Abstract
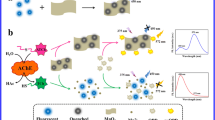
A novel fluorescent strategy has been developed by using an enzymatic reaction modulated DNA assembly on graphitic carbon nitride nanosheets (CNNS) for the detection of acetylcholinesterase (AChE) activity and its inhibitors. The two-dimensional and ultrathin-layer CNNS-material was successfully synthesized through a chemical oxidation and ultrasound exfoliation method. Because of its excellent adsorption selectivity to ssDNA over dsDNA and superior quenching ability toward the fluorophore labels, CNNS were employed to construct a sensitive fluorescence sensing platform for the detection of AChE activity and inhibition. The detection was based on enzymatic reaction modulated DNA assembly on CNNS, which involved the specific AChE-catalyzed reaction-mediated DNA/Hg2+ conformational change and subsequent signal transduction and amplification via hybridization chain reaction (HCR). Under the excitation at 485 nm, the fluorescence signal from 500 to 650 nm (λmax = 518 nm) of the developed sensing system was gradually increased with increasing concentration of AChE. The quantitative determination range of AChE is from 0.02 to 1 mU/mL and the detection limit was 0.006 mU/mL. The developed strategy was successfully applied to the assay of AChE in human serum samples, and can also be used to effectively screen AChE inhibitors, showing great promise providing a robust and effective platform for AChE-related diagnosis, drug screening, and therapy.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Acetylcholinesterase (AChE) plays a pivotal role in maintaining the normal function of the cholinergic nerves in the central and peripheral nervous system [1,2,3]. Numerous research have provided evidence that the abnormal content of acetylcholine caused by AChE is closely associated with the occurrence and development of neurodegenerative diseases such as Alzheimer’s disease (AD), myasthenia gravis, depression, and Parkinson’s disease [4, 5]. Recently, AChE has been considered as an early biomarker and significant therapeutic target for the development of new drugs for the treatment of dementia [6]. For example, some AChE inhibitors that can affect AChE activity and enhance cholinergic neurotransmission were widely used in the AD therapies [7, 8]. Therefore, the development of efficient methods for specific detection of AChE activity and screening its potential inhibitors were not only very crucial for elucidating the function of AChE in complicated living systems but also for providing useful tools to AChE-related therapy and biomedical research.
Until now, considerable efforts have been made to develop efficient strategies to monitor the enzymatic activity of AChE and screen its potential inhibitors. The most widely used method for AChE activity is the colorimetric detection by using Ellman’s reagent [9]. However, such methods may display some shortcomings such as low sensitivity, false-positive effects [10]. In recent years, various kinds of analytical techniques, including colorimetric methods [11,12,13], chemiluminescence [14], surface-enhanced Raman scattering (SERS) [15], fluorescence detection [16,17,18,19], and electrochemical analysis [20], have been developed for the sensing of AChE. In addition, advanced materials such as gold nanoparticles [21], quantum dots [22], single-walled carbon nanotubes (SWCNTs) [23], macromolecule polymers [24], and metal-organic frameworks (MOFs) [25] have also been utilized to construct novel strategies for AChE activity measurement. These approaches, however, still suffered from certain drawbacks, such as lack of sufficient sensitivity and specificity, the assistance of other enzymes for signal amplification and readout, and the requirement of expensive materials or instruments. Hence, the development of a new, sensitive, selective, and cost-effective method for the detection of AChE and its inhibitors is highly desirable.
Graphitic carbon nitride nanosheets (CNNS), a metal-free two-dimensional (2D) layered framework, belong to a type of rising star in the carbon-based nanomaterials family [26, 27]. The structure of CNNS is composed of s-triazine or tri-s-triazine formed by alternating C−N atoms instead of C−C atoms through sp2 hybridization, in which weak van der Waals, forces exists and the layer distance is about 3.3 nm [28]. In general, the CNNS can be easily prepared through the pyrolysis of low cost nitrogen-rich precursors, including melamine, cyanamide, dicyandiamide, urea, and thiourea, and then exfoliated into ultrathin nanosheets with various sizes. Excitingly, the CNNS were found to possess unique physicochemical properties, such as excellent stability and preferable biocompatibility, high surface-to-volume ratio, facilely accessible functional sites on the basal planes, and inertness to environmental influences, which enable them suitable for biosensor, optoelectronic, and catalytic applications [29,30,31,32,33]. Moreover, like other carbon-based nanomaterials, CNNS have been shown to selectively adsorb single-stranded DNA (ssDNA) but not double-stranded DNA (dsDNA), which can be used to design fluorescence sensors for nucleic acids by combining their ability to quench fluorophores [34, 35].
Herein, we report a novel fluorescent strategy for sensitive detection of AChE activity and for screening of its potential inhibitors. To the best of our knowledge, it is the first time that a highly effective enzymatic reaction modulated DNA assembly on CNNS has been developed for the assay of AChE activity and inhibition. Because of the ease of operation, satisfying sensitivity and specificity, this developed fluorescence sensing system will afford a useful platform for the detection of AChE activity and screening of its inhibitors.
Experimental section
Chemicals and materials
HPLC-purified oligonucleotides (Table S1) were synthesized by Shanghai Sangon Biotech Co., Ltd. (Shanghai, China). Acetylcholinesterase (AChE), acetylthiocholine chloride (ATCh), bovine albumin (BSA), glucose oxidase (GOX), lysozyme (Lys) ribonuclease (RNase), and horseradish peroxidase (HRP) were purchased from Sigma-Aldrich. 5,5′-dithiobis (2-nitro-benzoic acid) (DTNB), tacrine, dicyanamide, tris (hydroxymethyl) aminomethane (Tris), and magnesium sulfate (MgSO4) were gained from Aladdin Chemistry Co., Ltd. (Shanghai, China). All other reagents were of analytical grade. The human serum samples were obtained from the First Affiliated Hospital of Guangxi Medical University (Nanning, China). The double-distilled water, which was purified by a Milli-Q system (Millipore, Bedford, MA) with an electric resistance >18.25 MΩ·cm, was used for preparing needed solutions.
Apparatus and characterization
The fluorescence measurements were carried out on an FL-8500 fluorescence spectrometer (Perkin Elmer, USA). The transmission electron microscope (TEM) images were measured using a field-emission high-resolution 2100F TEM (JEOL, Japan). The morphology of the nanosheets was examined by field-emission scanning electron microscopy (SEM, S-4800, Hitachi, Japan). The atomic force microscopy (AFM) image was taken on means of Bruker Bioscope system (Bruker, USA). Zeta potential measurement was conducted using a Malvern Zetasizer 90 particle size and zeta potential analyzer (Malvern, UK). X-ray diffraction (XRD) patterns were recorded with a MiniFlex 600 X-ray diffractometer (Rigaku, Japan). The X-ray photoelectron spectroscopy (XPS) analysis was detected using an X-ray photoelectron spectroscope (Thermo Scientific, USA). Fourier-transform infrared (FT-IR) experiments were performed on a Spectrum Two FT-IR Spectrometer (Perkin Elmer, USA).
Preparation of two-dimensional (2D) CNNS
The CNNS were prepared according to our previously methods with some modification [34, 36]. Specific preparation produce was described in the Electronic Supporting Material.
Determination of AChE activity and inhibitor
For AChE activity detection, I-DNA and Hg2+ ions were first mixed with a molar ratio of 1:4 to obtain I-DNA/Hg2+ hairpin probe owing to the formation of T-Hg2+-T base pair in the stem region. Then, 2 μL of 5 μM ATCh, 2 μL of different concentrations of AChE, and 26 μL reaction buffer (5 mM Tris-HCl, 5 mM MgSO4, pH 8.0) were incubated at 37 °C for 30 min. After that, 2 μL of 1 μM I-DNA/Hg2+ hairpin probe, 2 μL of 5 μM H1, and 2 μL of 5 μM H2 were added into the reaction system and incubated for 60 min at 37 °C. Thereafter, 6 μL of 2 mg/mL CNNS solution was added into the above mixtures for 10 min at room temperature with a total volume of 200 μL. Finally, the fluorescence analysis of the samples was carried out. The fluorescence emission spectra of samples were collected from 500 to 650 nm (λmax = 518 nm) with a 485-nm excitation wavelength.
To study the inhibition of tacrine on AChE activity, 2 μL of different concentrations of tacrine and 2 μL of 0.1 U/mL AChE were mixed in reaction buffer at 37°C for 30 min. Then, the following experimental procedures were the same as those used for the AChE activity assay.
Detection of AChE activity in real samples
Human serum samples were chosen as actual samples to evaluate the practicability of the sensing strategy. First, the serum samples were obtained by centrifuging the whole blood samples at 3500 rpm for 5 min. Then, 20 μL of the human serum samples was mixed with 20 μL different concentrations of AChE and was diluted to 20 mL with 20 mM Tris-HCl buffer (pH 7.4). The final concentrations of added AChE were 0, 0.1, 0.2, 0.4, 0.8, and 1.0 mU/mL in the fluorescence analysis samples, respectively. Subsequently, the determination of AChE in the human serum samples was carried out using the procedure as described in the above.
Results and discussion
Characterization of two-dimensional (2D) CNNS
The main synthetic process of CNNS has already been described in our previous research works [34, 36]. Results from TEM (Fig. 1A) and SEM (Fig. S1) revealed that the as-obtained CNNS showed a 2D planar sheet structure with a mean length of about 110 nm and width of 95 nm. Moreover, the as-prepared CNNS are positively charged, with a zeta potential up to +29.2 mV (Fig. S2). The high surface charge endows the CNNS with excellent stability and can stand still for several weeks without any aggregation, which is favorable for further biomedical and biosensing applications. The AFM image and section analysis for CNNS (Figs. 1B and 1C) disclosed that the thickness of these nanosheets is ∼1.5 nm, suggesting they mainly comprised of a single layer. The XRD pattern presented a strong diffraction peak located at 27.5° (d = 0.32 nm), corresponding to the typical graphitic interlayer stacking (002) peak of CNNS. The other weak peak is (100) phase at 13.1° (d =0.67 nm), which can be ascribed to the periodic in-planar structure of the tri-s-triazine ring. Compared with bulk g-C3N4, the peak of CNNS located at 27.5° sharply decreased, and the peak located at 13.1° almost disappeared, implying that few-layered structure of CNNS after exfoliation (Fig. 1D) [37].
The surface structure and composition of the as-prepared CNNS were then investigated. The FT-IR spectrum (Fig. 1E) of the CNNS exhibits a distinct absorption peak at 814 cm−1, which was due to the vibration of the triazine ring. Furthermore, the absorption peaks around 1000 and 1800 cm−1 are assigned to the characteristic stretching band of C-N heterocycles (C−N(−C)−C and C−NH−C) [38]. For a better insight, XPS measurement was performed. As shown in Fig. 1F, the CNNS contain C, N, O three elements, and that the atomic ratio of N/C was calculated to be 1.35, closing to the stoichiometric value of 1.33. The relatively high oxygen content within CNNS may be caused by chemical oxidation and ultrasonic exfoliation. XPS analysis of the C 1s spectrum reveals two peaks at 284.4 and 287.8 eV, which are identified as the sp3-bonded carbon with oxygen (C−OH) and sp2-bonded carbon (N−C=N), respectively (Fig. 1G). The high-resolution N 1s spectrum is shown in Fig. 1H. It can be deconvoluted into four peaks, mainly located at 398.1, 399.2, 400.3, and 404.7 eV, demonstrating the presence of sp2-hybridized nitrogen (C=N−C), C−NH bond and sp3-hybridized nitrogen (N−[C]3), quaternary N bonded to three C in the aromatic cycles and terminal nitrate groups in heterocycles. What’s more, the peaks at around 531.0 and 532.1 eV in the XPS spectrum of O 1s belong to N−C=O and C−OH groups of CNNS, respectively (Fig. 1I) [39]. It can be seen that the surface components of the CNNS determined by the XPS were in good agreement with FT-IR results. Taken together, these results demonstrated that the CNNS has been successfully prepared.
Design principle
As illustrated in Scheme 1A, CNNS were firstly obtained by directly using chemical oxidation and ultrasound method to handle the bulk graphitic carbon nitride (g-C3N4) with pyrolysis of dicyanamide under high temperature. Then, the assay of AChE activity and its inhibitors were performed through enzymatic reaction modulated DNA assembly on CNNS (Scheme 1B).
In this rationally designed sensing system for AChE, three DNA probes (I-DNA, FAM-H1, and H2, Table S1), Hg2+, acetylthiocholine (ATCh), and CNNS were mainly involved. The I-DNA probe that containing eight thymin (T) bases can hybridize with 4 equiv of Hg2+ to form a stem-loop hairpin structure (I-DNA with T-Hg2+-T) through Hg2+-mediated T-T base pairing. When the analytical target AChE was introduced, the analogue of acetylcholine─ATCh can be catalyzed and decomposed to thiocholine (TCh) that containing the chemically reactive group (-SH). The resulting thiol products can bind to with Hg2+ and effectively captures Hg2+ from the hairpin-shaped I-DNA, owing to the higher affinity interaction between TCh and Hg2+. Accompanied by the release of Hg2+ triggered by AChE hydrolysis products, the hairpin I-DNA will be liberated into a single-stranded conformation, revealing its 3′-terminal, and thus generate a long dsDNA products via hybridization chain reaction (HCR) [40, 41], which was triggered by I-DNA for alternating hybridization between FAM-H1 and H2. Notably, the FAM-labeled HCR product has a relative rigid conformation, which remains highly fluorescent upon addition of CNNS because of the weak assembly of dsDNA on CNNS. On the contrary, the catalytic hydrolysis of AChE is blocked in the presence of AChE inhibitor, and thus less TCh is generated, resulting in I-DNA with T-Hg2+-T maintained their initial hairpin configurations in the detection system. In this case, the hairpin structured I-DNA disables the assembly of the HCR product. As a result, the unreacted hairpin DNA probes, FAM-H1 and H2, can be adsorbed and assembled on the CNNS surface through their sticky ends and loops, delivering a very weak fluorescence signal due to the super quenching ability of CNNS toward the fluorescence of FAM labels. Therefore, the concentrations of AChE can be detected by the change of fluorescence signal.
Feasibility study of AChE detection
With the as-prepared CNNS, we then test their feasibility as a fluorescent sensing platform for AChE detection. As illustrated in Fig. 2A, the mixture sample of H1 and H2 was found to exhibit a high fluorescence signal. After 60 μg/mL CNNS was introduced into the sample, a remarkable fluorescence decrease (> 92%) was obtained. This was ascribed to the fact that the strong adsorption of the hairpin DNA on the CNNS surfaces, which resulted in the proximity of fluorescein to CNNS and effective quenching of the fluorescence of the dye through photoexcited electrons transfer from the fluorophore to CNNS [34, 35]. In contrast, in the presence of 10 nM I-DNA, the response signal recovered even after CNNS were added into the solution, testifying that I-DNA was able to initiate the hybridization chain amplification with H1 and H2. In addition, if Hg2+ was further added, which could bind with I-DNA to form a hairpin structure, preventing the trigger of HCR, and thus the response signal remained almost unchanged. However, when reaction solution of AChE with its substrate ATCh was added to the mixture of Hg2+/I-DNA/H1/H2@CNNS, a significant fluorescence enhancement up to ∼8-fold was obtained. Moreover, in the comparative experiment, after only incubation of AChE or ATCh with the mixture, no obvious fluorescence response can still be observed. These data indicated that the hydrolysis product of ATCh by AChE inhibited the binding of Hg2+ with I-DNA, resulting in the trigger of HCR and providing a sensitive detection of AChE.
A The fluorescence spectral response obtained by incubating different probes. The concentrations of H1 and H2 were 50 nM, I-DNA was 10 nM, Hg2+ was 40 nM, CNNS was 60 μg/mL, ATCh was 50 nM, and AChE was 3 mU/mL. B Agarose gel electrophoresis images. Lane 1, 0.2 μM I-DNA; lane 2, 1 μM H1; lane 3, 1 μM H2; lane 4, 1 μM H1 + 1 μM H2; lane 5, 0.2 μM I-DNA + 1 μM H1 + 1 μM H2; lane 6, 0.2 μM I-DNA + 0.8 μM Hg2+ + 1 μM H1 + 1 μM H2; lane 7, 0.5 μM ATCh + 30 mU/mL AChE + 0.2 μM I-DNA + 0.8 μM Hg2+ + 1 μM H1 + 1 μM H2
Further gel electrophoresis analysis was performed to investigate the AChE mediated hybridization chain assembly (Fig. 2B). Three clear bands were observed for I-DNA, H1, and H2, respectively (lanes 1, 2, and 3). The mixture of two metastable H1 and H2 hairpin probes did not show any new band (lane 4), verifying a low back-ground of the HCR amplification. When incubating H1 and H2 with I-DNA, a new ladder-like band with higher molecular weights appeared, implying the initiation of the hybridization chain assembly by I-DNA (lane 5). Furthermore, we found that only a negligible ladder-like band was obtained when the same system was further treated by Hg2+ (lane 6), which indicated that Hg2+ can efficiently inhibit the occurrence of HCR. However, upon addition of the reaction solution of AChE with its substrate ATCh to the mixture of Hg2+/I-DNA/H1/H2, an obvious ladder-like HCR band was observed (lane 7), which is similar with the direct incubation of I-DNA with H1 and H2 (lane 5). Collectively, these results demonstrated the great potential of the proposed biosensing strategy for AChE detection.
Analytical performance of the fluorescence AChE assay
After verifying the feasibility, the analytical performance of developed fluorescence sensing system response to AChE was then evaluated. To achieve best performance for AChE, we first optimized several important parameters, including the concentration of CNNS, Hg2+, ATCh, and incubation time (Figs. S3 to S6). Accordingly, 60 μg/mL CNNS, 40 nM Hg2+, 0.05 μM ATCh, and 60 min reaction time were chosen as the optimal experimental conditions. Under the optimized experimental conditions, the fluorescence emission spectra of the sensing system in the presence of AChE at varying concentrations were measured. For the fluorescence measurement of AChE, after incubating different concentrations of AChE with its substrate ATCh, the resulting solution was then added to the mixture of Hg2+/I-DNA/H1/H2@CNNS and incubated at 37 °C for another 60 min. We found that the fluorescence signal was gradually increased with the increasing concentration of AChE (Fig. 3A). As can be seen in Fig 3B, there exhibited an excellent linear relationship between the fluorescence peak values at 518 nm and concentration of AChE ranging from 0.02 to 1.0 mU/mL (R2 = 0.996, inset in Fig. 3B). The detection limit (3σ/slope) was calculated to be as low as 0.006 mU/mL, which was comparable or better than most of existing assay methods especially those fluorescence assays for AChE detection (Table S2). Though the sensitivity is inferior to some methods, the CNNS adopted in this strategy has several advantages including cost-efficient, excellent stability, low biotoxicity, and outstanding quenching capacity. Moreover, the construction of this sensing system is easy-to-operate.
A Fluorescence spectral responses of CNNS-based sensing system to AChE with different concentrations. B Fluorescence peak intensity at 518 nm versus the concentration of AChE (Ex = 485 nm). Inset: linear plot of the fluorescence peak intensity at 518 nm against the concentration of AChE in the range from 0.02 to 1.0 mU/mL. The error bars represent the standard deviation of three measurements
In addition, to evaluate the selectivity of the developed sensing strategy, various potential interfering substances including glucose oxidase (GOX), lysozyme (Lys), bovine serum albumin (BSA), ribonuclease (RNase), and horseradish peroxidase (HRP) were tested under the same conditions. As shown in Fig. 4, compared with the target AChE, none of these proteins gave any noticeable fluorescence response, which clearly suggests that these proteins did not interfere with the assay, and developed sensing strategy has excellent selectivity for AChE detection. This excellent selectivity of our method can be attributed to the enzymatic reaction specificity of AChE and high affinity between the hydrolysis product of AChE and Hg2+. More importantly, the high specificity of this sensing method also provided the potential for the detection of AChE activity in complex real samples.
Detection of AChE in real samples
Next, we further explored the practical applicability of the developed fluorescence sensing system. The AChE in human serum samples were measured using proposed strategy through the standard addition method. Different concentrations of AChE were added in human serum sample to prepare the spiked samples. As shown in Table 1, the recoveries were in the range from 98.2 to 104.8% with an RSD of less than 4.0%. Besides, it is also clearly found that the detection results of AChE in human serum samples obtained by our sensing system were in accord with those data tested by standard Ellman’s method. These observations disclosed that the established method is applicable for AChE detection in complex real samples with satisfactory reliability.
AChE inhibitor assay
To demonstrate whether the developed sensing strategy could be also used for the screening of AChE inhibitors, a well-known AChE inhibitor tacrine that used for treatment of Alzheimer’s disease [42] was chosen as the case of study. In this case, 2 μL of tacrine at different concentrations was first mixed with 0.1 U/mL AChE for 30 min, and then incubated with the reaction solution of the sensing system. It was found that the fluorescence intensities decreased with the addition of increasing concentrations of the tacrine, clearly indicating the inhibitory capacity for AChE activity (Fig. 5). The corresponding IC50 value (the inhibitor concentration required for 50% inhibition of the enzyme activity) of tacrine was estimated to be 7.2 nM, which was comparable with those obtained by other AChE assays [43]. The above-mentioned results confirmed that our proposed sensing method possess great potential to screen AChE inhibitors and provides a broad prospect for AChE-based drug discovery.
The inhibition analysis of tacrine on AChE activity. A Fluorescence spectra of the biosensing platform at various concentrations of tacrine. B Calibration curve corresponding to the fluorescence intensity as a function of tacrine concentration. The error bars represent the standard deviation of three measurements
Although this established method for sensing AChE activity and screening its inhibitors displays many remarkable features, it still had some limitations. One major disadvantage of the method was that strong oxidizing reagents were adopted for the synthesis of CNNS and the preparation procedure is time-consuming, which usually required chemical oxidation and ultrasound exfoliation for more than 10 h. Additionally, hybridization chain reaction (HCR) usually need a relatively long reaction time to generate a significantly amplified readout signal, which can be ascribed to its slow reaction kinetics [44]. Therefore, our future research work will focus on the development of green and rapid methods for the preparation of CNNS. Moreover, the optimal sequences of DNA probes and the experimental conditions (such as reaction temperature, reaction buffer, and the concentration of metal ions) will further be investigated to circumvent the limitation.
Conclusion
In this work, the ultrathin CNNS was successfully synthesized to demonstrate excellent selective recognition toward dsDNA over ssDNA and further utilized to establish a fluorescence sensing platform for the assay of AChE activity and inhibition. Combining the specific AChE-catalyzed reaction-mediated DNA/Hg2+ conformational change and well-defined signal transduction and amplification via HCR, the constructed sensing platform enables sensitive and selective detection of AChE. Moreover, the developed strategy was successfully applied to the assay of AChE in real complex samples and could efficiently perform the screening of the AChE inhibitor. Given the method’s merits including ease of operation, satisfying sensitivity and specificity, our proposed assay system will hold certain practicability and application potential in the clinical diagnosis of AChE-related disease and drug screening.
References
Soreq H, Seidman S (2001) Acetylcholinesterase-new roles for an old actor. Nat Rev Neurosci 2:294–302. https://doi.org/10.1038/35067589
Behra M, Cousin X, Bertrand C, Vonesch JL, Biellmann D, Chatonnet A, Strahle U (2002) Acetylcholinesterase is required for neuronal and muscular development in the zebrafish embryo. Nat Neurosci 5:111–118. https://doi.org/10.1038/nn788
Silman I, Sussman JL (2017) Recent developments in structural studies on acetylcholinesterase. J Neurochem 142:19–25. https://doi.org/10.1111/jnc.13992
Fedorova TD, Seidelin LB, Knudsen K, Schacht AC, Geday J, Pavese N, Brooks DJ, Borghammer P (2017) Decreased intestinal acetylcholinesterase in early Parkinson disease: an 11C-donepezil PET study. Neurology 88:775–781. https://doi.org/10.1212/WNL.0000000000003633
Mineur YS, Obayemi A, Wigestrand MB, Fote GM, Calarco CA, Li AM, Picciotto MR (2013) Cholinergic signaling in the hippocampus regulates social stress resilience and anxiety- and depression-like behavior. Proc Natl Acad Sci USA 110:3573–3578. https://doi.org/10.1073/pnas.1219731110
Richter N, Beckers N, Onur OA, Dietlein M, Tittgemeyer M, Kracht L, Neumaier B, Fink GR, Kukolja J (2018) Effect of cholinergic treatment depends on cholinergic integrity in early Alzheimer’s disease. Brain 141:903–915. https://doi.org/10.1093/brain/awx356
Costanzi S, Machado JH, Mitchell M (2018) Nerve agents: what they are, how they work, how to counter them. ACS Chem Neurosci 9:873–885. https://doi.org/10.1021/acschemneuro.8b00148
Savelieff MG, Nam G, Kang J, Lee HJ, Lee M, Lim MH (2019) Development of multifunctional molecules as potential therapeutic candidates for Alzheimer’s disease, Parkinson’s disease, and amyotrophic lateral sclerosis in the last decade. Chem Rev 119:1221–1322. https://doi.org/10.1021/acs.chemrev.8b00138
Ellman GL, Courtney KD, Andres V Jr, Featherstone RM (1961) A new and rapid colorimetric determination of acetylcholinesterase activity. Biochem Pharmacol 7:88–95. https://doi.org/10.1016/0006-2952(61)90145-9
Tian T, Li X, Cui J, Li J, Lan Y, Wang C, Zhang M, Wang H, Li G (2014) Highly sensitive assay for acetylcholinesterase activity and inhibition based on a specifically reactive photonic nanostructure. ACS Appl Mater Interfaces 6:15456–15465. https://doi.org/10.1021/am504065v
Liu Y, Wei X, Chen J, Yu YL, Wang JH, Qiu H (2022) Acetylcholinesterase activity monitoring and natural anti-neurological disease drug screening via rational design of deep eutectic solvents and CeO2-Co(OH)2 nanosheets. Anal Chem 94:5970–5979. https://doi.org/10.1021/acs.analchem.2c00428
Liu DM, Xu B, Dong C (2021) Recent advances in colorimetric strategies for acetylcholinesterase assay and their applications. TrAC Trend Anal Chem 142:116320. https://doi.org/10.1016/j.trac.2021.116320
Cai Y, Zhu H, Zhou W, Qiu Z, Chen C, Qileng A, Li K, Liu Y (2021) Capsulation of AuNCs with AIE effect into metal–organic framework for the marriage of a fluorescence and colorimetric biosensor to detect organophosphorus pesticides. Anal Chem 93(19):7275–7282. https://doi.org/10.1021/acs.analchem.1c00616
He L, Jiang ZW, Li W, Li CM, Huang CZ, Li YF (2018) In situ synthesis of gold nanoparticles/metal-organic gels hybrids with excellent peroxidase-like activity for sensitive chemiluminescence detection of organophosphorus pesticides. ACS Appl Mater Interfaces 10:28868–28876. https://doi.org/10.1021/acsami.8b08768
Chen Y, Zhao W, Si J, Zheng Y, Tan H, Meng F, Yang G, Gu Y, Qu L (2022) Highly selective SERS detection of acetylcholinesterase in human blood based on catalytic reaction. Anal Chim Acta 1232:340495. https://doi.org/10.1016/j.aca.2022.340495
Wang X, Li P, Ding Q, Wu C, Zhang W, Tang B (2019) Observation of acetylcholinesterase in stress-induced depression phenotypes by two-photon fluorescence imaging in the mouse brain. J Am Chem Soc 141:2061–2068. https://doi.org/10.1021/jacs.8b11414
Li Y, Liang H, Lin B, Yu Y, Wang Y, Zhang L, Cao Y, Guo M (2021) A ratiometric fluorescence strategy based on inner filter effect for Cu2+-mediated detection of acetylcholinesterase. Microchim Acta 188:385. https://doi.org/10.1007/s00604-021-05044-0
Wu X, An JM, Shang J, Huh E, Qi S, Lee E, Li H, Kim G, Ma H, Oh MS, Kim D, Yoon J (2020) A molecular approach to rationally constructing specific fluorogenic substrates for the detection of acetylcholinesterase activity in live cells, mice brains and tissues. Chem Sci 11:11285–11292. https://doi.org/10.1039/d0sc04213g
Chen S, Li Z, Huang Z, Ji Q (2021) Investigation of efficient synergistic and protective effects of chitosan on copper nanoclusters: construction of highly active and stable nanozyme for colorimetric and fluorometric dual-signal biosensing. Sens Actuators, B Chem 332:129522. https://doi.org/10.1016/j.snb.2021.129522
Wan Y, Wang H, Zhang L, Chen Y, Li S, Zhou J, Zhang Q, Xia L (2022) Highly stable acetylcholinesterase electrochemical biosensor based on polymerized ionic liquids microgel for pesticides detection. Microchim Acta 189:300. https://doi.org/10.1007/s00604-022-05383-6
Zhang J, Zheng W, Jiang X (2018) Ag+-gated surface chemistry of gold nanoparticles and colorimetric detection of acetylcholinesterase. Small 14:1801680. https://doi.org/10.1002/smll.201801680
Pei T, He Y, Wang Y, Song G (2021) Fluorine-free synthesis of Ti3C2 MQDs for smartphone-based fluorescent and colorimetric determination of acetylcholine- esterase and organophosphorus pesticides. Microchim Acta 189:1–9. https://doi.org/10.1007/s00604-021-05115-2
Loewenthal D, Kamber D, Bisker G (2022) Monitoring the activity and inhibition of cholinesterase enzymes using single-walled carbon nanotube fluorescent sensors. Anal Chem 94:14223–14231. https://doi.org/10.1021/acs.analchem.2c02471
Sgobbi LF, Machado SA (2018) Functionalized polyacrylamide as anacetyl- cholinesterase-inspired biomimetic device for electrochemical sensing of organophosphorus pesticides. Biosens Bioelectron 100:290–297. https://doi.org/10.1016/j.bios.2017.09.019
Li S, Wei Z, Xiong L, Xu Q, Yu L, Xiao Y (2022) In situ formation of o-phenylenediamine cascade polymers mediated by metal-organic framework nanozymes for fluorescent and photothermal dual-mode assay of acetylcholin- esterase activity. Anal Chem 94:17263–17271. https://doi.org/10.1021/acs.analchem.2c04218
Liao G, He F, Li Q, Zhong L, Zhao R, Che H, Gao H, Fang B (2020) Emerging graphitic carbon nitride-based materials for biomedical applications. Prog Mater Sci 112:100666. https://doi.org/10.1016/j.pmatsci.2020.100666
Ahmad T, Khan S, Rasheed T, Ullah N (2022) Graphitic carbon nitride nanosheets as promising candidates for the detection of hazardous contaminants of environmental and biological concern in aqueous matrices. Microchim Acta 189:426. https://doi.org/10.1007/s00604-022-05516-x
Chen L, Song J (2017) Tailored graphitic carbon nitride nanostructures: synthesis, modification, and sensing applications. Adv Funct Mater 27:1702695. https://doi.org/10.1002/adfm.201702695
Wu G, Hu Y, Liu Y, Zhao J, Chen X, Whoehling V, Plesse C, Nguyen GT, Vidal F, Chen W (2015) Graphitic carbon nitride nanosheet electrode-based high-performance ionic actuator. Nat Commun 6:7258. https://doi.org/10.1038/ncomms8258
Wang Y, Liu X, Zheng C, Li Y, Jia S, Li Z, Zhao Y (2018) Tailoring TiO2 nanotube-interlaced graphite carbon nitride nanosheets for improving visible-light-driven photocatalytic performance. Adv Sci 5:1700844. https://doi.org/10.1002/advs.201700844
Gupta N, Todi K, Narayan T, Malhotra BD (2022) Graphitic carbon nitride-based nanoplatforms for biosensors: design strategies and applications. Mater Today Chem 24:100770. https://doi.org/10.1016/j.mtchem.2021.100770
Hoh HY, Zhang Y, Zhong YL, Bao Q (2021) Harnessing the potential of graphitic carbon nitride for optoelectronic applications. Adv Opt Mater 9:2100146. https://doi.org/10.1002/adom.202100146
Zhao H, Jiang Z, Xiao K, Sun H, Chan HS, Tsang TH, Yang S, Wong PK (2021) Photo-assisted separation of noble-metal-free oxidation and reduction cocatalysts for graphitic carbon nitride nanosheets with efficient photocatalytic hydrogen evolution. Appl Catal B Environ 280:119456. https://doi.org/10.1016/j.apcatb.2020.119456
Xiang MH, Li N, Liu JW, Yu RQ, Jiang JH (2020) A tumour mRNA-triggered nanoassembly for enhanced fluorescence imaging-guided photodynamic therapy. Nanoscale 12:8727–8731. https://doi.org/10.1039/d0nr00941e
Hu K, Zhong T, Huang Y, Chen Z, Zhao S (2015) Graphitic carbon nitride nanosheet-based multicolour fluorescent nanoprobe for multiplexed analysis of DNA. Microchim Acta 182:949–955. https://doi.org/10.1007/s00604-014-1412-5
Liu JW, Wang YM, Zhang CH, Duan LY, Li Z, Yu RQ, Jiang JH (2018) Tumor-targeted graphitic carbon nitride nanoassembly for activatable two-photon fluorescence imaging. Anal Chem 90:4649–4656. https://doi.org/10.1021/acs.analchem.7b05192
Zhang X, Xie X, Wang H, Zhang J, Pan B, Xie Y (2013) Enhanced photoresponsive ultrathin graphitic-phase C3N4 nanosheets for bioimaging. J Am Chem Soc 135:18–21. https://doi.org/10.1021/ja308249k
Niazi S, Khan IM, Yu Y, Pasha I, Lv Y, Mohsin A, Mushtaq BS, Wang Z (2020) A novel fluorescent aptasensor for aflatoxin M1 detection using rolling circle amplification and g-C3N4 as fluorescence quencher. Sensor Actuat B Chem 315:128049. https://doi.org/10.1016/j.snb.2020.128049
Guo X, Yue G, Huang J, Liu C, Zeng Q, Wang L (2018) Label-free simultaneous analysis of Fe(III) and ascorbic acid using fluorescence switching of ultrathin graphitic carbon nitride nanosheets. ACS Appl Mater Interfaces 10:26118–26127. https://doi.org/10.1021/acsami.8b10529
Chen HJ, Hu Y, Yao P, Ning D, Zhang YP, Wang ZG, Liu SL, Pang DW (2021) Accurate and efficient lipoprotein detection based on the HCR-DNAzyme platform. Anal Chem 93:6128–6134. https://doi.org/10.1021/acs.analchem.0c05322
Kim J, Shim JS, Han BH, Kim HJ, Park J, Cho IJ, Kang SG, Kang JY, Bong KW, Choi N (2021) Hydrogel-based hybridization chain reaction (HCR) for detection of urinary exosomal miRNAs as a diagnostic tool of prostate cancer. Biosens Bioelectron 192:113504. https://doi.org/10.1016/j.bios.2021.113504
Wang Y, Xue Y, Zhao Q, Wang S, Sun J, Yang X (2022) Colorimetric assay for acetylcholinesterase activity and inhibitor screening based on metal-organic framework nanosheets. Anal Chem 94:16345–16352. https://doi.org/10.1021/acs.analchem.2c03290
Zhang Y, Cai Y, Qi Z, Lu L, Qian Y (2013) DNA-templated silver nanoclusters for fluorescence turn-on assay of acetylcholinesterase activity. Anal Chem 85:8455–8461. https://doi.org/10.1021/ac401966d
Dirks RM, Pierce NA (2004) Triggered amplification by hybridization chain reaction. Proc Natl Acad Sci USA 101:15275–15278. https://doi.org/10.1073/pnas.0407024101
Funding
This work was supported by the National Nature Science Foundation of China (No. 22266007), the Basic Ability Enhancement Program for Young and Middle-aged Teachers of Guangxi (No. 2022KY0098), and the Project of Advanced Innovation Teams and Xinghu Scholars Program of Guangxi Medical University (No. 02304001020X).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, T., Qin, Y., Wang, B. et al. Enzymatic reaction modulated DNA assembly on graphitic carbon nitride nanosheets for sensitive fluorescence detection of acetylcholinesterase activity and inhibition. Microchim Acta 190, 268 (2023). https://doi.org/10.1007/s00604-023-05850-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-023-05850-8