Abstract
Background
In clinical practice, rectal cancer (RC) is classified according to tumor location. However, RC’s genetic characteristics according to tumor location remain unclear. Therefore, we aimed to compare RC’s genetic characteristics according to tumor location.
Methods
In 611 patients with surgically resected RC, we performed genetic analyses and compared the results between low and other RCs. Low RC was defined according to the European Society for Medical Oncology (ESMO) guidelines and Japanese Classification of Colorectal, Appendiceal, and Anal Carcinoma (JCCRC).
Results
KRAS mutation accumulation was significantly higher in low RC under the ESMO classification. Gene expression levels significantly differed between the groups for CTNNB1, KRAS, and ERBB2, under the ESMO classification and for TP53, KRAS, and ERBB2 under the JCCRC. Under the JCCRC, low RC had a significantly higher prevalence of fusion genes, such as EIF3E-RSPO2, PTPRK-RSPO3, and VTI1A-TCF7L2. Consensus molecular subtype (CMS) distribution was significantly different between the groups under both classifications. In particular, low RC had lower and higher frequencies of CMS2 and CMS4, respectively. CMS2 and CMS4 frequencies in low RC were 14.8% and 41.5% under the ESMO classification and 14.5% and 41.6% under the JCCRC, respectively. Multivariate Cox regression analysis demonstrated that pT3-4, pN1-2, and CMS4 were associated with poor relapse-free survival.
Conclusions
Low RC exhibited distinct genetic characteristics from other RCs. In particular, CMS4 was more frequent in low RC and was a risk factor for poor prognosis. These findings potentially avail further information regarding tumor biology and could lead to improvements in RC treatment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Large-scale genetic analyses of colorectal cancer (CRC) have recently been conducted [1, 2]. Several studies have revealed the genetic characteristics of CRC according to tumor location. When the tumors were classified as right- or left-sided, the prevalence of BRAF, KRAS, CTNNB1, SMAD4, and PIK3CA mutations as well as microsatellite instability were more common in right-sided tumors [3,4,5]. Right-sided tumors are associated with CpG island methylator phenotype [6] and consensus molecular subtype (CMS) 1 [2] and, while the TP53 mutation rate is higher in left-sided tumors [4]. Even after classifying CRC as right/left-sided, the genetic characteristics and prognosis differed according to tumor location on each side [4]. These data have been used for the selection of anti-tumor agents and for predicting CRC prognosis in clinical practice.
Generally, rectal cancer (RC) is classified into types according to tumor location in daily clinical practice. Lower RC has been reported to exhibit more aggressive behavior than upper RC [7]. Lower RC requiring rectal amputation has also been reported to have a poorer prognosis than other RCs [8, 9]. However, most large-scale genetic analyses of CRC classified RC as a left-sided tumor or as a single group, and the genetic characteristics of RC by tumor location have not been fully investigated, compared with those of colon cancer.
Here, we performed comprehensive genetic profiling of primary RC in a large Japanese cohort. We categorized RC by tumor location according to the general rules of the European Society for Medical Oncology (ESMO) guidelines [10] and Japanese Classification of Colorectal, Appendiceal, and Anal Carcinoma (JCCRC) [11]. We compared the genetic characteristics of low RC with those of other RCs. The identification of clinically significant differences could provide further information regarding tumor biology and lead to improvements in the treatment of RC.
Methods
Ethical statement
In 2014, the Shizuoka Cancer Center initiated Project HOPE (High-tech Omics-based Patient Evaluation) to investigate the biological characteristics of cancer, and the present study used selected data from this project [12, 13]. In Project HOPE, various tumor types, which were surgically resected at Shizuoka Cancer Center Hospital, were evaluated by multiomics-based analyses. This project was conducted at a single institution and designed according to the “Ethical Guidelines for Human Genome and Genetic Analysis Research,” revised in 2013. Informed consent was obtained from all patients and the Institutional Review Board of Shizuoka Cancer Center approved all aspects of this study (authorization no. 25-33).
Patient selection and study design
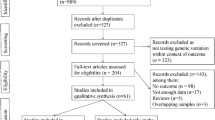
Patients with all types of cancer who underwent surgery at Shizuoka Cancer Center Hospital and were able to supply fresh cancer tissues of sufficient quantity were candidates for Project HOPE. Patients whose pathological diagnosis could be affected by the removal of cancer tissue were excluded from Project HOPE. Patients who underwent surgery for primary RC between February 2014 and March 2019 and were analyzed in Project HOPE were included in the present study. Patients who had a tumor at a distance exceeding 15 cm from the anal verge (AV), underwent preoperative treatments such as chemotherapy and/or radiotherapy, had squamous cell carcinoma or undifferentiated carcinoma, and provided samples from two or more colorectal tumors were subsequently excluded. Thus, 611 patients were eligible for this study. All tumors were pathologically diagnosed as adenocarcinomas. The clinicopathological and genetic characteristics of these RCs were retrospectively investigated.
Classification of the rectum
The localization of RC was classified according to the ESMO guidelines [10] and JCCRC general rules [11]. In the ESMO guidelines, RC is defined as a tumor located at a distance ≤ 15 cm from the AV. Further, RC is classified according to the tumor’s specific distance from the AV as follows: low, ≤ 5 cm; mid, > 5–10 cm; and high, > 10–15 cm. According to the JCCRC, the rectum is classified according to anatomical landmarks as follows: lower rectum (Rb), below the level of the peritoneal reflection; upper rectum (Ra), above the peritoneal reflection level and up to the lower margin of the second sacral vertebra; and rectosigmoid (RS), above the lower margin of the second sacral vertebra and up to the level of the sacral promontory. We classified tumors into RS-Ra or Rb (low) RC, based on the level of the lower border of the tumor.
Sample preparation
Tumors and their surrounding normal tissue weighing ≥ 100 mg were dissected from fresh surgical specimens. Peripheral blood was collected as a control for whole-exome sequencing (WES). For DNA analysis, clinical samples were frozen in liquid nitrogen before DNA extraction. DNA was extracted from these samples using a QIAamp DNA Blood MINI Kit (Qiagen, Venlo, Netherlands). DNA was quantified using a NanoDrop spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA) and Qubit 2.0 Fluorometer (Thermo Fisher Scientific). For RNA analysis, tissue samples were immersed in an RNAlater solution (Thermo Fisher Scientific) and stored at 4 °C before RNA extraction.
Whole-exome sequencing
Detailed protocols have previously been described [13,14,15]. Briefly, WES was performed on an Ion Proton System (Thermo Fisher Scientific) using the Ion Ampliseq Exome kit (Thermo Fisher Scientific). The WES exome library was constructed using the Ion Torrent AmpliSeq RDY Exome Kit (Thermo Fisher Scientific). Somatic mutations were identified by comparing tumor data with the corresponding blood samples.
Gene expression profiling (GEP)
GEP analysis was performed as described previously [13, 16]. Total RNA was extracted from approximately 10 mg of tumor tissue using the miRNeasy Mini Kit (Qiagen, Hilden, Germany). RNA samples with an RNA integrity number ≥ 6 were used in GEP analysis. Briefly, a total of 100 ng RNA was amplified and fluorescently labeled. These samples were hybridized to a SurePrint G3 Human Gene Expression 8 × 60 K v2 Microarray (Agilent Technologies, Santa Clara, CA, USA). Raw microarray data were normalized using GeneSpring GX software (Agilent Technologies). CMS classification was evaluated using the R package CMS caller [17].
Detection of fusion genes
A detailed protocol for the detection of fusion genes has been described previously [18]. Briefly, we constructed an in-house library that targeted total RNA to detect 491 known fusion genes. Thereafter, breakpoint sequencing was performed using a next-generation sequencer to determine the sequence of the fusion gene.
Outcome variables
Data on patients, pathological findings, genetic characteristics, and postoperative prognosis were collected. The tumor stage was defined according to tumor node metastasis classification [19]. These outcomes were compared between low and other RCs. As regards long-term outcomes, overall survival (OS) and relapse-free survival (RFS) rates were evaluated. Long-term outcomes were analyzed in cases that could be classified as CMS1–4 as well as those in which surgery was performed between February 2014 and December 2017 to exclude cases with short observation periods. Cases with pStage IV and with synchronous or metachronous malignancies were also excluded (Supplemental Fig. 1).
Statistical analysis
Fisher’s exact test or the chi-squared test was used to assess categorical variables, and Mann–Whitney U tests were used to compare continuous variables between the two groups. OS and RFS rates were calculated from the time of surgery using the Kaplan–Meier method and compared using the log-rank test. Univariate and multivariate analyses of the factors influencing RFS were performed using a Cox proportional hazard regression model. Risk factors with p values < 0.10 in univariate analysis were included in the multivariate analysis. Differences were considered statistically significant at a p-value < 0.05. On evaluating CMS classification using the CMS caller, q < 0.05 was considered statistically significant. All analyses, except for CMS classification, were performed using BellCurve for Excel (version 2.15; Social Survey Research Information Co., Ltd., Tokyo, Japan).
Results
Baseline characteristics
A total of 611 patients with RC were investigated, of whom 188 (30.8%) and 319 (52.2%) were classified as having low RC according to the ESMO and JCCRC classifications, respectively. Table 1 summarizes the baseline characteristics of the patients according to tumor location. The rate of locally advanced RC was relatively high, with 426 (69.7%) cases of pT3 or T4 and 313 (51.2%) with lymph node metastases.
Mutation accumulation
Table 2 shows the mutation accumulation of key genes in CRC, such as APC, TP53, SMAD4, CTNNB1, PIK3CA, PTEN, KRAS, NRAS, and BRAF, according to tumor location. Under the ESMO classification, only KRAS mutation accumulation was significantly higher in low RC than in high-mid RCs. However, under the JCCRC, the mutation accumulation of these genes was similar between low and other RCs.
Gene expression profiling
GEP data were available for 607 patients. Gene expression analysis was performed on nine genes examined for mutation accumulation and an additional gene, ERBB2. Under the ESMO classification, the expression levels of CTNNB1, KRAS, and ERBB2 differed significantly according to tumor location (Fig. 1a). Under the JCCRC, the expression levels of TP53, KRAS, and ERBB2 differed significantly according to tumor location (Fig. 1b).
Prevalence of fusion genes
Fusion gene expression was examined in 601 patients. Table 3 shows fusion gene expression details according to tumor location. Overall, 14 (2.3%) patients had fusion genes, of whom 13 (92.9%) exhibited RSPO-related fusion genes: two cases of EIF3E-RSPO2 and 11 of PTPRK-RSPO3. Only under the JCCRC fusion genes were found to be significantly more in the low RC group.
Distribution of CMS
CMS was analyzed in 601 patients (Table 4). One hundred and nine (18.1%) cases were unclassifiable. Under both classifications, the distribution of CMS was significantly different between low and other RCs. In particular, low RC had a lower rate of CMS2 and a higher rate of CMS4. Under the ESMO classification, the frequencies of CMS2 and CMS4 were 27.0% and 32.5% in high-mid tumors and 14.8% and 41.5% in low tumors, respectively. Under the JCCRC, the frequencies of CMS2 and CMS4 were 32.6% and 28.5% in RS-Ra tumors and 14.5% and 41.6% in Rb (low) tumors, respectively. The mutation accumulation of APC and TP53 in CMS4 tumors was 81.6% and 77.4%, respectively, and these figures were relatively higher than those reported previously [2]. In addition, the distribution of CMS was significantly associated with the pStage (Supplemental Table 1). As the tumor progressed, the frequency of CMS4 increased.
Genetic characteristics according to sex and histology
Under the ESMO classification, the frequency of poorly differentiated or mucinous tumors tended to be higher in low RC, and under the JCCRC, males were significantly more common in the low RC (Table 1). According to sex, the gene expression levels of PIK3CA, KRAS, BRAF, and ERBB2 (Supplemental Fig. 2a), and the distribution of CMS (Supplemental Table 2) differed significantly. According to histology, the mutation accumulation of APC and BRAF (Supplemental Table 3), and the gene expression level of SMAD4 and PTEN (Supplemental Fig. 2b) differed significantly. The distribution of CMS tended to differ, although it was not statistically significant (Supplemental Table 2). The prevalence of fusion genes was similar according to sex and histology (Supplemental Table 4).
Long-term outcomes by CMS classification
Thereafter, we investigated the association between CMS and long-term outcomes because the distribution of CMS differed clearly according to tumor location in both classifications. Long-term outcomes were analyzed in pStage 0–III cases that could be classified as CMS1–4 and in which surgery was performed between February 2014 and December 2017 (Supplemental Fig. 1). Tumor stages were similar among the four groups (Supplemental Table 5). Figure 2 shows the OS and RFS rates for each type of CMS. There was no significant difference in OS between the groups (Fig. 2a). RFS was significantly different between the groups, and CMS4 had a particularly poor prognosis (Fig. 2b). Multivariate Cox regression survival analysis revealed that pT3-4, pN1-2, and CMS4 were independently associated with poor RFS (Table 5). Further analyses of RFS by pStage and CMS revealed that in Stages 0–II, RFS tended to be worse in CMS4, although this outcome was not statistically significant (Fig. 2c). In Stage III, RFS was significantly worse in CMS4 than in CMS1–3 (Fig. 2d).
Discussion
This study demonstrated significant differences in genetic characteristics between low and other RCs. Most previous studies did not categorize RC based on location, although RC is always classified according to tumor location in daily clinical practice. To the best of our knowledge, this is the first study to compare the genetic characteristics of low RC with those of other RCs. Because this was a single-center study, detailed clinical data were available, and a detailed classification of RC was possible. Furthermore, in Western countries, the standard treatment for locally advanced RC is total mesorectal excision with neoadjuvant chemoradiotherapy (nCRT), whereas nCRT is not the standard treatment in Japan [20]. At our institution, the indications for nCRT are < 10% of all RC surgeries [21], and tumors treated with nCRT were excluded from this study. Thus, the present data are from a large number of surgically resected primary RC cases, which were not modified by preoperative treatments. The foregoing constitutes the strengths of this study.
In this study, the mutation accumulation and expression levels of key oncogenes, expression of fusion genes, and distribution of CMS were compared between low and other RCs. We used two classifications in which the location of the rectal tumor was categorized. Regarding mutation accumulation, only KRAS mutation accumulation was significantly higher in low RC under the ESMO classification (Table 2). We detected significant differences in the expression levels of several genes. However, these differences were exclusively observable under one classification (CTNNB1 and TP53), or the difference was not considerably large, even when it was identifiable under both classifications (KRAS and ERBB2) (Fig. 1). The prevalence of fusion genes was significantly higher in low RC, only under the JSCCR (Table 3). In contrast, the distribution of CMS was different according to tumor location under both classifications (Table 4). The CMS system is one of the most robust classifications for CRC, in accordance with gene signatures [2]. The CMS classification system consists of the following four subtypes with distinct biological and molecular characteristics: CMS1 (microsatellite instability immune), CMS2 (canonical), CMS3 (metabolic), and CMS4 (mesenchymal). Understanding these subsets is expected to influence treatment approaches and improve clinical outcomes. In this study, in particular, low RC exhibited a lower frequency of CMS2 and higher frequency of CMS4 than other RCs. Although CMS4 frequency is reportedly higher in cases with more advanced stages (III and IV) [2, 22], as confirmed by our study (Supplemental Table 1), the rates of lymph node and distant metastases were similar between low and other RCs in the present study (Table 1). Therefore, the differences in CMS distribution may be due to tumor location. CRC with CMS4 has previously been reported to have worse OS and RFS [2]. Here, even when the study population was limited to RC, CMS4 was found to be a risk factor for poor RFS after surgical resection (Table 5), suggesting the significance of the CMS classification system in predicting the prognosis of RC after surgery. Lower (extraperitoneal) tumors have previously been reported to exhibit more aggressive behavior than upper tumors [7]. The higher frequency of CMS4 in low RC may be one of the reasons for this finding.
CMS4 was found to be significantly more common in patients with low RC, and it proved to be a risk factor for poor prognosis. Treatment strategies based on these findings may improve the prognosis of RC. The therapeutic effects of cytotoxic drugs and molecular target agents have recently been reported to differ in accordance with CMS [23]. For example, concerning first-line chemotherapy, patients with CMS2/3 benefited from combination chemotherapy with bevacizumab compared with those with CMS1/4 [24]. In RAS wild-type cases, progression-free survival and OS in CMS4 were significantly better in patients treated with 5-fluorouracil/levofolinate/irinotecan (FOLFIRI) cetuximab than in those treated with FOLFIRI bevacizumab, while these benefits were not observed in CMS1/2/3 [25]. Irinotecan-based regimens were significantly superior to L-OHP-based regimens in patients with CMS4 CRC [26]. In high-mid or RS-Ra RC cases, tumors classified as CMS2 or CMS4 each accounted for approximately 30% of cases. In contrast, in low RC, CMS2 tumors accounted for approximately15% of cases only, whereas CMS4 tumors accounted for > 40% of cases (Table 4). These findings suggest the importance of drug selection targeting CMS4 in the treatment of low RC. If tumor CMS could be classified before chemotherapy by tumor biopsy, it may lead to the selection of more appropriate drugs. However, this is difficult because CMS classification using biopsy samples is significantly less reliable because of intratumor heterogeneity compared with that using resection samples [27]. The present data on CMS distribution by tumor location may help in the selection of therapeutic agents for RC cases in which the acquisition of resection samples is difficult (i.e., unresectable or preoperative cases). Total neoadjuvant therapy (TNT) has been developed for the preoperative treatment of locally advanced RC [28]. There was no change in the therapeutic drugs used in TNT according to tumor location. However, it may be preferable to alter the drugs according to tumor location based on CMS distribution. The relationship between CMS classification and drug efficacy in preoperative treatment remains unclear and will be the subject of further study.
In this study, we used two representative classifications that categorize RC according to location. Inconsistencies exist between the ESMO guidelines and the JSCCR [29]. From a molecular genetic perspective, it is unclear which classification makes more sense. In our opinion, if the one classification was able to distinguish clinically significant genetic differences, then this classification may be more appropriate for RC than the other classification, which cannot make that distinction. Here, we also found some genetic differences other than CMS distribution, according to tumor location in each classification. If clinical significance could be found in any of these differences, then the classification which reveals that difference may be more suitable. For example, the prevalence of fusion genes was significantly higher in low RC only under the JSCCR (Table 3). Of the 14 fusion gene cases, 13 (92.9%) had low RC. Furthermore, among these 13 cases, 11 (84.6%) were PTPRK-RSPO3, and PTPRK-RSPO3 was exclusively confirmed in low RC. Several therapeutic agents against CRC with PTPRK-RSPO3 have previously been reported [30,31,32]. If such therapies are availed for clinical use, it would be worthwhile to classify low RC according to the JSCCR rules for evaluating the expression of PTPRK-RSPO3 only in them. Tumors with fusion genes have already emerged as important therapeutic targets in lung cancer treatment [33], and the development of a specific treatment for CRC with fusion genes is anticipated.
This study had certain limitations. First, data on small tumors are scarce. Small tumors tended to be excluded from Project HOPE because the pathological diagnoses might have been affected by the removal of sample tissue. Second, the present data are from Japanese patients; hence, the generalizability of the results to other races remains uncertain. To further validate the present results, data on small tumors and other races are required. Third, the significance of the cases that could not be categorized by the CMS classification system remains unclear, and differences in tumor location possibly exist. Fourth, there were some differences in baseline characteristics of the patients according to the location of the tumor. The differences in genetic characteristics according to sex and histology might affect the present results about genetic characteristics of RC according to tumor location.
In conclusion, we found significant differences in genetic characteristics between low and other RCs. In particular, CMS distribution was significantly different, and CMS4, which was more frequent in low RC, was a risk factor for poor prognosis after surgical resection. In the future, it will be necessary to translate these data into clinical practice, and further investigation will enable physicians to provide optimal and personalized treatment options for patients with RC.
References
Cancer Genome Atlas N. Comprehensive molecular characterization of human colon and rectal cancer. Nature. 2012;487:330–7.
Guinney J, Dienstmann R, Wang X, et al. The consensus molecular subtypes of colorectal cancer. Nat Med. 2015;21:1350–6.
Loupakis F, Yang D, Yau L, et al. Primary tumor location as a prognostic factor in metastatic colorectal cancer. J Natl Cancer Inst. 2015;107:dju427.
Loree JM, Pereira AAL, Lam M, et al. Classifying colorectal cancer by tumor location rather than sidedness highlights a continuum in mutation profiles and consensus molecular subtypes. Clin Cancer Res. 2018;24:1062–72.
Tran B, Kopetz S, Tie J, et al. Impact of BRAF mutation and microsatellite instability on the pattern of metastatic spread and prognosis in metastatic colorectal cancer. Cancer. 2011;117:4623–32.
Cha Y, Kim KJ, Han SW, et al. Adverse prognostic impact of the CpG island methylator phenotype in metastatic colorectal cancer. Br J Cancer. 2016;115:164–71.
Benzoni E, Terrosu G, Bresadola V, et al. Analysis of clinical outcomes and prognostic factors of neoadjuvant chemoradiotherapy combined with surgery: intraperitoneal versus extraperitoneal rectal cancer. Eur J Cancer Care (Engl). 2006;15:286–92.
Law WL, Foo DC. Surgical results and oncologic outcomes for rectal cancer with tailored mesorectal excision over two decades. World J Surg. 2016;40:1500–8.
Lee KH, Kim JC, Kim JY, et al. Oncologic results and prognostic predictors of patients with locally advanced rectal cancer showing ypN0 after radical surgery following neoadjuvant chemoradiotherapy. Int J Colorectal Dis. 2015;30:1041–50.
Schmoll HJ, Van Cutsem E, Stein A, et al. ESMO Consensus Guidelines for management of patients with colon and rectal cancer. A personalized approach to clinical decision making. Ann Oncol. 2012;23:2479–516.
Japanese Society for Cancer of the C, Rectum. Japanese Classification of colorectal, appendiceal, and anal carcinoma. J Anus Rectum Colon. 2019;3:175–95.
Yamaguchi K, Urakami K, Ohshima K, et al. Implementation of individualized medicine for cancer patients by multiomics-based analyses-the Project HOPE. Biomed Res. 2014;35:407–12.
Nagashima T, Yamaguchi K, Urakami K, et al. Japanese version of The Cancer Genome Atlas, JCGA, established using fresh frozen tumors obtained from 5143 cancer patients. Cancer Sci. 2020;111:687–99.
Shimoda Y, Nagashima T, Urakami K, et al. Integrated next-generation sequencing analysis of whole exome and 409 cancer-related genes. Biomed Res. 2016;37:367–79.
Nagashima T, Shimoda Y, Tanabe T, et al. Optimizing an ion semiconductor sequencing data analysis method to identify somatic mutations in the genomes of cancer cells in clinical tissue samples. Biomed Res. 2016;37:359–66.
Ohshima K, Hatakeyama K, Nagashima T, et al. Integrated analysis of gene expression and copy number identified potential cancer driver genes with amplification-dependent overexpression in 1,454 solid tumors. Sci Rep. 2017;7:641.
Eide PW, Bruun J, Lothe RA, et al. CMScaller: an R package for consensus molecular subtyping of colorectal cancer pre-clinical models. Sci Rep. 2017;7:16618.
Urakami K, Shimoda Y, Ohshima K, et al. Next generation sequencing approach for detecting 491 fusion genes from human cancer. Biomed Res. 2016;37:51–62.
Brierley JD, Gospodarowicz MK, Wittekind C. TNM classification of malignant tumours. Wiley; 2017.
Hashiguchi Y, Muro K, Saito Y, et al. Japanese Society for Cancer of the Colon and Rectum (JSCCR) guidelines 2019 for the treatment of colorectal cancer. Int J Clin Oncol. 2020;25:1–42.
Yamaguchi T, Kinugasa Y, Shiomi A, et al. Short- and long-term outcomes of robotic-assisted laparoscopic surgery for rectal cancer: results of a single high-volume center in Japan. Int J Colorectal Dis. 2018;33:1755–62.
Fontana E, Eason K, Cervantes A, et al. Context matters-consensus molecular subtypes of colorectal cancer as biomarkers for clinical trials. Ann Oncol. 2019;30:520–7.
Sawayama H, Miyamoto Y, Ogawa K, et al. Investigation of colorectal cancer in accordance with consensus molecular subtype classification. Ann Gastroenterol Surg. 2020;4:528–39.
Mooi JK, Wirapati P, Asher R, et al. The prognostic impact of consensus molecular subtypes (CMS) and its predictive effects for bevacizumab benefit in metastatic colorectal cancer: molecular analysis of the AGITG MAX clinical trial. Ann Oncol. 2018;29:2240–6.
Stintzing S, Wirapati P, Lenz HJ, et al. Consensus molecular subgroups (CMS) of colorectal cancer (CRC) and first-line efficacy of FOLFIRI plus cetuximab or bevacizumab in the FIRE3 (AIO KRK-0306) trial. Ann Oncol. 2019;30:1796–803.
Okita A, Takahashi S, Ouchi K, et al. Consensus molecular subtypes classification of colorectal cancer as a predictive factor for chemotherapeutic efficacy against metastatic colorectal cancer. Oncotarget. 2018;9:18698–711.
Alderdice M, Richman SD, Gollins S, et al. Prospective patient stratification into robust cancer-cell intrinsic subtypes from colorectal cancer biopsies. J Pathol. 2018;245:19–28.
Yoo RN, Kim HJ. Total neoadjuvant therapy in locally advanced rectal cancer: role of systemic chemotherapy. Ann Gastroenterol Surg. 2019;3:356–67.
Tanaka A, Sadahiro S, Suzuki T, et al. A comparison of the localization of rectal carcinomas according to the general rules of the Japanese classification of colorectal carcinoma (JCCRC) and Western guidelines. Surg Today. 2017;47:1086–93.
Madan B, Ke Z, Harmston N, et al. Wnt addiction of genetically defined cancers reversed by PORCN inhibition. Oncogene. 2016;35:2197–207.
Han T, Schatoff EM, Murphy C, et al. R-Spondin chromosome rearrangements drive Wnt-dependent tumour initiation and maintenance in the intestine. Nat Commun. 2017;8:15945.
Storm EE, Durinck S, de Sousa e Melo F, et al. Targeting PTPRK-RSPO3 colon tumours promotes differentiation and loss of stem-cell function. Nature. 2016;529:97–100.
Saito M, Suzuki H, Kono K, et al. Treatment of lung adenocarcinoma by molecular-targeted therapy and immunotherapy. Surg Today. 2018;48:1–8.
Acknowledgements
We thank the members of the Shizuoka Cancer Center Hospital and Research Institute for their support.
Funding
This research was supported by the Shizuoka Prefectural Government, Japan.
Author information
Authors and Affiliations
Contributions
Conceptualization and study design: HH, AS, and KH; data acquisition: HH, AS, KH, HK, SM, YY, and TN; analysis and interpretation of data: HH and KH; drafting of the manuscript: HH and KH; study supervision: TN, KO, KU, YA, and KY.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hino, H., Shiomi, A., Hatakeyama, K. et al. Comprehensive genetic characterization of rectal cancer in a large cohort of Japanese patients: differences according to tumor location. J Gastroenterol 57, 476–485 (2022). https://doi.org/10.1007/s00535-022-01875-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00535-022-01875-7