Abstract
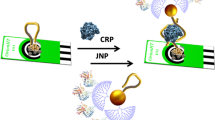
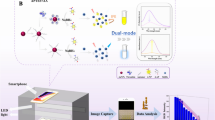
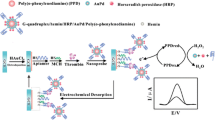
An ultra-efficient biocatalytic peroxidase-like Au-based single-atom nanozyme (Au-SAzymes) has been synthesized from isolated Au atoms on black nitrogen doped carbon (Au–N-C) using a simple complexation-adsorption-pyrolysis method. The atomic structure of AuN4 centers in black carbon was revealed by combined high-resolution transmission electron microscopy/high-angle annular dark-field scanning transmission electron microscopy. The Au-SAzymes showed a remarkable peroxidase activity with 1.7 nM as Michaelis–Menten constant, higher than most previously reported SAzyme activity. Density functional theory and Monte Carlo calculations revealed the adsorption of H2O2 on AuN4 with formation of OH* and O*. Molecular recognition was greatly enhanced via label-free integration of thiol-terminal aptamers on the surface of single Au atoms (Aptamer/Au-SAzyme) to design off–on ultrasensitive aptananozyme-based sensor for detecting thrombin and CRP with 550 pM and 500 pg mL−1 limits of detection, respectively. The Aptamer/Au-SAzyme showed satisfactory accuracy and precision when applied to the serum and plasma of COVID-19 patients. Due to the maximum Au atom utilization, approximately 3636 samples can be run per 1 mg of gold, highlighting the commercialization potential of the developed Aptamer/Au-SAzyme approach.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nanozymes, nanomaterials possessing inherent enzyme-like characteristics, have garnered significant interest in recent years due to their potential to overcome the limitations of natural enzymes, such as low stability, high cost, and challenging storage [1, 2]. This is due to their growing applications in biosensing [3, 4], imaging [5], therapy [6], and environmental remediations [7, 8]. Compared with natural enzymes, enzyme mimics stand out due to their distinct advantages such as great stability, flexibility, easy storage, and easy production [9]. In this area, the primary obstacle is the limited catalytic efficiency of enzyme mimics, especially peroxidase mimics [10, 11].
Nanozymes are nano-sized materials exhibiting catalytic activity mimicking the functions of natural enzymes across various catalytic reactions, such as peroxidase, oxidase, catalase, and superoxide dismutase [2, 12,13,14]. Nanozymes showed higher advantageous over natural enzymes due to shelf-life stability, low-cost production, flexibility in tailoring, and easy functionalization [15]. Metal nanostructures have been used effectively as nanozyme starting from the first observation in Fe3O4 [2, 14,15,16]. Nevertheless, the catalytic efficacy of nanozymes relies on multiple factors, one of them is the size of the particle [13, 16,17,18,19].
Recently, single-atom catalysts (SACs) have come to the forefront as a new field of heterogeneous catalysis in which different catalytic reactions can be carried out via a single atom on a surface [20,21,22]. SACs represent the smallest possible size for metal particles, featuring isolated metal atoms dispersed individually on supports. SACs optimize the utilization efficiency of metal atoms. Additionally, it is essential to have an appropriate substrate to prevent the aggregation of these isolated single atoms. Single atoms possess high surface energy, and maintaining their uniform dispersion is crucial for the catalytic stability of SACs [23]. Due to their distinct geometric structures, uniform active sites, and remarkable activity and selectivity, SACs can amalgamate the benefits inherent in both homogeneous and heterogeneous catalysts. Additionally, they exhibit high efficiency in atom utilization, excellent activity, and remarkable recyclability [24,25,26].
Following the ground-breaking finding of SACs by Zhang et al., [27] numerous reports were published using SACs for various applications, such as hydrogen evolution [28], oxygen reduction [28], and carbon dioxide reduction [29]. SACs with structure M–N-C (refers to metals, Pt, Co, Fe, Mn, etc.) have similar M-Nx sites like natural metalloenzymes. Therefore, it is proposed that M–N-C SACs can behave as bioinspired single-atom enzymes (SAzymes) to mimic the composition and catalytic activity of natural enzymes [30]. However, comparing to the use of SAC in other applications, yet, not much SACs are reported as biocatalyst (or SAzyme). Zhu et al. [31] prepared Fe–N-C SAzymes that showed peroxidase-mimic catalytic activity using FeCl3, glucose, and dicyandiamide as precursors under high temperature calcination. The Fe–N–C SAzymes were used for colorimetric detection of H2O2 in via using chromogenic 3,3′,5,5′-tetramethylbenzidine (TMB) substrate, demonstrating the acceptable specificity and sensitivity. This study provides evidence that the active sites of Fe–N-C SAzymes, which include atomically dispersed FeN4, play a critical role for peroxides-like action. SAzyme have shown high sensitivity due to extraordinary catalytic activity of single atoms, however, for biochemical sensing in real matrices, one should think about improving the selectivity. For this reason, combining SAzyme with aptamer would be an excellent choice to synergize the merits.
The global COVID-19 pandemic, triggered by the highly transmissible SARS-CoV-2 virus, manifests in respiratory symptoms of diverse severity. Initially identified in Wuhan, China, in December 2019, the virus rapidly spread worldwide, leading to a declared global pandemic in 2020 [32,33,34,35]. Numerous biomarkers have been associated with severe cases of COVID-19, suggesting an immunochemical profile consistent, elevation of proinflammatory cytokines, particularly interleukin (IL)-6 and tumor necrosis factor-α (TNF-α) [36,37,38], ferritin [39], and C-reactive protein (CRP) [40, 41]. Those have all been linked to severe COVID-19 and can be used to measure disease severity [42]. Moreover, the (in)-direct effects of the viral illness have been linked to a number of cardiovascular [43, 44], hematologic [45, 46], and thrombotic problems [47].
Thrombin is an important and unique biomolecule with both procoagulant and anticoagulant properties [41, 48,49,50]. It has a multifunctional protein that serves as an inhibitor, activator, and regulator of the cellular coagulation process [51, 52] while CRP is a protein produced by the liver in a response to inflammation [53, 54], and it has a pentameric structure made up of five identical 23-kDa globular subunits [55]. During the acute phase of an inflammatory or viral condition, interleukin-6 (IL-6) primarily induces the production of CRP by influencing the gene responsible for CRP synthesis [56,57,58]. Detection of thrombin and CRP is very important among biomarkers of diagnosis of Covid-19.
There are numerous techniques and methods to determine thrombin and CRP, including electrochemical [59, 60], colorimetric [61, 62], and fluorescence [63,64,65,66,67]. Common and benchmark procedures like radio-assay- and immunoassay-based techniques are also in use [68, 69]. In clinical diagnostics, having a dependable, sensitive, selective, stable, and cost-effective method for detecting thrombin and CRP is crucial, particularly for on-site detection and diagnosis. Enzyme-linked immunosorbent assay (ELISA) has always been the “the best standard way” for quantification of CRP [70,71,72,73]. ELISA can produce false positive results due to cross-reactivity, may not be sensitive or specific enough, is time-consuming and expensive, and has a limited dynamic range. It is important to be aware of these limitations when using ELISA for diagnostic or research purposes. Thus, one should think of simple, low-cost, highly selective, sensitive, robust, and fast-response protocol for detection of CRP and thrombin.
Aptamers are single-stranded oligonucleotides (DNA or RNA), with a short length that attach to certain molecules. Due to the shape-forming ability of single-stranded oligonucleotides, they can bind to their targets with tremendous affinity and selectivity [74,75,76,77,78] the attractions of aptamer have included a broad range of target molecules, robust and strong binding capacity, high stability, long shelf-life, simplicity in synthesis, and easy modification [79,80,81].
In the present work, we use a complexation-adsorption-pyrolysis strategy to synthesize isolated Au-SAzyme on N-doped carbon. The resulting Au-SAzyme exhibited remarkably efficient peroxidase activity. To gain deeper insights, we investigated the peroxidase mechanism through theoretical calculations. Subsequently, thiol-terminal aptamers (ssDNA or RNA) were immobilized on the dispersed Au-SAzymes to inhibit the peroxidase enzymatic activity, resulting in the creation of Aptamer/Au-SAzyme hybrids. A biochemical sensor based on Aptamer/Au-SAzyme was utilized for detection of thrombo-inflammatory COVID-19 biomarkers, thrombin, and CRP. To the best of our knowledge, this is the first report on mimetic enzyme activity of isolated Au atoms combined with ssDNA/RNA. Scheme 1 shows the preparation roadmap of Au-SAzyme and Aptamer/Au-SAzyme.
Experimental section
Chemicals
Chloroauric acid trihydrate (HAuCl4·3H2O) was purchased from Sigma Aldrich Co. (Hong Kong, China). Thiol-modified aptamer for thrombin was synthesized and purified by Generay biotech Co. Ltd. (Shanghai, China) with sequences for ssDNA (5-(SH)-(CH2)6-CCA TCT CCA CTT GGT TGG TGT GGT TGG-3) specific for thrombin. The sequence RNA strand is (5'-SH-(CH2)6-GCC UGU AAG GUG GUC GGU GUG GCG AGU GUG UUA GGA GAG AUU GC-3) specific for CRP. Thrombin protein extracted from human plasma, C-reactive protein (CRP), Dimethyl sulfoxide (DMSO), Dimethylformamide (DMF), 3, 3′ 5, 5’-Tetramethylbenzidine (TMB) were all purchased from Merck-Sigma-Aldrich (Baden-Württemberg, Germany) and used as received. 1, 10 phenanthroline monohydrate and hydrogen peroxide (H2O2, 30%) were bought from Biochem chemopharma Co.Ltd. (Biochem, ZA Cosne sur Loire, France). TE buffer (Tris–EDTA, pH 8.0) was bought from EMB Corporation (EMB Co., USA). PBS (phosphate buffer saline, pH 7) was ordered from (CDH Co. Ltd. Delhi, India).
Instruments
All UV–Vis absorption spectra and enzyme kinetics data were performed by a PG spectrophotometer (PG, UK). X-ray photoelectron spectroscopy (XPS) spectra were recorded using Thermo ESCALAB 250XI (Thermo Fisher, USA). X-ray diffraction (XRD) spectra were acquired by a D8 ADVANCE (Bruker, Germany). Raman spectra were recorded using a Renishaw 1000 micro-Raman system (Renishaw, UK). The aberration‐corrected high‐angle annular dark‐field scanning transmission electron microscopy (HAADF‐STEM) was performed by a Titan G2-600 (FEI, USA).
Preparation of Au-SAzyme
The Au-SAzyme biocatalyst was prepared according to literature with some modifications [82]. In brief, a mixture of 0.076 g of HAuCl4·3H2O and 0.15 g of 1,10-phenanthroline monohydrate was combined and dissolved in 2.0 mL of dimethyl sulfoxide (DMSO), with stirring for about 20 min at room temperature. Following this, carbon black (69.6 mg) was introduced into the solution, and the resultant mixture underwent heating in water bath at 60 °C for 5 h with continuous stirring. The resulting dispersion was later subjected to heating at 80 °C in air for 12 h to remove the DMSO, resulting in the formation of a black solid product. The produced black solid was gently grounded using a mortar and pestle, then moved into a ceramic crucible and positioned inside a tube furnace for pyrolysis. The black product underwent pyrolysis under nitrogen gas at 600 °C for a duration of 5 h.
Integration of aptamer on Au-SAzyme surface
Au-SAzyme- thrombin binding ssDNA: To prepare a 100 µg/mL solution of dispersed Au-SAzyme, 0.01 g of Au-SAzyme powder was dispersed in 100 mL of deionized water. Subsequently, 1.0 mL of the 100 µg/mL Au-SAzyme solution was combined with 0.5 mL of 1.0 µM thiolate terminal single-stranded DNA, stirred for 15 min, and then incubated for 1 h at room temperature.
Au-SAzyme- CRP binding RNA: 1.0 mL of 100 µg.mL−1 of Au-SAzyme was mixed with 1.0 mL of 0.46 µM thiolate terminal RNA (same as ssDNA) and stirred for 15 min, followed by incubation for 1 h at room temperature.
Colorimetric analysis
For the thrombin colorimetric analysis: 100 µL of the mixture of Au-SAzyme/ssDNA probe was taken and mixed with certain amount of the thrombin and 100 µL of 5 mM of TMB, 100 µL of 30 mM H2O2 added the volume was completed to 1 mL by buffer pH 4.
For the CRP colorimetric analysis: same as for thrombin analysis, 100 µL of the mixture of Au-SAzyme/RNA probe was taken and mixed with certain amount of CRP and 100 µL of 5 mM of TMB, 100 µL of 30 mM H2O2 added the volume was completed to 1 mL by buffer pH 4.
DFT and Monte Carlo simulations
Ab-initio calculations were conducted utilizing plane-wave basis and pseudopotential methods within the framework of Density Functional Theory (DFT). All computations were performed using the Dmol3 code and adsorption locator in material studio provided by BIOVIA. Nonlocal-functional generalized gradient approximations (GGA) have been applied. A 4 × 4 × 2 k-point Monkhorst–Pack grid was used to optimize the structure, as well as the SCF tolerance was 1*10–6 eV/atom. For thermodynamic properties calculation 3 × 3x1 k-point Monkhorst–Pack grid was applied, double numerical plus polarization and double numerical plus d-functions basis sets were used respectively. Monte Carlo (MC) simulations were performed to acquire more information about the interaction between the AuN4@C surface with hydrogen peroxide and reactive oxygen molecules. The absorption sites and energy (Eads) were calculated using adsorption locator code and universal force field integrated which carried out with Monte Carlo method. We have established that the entire system was in equilibrium until temperature and energy were equal. For this reason, the full simulation was carried out with AuN4@C (001) slab model with a lattice parameter of a = b = 29Ǻ and c = 20Ǻ in order to avoid the effect of interaction between layers.
Results and discussion
Characterizations
Figure 1A shows Raman spectrum of the Au-SAzyme biocatalyst. Two well-resolved peaks are shown in the spectrum locating at 1350 and 1500 cm−1, which are normally assigned to D and G bands. The G-band (1597 cm−1) is related to stretching Eigen mode of the C–C bond in graphitic structure, and indicative of a typical sp2-hybridized carbon network. The D band at 1350 cm−1 arises from a hybridized vibrational mode linked to graphene edges, signifying the existence of structural disorder in the graphene [83, 84].
The X-ray powder diffractometer (XRD) analysis of the as-synthesized materials confirmed that the sample synthesized with Au-SAzyme exhibits a graphitic packing structure. (Fig. 1B) [82, 85] The determination of the composition and electronic state of surface elements in the Au-SAzyme catalyst was initially conducted through X-ray photoelectron spectroscopy (XPS). The survey spectrum (depicted in Fig. 1C) exhibits distinct peaks corresponding to C, N, and Au elements. In the Au-SAzyme, carbon atoms are predominantly found in sp2-hybridized structure. (Fig. 1F). The Au XPS spectra were recorded under these optimized conditions (Fig. 1D). The high-resolution Au-4f XPS. A thorough analysis of the XPS spectra unveiled a notable presence at a binding energy (BE) of 83.38 eV, a value typically associated with Au0.On the contrary, also the Au3+ species, located at 86.90, eV. The other two peaks in 83.13 and 86.76 are related to Au* [86]. These peaks confirm presence of are a mixture of Au3+, Au0* and Au0.
The high-resolution N 1 s spectrum (Fig. 1E) exhibits three peaks at 399.23, and 397.86, 397.02 eV, related to C = N, C-N–C, and C-NH2, respectively [87, 88]. Carbon atoms in Au-SAzyme are mainly present in sp2 -hybridized structure (Fig. 1F). According to the convoluted spectrum of C, two small peaks at 285.67 and 283.93 eV can be assigned to the C 1 s orbital of C = C and C-N respectively [89, 90].
Figure 2 illustrates the verification of the unique presence of individual gold atoms on the nitrogen-doped carbon platform using HAADF-STEM and the corresponding EDX mapping.
A–E Energy dispersive X-ray spectroscopy elemental mapping results of Au-SAzyme, indicating the Au (yellow), C (red), N (blue) elements homogeneously distributed in pyrolyzed sample. F, G HAADF-STEM image of Au-SAzyme, suggesting Au single atoms was atomically dispersed on the substrate at 5 nm and 2 nm respectively
To determine the elemental composition, EDX analysis was carried out on the Au-SAzyme, as shown in Fig. 2A-E, as well as area was 10 nm of mapping on Au- SAzyme. In the EDX analysis, it is shown that the co-existence of Au, C, and N corroborates the composite of gold single atoms hosted on N-doped carbon, derived by the controlled pyrolysis of Au-complex with 1,10-phenanthroline adsorbed on charcoal, under N2 gas at 600 °C formation. HAADF-STEM was performed to observe dispersed isolated atoms (Fig. 2F and 2G), and is obvious that Au atoms are well-dispersed and isolated as indicated by are indicated by red circles, and they size are around 0.2 nm which is exactly close to atomic size of gold [91, 92]. Thus, HAADF-STEM images combined with EDX mapping confirmed the formation of isolated Au atoms.
Enzymatic activity of Au-SAzyme
Following formation of isolated Au atoms on the N-doped carbon platform, enzymatic activity was evaluated. Figure 3A (blue line) shows the enzymatic activity of Au-SAzyme (10 µg/mL) using TMB as a chromogenic color in the presence of H2O2. Au-SAzyme showed remarkable excellent peroxidase-like activities. The best concentrations of TMB (substrate) were studied and it was between 0.16 and 1.6 mM for 10 µg mL−1 of Au-SAzyme as shown in Figure S1 (supporting information). Furthermore, kinetic parameters of peroxidase property of Au-SAzyme were investigated. For an enzyme activity, the Michaelis–Menten saturation curve illustrates the link between substrate concentration and reaction rate. The kinetic parameters explain this relationship [93, 94]. The SAzyme’s activities with different substrate concentrations were plotted, and the Michaelis–Menten’s constants (Km) and the maximal reaction velocity (Vmax) were determined using the equation 1/ν = Km/Vmax · [S] + 1/Vmax, where ν denotes the initial velocity, [S] represents the substrate concentration, and Vmax is the maximum reaction velocity. As it shows in Fig. 3B, the value of Vmax was 5.5 nM min−1, and Km was 1.7 nM. The Km value obtained is exceptionally minute, and to the best of our knowledge, it represents the lowest reported in the literature (Table 1).
A The enzymatic activity of the Au-SAzyme and the carbon@nitrogen as a blank in presence of TMB and H2O2. B The kinetic study of Au-SAzyme which include the Michaelis–Menten and Lineweaver–Burk plot for Au-SAzyme. C, D Schematic presentation of peroxidase on AuN4. E Energy distribution diagram from DFT calculation. F Energy level representation from Monte Carlo simulation
Proposed mechanism of the efficient peroxidase
Highly catalytic peroxidase was investigated based on the DFT and Monte’ Carlo calculation as performed in the literature [101,102,103,104]. As shown in Fig. 3, the first step is adsorption of H2O2 on the coordinately unsaturated AuN4 site. At first, as shown in Fig. 3A to C to F, H2O2 molecule was absorbed on Au–N4 active sites (i). The energy of adsorption was 2.266 eV based on DFT (Fig. 3E). The activated H2O2 molecules easily dissociated by homolytic path, resulting in the generation of 2OH* (ii). Step ii with energy of 2.024 eV for cleavage of H2O2 to 2OH*. Here the OH* can desorb and oxidize TMB to oxTMB. Another probable step (step iii) is formation of H2O and singlet oxygen (O*) with desorption of O* to oxidize TMN to oxTMB (iv) as shown in Fig. 3D-E.
Applications
The catalytic activity of peroxidase mimic was found by oxidizing colorless chromogenic TMB to blue ox-TMB in presence of H2O2. When the S- terminal ssDNA was added to the Au-SAzyme, due to the strong gold thiol bonds (Au–S), the gold single atoms were covered and lost their peroxidase catalytic activity. Later, when thrombin was added, and due to the high affinity of the ssDNA to the thrombin, thrombin-ssDNA was formed and the Au–S bond was weakened, then the blue color was restored within 15–20 min the (Fig. 4A). For the quantitative determination of the thrombin, different concentrations of standard thrombin were added (0.7, 1.5, 2, 5, 10, 20, 50, 100 nM). The increase of blue color intensity was noted with an increase in thrombin concentration (see Fig. 4B). A linear correlation was established between absorbance and thrombin concentration, revealing a remarkably low limit of detection (LOD) at 550 pM and a limit of quantification (LOQ) at 1.8 nM, as illustrated in Fig. 4C.
Au-SAzyme responses: A and D blue lines: Au-SAzyme signal 0.5 mM TMB, 0.3 mM H2O2 and 10 µg/mL of Au-SAzyme; black line: integration with 0.5 mL of 1.0 µM ssDNA and 1.0 mL of 0.46 µM RNA; red line: signal recovery by thrombin and CRP respectively. B, E UV–Vis absorption spectra of oxidation of TMB in the presence of Au-SAzyme/ssDNA verse thrombin at various concentrations, Au-SAzyme /RNA verse different concentrations of CRP respectively. C, F A graph plotting the absorbance at 652 nm against the concentrations of thrombin and CRP, respectively, with error bars depicting the standard deviation from three independent measurements
For the CRP analysis, like thrombin analysis mentioned above, the blue colored ox-TMB via Au-SAzyme was converted to colorless with adding thiol terminal RNA to the solution but a rapid color recovery has been detected with in adding CRP to the Au-SAzyme/RNA probe with 6–10 min (Fig. 4D). Quantification analysis was conducted with various standard CRP concentrations of CRP from 0.1 to 50.0 ng mL−1. The blue color was directly increases with increase of CRP concentrations. The recovery will be close to the Au-SAzyme signal (Fig. 4E). A linear relationship was obtained between the absorbance and CRP concentrations with very small LOD as 500 pg mL−1, and LOQ was 165,000 pg mL−1 with S/N ratio = 3. The LOD and LOQ were established using the criteria of Signal-to-Noise (S/N) ratios of 3 and 10, respectively. The signal (S) was determined relative to the standard deviation (SD) from ten measurements, taken from the current intensity of the lowest concentrations in the calibration, which were 0.7 nM for thrombin and 0.1 ng mL−1 for CRP, respectively (Fig. 4F).
Selectivity study
As the human sera from COVID-19 patients serve as the matrix for the proposed probe, the typical ions, biomolecules, and proteins present in the serum matrix are considered. The possible interferents’ concentration were 100 times more than that of the target species, thrombin, and CRP should be examined (Fig. 5). In this study, the thrombin concentration was 10 nM, other interferences were 1 µM. CRP was 10 ng mL−1, other interferents were 1.0 µg mL−1. All selectivity experiments were performed in the presence of 0.5 mM TMB, 0.3 mM H2O2. The absorbance intensity of thrombin and CRP was notably high compared to that of other proteins and metals.
The selectivity tests have shown that the aptanoanzyme-based ssDNA is a highly selective probe to thrombin (Fig. 5A). Similarly, the aptanoanzyme-based RNA probe is highly selective for the CRP (Fig. 5B). These findings show that the Au-Sazyme/ssDNA and Au-SAzyme/RNA sensor has a high thrombin and CRP selectivity respectively, making it a suitable probe for a human serum fluid sample.
Thrombin and CRP detection
The developed Aptamer/Au-SAzyme biosensor was utilized for individuals diagnosed with COVID-19. Serum and plasma samples from COVID-19 patients were gathered from the emergency and intensive care units at Hospital of Shahid-Hemn in the Slemani City, Kurdistan Region, Iraq. The collection of serum samples adhered to the ethical standards and regulations set by the hospital. The hospital administration and the patients themselves were duly informed. Standard concentrations of thrombin or CRP were introduced into the plasma and serum samples from the patients. The plasma samples underwent a 1000-fold dilution for thrombin determination, while the serum samples were diluted by a factor of 10, all serum samples were diluted using PBS reagent and underwent measurements in triplicate (n = 3).
Table 2 shows spiking recoveries for both COVID-19 biomarkers (thrombin and CRP) ranged for colorimetric assay from 90.5 to 98.9% for thrombin, and for CRP the spiking ranges was 89.6–103.2%, with RSD 0.45–0.81%, and 0.70–2.38% for thrombin and CRP respectively. The intra-assay precision achieves an RSD of 2% (n = 3) within the same day, and the inter-assay precision (reproducibility) is 2% (n = 3) across different days.
Commercialization potential of the Au-SAzyme-based aptasensor
Due to maximum atom utilization, SAzymes are expected to have a great potential for commercialization. At first, 33 mg of Au was taken (from HAuCl4. 4H2O), and 0.6 g of Au-SAzyme was obtained after pyrolysis. Then, 0.01 g of the Au-SAzyme was then dispersed in 100 mL of water to prepared stock Au-SAzyme solution. 1.0 mL of Au-SAzyme stock solution was mixed with 1 mL of aptamer to make the probe. Form the probe, 100 µL was used for running the tests analysis. Scheme 2 shows the consumption rout of gold from starting precursor to the running test.
Based on this calculation, using 33 mg of gold, one can test 120,000 sample of a biomarker. This means for 1 g of gold approximately 3,636,000 samples can be analyzed. This assigns for highly valuable and economic of our SAzyme-based aptasensor for real applications. This ascribed to the maximum atom utilization of the catalyst and high peroxidase activity. Thus, we can confidently state that our probe is suitable commercialization purpose.
Conclusions
Highly efficient peroxidase mimic from Au dispersed single atoms on black carbon was synthesized using very cost-effective strategy. Highly selective and sensitive biocatalyst of well-dispersed Au single atoms were prepared and used as mimicking enzymes (SAzymes). Simple and high-yield synthetic method based on complexation-adsorption-pyrolysis was used successfully. HAAD-STEM showed the Au atoms are well-dispersed and isolated on the carbon platform. The Au-SAzyme showed remarkable low Michaelis–Menten constant (KM) of 1.7 nM. Thiolate aptamers for thrombin and CRP combined with SAzyme enhanced the molecular recognition selectivity and showed remarkable results thrombin and CRP detection in human serum with COVID-19 patients. LOD as small as 550 pM and as 500 pg.mL−1 were calculated for thrombin and CRP, respectively. Interestingly, our Aptamer/Au-SAzyme probe is very economic and one can commercialize it as approximately 360,000 tests can be done with 1.0 g of gold.
References
Chang B, Zhang L, Wu S, Sun Z, Cheng Z (2022) Engineering single-atom catalysts toward biomedical applications. Chem Soc Rev 51:3688
Jiao L et al (2020) When nanozymes meet single-atom catalysis. Angew Chem – Int Ed 59:2565–2576
Jiao L et al (2021) Single-atom catalysts boost signal amplification for biosensing. Chem Soc Rev 50:750–765
Wu Y et al (2020) Cascade reaction system integrating single-atom nanozymes with abundant Cu sites for enhanced biosensing. Anal Chem 92:3373–3379
Zhang T, Walsh AG, Yu J, Zhang P (2021) Single-atom alloy catalysts: structural analysis, electronic properties and catalytic activities. Chem Soc Rev 50:569–588
Lu X, Gao S, Lin H, Shi J (2021) Single-atom catalysts for nanocatalytic tumor therapy. Small 17:2004467
Xiang H, Feng W, Chen Y (2020) Single-atom catalysts in catalytic biomedicine. Adv Mater 32:1905994
Su R, Zhang H, Chen F, Wang Z, Huang L (2022) Applications of single atom catalysts for environmental management. Int J Environ Res Public Health 19:11155
Wang H, Wan K, Shi X (2019) Recent advances in nanozyme research. Adv Mater 31:1–10
Li Z et al (2018) Peroxidase-mimicking nanozyme with enhanced activity and high stability based on metal–support interactions. Chem–A Eur J 24:409–415
Thangudu S, Su C-H (2021) Peroxidase mimetic nanozymes in cancer phototherapy: Progress and perspectives. Biomolecules 11:1015
Yan X, Gao L (2020) Nanozymology: An Overview. In: Yan X (eds) Nanozymology. Nanostructure Science and Technology. Springer, Singapore. https://doi.org/10.1007/978-981-15-1490-6_1
Pei J et al (2020) Single-atom nanozymes for biological applications. Biomater Sci 8:6428–6441
Unnikrishnan B, Lien C-W, Chu H-W, Huang C-C (2021) A review on metal nanozyme-based sensing of heavy metal ions: Challenges and future perspectives. J Hazard Mater 401:123397
Huang L, Chen J, Gan L, Wang J, Dong S (2019) Single-atom nanozymes. Sci Adv 5:eaav5490
Jiao L et al (2019) Fe-N-C Single-atom nanozymes for the intracellular hydrogen peroxide detection. Anal Chem 91:11994–11999
Ding H, Hu B, Zhang B et al (2021) Carbon-based nanozymes for biomedical applications. Nano Res 14:570–583. https://doi.org/10.1007/s12274-020-3053-9
Zhang R, Yan X, Fan K (2021) Nanozymes inspired by natural enzymes. Acc Mater Res 2:534–547
Yang X-F et al (2013) Single-atom catalysts: a new frontier in heterogeneous catalysis. Acc Chem Res 46:1740–1748
Samantaray MK et al (2019) The comparison between single atom catalysis and surface organometallic catalysis. Chem Rev 120:734–813
Zhang H, Liu G, Shi L, Ye J (2018) Single-atom catalysts: emerging multifunctional materials in heterogeneous catalysis. Adv Energy Mater 8:1701343
Xu H, Zhao Y, Wang Q, He G, Chen H (2022) Supports promote single-atom catalysts toward advanced electrocatalysis. Coord Chem Rev 451:214261
Kaiser SK, Chen Z, Faust Akl D, Mitchell S, Pérez-Ramírez J (2020) Single-atom catalysts across the periodic table. Chem Rev 120:11703–11809
Zhang J et al (2018) Single-atom Au/NiFe layered double hydroxide electrocatalyst: probing the origin of activity for oxygen evolution reaction. J Am Chem Soc 140:3876–3879
Li J et al (2018) Atomically dispersed manganese catalysts for oxygen reduction in proton-exchange membrane fuel cells. Nat Catal 1:935–945
Wu W, Huang L, Wang E, Dong S (2020) Atomic engineering of single-atom nanozymes for enzyme-like catalysis. Chem Sci 11:9741–9756
Qiao B et al (2011) Single-atom catalysis of CO oxidation using Pt1/FeOx. Nat Chem 3:634–641
Chen W et al (2017) Rational design of single molybdenum atoms anchored on N-doped carbon for effective hydrogen evolution reaction. Angew Chem Int Ed 56:16086–16090
Zhao C et al (2017) Ionic exchange of metal–organic frameworks to access single nickel sites for efficient electroreduction of CO2. J Am Chem Soc 139:8078–8081
Martinelli LM, Carucci A, Payano VJH et al (2023) Translational comparison of the human and mouse yolk sac development and function. Reprod Sci 30:41–53. https://doi.org/10.1007/s43032-022-00872-8
Jiao L et al (2019) Fe–N–C single-atom nanozymes for the intracellular hydrogen peroxide detection. Anal Chem 91:11994–11999
Pokhrel S, Chhetri R (2021) A literature review on impact of COVID-19 pandemic on teaching and learning. Higher Education Future 8:133–141
Castagnoli R et al (2020) Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection in children and adolescents: a systematic review. JAMA Pediatr 174:882–889
Tan HW, Xu Y, Lau ATY (2020) Angiotensin-converting enzyme 2: the old door for new severe acute respiratory syndrome coronavirus 2 infection. Rev Med Virol 30:e2122
Ganatra S et al (2020) Management of cardiovascular disease during coronavirus disease (COVID-19) pandemic. Trends Cardiovasc Med 30(6):315–325. https://doi.org/10.1016/j.tcm.2020.05.004
Que Y et al (2022) Cytokine release syndrome in COVID-19: a major mechanism of morbidity and mortality. Int Rev Immunol 41:217–230
Hu B, Huang S, Yin L (2021) The cytokine storm and COVID-19. J Med Virol 93:250–256
Patel S, Saxena B, Mehta P (2021) Recent updates in the clinical trials of therapeutic monoclonal antibodies targeting cytokine storm for the management of COVID-19. Heliyon 7:e06158
Kaushal K et al (2022) Serum ferritin as a predictive biomarker in COVID-19. A systematic review, meta-analysis and meta-regression analysis. J Crit Care 67:172–181
Mosquera-Sulbaran JA, Pedreañez A, Carrero Y, Callejas D (2021) C-reactive protein as an effector molecule in Covid-19 pathogenesis. Rev Med Virol 31:e2221
Chaudhary R et al (2021) Thromboinflammatory biomarkers in COVID-19: systematic review and meta-analysis of 17,052 patients. Mayo Clin Proc: Innov, Qual Outcomes 5:388–402
Iwamura APD et al (2021) Immunity and inflammatory biomarkers in COVID-19: a systematic review. Rev Med Virol 31:e2199
Chung MK et al (2021) COVID-19 and cardiovascular disease: from bench to bedside. Circ Res 128:1214–1236
Talasaz AH, Kakavand H, Van Tassell B et al (2021) Cardiovascular Complications of COVID-19: Pharmacotherapy Perspective. Cardiovasc Drugs Ther 35:249–259. https://doi.org/10.1007/s10557-020-07037-2
Korompoki E et al (2022) Late-onset hematological complications post COVID-19: an emerging medical problem for the hematologist. Am J Hematol 97:119–128
Papakonstantinou E et al (2021) Haematological malignancies implications during the times of the COVID-19 pandemic. Oncol Lett 22:1–7
Mungmunpuntipantip R, Wiwanitkit V (2022) COVID-19, neurovascular thrombotic problem and short summary on blood coagulation disorder: a brief review. Egypt J Neurol, Psychiatr Neurosurg 58:1–5
Aliter KF, Al-Horani RA (2021) Thrombin inhibition by argatroban: potential therapeutic benefits in COVID-19. Cardiovasc Drugs Ther 35:195–203
Sriram K, Insel PA (2021) Inflammation and thrombosis in COVID-19 pathophysiology: proteinase-activated and purinergic receptors as drivers and candidate therapeutic targets. Physiol Rev 101:545–567
Ali GK, Omer KM (2022) Nanozyme and stimulated fluorescent Cu-based metal–organic frameworks (Cu-MOFs) functionalized with engineered aptamers as a molecular recognition element for thrombin detection in the plasma of COVID-19 patients. ACS Omega 7:36804–36810
Di Cera E (2008) Thrombin. Mol Aspects Med 29:203–254
Mann KG (2003) Thrombin formation. Chest 124:4S-10S
Hurlimann J, Thorbecke GJ, Hochwald GM (1966) The liver as the site of C-reactive protein formation. J Exp Med 123:365–378
Eklund CM (2009) Proinflammatory cytokines in CRP baseline regulation. Adv Clin Chem 48:111–136
Meurman J, Celec P, Gutierrez AM, Sjöwall C (2022) Pronounced diurnal pattern of salivary C-reactive protein (CRP) with modest associations. Diagn Ther Appl Pentraxin Pentraxin-Assoc Proteins
Eisenhardt SU, Thiele JR, Bannasch H, Stark GB, Peter K (2009) C-reactive protein: how conformational changes influence inflammatory properties. Cell Cycle 8:3885–3892
Agrawal A (2005) CRP after 2004. Mol Immunol 42:927–930
Ali GK, Omer KM (2022) Ultrasensitive aptamer-functionalized Cu-MOF fluorescent nanozyme as an optical biosensor for detection of C-reactive protein. Anal Biochem 114928. https://doi.org/10.1016/j.ab.2022.114928
Kotru S et al (2021) Electrochemical sensing: A prognostic tool in the fight against COVID-19. TrAC, Trends Anal Chem 136:116198
Thuerlemann C, Haeberli A, Alberio L (2009) Monitoring thrombin generation by electrochemistry: development of an amperometric biosensor screening test for plasma and whole blood. Clin Chem 55:505–512
Mahmoud M, Ruppert C, Rentschler S, Laufer S, Deigner H-P (2021) Combining aptamers and antibodies: lateral flow quantification for thrombin and interleukin-6 with smartphone readout. Sens Actuators, B Chem 333:129246
Nagy-Simon T, Hada A-M, Suarasan S, Potara M (2021) Recent advances on the development of plasmon-assisted biosensors for detection of C-reactive protein. J Mol Struct 1246:131178
Mallakpour S, Azadi E, Hussain CM (2021) Fight against COVID-19 pandemic with the help of carbon-based nanomaterials. New J Chem 45:8832–8846
Maddali H, Miles CE, Kohn J, O’Carroll DM (2021) Optical biosensors for virus detection: prospects for SARS-CoV-2/COVID-19. ChemBioChem 22:1176–1189
Sh. Mohammed Ameen S, Sher Mohammed NM, Omer KM (2022) Visual monitoring of silver ions and cysteine using bi-ligand Eu-based metal organic framework as a reference signal: Color tonality. Microchem J 181:107721
Mohammed Ameen SS, Sher Mohammed NM, Omer KM (2023) Ultra-small highly fluorescent zinc-based metal organic framework nanodots for ratiometric visual sensing of tetracycline based on aggregation induced emission. Talanta 254:124178
Mohammed Ameen SS, Qasim FO, Alhasan HS, Hama Aziz KH, Omer KM (2023) Intrinsic dual-state emission zinc-based MOF rodlike nanostructures with applications in smartphone readout visual-based detection for tetracycline: MOF-Based Color Tonality. ACS Appl Mater Interfaces 15:46098–46107
Wang C, Sun Y, Zhao Q (2020) A sensitive thrombin-linked sandwich immunoassay for protein targets using high affinity phosphorodithioate modified aptamer for thrombin labeling. Talanta 207:120280
Kurantsin-Mills J, Ofosu FA, Safa TK, Siegel RS, Lessin LS (1992) Plasma factor VII and thrombin–antithrombin III levels indicate increased tissue factor activity in sickle cell patients. Br J Haematol 81:539–544
Kushner I, Somerville JA (1970) Estimation of the molecular size of C-reactive protein and Cx-reactive protein in serum. Biochim Biophys Acta (BBA)-Protein Struct 207:105–114
Algarra M, Gomes D, da Silva JCGE (2013) Current analytical strategies for C-reactive protein quantification in blood. Clin Chim Acta 415:1–9
White D et al (2021) Evaluation of COVID-19 coagulopathy; laboratory characterization using thrombin generation and nonconventional haemostasis assays. Int J Lab Hematol 43:123–130
Ong DSY, de Man SJ, Lindeboom FA, Koeleman JGM (2020) Comparison of diagnostic accuracies of rapid serological tests and ELISA to molecular diagnostics in patients with suspected coronavirus disease 2019 presenting to the hospital. Clin Microbiol Infect 26:1094-e7
Adachi T, Nakamura Y (2019) Aptamers: A review of their chemical properties and modifications for therapeutic application. Molecules 24:4229
Mascini M (2009) Aptamers in bioanalysis. Wiley
Kong HY, Byun J (2013) Nucleic acid aptamers: new methods for selection, stabilization, and application in biomedical science. Biomol Ther 21:423
Ospina-Villa JD et al (2018) Advances on aptamers against protozoan parasites. Genes 9:584
Cho EJ, Lee J-W, Ellington AD (2009) Applications of aptamers as sensors. Annu Rev Anal Chem 2:241–264
Zhang Y, Lai BS, Juhas M (2019) Recent advances in aptamer discovery and applications. Molecules 24:941
Ali GK, Omer KM (2022) Molecular imprinted polymer combined with aptamer (MIP-aptamer) as a hybrid dual recognition element for bio(chemical) sensing applications. Rev Talanta 236:122878
Hong P, Li W, Li J (2012) Applications of aptasensors in clinical diagnostics. Sensors 12:1181–1193
Yang H et al (2019) A universal ligand mediated method for large scale synthesis of transition metal single atom catalysts. Nat Commun 10:1–9
Lee AY et al (2021) Raman study of D* band in graphene oxide and its correlation with reduction. Appl Surf Sci 536:147990
Brown SDM, Jorio A, Dresselhaus MS, Dresselhaus G (2001) Observations of the D-band feature in the Raman spectra of carbon nanotubes. Phys Rev B 64:73403
Liu J et al (2017) Palladium–gold single atom alloy catalysts for liquid phase selective hydrogenation of 1-hexyne. Catal Sci Technol 7:4276–4284
Carter JH et al (2019) Enhanced activity and stability of gold/ceria-titania for the low-temperature water-gas shift reaction. Front Chem 7:443
Hellgren N, Haasch RT, Schmidt S, Hultman L, Petrov I (2016) Interpretation of X-ray photoelectron spectra of carbon-nitride thin films: new insights from in situ XPS. Carbon 108:242–252
Vilian ATE et al (2020) Improved conductivity of flower-like MnWO4 on defect engineered graphitic carbon nitride as an efficient electrocatalyst for ultrasensitive sensing of chloramphenicol. J Hazard Mater 399:122868
Luo Z et al (2011) Pyridinic N doped graphene: synthesis, electronic structure, and electrocatalytic property. J Mater Chem 21:8038–8044
Wang Y et al (2021) Tautomeric molecule acts as a “sunscreen” for metal halide perovskite solar cells. Angew Chem 133:8755–8759
Viljas JK, Cuevas JC (2007) Role of electronic structure in photoassisted transport through atomic-sized contacts. Phys Rev B 75:75406
Biggs S, Mulvaney P (1994) Measurement of the forces between gold surfaces in water by atomic force microscopy. J Chem Phys 100:8501–8505
Schnell S (2014) Validity of the Michaelis-Menten equation - steady-state or reactant stationary assumption: That is the question. FEBS J 281:464–472
Rodriguez J-MG, Hux NP, Philips SJ, Towns MH (2019) Michaelis-Menten graphs, lineweaver–Burk plots, and reaction schemes: investigating introductory biochemistry students’ conceptions of representations in enzyme kinetics. J Chem Educ 96:1833–1845
Ji S et al (2021) Matching the kinetics of natural enzymes with a single-atom iron nanozyme. Nat Catal 4:407–417
Gao L et al (2007) Intrinsic peroxidase-like activity of ferromagnetic nanoparticles. Nat Nanotechnol 2:577–583
Jiao L et al (2020) Densely isolated FeN4 sites for peroxidase mimicking. ACS Catal 10:6422–6429
Shao B et al (2022) Mn-doped single atom nanozyme composited Au for enhancing enzymatic and photothermal therapy. J Colloid Interface Sci 628:419–434
Lin Z et al (2020) A facile route for constructing Cu–N–C peroxidase mimics. J Mater Chem B 8:8599–8606
Chen Y et al (2021) Thermal atomization of platinum nanoparticles into single atoms: an effective strategy for engineering high-performance nanozymes. J Am Chem Soc 143:18643–18651
Chen Y et al (2018) Enhanced oxygen reduction with single-atomic-site iron catalysts for a zinc-air battery and hydrogen-air fuel cell. Nat Commun 9:5422
Deng D et al (2015) A single iron site confined in a graphene matrix for the catalytic oxidation of benzene at room temperature. Sci Adv 1:e1500462
Xu B et al (2022) A bioinspired five-coordinated single-atom iron nanozyme for tumor catalytic therapy. Adv Mater 34:2107088
Xu B et al (2019) A single-atom nanozyme for wound disinfection applications. Angew Chem 131:4965–4970
Acknowledgements
This work was supported and funded by the Deanship of Scientific Research at Imam Mohammad Ibn Saud Islamic University (IMSIU) (grant number IMSIU-RG23097).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ali, G.K., Algethami, F.K. & Omer, K.M. Gold single atom-based aptananozyme as an ultrasensitive and selective colorimetric probe for detection of thrombin and C-reactive protein. Microchim Acta 191, 59 (2024). https://doi.org/10.1007/s00604-023-06147-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-023-06147-6