB3 and CREB1 gene polymorphisms combined with negative life events increase susceptibility to major depression in a Chinese Han population. PLoS One 12:e0170994. https://doi.org/10.1371/journal.pone.0170994
Malhi GS, Mann JJ (2018) Depression. The Lancet. 392:2299–312. https://doi.org/10.1016/S0140-6736(18)31948-2
Marchese E, Di Maria V, Samengo D, Pani G, Michetti F, Geloso MC (2017) Post-natal deletion of neuronal cAMP responsive-element binding (CREB)-1 promotes pro-inflammatory changes in the mouse hippocampus. Neurochem Res 42:2230–2245. https://doi.org/10.1007/s11064-017-2233-9
Murphy GM, Sarginson JE, Ryan HS, O’Hara R, Schatzberg AF, Lazzeroni LC et al (2013) BDNF and CREB1 genetic variants interact to affect antidepressant treatment outcomes in geriatric depression. Pharmacogenet Genom 23:301–313. https://doi.org/10.1097/FPC.0b013e328360b175
O’Dushlaine C, Ripke S, Ruderfer DM, Hamilton SP, Fava M, Iosifescu DV et al (2014) Rare copy number variation in treatment-resistant major depressive disorder. Biol Psychiatry. 76:536–41. https://doi.org/10.1016/j.biopsych.2013.10.028
Perlis RH, Moorjani P, Fagerness J, Purcell S, Trivedi Madhukar H, Fava M et al (2008) Pharmacogenetic analysis of genes implicated in rodent models of antidepressant response: association of TREK1 and treatment resistance in the STAR(*)D study. Neuropsychopharmacology. 33(12):2810–9. https://doi.org/10.1038/npp.2008.6
Rafa-Zabłocka K, Kreiner G, Bagińska M, Nalepa I (2018) Selective depletion of CREB in serotonergic neurons affects the upregulation of brain-derived neurotrophic factor evoked by chronic fluoxetine treatment. Front Neurosci 12:637. https://doi.org/10.3389/fnins.2018.00637
Rosenblat JD, McIntyre RS, Alves GS, Fountoulakis KN, Carvalho AF (2015) Beyond monoamines-novel targets for treatment-resistant depression: a comprehensive review. Curr Neuropharmacol 13(5):636–655. https://doi.org/10.2174/1570159X13666150630175044
Schosser A, Serretti A, Souery D, Mendlewicz J, Zohar J, Montgomery S et al (2012) European group for the study of resistant depression (GSRD)—where have we gone so far: review of clinical and genetic findings. Eur Neuropsychopharmacol 22:453–468. https://doi.org/10.1016/j.euroneuro.2012.02.006
Seib C, Whiteside E, Voisey J, Lee K, Alexander K, Humphreys J et al (2016) Stress, COMT polymorphisms, and depressive symptoms in older Australian women: an exploratory study. Genet Test Mol Biomarkers 20:478–481. https://doi.org/10.1089/gtmb.2015.0028
Serretti A, Chiesa A, Calati R, Massat I, Linotte S, Kasper S et al (2011) A preliminary investigation of the influence of CREB1 gene on treatment resistance in major depression. J Affect Disord 128:56–63. https://doi.org/10.1016/j.jad.2010.06.025
Strawbridge R, Hodsoll J, Powell TR, Hotopf M, Hatch SL, Breen G et al (2019) Inflammatory profiles of severe treatment-resistant depression. J Affect Disord 246:42–51. https://doi.org/10.1016/j.jad.2018.12.037
Tang Z, Zhang S, Guo D, Wang H (2020) Association between COMT gene Val108/158Met and antidepressive treatment response: a meta-analysis. Gene 15(734):144333. https://doi.org/10.1016/j.gene.2020.144333
Taranu A, Asmar KE, Colle R, Ferreri F, Polosan M, David D et al (2017) The Catechol-O-methyltransferase Val(108/158)Met genetic polymorphism cannot be recommended as a biomarker for the prediction of venlafaxine efficacy in patients treated in psychiatric settings. Basic Clin Pharmacol Toxicol 121:435–441. https://doi.org/10.1111/bcpt.12827
Tenhunen J (1996) Characterization of the rat Catechol-O-Methyltransferase gene proximal promoter: identification of a nuclear protein–DNA interaction that contributes to the tissue-specific regulation. DNA Cell Biol 15:461–473. https://doi.org/10.1089/dna.1996.15.461
Tsai SJ, Gau YTA, Hong CJ, Liou YJ, Yu YWY, Chen TJ et al (2009) Sexually dimorphic effect of catechol-O-methyltransferase val158met polymorphism on clinical response to fluoxetine in major depressive patients. J Affect Disord 113:183–187. https://doi.org/10.1016/j.jad.2008.04.017
Utge S, Soronen P, Partonen T, Loukola A, Kronholm E, Pirkola S et al (2010) A population-based association study of candidate genes for depression and sleep disturbance. Am J Med Genet 153B:468–476. https://doi.org/10.1002/ajmg.b.31002
Wang P, Yang Y, Yang X, Qiu X, Qiao Z, Wang L, Ma J (2015) CREB1 gene polymorphisms combined with environmental risk factors increase susceptibility to major depressive disorder (MDD). Int J Clin Exp Pathol 8(1):906
Wolk DM, Clark AE (2018) Matrix-assisted laser desorption time of flight mass spectrometry. Clin Lab Med 38(3):471–486. https://doi.org/10.1016/j.cll.2018.05.008
Xiao X, Zhang C, Grigoroiu-Serbanescu M, Wang L, Li L, Zhou D et al (2018) The cAMP responsive element-binding (CREB)-1 gene increases risk of major psychiatric disorders. Mol Psychiatry 23:1957–1967. https://doi.org/10.1038/mp.2017.243
Xing J, Ginty DD, Greenberg ME (1996) Coupling of the RAS-MAPK pathway to gene activation by RSK2, a growth factor-regulated CREB kinase. Science. 273(5277):959–63. https://doi.org/10.1126/science.273.5277.959
Acknowledgements
We thank all the study participants for their cooperation. These sources had no further role in this study design, data collection, and statistical analysis, drafting of the report, and submitting the paper for publication. Thanks to Jaelin Rippe for contributing to this article.
Author information
Authors and Affiliations
Contributions
Study concept and design: JL, SL. Acquisition of data: SL, LN, YQ, YM, SL, NG, HC. Statistical analysis: YW, SL. Drafting of the manuscript: YW, SL, JL. Study supervision: JL.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wang, Y., Li, S., Niu, L. et al. Polymorphisms of COMT and CREB1 are associated with treatment-resistant depression in a Chinese Han population. J Neural Transm 129, 85–93 (2022). https://doi.org/10.1007/s00702-021-02415-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00702-021-02415-y