Abstract
A sensitive and specific bioassay based on cyclic enzymatic amplification was developed for determination of microRNA (miRNA) by taking advantage of the exodeoxyribonuclease activity of T7 exonuclease (T7 Exo). In the presence of miRNA, DNA/RNA duplexes are formed by hybridization of miRNA and capture probes (Cp). Then, the Cp is digested to release miRNA into the next amplification cycle assisted by T7 Exo. This leads to the digestion of numerous Cp molecules. The broken Cp does no longer hybridize with hairpin probes (Hp) to unveil G-quadruplex DNAzyme (GDNAs). However, in the absence of miRNA, the Hp hybridizes with Cp to unveil GDNAs. The generated GDNAs form assemblies with hemin to form the G-quadruplex/hemin DNAzyme complex which is capable of catalyzing the oxidation of the substrate ABTS by H2O2. Upon cyclic enzymatic amplification, the output signal is reduced accordingly, this resulting in a “signal-off” signal best acquired at a wavelength of 418 nm. The lower detection limit is 0.69 pM (at an S/N ratio of 3). The assay involves efficient signal amplification, is homogeneous and isothermal, and enables visual detection. It provides a simple, rapid and sensitive platform for use in clinical diagnostics.

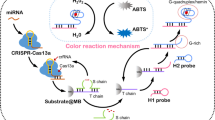
Schematic of a method for colorimetric and visual detection of miRNA using T7 exonuclease-based cycling signal amplification.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
MicroRNAs (miRNAs) are a class of small (18–23 nucleotides), endogenous, single-stranded RNA molecules that regulate the expression of genes in many cellular processes [1, 2]. The unnatural expression of miRNA may lead to serious diseases, such as cancers, nervous diseases, and diabetes [3–5], acting as significant signaling molecules in disease development and progression [6, 7]. Thus, the quantitative analysis of miRNA is urgently needed as novel biomarkers [8]. However, given the unique characteristics of miRNAs, including short-length, low abundance, and sequence homology among family members, sensitive detection of miRNAs has remained a great challenge [9].
To date, a variety of miRNA detection methods have been reported, including traditional northern blotting, microarray and real-time PCR. Among these methods, northern blotting are taken as standard method for the detection of MicroRNA, but the disadvantages of low sensitivity and sophisticated procedures may limit its applications [10]. In spite of the excellent sensitivity and widespread applications of PCR in routine miRNA detection, real-time PCR is hindered by strict control of temperature cycling for amplification detection and false positive result [11]. Microarray is considered as an ideal platform for profiling miRNAs on a global scale because of its exceptional multiplexing capacity microarray technology, but this method is restricted by large sample size, fluorescent labeling and time-consuming hybridization time [12]. In addition, these methods are not applicable for point of care testing (POCT), which is highly required for treating clinical emergent events in rapid diagnosis.
To explore alternative methods for detection of miRNA has attracted more and more attention and thus different biosensors have been fabricated, such as surface plasmon resonance(SPR) [13] and electrochemical [14], fluorescent [15], and colorimetric biosensors [16]. However, many of these methods still require enhanced sensitivity, some involve laborious experiment operations, and other methods may rely on advanced instruments and fluorescent labeling. Of the above mentioned methods, homogenous colorimetric biosensors exhibits particularly attractive merits of low cost, ease of use and visual detection, thus holding great promise as a potential point-of-care diagnostic tool for miRNA detection [17–19]. In addition, to further overcome the shortcoming of low sensitivity, an emerging research field of isothermal nucleic acid amplification has been found growing attention due to simplicity and high amplification efficiency, such as rolling circle amplification (RCA) [20–22], signal mediated amplification of RNA technology (SMART) [23], exponential isothermal amplification reaction (EXPAR) [24, 25] and cyclic enzymatic amplification method (CEAM) [26].
Of the above amplification techniques, most of these methods suffer from complicated design of primers or time-consuming operation. CEAM, a recently developed method, possesses the superior features of operational simplicity, high amplification efficiency and low cost in nucleic acid detection. The cyclic enzymatic amplification method based on nucleases, in which one target leads to many cycles of target dependent nuclease cleavage of reporter probes attains output signal amplification [26, 27]. Moreover, T7 Exo, as a sequence independent nuclease, can catalyze the stepwise removal of 5′ mononucleotides from the 5′-bluntor recessed terminus of RNA/DNA hybrids or double stranded DNA [28–31]. Compared with previous nicking endonuclease assisted cyclic amplification, no specific recognition site was required for T7 Exo, facilitating the DNA sequence design. Importantly, in contrast to another widely used Exo III-assisted cyclic amplification, T7 Exo can provide a low background signal [32, 33]. Therefore, CEAM with the assistance of T7 Exo not only enhances detection sensitivity, flexibility and simplicity but also reduces the background signal.
Thus, a variety of biosensing strategies based on T7 Exo have been developed, including fluorescent, electrochemical and electrochemiluminescence biosensing strategy [27, 33, 34]. However, as the best of our knowledge, a simple homogeneous colorimetric biosensor, allowing to be visualized with the bare eye, has not been developed. Therefore, it is of great significance to develop a colorimetric biosensing strategy based on T7 Exo assisted amplification for rapid and visual detection of miRNA.
Based on these approving properties, we fabricated a simple “signal-off” colorimetric biosensor for ultrasensitive and selective detection of miRNA-21using signal amplification by T7 Exo assisted target recycling in this work. With the cyclic digestion of T7 Exo, the amount of capture probe (Cp) that hybridized with hairpin probe (Hp) to unveil GDNAs and form G-quadruplex/hemin DNAzyme for signal output in the presence of hemin was accordingly reduced, resulting in a “signal-off” colorimetric biosensing strategy. The biosensing strategy exhibits the merits of homogeneous and isothermal assay system, and excellent analytical performance towards miRNA-21, offering a high potential detection strategy for miRNA-21 determination in clinical molecular diagnostics.
Experimental
Reagents and materials
T7 Exo and 10 × NEbuffer 4 (500 mM Potassium acetate, 200 mM Tris-acetate buffer, 100 mM magnesium acetate, 10 mM dithiothreitol, pH 7.9 at 25°C) were obtained from New England Biolabs (USA). Hemin and 2, 2′-amino-di (2-ethyl-benzothiazoline sulphonic acid-6) ammonium salt (ABTS2−) were obtained from Sigma-Aldrich (St Louis, MO, USA, http://www.sigmaaldrich.com/). MiRNA-21 was purchased from TaKaRa Biotech. Inc. (Dalian, China, http://www.takara.com.cn/). All oligonucleotides used in the experiment were synthesized by Sangon Biotechnology Co. Ltd. (Shanghai, China, http://www.sangon.com/). The corresponding nucleotide sequences of the oligonucleotides were shown in Table 1. All oligonucleotides were dissolved in tris-ethylene diaminetetraacetic acid (TE) buffer (pH 8.0, 10 mM Tris-HCl, 1 mM EDTA) and stored at −20°C, which were diluted in appropriate buffer prior to use.
The stock solution of 1.0 μg⋅μL−1 total RNA extracted from breast adenocarcinoma (MCF-7) cells was purchased from Ambion (California, USA). All solutions and deionized water used were treated with diethyprocarbonated (DEPC) to protect from RNase degradation. Ultrapure water from a Millipore water purification system (≥18 MΩ⋅cm, Milli-Q, Millipore) was used in all experiments. The stock solution of hemin (5 mM) was diluted with 25 mM 4-(2-hydroxyethyl)-1-piperazineëthanesulfonic acid (HEPES) (pH 8.0, 20 mM KCl, 200 mM NaCl, 0.05% Triton X-100 and 1% dimethyl sulfoxide (DMSO)).
Apparatus
A UV-visible spectrophotometer (UV-2550, Shimadzu, Kyoto, Japan, http://www.shimadzu.com/) was used to detect the UV-vis absorption. The concentrations of DNA suspensions were measured by ultraviolet spectrophotometry using a NanoDrop 1000 spectrophotometer (Thermo Scientific, Wilmington, DE, USA, http://www.nanodrop.com/). And the gel electrophoresis was performed on the DYY-6C electrophoresis analyzer (Liuyi Instrument Company, China) and imaged on a Bio-rad ChemDoc XRS (Bio-Rad Laboratories, USA).
Analytical protocol
First, 5 μL of different concentrations of miRNA-21 (10 pM to 100 nM) and 5 μL of 400 nM capture probe (Cp) were added into 25 μL of TNaK buffer (20 mM Tris, 140 mM NaCl, 20 mM KCl, pH 7.5), and then the reaction mixture was incubated at 37°C for 45 min. Next, 10 μL of 10 × NEBuffer, 55 μL of DEPC water and 0.05 U of T7 Exo were added into the mixture and incubated at 37°C for 45 min. After that, 5 μL of 500 nM hairpin probe (Hp) was added into the resulting mixture and incubated at 37°C for 30 min. At last, 110 μL of HEPES buffer and 3 μL of 200 μM hemin were added into the resulting solution and incubated for 30 min at room temperature.
To measure the UV-vis absorption, 220 μL of 4 mM ABTS2− and 0.6 μL of 32 mM H2O2 as the DNAzyme substrate were added into the reaction mixture, followed by incubating at room temperature for 5 min. The absorbance of the mixture was then measured using UV-vis spectrophotometer in the wavelength range from 500 to 400 nm. The reaction rate was monitored at 418 nm.
Gel electrophoresis analysis
The product solution was subjected to electrophoresis analysis on the 3% agarose gel electrophoresis (AGE). Electrophoresis was carried out in 1 × TBE buffer (89 mM Tris-boric acid, 2 mM EDTA, pH 8.3) at a 120 V constant voltage for 25 min. After that, the gels were visualized via gel image system.
Results and discussion
Principle of the biosensing strategy
The principle of the biosensing strategy for miRNA-21 detection is depicted in Scheme 1. The assay system consists of the signal transducer of hairpin DNA, T7-Exo assisted amplification and DNAzyme signal readout steps. The signal transducer of Hp DNA is composed of three domains, including Cp recognition domain (in green), G-rich DNA domain (in blue) and a sequence domain (in black) served as blocker of G-rich DNA. In the absence of miRNA, Cp hybridizes with the Hp recognition domain of hairpin DNA and free G-rich DNA, which assembled with hemin and fold into an active conformation. The active G-quadruplex/hemin complex exhibits superior peroxidase-like activity and thereby catalyzes the conversion of the colorless substrate ABTS2− to the green product ABTS− in the presence of H2O2, generating a colorimetric signal change. In the presence of miRNA, DNA/RNA duplexes are formed by the hybridization of miRNA and Cp. With the assistance of T7 Exo, Cp is digested and miRNA was released into additional cycle, resulting in the digestion of a large amount of Cp. The broken Cp does not hybridize with Hp to unveil GDNAs from Hp. Thus, with the cyclic enzymatic amplification, the output signal reduces accordingly, achieving a simple “signal-off” colorimetric detection. In addition, the homogenous and isothermal assay system together with excellent amplification efficiency enables the designed colorimetric biosensing strategy to be simplicity and rapidity for the detection of miRNA-21. Thus, this simple and sensitive detection method provides a promising tool for miRNA-21 determination in clinical molecular diagnostics.
Verification of the signal amplification strategy
To verify the feasibility of the signal amplification strategy for the detection of miRNA-21, colorimetric detection was performed with or without the signal amplification strategy. As shown in Fig. 1A, only addition of Hp, a little UV-vis absorption was obtained (line e) because G-quadruplex was blocked in the hairpin structure. However, with addition of equal amount of Cp to the reaction solution containing Hp, Hp was fully opened by Cp, resulting in an increased signal and maximum value (line a). Furthermore, only T7 Exo was added into the assay system containing Hp and Cp, the observed signal value (line b) had no obvious change in comparison to the maximum value (line a). This result is attributed to the lack of T7 Exo-assisted cyclic digestion of Cp in the absence of miRNA-21. Subsequently, when miRNA-21 was added into the system containing Hp and Cp, the signal reduced to a certain extent, due to reduction of Cp which combines with miRNA-21 and opens the DNA/RNA double-stranded (line c). However, when T7 Exo was added into the assay system containing Cp, Hp and miRNA-21, the signal value was obviously reduced, indicating Cp in DNA/RNA double-stranded is stepwise digested by T7 Exo (line d), thus achieving a “signal off” colorimetric biosensing system.
(A) UV-vis absorption spectra of the signal amplification strategy: (a) Capture probe and hairpin probe; (b) Capture probe, hairpin probe and T7 exonuclease; (c) Capture probe, miRNA-21 and hairpin probe; (d) Capture probe, miRNA-21, T7 exonuclease and hairpin probe; (e) Only hairpin probe. (B) Gel electrophoresis images of the amplification products. Lanes 1, 500 bp DNA marker; lane 2, capture probe and hairpin probe; lane 3, capture probe, hairpin probe and T7 exonuclease; lane 4, capture probe, miRNA-21 and hairpin probe; lane 5, capture probe, miRNA-21, T7 exonuclease and hairpin probe
To further verify the feasibility of the biosensing strategy, the cycling amplification assay was demonstrated by testing the products of each reaction with agarose electrophoresis (Fig. 1B). Lane 2 represented the mixture of hairpin probe and capture probe, which showed a band with slower migration due to the assembly of capture probe and hairpin probe. With addition of T7 Exo to the mixture in lane 2, the band in lane 3 exhibited no change. Because capture probe is not digested by T7 Exo in the absence of target miRNA-21, and then capture probe hybridize with hairpin probe, resulting in a same band as that in lane 2. Furthermore, the addition of target miRNA to the mixture in lane 2 led to a lighter band in lane 4 on the reason that the capture probe is partly hybridized by target miRNA. However, when the mixture of lane 4 was added by T7 Exo, the assembled band of capture probe and hairpin probe disappeared. This is ascribed to the cycling digestion of capture with the assistance of T7 Exo in the presence of miRNA (lane 5). Thus, the cycling amplification proceeds as design.
Optimization of probe preparation and detection conditions
To obtain the highest biosensing performance, several important conditions were optimized. On account of the significance of Cp for signal amplification, the Cp concentration was firstly optimized. As shown in Fig. S1A, with the increasing concentrations of Cp, the response value rose and reached a plateau at 400 nM, which was designated as the appropriate concentration. HP was employed as signal transducer of the colorimetric method, and therefore its optimum concentration was optimized and obtained at 500 nM (Fig. S1B). T7 Exo participated in cyclical amplification process, and the effect of variable concentrations of T7 Exo on amplification reaction was investigated in Fig. S1C. At low concentrations, the signal showed the maximum response due to the saturated binding, and the UV-vis absorption was reduced with the increase of T7 Exo concentration accordingly with a stable trend at 0.05 U, which was used in the following experiment. The signal amplification catalyzed by DNAzyme was influenced by H2O2 volume. Hence, its significant volume was also investigated and selected at 0.6 μL (32 mM) (Fig. S1D).
Analytical performance
Under the optimal experimental conditions, the performance of the biosensing strategy was investigated by detecting different concentrations of target miRNA. As presented in Fig. 2, UV-vis absorption increased with the reduced concentrations of target miRNA, and there was a good linear relationship between the colorimetric responses and the logarithm of miRNA-21 concentration in the range of 10 pM to 100 nM. The resulting linear equation is Y = 0.198–0.052 lgX (X is the concentration of miRNA-21 and Y is the value of absorbance) with a correlation coefficient of 0.9995. Additionally, the detection limit was estimated to be 0.69 pM (S/N = 3). Compared with the reported colorimetric methods listed in Table 2, the biosensing strategy exhibits great improvement in the analytical performance without any special nanomaterials. These results come from the high efficient amplification of T7 Exo and the cascade DNAzyme catalysis.
Specificity and reproducibility of the biosensing strategy
To investigate the specificity of the biosensing strategy, we compared the colorimetric response to miR-21 with responses to other three different oligonucleotides, including single-base mismatched sequences at different mutation positions, double-base mismatched sequence and non-complementary oligonucleotides at the same concentration of 100 nM. As shown in Fig. 3, the colorimetric response to full-complementary sequence was much lower than those of single-base mismatched sequences, double-base mismatched sequence, non-complementary sequence and blank control, demonstrating that the biosensing system effectively discriminates different oligonucleotides sequences and displays excellent specificity.
Interference of biological samples on miRNA detection
Besides sensitivity and specificity, interference of complex matrix on the biosensing strategy is another important feature need to be investigated. Total RNA samples (200 ng) extracted from breast adenocarcinoma (MCF-7) cells was firstly measured. According to the calibration curve shown in Fig. 4, the amount of miRNA-21 in the total RNA sample was estimated to be 21 fmol per 200 ng (RSD = 3.8%, n = 5). Then, 30 fmol synthetic miRNA-21 was spiked into 200 ng of MCF-7 total RNA, and the amount of miRNA-21 in the spiked sample was estimated to be 52 fmol (RSD = 2.7%, n = 5) with a recovery ratio of 98.1%, suggesting that the sensing method did not compromise in complex matrix.
Conclusions
In summary, this work has fabricated a simple colorimetric biosensing strategy for highly sensitive and specific detection of target miRNA-21 using T7 Exo-based signal amplification. The achieved biosensing system shows high sensitivity and good specificity for the determination of target miRNA-21 with a low detection limit of 0.69 pM. This strategy not only takes the advantage of high efficient amplification feature but also specific amplification feature of T7 Exo. Besides, the biosensing performance does not compromise in complex matrix. Notably, the colorimetric strategy demonstrates the distinct advantages including the simplicity in probe design, homogeneous and isothermal assay system as well as visual detection. Therefore, the colorimetric strategy provides a potential alternative platform for the detection of miRNA-21 in biomedical research and early clinical diagnosis.
References
Wen YQ, XU Y, Mao XH, Wei YL, Song HY, Chen N, Huang Q, Fan CH, Li D (2012) DNAzyme-based rolling-circle amplification DNA machine for ultrasensitive analysis of microRNA in drosophila larva. Anal Chem 84(18):7764–7669
Wang M, Yang Z, Guo Y, Wang X, Yin H, Ai S (2015) Visible-light induced photoelectrochemical biosensor for the detection of microRNA based on Bi2S3 nanorods and streptavidin on an ITO electrode. Microchim Acta 182(1–2):241–248
Shi HB, Xu J, Zhang GD, Xu LD, Li CQ, Wang L, Zhao Z, Jiang W, Guo Z, Li X (2013) Walking the interactome to identify human miRNA-disease associations through the functional link between miRNA targets and disease genes. BMC SystBiol 7(1):1–12
Li DD, Cheng W, Yan YR, Zhang Y, Yin YB, Ju HX, Ding SJ (2016) A colorimetric biosensor for detection of attomolar microRNA with a functional nucleic acid-based amplification machine. Talanta 146:470–476
Deng RJ, Tang LH, Tian QQ, Wang Y, Lin L, Li JH (2014) Toehold-initiated rolling circle amplification for visualizing individual microRNAs in situ in single cells. Angew Chem Int Ed 53:2389–2393
Wang GL, Zhang CY (2012) Sensitive detection of microRNAs with hairpin probe-based circular exponential amplification assay. Anal Chem 84(16):7037–7042
Shah P, Thulstrup PW, Cho SK, Bhang YJ, Ahn JC, Choi SW, Yang SW (2014) In-solution multiplex miRNA detection using DNA-templated silver nanocluster probes. Analyst 139(9):2158–2166
Cho WC (2010) MicroRNAs: potential biomarkers for cancer diagnosis, prognosis and targets for therapy. Int J Biochem Cell B 42(8):1273–1281
Li Y, Liang L, Zhang CY (2013) Isothermally sensitive detection of serum circulating miRNAs for lung cancer diagnosis. Anal Chem 85(23):11174–11179
Várallyay E, Burgyán J, Havelda Z (2008) MicroRNA detection by northern blotting using locked nucleic acid probes. Nat Protoc 3(2):190–196
Duan R, Zuo X, Wang S, Quan X, Chen D, Chen Z, Xia F (2013) Lab in a tube: ultrasensitive detection of microRNAs at the single-cell level and in breast cancer patients using quadratic isothermal amplification. J Am Chem Soc 135(12):4604–4607
Nagl S, Schaeferling M, Wolfbeis OS (2005) Fluorescence analysis in microarray technology. Microchim Acta 151(1–2):1–21
Zhang DC, Yan YR, Cheng W, Zhang W, Li Y, Ju HX, Ding SJ (2013) Streptavidin enhanced surface plasmon resonance biosensor for highly sensitive and specific detection of microRNA. Microchim Acta 180:397–403
Liu H, Bei X, Xia Q, Fu Y, Zhang S, Liu M, Yang Y (2016) Enzyme-free electrochemical detection of microRNA-21 using immobilized hairpin probes and a target-triggered hybridization chain reaction amplification strategy. Microchim Acta 183(1):297–304
Zuo X, Xia F, Xiao Y, Plaxco KW (2010) Sensitive and selective amplified fluorescence DNA detection based on exonuclease III-aided target recycling. J Am Chem Soc 132:1816–1818
Xu M, He Y, Gao Z, Chen G, Tang D (2015) Isothermal cycling and cascade signal amplification strategy for ultrasensitive colorimetric detection of nucleic acids. Microchim Acta 182(1):449–454
Huang KW, Yu CJ, Tseng WL (2010) Sensitivity enhancement in the colorimetric detection of lead (II) ion using gallic acid-capped gold nanoparticles: improving size distribution and minimizing interparticle repulsion. Biosens Bioelectron 25(5):984–989
Fu XL, Chen LX, Li JH, Lin M, You HY, Wang WH (2012) Label-free colorimetric sensor for ultrasensitive detection of heparin based on color quenching of gold nanorods by graphene oxide. Biosens Bioelectron 34(1):227–231
Martin A, Palomino JC, Portaels F (2005) Rapid detection of of loxacin resistance in mycobacterium tuberculosis by two low-cost colorimetric methods: resazurin and nitrate reductase assays. J Clin Microbiol 43(4):1612–1616
Bi S, CuiY LL (2013) Dumbbell probe-mediated cascade isothermal amplification: a novel strategy for label-free detection of microRNAs and its application to real sample assay. Anal Chim Acta 760:69–74
Wang F, Lu CH, Liu X, Freage L, Willner I (2014) Amplified and multiplexed detection of DNA using the dendritic rolling circle amplified synthesis of DNAzyme reporter units. Anal Chem 86(3):1614–1621
Cheng Y, Zhang X, Li Z, Jiao X, Wang Y, Zhang Y (2009) Highly sensitive determination of microRNA using target-primed and branched rolling-circle amplification. Angew Chem Int Ed 48(18):3268–3272
Hall MJ, Wharam SD, Weston A, Cardy DLN, Wilson WH (2002) Use of signal-mediated amplification of RNA technology (SMART) to detect marine cyanophage DNA. BioTechniques 32(3):604–611
Jia H, Li Z, Liu C, Cheng Y (2010) Ultrasensitive detection of microRNAs by exponential isothermal amplification. Angew Chem Int Ed 49(32):5498–5501
Nie J, Zhang DW, Tie C, Zhou YL, Zhang XX (2014) G-quadruplex based two-stage isothermal exponential amplification reaction for label-free DNA colorimetric detection. Biosens Bioelectron 56:237–242
Xuan F, Luo X, Hsing IM (2012) Ultrasensitive solution-phase electrochemical molecular beacon-based DNA detection with signal amplification by exonuclease III-assisted target recycling. Anal Chem 84(12):5216–5220
Wang M, Fu Z, Li B, Zhou Y, Yin H, Ai S (2014) One-step, ultrasensitive, and electrochemical assay of microRNAs based on T7 exonuclease assisted cyclic enzymatic amplification. Anal Chem 86(12):5606–5610
Wu ZK, Zhou DM, Wu Z, Chu X, Yu RQ, Jiang JH (2015) Single-base mismatch discrimination by T7 exonuclease with target cyclic amplification detection. Chem Commun 51(14):2954–2956
Shinozaki K, Tuneko O (1978) T7 gene 6 exonuclease has an RNase H activity. Nucleic Acids Res 5(11):4245–4262
Kerr C, Sadowski PD (1972) Gene 6 exonuclease of bacteriophage T7 purification and properties of the enzyme. J Biol Chem 247(1):305–310
Ma Y, Chen L, Zhang L, Liao S, Zhao J (2015) A sensitive strategy for the fluorescence detection of DNA methyltransferase activity based on the graphene oxide platform and T7 exonuclease-assisted cyclic signal amplification. Analyst 140(12):4076–4082
Wang HB, Wu S, Chu X, Yu RQ (2012) A sensitive fluorescence strategy for telomerase detection in cancer cells based on T7 exonuclease-assisted target recycling amplification. Chem Commun 48(47):5916–5918
Cui L, Zhu Z, Lin NH, Zhang HM, Guan ZC, Jamesá Yang C (2014) A T7 exonuclease-assisted cyclic enzymatic amplification method coupled with rolling circle amplification: a dual-amplification strategy for sensitive and selective microRNA detection. Chem Commun 50(13):1576–1578
Zhang H, Li M, Li C, Guo Z, Dong H, Wu P, Cai C (2015) G-quadruplex DNAzyme-based electrochemiluminescence biosensing strategy for VEGF165 detection: combination of aptamer-target recognition and T7 exonuclease-assisted cycling signal amplification. Biosens Bioelectron 74:98–103
Shi HY, Yang L, Zhou XY, Bai J, Gao J, Jia HX, Li QG (2017) A gold nanoparticle-based colorimetric strategy coupled to duplex-specific nuclease signal amplification for the determination of microRNA. Microchim Acta 2016:1–7
Oishi M, Sugiyama S (2016) An efficient particle-based DNA circuit system: catalytic disassembly of DNA/PEG-modified gold nanoparticle-magnetic bead composites for colorimetric detection of miRNA. Small 12(37):5153–5158
Wang Q, Li RD, Yin BC, Ye BC (2015) Colorimetric detection of sequence-specific microRNA based on duplex-specific nuclease-assisted nanoparticle amplification. Analyst 140(18):6306–6312
Miao X, Ning X, Li Z (2016) Cheng Z (2016) sensitive detection of miRNA by using hybridization chain reaction coupled with positively charged gold nanoparticles. Sci Rep 6:32358
Acknowledgements
This work was supported by the National Natural Science Foundation of China (81572080, 81101638), Natural Science Foundation Project of Chongqing (cstc2014kjrc-qnrc10001), and Achievement Transfer Project of Institutions of Higher Education in Chongqing (KJZH14205).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflict of interest. This article does not contain any studies with animals performed by any of the authors.
Additional information
Ye Sang and Yongjie Xu are contributed equally to this work.
Electronic supplementary material
ESM 1
(DOC 330 kb)
Rights and permissions
About this article
Cite this article
Sang, Y., Xu, Y., Xu, L. et al. Colorimetric and visual determination of microRNA via cycling signal amplification using T7 exonuclease. Microchim Acta 184, 2465–2471 (2017). https://doi.org/10.1007/s00604-017-2238-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00604-017-2238-8