Abstract
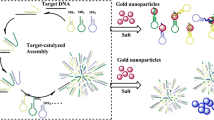
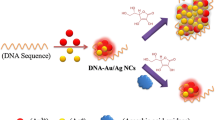
The authors describe a dual (colorimetric and fluorometric) detection scheme for microRNA. The method is based on the use of fluorescent DNA-modified silver nanoclusters (DNA-AgNCs) and gold nanoparticles (AuNPs). The DNA-AgNCs were linked to AuNPs via the interaction of ssDNA-AgNC probes with AuNPs. This led to quenching of the fluorescence of the DNA-AgNCs (best measured at excitation/emission peaks of 370/450 nm) and also prevents salt-induced aggregation of the AuNPs. Upon addition of microRNA, the DNA on the AgNCs hybridizes with microRNA. This led to the formation of a DNA-AgNC/miRNA hetero-duplex and increase of the distance to the AuNPs. Quenching was suppressed and fluorescence was restored. The presence of microRNA also affected salt-induced particle aggregation as reflected by a color change from red to purple that is visible with bare eyes. The visual detection limit for microRNA is 0.6 nM, and the fluorometric detection limit is 0.4 pM.

Schematic of the colorimetric and fluorometric assay for microRNA (miRNA). It is based on the use of fluorescent DNA-modified silver nanoclusters (AgNCs) and gold nanoparticles (AuNPs).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
MicroRNAs (microRNAs) are a class of evolutionally conserved, endogenous small RNAs (approximately 19–23 nucleotides). These non-coding RNAs act as gene expression regulators in animals, plants, viruses and humans [1]. MicroRNAs play an important role in cancer initiation and progression, and can be ordered as oncogenes (oncomicroRNAs) and tumor-suppressor genes [2, 3]. Some microRNAs are oncomicroRNA and are associated with breast cancer. In the blood plasma of patients with breast cancer, tumor-suppressing microRNAs are low, while oncomicroRNAs are abundant [4]. Levels of these microRNAs are commonly measured and used to characterize treatment response [5]. MicroRNA-155 has been identified as a member of the oncomicroRNA, which is over-expressed in many human cancers such as breast cancer. Suppressor of cytokine signaling 1 (SOCS1) is a conserved target gene for microRNA-155 in human breast cancer. Researchers have shown that SOCS1 is negatively regulated by microRNA-155 [2, 6]. Hence, this is an important step to develop efficient and reliable methods for the detection of microRNAs and understanding its functions in different regulatory mechanisms, which influence the progress of microRNA-based therapies in diagnostic assessments at molecular level and new targets in drug discovery.
Conventional microRNA detection strategies include Northern blotting, microarray-based detection, qRT-PCR methods and others. Each of these strategies has its own limitations. For example, Northern blotting analysis which uses a radiolabeled probe with low detection efficiency is very complex; while, RT-PCR and microarrays require specialized reagents and equipment. Due to these limitations, the development of more efficient detection techniques for the species is important and vital [7].
Various strategies have been developed for the detection of mature microRNA detection based on nanotechnology. A number of these methods which have been suggested in recent years are based on fluorescence, electrochemistry, chemiluminescence, surface plasmon resonance imaging (SPRI) and surface-enhanced Raman scattering (SERS) [7,8,9,10]. Each set of strategies has its own strengths and weaknesses and as such, these requirements should be carefully assessed to develop a successful microRNA research strategy. To establish a simple read-out and high sensitive method that does not require costly equipment, researchers have focused on fluorescence-based detection assays and colorimetric approaches [11].
Nanomaterials have encouraged and enabled researchers to develop new detection methods for biomedical applications. Very fine nano-particles (Sizes down to <2 nm), frequently referred to as nano-clusters (NCs), have received increasing attention as promising substitutes for conventional fluorophores [12, 13]. As bare NCs are unstable in aqueous solution, they need to be synthesized using a number of ligands as encapsulation agents such as thiolates, phosphines, and dendrimers. However, the uses of biomolecules, such as proteins, amino acids and DNA, as functional ligands for NC synthesis and bioconjugation, are rather recent developments [14,15,16,17,18,19,20,21]. Nano-clusters have unique properties such as chirality, ferromagnetism, photoluminescence, quantum behavior, and single-molecule optoelectronics [22]. Silver nano-clusters (AgNCs) have shown great promise in bimolecular labeling and biosensing due to their high fluorescence quantum yield, photostability, and excellent biocompatibility. Synthesis of AgNCs by using a DNA oligomer as the template is a time-saving and cost-effective method [23]. In this paper, gold nano-particles (AuNPs) have been used as colorimetric reporters for the detection of microRNA. The AuNPs were used as a colorimetric indicator due to their surface plasmon resonance (SPR), based on which dispersed AuNP suspensions had an optical absorption peak at around 520 nm. In the case of aggregated AuNP suspensions the absorption peak occurs at a longer wavelength, which can be described by the Mie theory [24, 25]. The central idea in this work is that DNA-AgNC/microRNA hetero-duplex and single stranded DNA-AgNCs have different electrostatic affinities to gold nano-particles [26,27,28,29,30]. Consequently we proposed a novel, label free dual-mode “light on” biosensor with colorimetric and fluorometric readouts for measuring the concentration of microRNA. The DNA-AgNC probes readily bind to the surface of AuNPs through strong attractive electrostatic interactions with citrate-coated AuNPs. Thus, the AuNPs were stabilized against salt-induced aggregation and fluorescence quenching of DNA-AgNCs probe happened due to the FRET phenomena. On the other hand, in the presence of microRNA, the DNA-AgNC probes preferred to hybridize with their specific microRNA and form a DNA-AgNC/microRNA hetero-duplex, which got distance from AuNPs and resulting in recovery of QDs fluorescence. In addition, the solution changed from a dispersion to an aggregation.
Experimental
Reagents and materials
Chloroauric acid (HAuCl4•4H2O) and sodium citrate (Na3C6H5O7) were purchased from Sigma-Aldrich. Silver nitrate (AgNO3) and sodium borohydride (NaBH4) were obtained from Merck, and all other commercially available substances were purchased from Aldrich, Merck and Acros, and used without further treatment. The oligonucleotides sequences in Table S1 were synthesized by Shanghai Generay Biotech Co (Shanghai, China). All other reagents were of analytical reagent grade. Ultra-pure water (Milli-Q plus, Millipore Inc., Bedford, MA) was used throughout the reactions.
Cells and cell culture human cell lines used in this study were MCF-7 cells (human breast cancer cell line) and HEK 293 cells (from normal human embryonic kidney cell line).
Apparatus
Absorption spectra were acquired using a Perkine Elmer lambda25 spectrometer and all the fluorescence measurements were carried out on a Perkin-Elmer LS55 luminescence spectrometer. TEM images were obtained with a transmission electron microscope (Zeiss, EM10C, 80 KV, Germany) on a copper grid.
Synthesis of gold nanoparticles
50 ml of an aqueous solution of hydrogen tetrachloroaurate (III) tetrahydrate (1 mM) was heated to boiling while being stirred in a round-bottom flask with a reflux condenser. 10 ml of trisodium citrate (38.8 mM) was then added into the solution rapidly. The solution was boiled again for another 10 min, when the color of the solution changed from yellow to red. The heating was stopped but the stirring continued until the mixture reached room temperature. The AuNPs solution was then stored at 4 °C.
TEM analysis was used to determine the diameter and dispersion state of the synthesized AuNPs. Using the extinction coefficient (2.7 × 108 M −1 cm−1) at 520 nm, the concentration of the AuNPs solution was calculated to be about 4.4 nM according to Beer’s law. The average diameter of the synthesized AuNPs was about 15 nm [31].
Synthesis of DNA-AgNCs
In a typical experiment, AgNO3 solution (1 mM, 6 μl) was added to DNA solution (100 μM, 15 μL) prepared in phosphate buffer (20 mM, 100 μl, pH 7.0) to obtain an Ag+-to-DNA molar ratio of 6:1. The mixture was incubated for 30 min in an ice bath and then reduced by introducing freshly prepared NaBH4 (1 mM, 6 μL) under vigorous shaking. After that, the mixture was reacted for 1 h in an ice bath. The molar ratio of Ag-DNA-NaBH4 in the solution was 6: 1: 6. The reaction mixture was kept at room temperature in the dark for 12 h before use. The AgNCs exhibited a fluorescence emission peak at 450 nm when excited at 370 nm [32].
Total RNA extraction
Cell samples were disrupted and total RNA was extracted from MCF-7 and HEK 293 cell lines using Cell Culture LysisReagent (CCLR). Approximately 2.0 × 106 cells were collected by low-speed centrifugation at 1000 rpm for 10 min. Culture medium was carefully removed and the pellet was washed twice with PBS. The PBS was carefully removed and 600 μl CCLR buffer was added. Cells were gently resuspended in CCLR buffer with a vortex and incubated for 20 min. Subsequently, 0.2 ml chloroform was added and the mixture was subjected to a 20 s vortex violently. Next, the mixture was centrifuged for 20 min at 13,000 rpm at 4 °C by refrigerated centrifugation. The same volume of isopropyl alcohol was added to the upper water which has been taken out and mixed evenly in −20 °C precipitation for the night. Afterward, the mixture was centrifuged for 20 min at 13,000 rpm at 4 °C, and then the supernatant was removed and precipitates were washed by 80% ethanol with DEPC water and centrifuged for 20 min at 13,000 rpm at 4 °C. The ethanol was volatilized (Letting the tube dry) and the purified RNA dissolved in an appropriate volume of DEPC water.
Assay procedure
Unmodified AuNPs (4.4 nM, 30 μl) were added to DNA-AgNCs solutions (100 μl) and incubated for 10 min at room temperature. After incubation, the fluorescent DNA-AgNC probes were adsorbed onto AuNPs based on the electrostatic interaction of single strand DNA-AgNC probe with AuNPs leading to the quenching of the DNA-AgNCs fluorescence. Then, this reaction solution (DNA-AgNCs@AuNPs) was added to the sample solutions which contained different concentrations of microRNA (15 μl) and were incubated for 1 h at 37 °C. Finally, salt (NaCl, 1 mM, 5 μl) was added to reach the aggregation of AuNPs and mixtures were incubated for 10 min. The fluorescent intensity and absorbance of the solution were measured by luminescence and UV-vis spectrometers. The principle of detection system is clearly illustrated in Scheme 1b.
Results and discussion
Characterizations of AuNPs and DNA-AgNCs
UV–Vis absorption spectroscopy and transmission electron microscopy (TEM) were used for characterization of the gold nano-particles. The morphology and size of AuNPs were determined through transmission electron microscopy (TEM) imaging (Fig. S1A), which show the nano-particles to be uniform spheres with an average diameter of about 15 nm. In addition, the dispersed AuNPs exhibit a red color with a strong absorption peak at 525 nm (Fig. S1B).
The DNA-AgNCs were primarily synthesized by sequestering AgNO3 with the DNA and then reducing the silver ions to Ag0 nano-clusters with NaBH4. The interaction of the Ag-cytosine bases allows for the formation of DNA-AgNCs with excellent optical properties [32]. To approve the production of AgNCs, the DNA-AgNCs were characterized through TEM images, fluorescence spectrometry and UV-Visible absorption spectra (Fig. S2A). The size of the AgNCs was characterized by TEM and as the results are shown in Fig. S2 B, the average diameter of AgNCs is about 2.0 nm.
General mechanism of dual-readout assay
In this work, AuNPs played two roles (as colorimetric reporters and quenchers). Firstly, the AuNPs behaved as colorimetric reporters enabling the colorimetric assay. Secondly, the AuNPs served as a strong quencher, which possess an extraordinary quenching efficiency over a broad range of wavelengths [24, 33, 34]. There are many reasons why AuNPs are excellent fluorescence quenchers over longer distances. For example, (1) since the spherical AuNPs have no defined dipole moments, as dye molecules do, the fluorescence resonance energy transfer (FRET) to the AuNP occurs for any orientation of the donor relative to the surface of the AuNPs. (2) Due to the size of the AuNPs, the FRET efficiency from a molecular donor to the AuNP acceptor declines with distance as compared to the case of a molecular acceptor. (3) The large absorption cross-section of AuNPs, especially near their plasmon resonance, enhances their performance as energy acceptors. (4) In contrast to other molecular acceptors, AuNPs may also quench fluorescence due to radiative rate suppression [35].
Accordingly, we developed a biosensor for the detection of microRNA based on FRET system, using DNA-AgNCs and AuNPs as the donor and acceptor, respectively. In this system, we used the AgNCs-based FRET approach for the detection of specific microRNA (microRNA-155) sequence. A specific single stranded DNA sequence was chosen as a template for the formation of DNA-AgNCs. It was concluded that the ssDNA-AgNCs sample caused a close proximity between DNA-AgNCs as the energy donor and AuNPs as the energy acceptor. This met the requirement for FRET, therefore the FRET signal was generated. Contrastingly there was no FRET signal in presence of microRNA-155 target since the hybridization led to dispersion of DNA-AgNCs and AuNPs. The fluorescence and absorption spectra of the DNA-AgNCs and AuNPs in buffer are shown in Fig. 1. As it is shown, the fluorescence emission of the DNA-AgNCs overlaps the absorption of spectrum of AuNPs very well. The sufficient spectral overlap between the donor and acceptor may guarantee an efficient energy transfer from DNA-AgNCs to AuNPs.
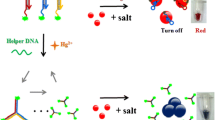
Scheme 2 depicts the principle mechanism for the proposed sensing system. Li and Rothberg [26] previously showed that single-stranded DNA (ssDNA) could readily adsorb on the particle surface to act as a stabilizer. Because the ssDNA is sufficiently flexible to partially uncoil its bases, they can be exposed to the gold nanoparticles. Under these conditions, the negative charge on the backbone is sufficiently distant so that attractive van der Waals forces between the bases and the gold nanoparticle are sufficient to cause ssDNA to stick to the gold. Therefore, the fluorescence signal of DNA-AgNCs was quenched after incubating with AuNPs (Fig. 2a). Yet upon the addition of microRNA, the DNA-AgNC/microRNA hetero-duplex got distance from the AuNPs, resulting in the recovery of NCs fluorescence (Fig. 2b), because proximity of negatively charged AuNPs induced a repulsive effect.
Fluorescence emission of DNA-AgNCs/AuNPs, (a) Quenching effect of AuNPs and (b) recovery of fluorescence signal after addition of microRNA. The absorption spectra of DNA-AgNCs/AuNPs; (a´) DNA-AgNCs/AuNPs in the presence of microRNAs, (b´) after the addition of the salt; and TEM image of AuNPs before (c) and after (b) addition of salt
As another function, the ssDNA stabilizes AuNPs against salt induced aggregation, whereas double-strand DNA does not. In the absence of microRNA, the dispersed AuNPs exhibited a red color with a strong absorption peak at 525 nm (Fig. 2a´). On the other hand, in the presence of microRNA, the AuNPs were aggregated and manifested by the dramatic decrease in the absorbance at 525 nm and the concomitant red-shift of the SPR absorption to 705 nm, which appeared as a purple solution (Fig. 2b´). To further confirm that the aggregation occurred, TEM imaging was used to test the feasibility of the colorimetric platform. The morphology and size of AuNPs were investigated by TEM imaging of gold nano-particles both before (Fig. 2c) and after (Fig. 2d) addition of microRNAs.
Analytical performance of DNA assay
To evaluate the DNA-AgNC-AuNP as a dual probe for assay of microRNA, the absorbance and fluorescence changes were measured in the presence of concentration ranges of 0.5 nM- 1 μM and 1pM – 5 μM, by UV-visible and fluorescence techniques, respectively. As shown in Fig. 3, the characteristic absorption peak of AuNPs at 525 nm gradually decreases and new peaks at 705 nm progressively strengthen with increasing the concentration of microRNA-155, with a color change from red to purple solution. Based on the results, good linear relationships were found between the absorption ratios (A705/A525) and the concentration of microRNA-155 in the range 1 nM-5 μM, with a regression equation of A = −0.203C + 2.42 (R2 = 0.961). The detection limit for microRNA-155 was determined to be 0.6 nM (S/N = 3).
a Absorption spectra of DNA-AgNCs/AuNPs, (b) the logarithmic plot for A705/A525 versus target microRNA155 concentration (c) and colorimetric changes after incubation with different concentrations of microRNAs (0.5 nM, 1 nM, 5 nM, 50 nM, 0.5 μM, 1 μM), Incubation time 10 min in phosphate buffer (The standard deviations from three replicate measurements is shown as the error bars)
As shown in Fig. 4 the emission peak at 450 nm gradually increases with increasing the concentration of microRNA-155 in the range 1 pM–5 μM. The results prove that in the presence of microRNA, the DNA-AgNC probes were hybridized to their specific target microRNA and do not bind onto the surface of AuNPs which led to a stronger fluorescence signal. The higher the microRNA concentration, the higher the observed fluorescence emission intensity. As a result, a linear relationship was obtained from the plot of fluorescence intensity as function of the concentration of microRNA-155 over the range of 1 pM −5 μM, with a regression equation of I = −4.50 C + 82.2 and C = (−log microRNA155) (R2 = 0.994). The detection limit for microRNA155 was estimated to be 0.4 pM according to the equation of LOD = 3SD/slope, which is lower than those reported in the case of previous nano-clusters [Table 1] [36,37,38,39,40,41,42,43].
Fluorescence recovery at different microRNA-155 concentration (1 pM, 10 pM, 50 pM, 100 pM, 0.5 nM, 1 nM, 5 nM, 50 nM, 0.5 μM, 1 μM, 5 μM,) λex of DNA-AgNCs = 370 nm; Incubation time 10 min, Inset: calibration curves for detection of microRNA (The standard deviations from three replicate measurements is shown as the error bars)
According to Table 1, the sensitivity of the analytical assay is also comparable to other methods based on nano-clusters, except for electrochemical methods and TAPNR amplification-based methods, which require complicated experimental steps such as labeling, immobilization, washing and enzyme amplification. However, compared with current sensors, the results in this work clearly indicate an excellent sensitivity for detection of microRNA, on the biosensing probe.
Selectivity
Further the selectivity of the developed system was evaluated by examining its fluorescence responses to three artificially synthesized microRNAs of microRNA-21, let-7a, microRNA-155 and a mixture of all those microRNAs at the same concentration of 5 nM alongside microRNA-155 specific sequence. As shown in Fig. 5, the fluorescence signal of the probe toward microRNA-155 is much stronger than that of other microRNAs, which are close to that of the blank sample. These results confirm the high selectivity of the probe for microRNA-155.
Analysis of microRNA-155 in total RNA sample
Finally, to test the feasibility of the method to determine miRNA in real biological samples, we investigated expression level of miR-155 in total RNA sample extracted from the MCF-7 as a human breast cancer cell lines and HEK 293 as a normal human embryonic kidney cell line. The results indicate that MCFcell lines had higher miR-155 expression level than HEK 293 cell lines (Fig. 6), which is in good accordance with quantitative real-time PCR (qRT-PCR; as a gold standard method) results. The relative intensity was evaluated and normalized to the expression of U6 small nuclear RNA (U6 snRNA). The relative expression was calculated by the equation: Fold change = 2−ΔΔCt.
Conclusion
In summary, this study describes the development of a dual-mode sensing platform that not only simply makes the visualization of microRNA with bare eyes possible, but it also provides a label-freeand rapid FRET sensing system. Additionally, the ability of the colorimetric assay to quantify miRNA levels in cell culture lysates from breast cancer cells was acceptable but not exactly the same as that of qRT-PCR. Although, this platform is not ultrasensitive, it′s the first report for microRNA detection based on energy transfer between NCs and AuNPs, with two different signals for the detection and quantification of miRNAs in biological samples without the use of expensive and complex equipment.
References
Dong H, Lei J, Ding L, Wen Y, Ju H, Zhang X (2013) MicroRNA: function, detection, and bioanalysis. Chem Rev 113:6207–6233
Gurses HE, Hatipoğlu OF, Gunduz M, Gunduz E. (2015) MicroRNAs as therapeutic targets in human breast Cancer. Intechopen, Chapter 5
Van Schooneveld E, Wildiers H, Vergote I, Vermeulen PB, Dirix L, Van Laere SJ (2015) Dysregulation of microRNAs in breast cancer and their potential role as prognostic and predictive biomarkers in patient management. Breast Cancer Res 17:21
Zhu J, Zheng Z, Wang J, Sun J, Wang P, Cheng X, Fu L, Zhang L, Wang Z, Li Z (2014) Different microRNA expression profiles between human breast cancer tumors and serum. Front Genet 5:149
Zhao SY, Wu Q, Gao F, Zhang CB, Yang XW (2012) Increased expression of MicroRNA-155 in the serum of women with early-stage breast Cancer. Lab Med 43(5)
Jiang S, Zhang HW, Lu MH, He XH, Li Y, Gu H, Liu MF, Wang ED (2010) MicroRNA-155 functions as an OncomicroRNA in breast cancer by targeting the suppressor of cytokine signaling 1 gene. Cancer Res 70:3119–3127
Tian T, Wang J, Zhou X (2015) A review: microRNA detection methods. Org Biomol Chem 13:2226–2238
Sang Y, Xu Y, Xu L, Cheng W, Li X, Wu J, Ding S (2017) Colorimetric and visual determination of microRNA via cycling signal amplification using T7 exonuclease. Microchim Acta 184:2465–2247
Xiong H, Zheng X (2017) Electrochemiluminescence based determination of micro-RNA using target-guided assembly of gold nanoparticles on an electrode modified with Nafion, carbon nanotubes and polyvinylpyrrolidone. Microchim Acta 184:1781–1789
Zeng K, Li H, Peng Y (2017) Gold nanoparticle enhanced surface plasmon resonance imaging of microRNA-155 using a functional nucleic acid-based amplification machine. Microchim Acta 184:2637–2644
Catuogno S, Esposito CL, Quintavalle C, Cerchia L, Condorelli G, de Franciscis V (2011) Recent advance in biosensors for microRNAs detection in Cancer. Cancers 3:1877–1898
Obliosca JM, Liu C, Yeh HC (2013) Fluorescent silver nano-clusters as DNA probes. Nanoscale 5:8443–8461
Biju V (2014) Chemical modifications and bioconjugate reactions of nanomaterials for sensing, imaging, drug delivery and therapy. Chem Soc Rev 43:744–764
Nirmal G, Zheng K, Xie J (2014) Bio-NCs – the marriage of ultrasmall metal nano-clusters with biomolecules. Nanoscale 6:13328–13347
Borghei YS, Hosseini M, Khoobi M, Ganjali MR (2016) Novel Fluorometric assay for detection of cysteine as a reducing agent and template in formation of copper nano-clusters. J Fluoresc 27:529–536
Borghei YS, Hosseini M, Ganjali MR, Hosseinkhani S (2018) A novel BRCA1 gene deletion detection in human breast carcinoma MCF-7 cells through FRET between quantum dots and silver nanoclusters. J Pharm Biomed Anal 152:81–88
Borghei YS, Hosseini M, Ganjali MR (2017) Detection of large deletion in human BRCA1 gene in human breast carcinoma MCF-7 cells by using DNA-silver Nanoclusters. Methods Appl Fluoresc 6:015001
Borghei YS, Hosseini M, Ganjali MR (2017) Fluorometric determination of microRNA via FRET between silver nanoclusters and CdTe quantum dots. Microchim Acta 184:4713–4721
Borghei YS, Hosseini M, Ganjali MR, Hosseinkhani S (2017) Label-free fluorescent detection of microRNA-155 based on synthesis of hairpin DNA-templated copper nanoclusters by etching (top-down approach). Sensors Actuators B Chem 248:133–139
Hosseini M, Ahmadi E, Borghei YS, Ganjali MR (2017) New fluorescence turn-on nanobiosensor for the detection of micro-RNA-21 based on a DNA–gold nanocluster. Methods Appl Fluoresc 5:015005 16
Hosseini M, Mohammadi S, Borghei YS, Ganjali MR (2017) Detection of p53 gene mutation (Single-Base mismatch) using a fluorescent silver Nano-clusters. J Fluoresc 27:1443–1448. https://doi.org/10.1007/s10895-017-2083-5.17
Lin CAJ, Lee CH, Hsieh JT, Wang HH, Li JK, Shen JL, Chan WH, Yeh HI, Chang WH (2009) Synthesis of fluorescent metallic Nano-clusters toward biomedical application: recent progress and present challenges. J Med Biolog Eng 29:276–283.18
Jeong J, Yoon B, Kwon YW, Choi D, Seob Jeong K (2017) Singly and doubly occupied higher quantum state in Nanocrystals. Nano Lett 17:1187–1193
Zhao D, Chen C, Lu L, Yang F, Yang X (2015) A dual-mode colorimetric and fluorometric “light on” sensor for thiocyanate based on fluorescent carbon dots and unmodified gold nano-particles. Analyst 140:8157–8164
Zhao W, Chiuman W (2007) Simple and rapid colorimetric biosensors based on DNA Aptamer and noncrosslinking gold nanoparticle aggregation. Chembiochem 8:727–731
Li H, Rothberg L (2004) Colorimetric detection of DNA sequences based on electrostatic interactions with unmodified gold nano-particles. PNAS 101:14036–14039
Liu Y, Yang Y, Zhao X, Liu Z, Li Y (2013) Responsive disassembly of the gold nanoparticle aggregates triggered by the competitive adsorption for lighting up the colorimetric sensing. Anal Methods 5:3242–3247
Li H, Rothberg LJ (2004) Label-free colorimetric detection of specific sequences in genomic DNA amplified by the polymerase chain reaction. J Am Chem Soc 126:10958–10961
Girigoswami A, Li T, Jun C, Young MH, Gyu Park H (2009) Gold nanoparticle-based label-free detection of BRCA1 mutations utilizing DNA ligation on DNA microarray. Nanosci Nanotechnol 9:1019–1024
Lee H, Kang T, Yoon KA, Lee SY, Joo SW, Lee K (2010) Colorimetric detection of mutations in epidermal growth factor receptor using gold nanoparticle aggregation. Biosens Bioelectron 25:1669–1674
Borghei YS, Hosseini M, Dadmehr M, Hosseinkhani S, Ganjali MR, Sheikhnejad R (2016) Visual detection of cancer cells by colorimetric aptasensor based on aggregation of gold nano-particles induced by DNA hybridization. Anal Chim Acta 904:92–97
Dadmehr M, Hosseini M, Hosseinkhani S, Ganjali MR, Sheikhnejad R (2015) Label free colorimetric and fluorimetric direct detection of methylated DNA based on silver nano-clusters for Cancer early diagnosis. Biosens Bioelectron 73:108–113
Ghosh D, Chattopadhyay N (2013) Gold Nano-particles: acceptors for efficient energy transfer from the Photoexcited Fluorophores. Optics Photonics J 3:18–26
He H, Dai J, Duan Z, Zheng B, Meng Y, Guo Y, Xiao D (2016) Unusual sequence length-dependent gold nano-particles aggregation of the ssDNA sticky end and its application for enzyme-free and signal amplified colorimetric DNA detection. Sci Rep 6:30878
Mayilo S, Kloster MA, Wunderlich M, Lutich A, Klar TA, Nichtl A, Rzinger KK, Stefani FD, Feldmann J (2009) Long-range fluorescence quenching by gold Nano-particles in a sandwich immunoassay for cardiac troponin T. Nano Lett 9:4558–4563
Borghei YS, Hosseini M, Ganjali MR (2017) Fluorescence based turn-on strategy for determination of microRNA-155 using DNA-templated copper nanoclusters. Microchim Acta 184:2671–2677
Li RD, Wang Q, Yin BC, Ye BC (2016) Enzyme-free detection of sequence-specific microRNAs based on nanoparticle-assisted signal amplification strategy. Biosens Bioelectron 77:995–1000
Shah P, Choi SW, Kim HJ, Cho SK, Bhang YJ, Ryu MY, Thulstrup PW, Bjerrum MJ, Yang SW (2016) Locking-to-unlocking system is an efficient strategy to design DNA/silver nano-clusters (AgNCs) probe for human microRNAs. Nucleic Acids Res 44:e57
Wang Z, Si L, Bao J, Dai Z (2015) A reusable microRNA sensor based on the electrocatalytic property of hetero-duplex templated copper nano-clusters. Chem Commun 51:6305–6307
Yang C, Shi K, Dou B, Xiang Y, Chai Y, Yuan R (2015) In situ DNA-Templated synthesis of silver Nano-clusters for ultrasensitive and label-free electrochemical detection of MicroRNA. ACS Appl Mater Interfaces 7:1188–1193
Shah P, Choi SW, Kim HJ, Cho SK, Thulstrup PW, Bjerrum MJ, Bhang YJ, Ahn JC, Yang SW (2015) DNA/RNA chimera templates improve the emission intensity and target the accessibility of silver nanocluster-based sensors for human microRNA detection. Analyst 140:3422–3430
Liu L, Jiang S, Wang L, Zhang Z, Xie G (2015) Direct detection of microRNA-126 at a femtomolar level using a glassy carbon electrode modified with chitosan, graphene sheets, and a poly(amidoamine) dendrimer composite with gold and silver nano-clusters. Microchim Acta 182:77–84
Cheng Y, Lei J, Chen Y, Ju H (2014) Highly selective detection of microRNA based on distance-dependent electrochemiluminescence resonance energy transfer between CdTe nanocrystals and Au nano-clusters. Biosens Bioelectron 51:431–436
Acknowledgments
The authors thank the research Council of University of Tehran (Grant 28645/01/02) and NBIC Research Center for financial support of this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no competing interests.
Electronic supplementary material
ESM 1
(DOC 474 kb)
Rights and permissions
About this article
Cite this article
Borghei, YS., Hosseini, M., Ganjali, M.R. et al. Colorimetric and energy transfer based fluorometric turn-on method for determination of microRNA using silver nanoclusters and gold nanoparticles. Microchim Acta 185, 286 (2018). https://doi.org/10.1007/s00604-018-2825-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-018-2825-3