Abstract
A lateral-flow immunochromatographic assay with excellent sensitivity and wide application potential is described. The bovine serum albumin (BSA) antibody was immobilized in the test line for universality, and preincubation was introduced for high method sensitivity. Carboxy-modified CdSe/ZnS core–shell nanoparticles were used as label, and the fluorescence peaking at 605 nm was detected. The fluorescence in the test line was negative against the relevant analyte content. The chloramphenicol (CAP) and the aflatoxin M1 (AFM1) in milk were detected using the same strip to validate the universality. After optimization, the detection limit for CAP is 10 pg·mL−1, which is three times less that of a conventional assay (30 pg·mL−1). The detection limit for AFM1 was 6 pg·mL−1, which was 13 times less than that of a conventional assay (8 pg·mL−1). The method was applied in the analysis of spiked milk samples. The performance was compared with that of the commercial ELISA kit, and good agreement was observed.

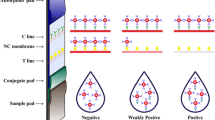
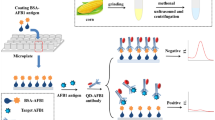
Schematic representation of the universal and sensitive combined immunochromatographic assay (USICA) and conventional immunochromatographic assay (TICA) of chloramphenicol (CAP) and aflatoxin M1.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chloramphenicol (CAP) is widely used in veterinary medicine, but excessive levels of CAP in human blood may lead to diseases such as leukemia, aplastic anemia, and gray baby syndrome [1, 2]. Thus, the use of CAP is banned in food-producing animals in many countries, including China, the USA, and the European Union, and a minimum required performance limit value is set at 0.3 × 10−6 g·kg−1 [3]. As a group I carcinogen and “milk toxin,” aflatoxin B1 (AFB1)-derived aflatoxin M1 (AFM1) is hazardous to humans exposed to contaminated milk [4,5,6]. The limits on AFM1 levels in milk and dairy products were set, and the sources of AFM1 contamination need to be monitored [7].
A paper-based lateral flow immunochromatographic assay (ICA) was developed for sensitive and economical on-site determination of residues, including CAP and AFM1 [8,9,10]. For better ICA detection efficiency, several signal labels, including time-resolved fluorescence [11], up-converting phosphors [12], magnetic nanoparticles [13], quantum dots (QDs) [14], and QD submicrobead (QB), have been developed and discussed. QB has attracted great interest for its unique properties, such as narrow fluorescent emission spectra, high quantum yield, high photochemical stability, and high-throughput detection [15,16,17,18,19]. Single or several distinguishable signal-providing labels are applied in a single ICA detection for multiplexing and usually involves the modification of the architecture of the ICA strip [20,21,22,23,24,25,26,27,28].
However, one ubiquitous characteristic of the existing ICA system is that target-specific antigen or antibody immobilized on the testing zone hinders the development of another crucial property: universality. Transient and largely uncontrolled immune recognition on the testing zone has led to insufficient reaction and limited sensitivity.
In this work, a universal and sensitive ICA (USICA) for CAP and AFM1 was established using the anti-BSA antibody as coating element, and a preincubation was introduced. BSA-modified CAP (CAP–BSA) exposes the binding site thoroughly in mixture and competes with CAP for the binding site on the anti-CAP antibody-modified QB in the preincubation step for high sensitivity. Two kinds of immune complexes, namely, QB–mAb–CAP and QB–mAb–CAP–BSA, were formed. The QB–mAb–CAP–BSA is conjugated with the anti-BSA antibody on the test zone by the BSA terminal, and the QB–mAb–CAP is captured by the anti-IgG antibody on the control zone in the USICA strip. The immune complex QB–mAb–CAP–BSA is recognized by the BSA terminal, which is nontarget-specific and brings the foundation for universality. The AFM1 is detected for validation of universality by using the BSA-modified AFM1 and the USICA of the same structure. The properties of the two USICAs are systematically summarized and compared with those of the ICA based on traditional structure (TICA). The applicability of the USICA is demonstrated by analyzing spiked milk samples and its performance compared with the commercial ELISA kit.
Experimental
Materials and instruments
CAP and AFM1 standards, 1-ethyl-3-(3-(dimethylamino)propyl)-carbodiimide hydrochloride (EDC), and N-hydroxysuccinimide (NHS) were supplied by the J&K Scientific Ltd. (Shanghai, China, www.jkchemical.com/Company-inf.aspx?language=ch). The CAP−BSA and AFM1 − BSA conjugates, the commercial ELISA kits for CAP and AFM1, and the anti-CAP, anti-AFM1, anti-BSA, and goat antimouse IgG antibodies were supplied by the Shandong Landu Bio-Science & Technology Co., Ltd. (Shandong, China, www.11467.com/qiye/45274356.htm). Carboxylic group-modified CdSe/ZnS core–shell QBs (emission at 605 ± 5 nm) were purchased from the Beijing Najing Biological Technology Co., Ltd. (Beijing, China, www.najingbio.com/). The protein stabilizer solution (Cat: PR-SS-002, Huzhou Yingchuang Biological Technology Co., Ltd., Huzhou, China, www.innoreagents.com) was used for QB–mAb storage. Nitrocellulose (NC) membrane and the sample and the absorbent pads were obtained from the Kinbio Tech. Co., Ltd. (Shanghai, China, www.goldbio.cn/article-item-288.html). All other reagents (analytical grade) were supplied by the National Pharmaceutical Group Chemical Reagent Co., Ltd. (Shanghai, China, www.sinopharm.com/1156.html). A fluorescent strip reader was supplied by the Beijing Najing Biological Technology Co., Ltd. (Beijing, China). The USICA and TICA strips were prepared by using the BioDot XYZ platform combined with a BioJet Quanti3000k dispenser and motion controller from BioDot (Irvine, CA) and cut by an automatic programmable cutter from the Shanghai Jinbiao Biotechnology Co., Ltd. (Shanghai, China, www.goldbio.cn/article-item-288.html). Pure water was prepared using the Elix-3 and the Milli-QA system (Millipore Co., Bedford, MA, USA, http://www.well-honor.com/goods1-202.html).
QB modification with anti-target mAb
Carboxy-modified QB was activated and modified using the antitarget mAb through the active ester method [29]. Details are described in the Electronic Supporting Material (ESM).
Fabrication of the USICA and the TICA systems
The two ICA systems were similar in structure. These systems contain three parts: (1) the NC membrane in the middle, (2) the glass fiber, and (3) absorption pads on both ends overlapping nearly 2 mm with NC membrane on the backing card. The USICA system had an anti-BSA antibody at 0.25 mg·mL−1 immobilized at a density of 3 μL·cm−1 in the NC membrane as test line, whereas the TICA system had a BSA-modified target (CAP–BSA) at a density of 3 μL·cm−1. The goat antimouse IgG antibodies (0.5 mg mL−1) were immobilized in both ICA systems at a density of 3 μL·cm−1 in the NC membrane as the control line. The formed ICA system was dried at 37 °C in a blasting drying trunk for 2 h, cut into strips (3.5 mm in width) by using the automatic cutter, and stored at 4 °C until use.
Quantitative procedure of the USICA and the TICA systems
Parameters, such as the amount of mAb modified on QB, pH value, ionic strength, the amount of antigen coated on T line, the amount of surfactant, dilution ratio of QB–mAb, and ICA reaction time, was optimized in the TICA system for best detection results. In addition to the parameters optimized in the TICA system, the amount of antigen used for preincubation, the amount of anti-BSA mAb immobilized on the T line, and the incubation time were optimized in the USICA system. The optimization was carried out in competitive inhibition mode, in which the target was spiked at the final concentrations of 0 and 1.25 ng·mL−1, and the parameter inducing the most obvious inhibition was selected. The analyte was dissolved in methanol, and the final methanol concentration was 5% (v/v) in the spiked samples. All experiments were performed in triplicate. The fluorescence intensity (FI) in the T line was inversely proportional to the analyte content, and a calibration plot was established on this basis.
AFM1- and CAP-spiked samples were pretreated. Trichloroacetic acid was added to the spiked milk samples to a final concentration of 10% (w/v). After thorough mixing and centrifugation, 2.5 μL NaOH (5 mol·L−1) was used per 100 μL supernatant for neutralizing. The insoluble materials that appeared during neutralizing were removed by centrifugation, and the neutral supernatant was used for ICA. The CAP- and AFM1-free milk samples, which were confirmed by LC–MS/MS, were collected from the local market. Accuracy and precision analyses were carried out. The universality was validated by detecting the CAP and the AFM1 using the USICA strip.
Comparative study with commercial ELISA kit
Spiked milk samples were determined using the USICA and the TICA strips and the commercial ELISA kit. Sample pretreatment and the detection procedure for the commercial ELISA kit was performed according to the manufacturer’s instructions. A correlation coefficient (R2) of the two methods was calculated through a fit plot.
Results and discussion
Figure 1 presents a schematic for USICA and TICA using CAP as an example. In USICA, preincubation was introduced for high sensitivity, in which the target (CAP) and the BSA-modified target (CAP–BSA) competed for the binding site on the QB–mAb under optimal conditions. The anti-BSA antibody was introduced in the test line (T line) for universal detection, and the nontarget-specific and BSA-containing complexes were conjugated. The intensity of QB in the T line was inversely proportional to the amount of the target, which was the basis of establishment of the novel USICA system. Meanwhile, in the TICA system, the immune recognition that occurred on the testing zone was transient and largely uncontrolled, and universal detection was not discussed.
Schematic for USICA and TICA. Preincubation was introduced for competitive recognition of high sensitivity, and the T line was coated with anti-BSA antibody for universal detection (a); the competitive recognition occurred at the T line in TICA (b); Excess modified QBs (e.g., QB–CAP–mAb) were captured by element in the C line
QB modification with antitarget mAb
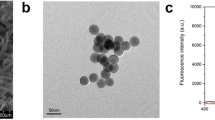
The morphology of the anti-AFM1 mAb-modified QB under a scanning electron microscope (SEM) is shown in Fig. 2a. A layer of material appeared on the surface (Fig. A insert) after modification. Fourier transform infrared (FTIR) spectroscopy was applied to characterize the result of modification. As shown in Fig. 2b, the characteristic absorption peaks corresponding to protein amide bands I (1641 cm−1) and II (1530 cm−1) existing in the QB–mAb proved successful modification. The anti-CAP mAb-modified QB was prepared using the same method and applied in the following experiment.
Property and analytical performance of the USICA and the TICA systems
In the TICA for small molecule detection, competitive recognition occurred on the test line. Target-specific antigen immobilized on the T line was adverse for full exposure of the binding site, and the transient and largely uncontrolled recognition on the T line negatively affected the degree of thorough recognition. The USICA system was established in this work to address these problems. Preincubation was introduced for high sensitivity. The target-specific antigen competed with the target for the binding site on the QB–mAb in the preincubation solution. The binding site was fully exposed in the solution, and competitive recognition was carried out in optimal conditions, including ample time. The anti-BSA antibody was introduced for universality. Two kinds of complexes were formed in preincubation, namely, QB–mAb–CAP and QB–mAb–CAP–BSA. Only the latter complexes were captured by the anti-BSA antibody through the BSA terminal in the complexes. The BSA in the target-specific antigen was nontarget-specific and brought the foundation for universality. Any analyte that can be modified with BSA and antibody-available may be detected by this method. CAP was detected using the USICA system under optimal conditions. For sensitivity comparison, CAP was also detected using the TICA system. The AFM1 was detected to validate the universality of USICA system, and the TICA system was used after systematic optimization. A calibration plot was established by plotting the B/B0 × 100% against the target amount in logarithm, where B and B0 represent the FI on the test line (FIT) with and without the presence of spiked analyte, respectively.
The LOD was defined as the concentration of the spiked target, at which 10% inhibition (IC10) of the total inhibition was reached. Optimal parameters for CAP and AFM1 using USICA and TICA are summarized in Table E1 (shown in ESM) After optimization, the LOD of USICA for CAP was nearly three times that of TICA (Table 1). The LOD for AFM1 improved about 13 times by using USICA compared with that using TICA. These results verify the high sensitivity and the universality of the USICA system. Compared with the recently reported nanomaterial-based optical methods for the determination of CAP and AFM1 in Table 1, the unique merit of USICA is its universality.
The specificity of the USICA and the TICA systems for CAP was investigated using the cross-reaction (CR) value. Similar results among structural analogs, such as thiamphenicol (TAP) and florfenicol (FF), and other contaminants, such as kanamycin (KNM), streptomycin (SPM), and ceftiofur (CTF), were observed. The CR value was calculated using the eq. CR% = [(IC50 analyte) / (IC50 analog)] × 100 [36]. Figure 3 shows that the structural analogs TAP (CR = 14.3% in USICA) and FF (CR = 2.3% in USICA) had obvious CR against CAP, and the other potential pollutants (KNM, SPM, and CTF) had CR values lower than 0.01. The specificity of the USICA and the TICA systems for AFM1 was also evaluated by CR. Results show that the structural analogs aflatoxin M2 (AFB1, CR = 58% in USICA), aflatoxin G2 (AFG2, CR = 55% in USICA), and aflatoxin B2 (AFG2, CR = 20% in USICA) had obvious CR against AFM1 and that the CR values of aflatoxin G1 (AFG1), aflatoxin B1 (AFB1), deoxynivalenol (DON), fumonisin B1 (FMB1), and zearalenone (ZEN) were negligible (< 0.01%). These results suggest that the USICA and the TICA system for CAP and AFM1 are specific. Although specific, analogs might fake the presence of the analyte, especially during the analysis of complex samples. Some specificity data were usually demonstrated for reference in actual use (Table 1).
The accuracy and precision of the USICA and the TICA systems were evaluated through the recovery of the intra- and inter-assay, and the results are summarized in Table 2. The analysis was carried out in triplicate at each spiked concentration. The intra-assay was completed within 1 day, and the inter-assay was performed continuously for 3 days. The average recoveries of the two ICA systems ranged from 85% to 116%, which is acceptable for ICA quantitative analysis [37].
Comparative study with a commercial ELISA kit
The performances of the USICA and the TICA systems for CAP or AFM1 were compared with that of the commercial ELISA kit to demonstrate reliability and practicability. For the USICA and the TICA systems, the spiked milk samples were pretreated using trichloroacetic acid, which was mentioned in the “quantitative procedure of the USICA and the TICA systems” section. For the ELISA, the sample pretreatment was performed according to the manufacturer’s instructions.
The results in Fig. 4 show that the USICA (R2 = 0.9854 for CAP and R2 = 0.9828 for AFM1) and the TICA (R2 = 0.9562 for CAP and R2 = 0.9664 for AFM1) systems were in good agreement with the ELISA method. In addition to their universality and high sensitivity, the USICA and the TICA systems were easier to operate and took no more than 40 min to complete one sample analysis. The traditional ELISA involved multiple incubation, washing, and coloring, which took nearly 90 min. The main limit of the established method is the increased susceptibility induced by the delicate preincubation. Each step should be performed exactly to address the side effect.
Correlation between the ICA systems (x-axis) and the ELISA (y-axis). The plot fits for CAP between USICA and ELISA (a) and between TICA and ELISA (b) and AFM1 between USICA and ELISA (c) and between TICA and ELISA (d). Eighteen blank samples were spiked with analyte standard solutions under different concentrations in each group
Conclusion
The USICA system combines the advantages of sensitivity and universality. The USICA system is beneficial for quantitative analysis due to its potential to provide a uniform ICA strip for different analytes. Any analyte can be detected using the USICA system if analyte-specific mAb and BSA-modified analytes are available. The core merit of the USICA is its universality, and the susceptibility induced by the delicate preincubation may be reduced by standard operations.
The availability of the ICA was improved by the USICA system, paving the way for its wider application. On the basis of this work, sensitive and universal detection systems for other analytes may be further investigated.
References
Abnous K, Danesh NM, Ramezani M, Emrani AS, Taghdisi SM (2016) A novel colorimetric sandwich aptasensor based on an indirect competitive enzyme-free method for ultrasensitive detection of chloramphenicol. Biosens Bioelectron 78:80–86
Duan Y, Wang LH, Gao ZQ, Wang HS, Zhang HX, Li H (2017) An aptamer-based effective method for highly sensitive detection of chloramphenicol residues in animal-sourced food using real-time fluorescent quantitative PCR. Talanta 165:671–676
Bangemann M (1994) Commission regulation (EC) no 1430/94, official journal of the European Communities. Brussels, Belgium
Van Egmond HP (2004) Natural toxins: risks, regulations and the analytical sit- uation in Europe. Anal Bioanal Chem 378:1152–1160
Guo XD, Wen F, Zheng N, Luo QJ, Wang HW, Wang H, Li SL, Wang JQ (2014) Development of an ultrasensitive aptasensor for the detection of aflatoxin B1. Biosens Bioelectron 56:340–344
Taherimaslak Z, Amoli-Diva M, Allahyary M, Pourghazi K (2014) Magnetically assisted solid phase extraction using Fe3O4 nanoparticles combined with enhanced spectrofluorimetric detection for aflatoxin M1 determination in milk samples. Anal Chim Acta 2014(842):63–69
Xu X, Liu XJ, Li YB, Ying YBA (2013) Simple and rapid optical biosensor for detection of Aflatoxin B1 based on competitive dispersion of gold Nanorods. Biosens Bioelectron 47:361–367
Song D, Yang R, Fang SY, Liu YP, Long F (2018) A FRET-based dual-color evanescent wave optical fiber aptasensor for simultaneous fluorometric determination of aflatoxin M1 and ochratoxin a. Microchim Acta 185:508–517
Scott AW, Guillermo LA, Kim ES, Carl WBIII, Clare ER, Joyce CB, Igor LM (2018) Detecting biothreat agents: from current diagnostics to developing sensor technologies. ACS Sens 3(10):1894–2024
Sharma R, Akshath US, Bhatt P, Raghavarao KSMS (2019) Fluorescent aptaswitch for chloramphenicol detection – quantification enabled by immobilization of aptamer. Sensors Actuators B Chem 290:110–117
Song X, Knotts M (2008) Time-resolved luminescent lateral flow assay technology. Anal Chim Acta 626:186–192
Corstjens PL, de Dood CJ, van der Ploeg-van Schip JJ, Wiesmeijer KC, Riuttamaki T, van Meijgaarden KE, Spencer JS, Tanke HJ, Ottenhoff TH, Geluk A (2011) Lateral flow assay for simultaneous detection of cellular and Humoral immune responses. Clin Biochem 44:1241–1246
Liu CY, Jia QJ, Yang CH, Qiao RR, Jing LH, Wang LB, Xu CL, Gao MY (2011) Lateral flow Immunochromatographic assay for sensitive pesticide detection by using Fe3O4 nanoparticle aggregates as color reagents. Anal Chem 83:6778–6784
Yang QH, Gong XQ, Song T, Yang JM, Zhu SJ, Li YH, Cui Y, Li YX, Zhang BB, Chang J (2011) Quantum dot-based Immunochromatography test strip for rapid, quantitative, and sensitive detection of alpha fetoprotein. Biosens Bioelectron 30:145–150
Shen J, Zhou YF, Fu F, Xu HY, Lv JF, Xiong YH, Wang A (2015) Immunochromatographic assay for quantitative and sensitive detection of hepatitis B virus surface antigen using highly luminescent quantum dot-beads. Talanta 142:145–149
Hu J, Zhang ZL, Wen CY, Tang M, Wu LL, Liu C, Zhu L, Pang DW (2016) Sensitive and quantitative detection of C- reaction protein based on immunofluorescent nanospheres coupled with lateral flow test strip. Anal Chem 88:6577–6584
Duan H, Chen XL, Xu W, Fu JH, Xiong YH, Wang A (2015) Quantum-DoT submicrobead-based immunochromatographic assay for quantitative and sensitive detection of zearalenone. Talanta 132:126–131
Zhou JY, Nie W, Chen YQ, Yang CJ, Gong L, Zhang C, Chen Q, He LD, Feng XY (2018) Quadruplex gold immunochromatographic assay for four families of antibiotic residues in milk. Food Chem 256:304–310
Cheng N, Song Y, Zeinhom MMA, Chang YC, Sheng L, Li H, Du D, Li L, Zhu MJ, Luo Y, Xu W, Lin Y (2017) Nanozyme-mediated dual immunoassay integrated with smartphone for use in simultaneous detection of pathogens. ACS Appl Mater Interfaces 9:40671–40680
Wang WW, Su XX, Ouyang H, Wang L, Fu ZF (2016) A novel immunochromatographic assay based on a time-resolved chemiluminescence strategy for the multiplexed detection of ractopamine and clenbuterol. Anal Chim Acta 917:79–84
Yen CW, de Puig H, Tam JO, Gómez-Márquez J, Bosch I, Hamad-Schifferli K, Gehrke L (2015) Multicolored silver nanoparticles for multiplexed disease diagnostics: distinguishing dengue, yellow fever, and Ebola viruses. Lab Chip 15:1638–1641
Wang CY, Hou F, Ma YC (2015) Simultaneous quantitative detection of multiple tumor markers with a rapid and sensitive multicolor quantum dots based immunochromatographic test strip. Biosens Bioelectron 68:156–162
Di Nardo F, Alladio E, Baggiani C, Cavalera S, Giovannoli C, Spano G, Anfossi L (2019) Colour-encoded lateral flow immunoassay for the simultaneous detection of aflatoxin B1 and type-B fumonisins in a single test line. Talanta 192:288–294
Sun YN, Yang JF, Yang SZ, Sang QB, Teng M, Li QM, Deng RG, Feng LL, Hu XF, Zhang GP (2018) Development of an immunochromatographic lateral flow strip for the simultaneous detection of aminoglycoside residues in milk. RSC Adv 8:9580–9586
Wang Q, Liu YC, Wang MY, Chen YJ, Jiang WA (2018) Multiplex immunochromatographic test using gold nanoparticles for the rapid and simultaneous detection of four nitrofuran metabolites in fish samples. Anal Bioanal Chem 410:223–233
Liu BB, Gong H, Wang YL, Zhang XS, Li P, Qiu YL, Wang LM, Hua XD, Guo YR, Wang MH, Liu FQ, Zhang CZ (2018) A gold immunochromatographic assay for simultaneous detection of parathion and triazophos in agricultural products. Anal Methods 10:422–428
Taranova NA, Berlina AN, Zherdev AV, Dzantiev BB (2015) ‘Traffic light’ immunochromatographic test based on multicolor quantum dots for the simultaneous detection of several antibiotics in milk. Biosens Bioelectron 63:255–261
Peng J, Wang YW, Liu LQ, Kuang H, Li A, Xu CL (2019) Multiplex lateral flow immunoassay for five antibiotics detection based on gold nanoparticle aggregations. RSC Adv 6:7798–7805
Qie ZW, Liu Q, Yan WL, Gao ZC, Meng W, Xiao R, Wang SQ (2019) Universal and ultrasensitive Immunochromatographic assay by using an antigen as a Bifunctional element and Antialbumin antibody on a test line. Anal Chem 91(15):9530–9537
Tu CY, Dai YY, Zhang Y, Wang WP, Wu L (2020) A simple fluorescent strategy based on triple-helix molecular switch for sensitive detection of chloramphenicol. Spectrochim Acta A Mol Biomol Spectrosc 224:11745–11751
Xie SL, Wen K, Wang SH, Wang JY, Peng T, Mari GM, Li JC, Wang ZH, Yu XZ, Jiang HY (2019) Quantitative and rapid detection of amantadine and chloramphenicol based on various quantum dots with the same excitations. Anal Bioanal Chem 411(10):2131–2140
Wang JY, Wang Q, Zheng YJ, Peng T, Yao K, Xie SL, Zhang XY, Xia X, Li JC, Jiang HY (2017) Development of a quantitative fluorescence-based lateral flow immunoassay for determination of chloramphenicol, thiamphenicol and florfeniclo in milk. Food Agric Immunol 29(1):56–66
Wang C, Peng J, Liu DF, Xing KY, Zhang GG, Huang Z, Cheng S, Zhu FF, Duan ML, Zhang KY, Yuan MF, Lai WH (2018) Lateral flow immunoassay integrated with competitive and sandwich models for the detection of aflatoxin M1 and Escherichia coli O157:H7 in milk. J Dairy Sci 101:8767–8777
Zhang XY, Wen K, Wang ZH, Jiang HY, Beier RC, Shen JZ (2016) An ultra-sensitive monoclonal antibody-based fluorescent microsphere immunochromatographic test strip assay for detecting aflatoxin M1 in milk. Food Control 60:588–595
Wu CH, Hu LM, Xia J, Xu GM, Luo K, Liu DF, Duan H, Cheng S, Xiong YH, Lai WH (2016) Comparison of immunochromatographic assays based on fluorescent microsphere and quantum-dot submicrobead for quantitative detection of aflatoxin M1 in milk. J Dairy Sci 100:2501–2511
Li CH, Luo W, Xu HY, Zhang Q, Xu H, Aguilar ZP, Lai WH, Wei H, Xiong YH (2013) Development of an Immunochromatographic assay for rapid and quantitative detection of Clenbuterol in swine urine. Food Control 34:725–732
Semenova V, Schiffer J, Steward-Clark E, Soroka S, Schmidt D, Brawner M, Lyde F, Thompson R, Brown N, Foster L (2012) Validation and Long term performance characteristics of a quantitative enzyme linked Immunosorbent assay (ELISA) for human anti-PA IgG. J Immunol Methods 376:97–107
Acknowledgments
We thank our colleagues for their contributions to this manuscript. This work was supported by the Major Infectious Diseases, such as the AIDS and Viral Hepatitis Prevention and Control Technology Major Projects (2018ZX10712-001), and the Major Projects (No. AWS16J020).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
An ESM on QB modification with antitarget mAb and the optimal parameters of CAP- and AFM1-spiked samples using different ICA systems (DOCX 21.7 kb)
Rights and permissions
About this article
Cite this article
Qie, Z., Yan, W., Gao, Z. et al. An anti-BSA antibody-based immunochromatographic assay for chloramphenicol and aflatoxin M1 by using carboxy-modified CdSe/ZnS core–shell nanoparticles as label. Microchim Acta 187, 10 (2020). https://doi.org/10.1007/s00604-019-4009-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-019-4009-1