Abstract
Significant evolution in deep learning took place in 2010, when software developers started using graphical processing units for general-purpose applications. From that date, the deep neural network (DNN) started progressive steps across different applications ranging from natural language processing to hyperspectral image processing. The convolutional neural network (CNN) mostly triggers the interest, as it is considered one of the most powerful ways to learn useful representations of images and other structured data. The revolution of DNNs in medical imaging (MI) came in 2012, when Li launched ImageNet, a free database of more than 14 million labeled medical images. This state-of-the-art work presents a comprehensive study for the recent DNNs research directions applied in MI analysis. Clinical and pathological analysis through a selected patch of most cited researches is introduced. It will be shown how DNNs are able to tackle medical problems: classification, detection, localization, segmentation, and automatic diagnosis. Datasets comprises a range of imaging technologies: X-Ray, MRI, CT, Ultrasound, PET, Fluorescene Angiography, and even photographic images. This work surveys different patterns of DNNs and focuses somehow on the CNN, which offers an outstanding percentage of solutions compared to other DNNs structures. CNN emphasizes image features and has well-known architectures. On the other hand, limitations beyond DNNs training and execution time will be explained. Problems related to data augmentation and image annotation will be analyzed among a multiple of high standard publications. Finally, a comparative study of existing software frameworks supporting DNNs and future research directions in the area will be presented. From all presented works it could be deduced that the use of DNNs in healthcare is still in its early stages, there are strong initiatives in academia and industry to pursue healthcare projects based on DNNs.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
1.1 Artificial intelligence (AI) from ideas to practice
The term AI was firstly mentioned in a workshop organized in a US non-famous university, Dartmouth College in 1956. The aim was to design a machine predicting human intelligence [1]. Machine learning booming started in the first decade of the twenty-first century due to the presence of powerful computer hardware and workstations. Machine learning, as a part of AI, is being successfully applied to academic or industrial problems. Such machines with powerful capabilities are exceeding human beings performance [2]. Advance in big data and computing power pushes AI from research to technology or from ideas to practice. Starting 2016, four AI perspectives appeared: research, teaching, media, and industry [3]. AI publications are growing by about 13% annually during the last 5 years. Five main clusters identify AI researches: search and optimization, natural language processing, computer vision, machine learning, and health care and medical imaging. When focusing on health care and medical image analysis, this number is widely increasing by 38% annually.
To conclude, AI is the main umbrella for machine learning and deep learning. Such a relation will be explained in the incoming sections. From my opinion, the move of researchers from academia to industry during the last 5 years could give a clear vision of how the number of AI applications is increasing tremendously in all disciplines.
1.2 Machine learning
There exist a wide range of open ended problems where it is difficult to find mathematical models; therefore solutions may depend on finding couples of examples with reasonable accuracy [4]. Such problems could be better solved using ML algorithms, where a set of sufficient examples (learning examples) are provided, then the machine will be able to take decisions regarding new examples (testing examples). Biological effects usually inspire ML techniques: neural networks, genetic algorithms, ants, decision trees, or particle swarm [5]. From year to year, ML is profiting from the increasing existence of digital information, the fast evolution in high performance computing facilities, and now from the possibility to be executed over cloud [6]. The artificial neural network (ANN) is the first ML technique inspired by the human neuronal synapse system.
1.3 Deep learning and conventional neural networks
Deep learning (DL) has changed forms and definitions slowly since 2008. DL comprises layers of nonlinear information processing in a hierarchical architecture for feature extraction, pattern analysis, and data classification [7]. Medical image classifications, computer vision, text-to-speech synthesis tools, and language translation are all highly annotated deep learning areas of research. MI analysis aided with DL still needs much effort, not only from computer scientists but also from physicians especially regarding lack of annotated image data and encouragement of automatic diagnosis systems. Though ANNs are designed to fit with different input data representations, DL networks are usually designed to cope with highly specific applications.
DL uses networks with larger number of layers, thus more parameters are needed to learn and converge. Parameters and weights tuning in DL require a compromise between training for minimal errors and overfitting; a situation that happens rarely in regular NNETs [8]. Comparing DL networks to conventional NNETs, DL have larger number of neurons, larger number of connections between neurons, and larger number of hidden layers. DL has several advantages, such as learning from the data itself, having state-of-the-art results in many domains, and outperforming humans in accuracy. However, to perform well, DL networks need high computation capabilities, High performance of H/W implementations, and significant amount of annotated training data. In my opinion, training model in DL seeks the best set of values for the network parameter vectors. This relation could be seen as a heuristic optimization problem targeting minimization of the loss function with respect to the network parameters. Minimization constrains are the tuning of network parameters towards the desired values.
2 Fundamentals of deep neural networks
The mostly used DL networks applied in MI analysis take one of the following architectures:
-
Convolutional Neural Networks (CNN)
-
Recurrent Neural Networks (RNN)
-
Restricted Boltzmann Machine (RBM)
2.1 Convolutional neural network (CNN)
A convolutional neural network (CNN) is classified as a supervised learning model that aims to learn higher-order features in the data via convolutions. The benefit of using a CNN is its ability to develop an internal representation of a two-dimensional signal. This allows the model to learn position and scale in variant structures in the data, which is important when working with images. CNNs, as shown in Fig. 1 were designed to map image data to an output variable. The first two layers seen in a CNN are the convolution layer and the affine (sub-sampling) layer. The convolution layer calculates the convolution between inputs to acquire feature maps. A nonlinear activation function is then used post convolution followed by a sub-sampling layer to reduce the dimension of feature maps through averaging. Following the sub-sampling is a set of ANN layers for classification, recognition, or decision purposes [9]. CNN is initially designed for image classification and nowadays is used for a variety of tasks. In a CNN, assume the input image dimension is L, Kernel size is K, thus the first convolution layer gives (L−K + 1) image sizes usually smaller than the original input. CNN is an outstanding tool for MI Analysis for many reasons: applying convolutional filters to learn image features, performing hierarchical feature extraction, which is useful while studying pathological images with different lesions, using a pooling layer that is able of averaging all acquired features and relating them to neighboring pixels.
Convolutional Neural Network [8]
2.2 Recurrent neural network (RNN)
Recurrent neural networks (RNNs) are designed using feedback signals to allow creating internal states or memories. These memories keep necessary information related to previous inputs (recurrent). This design makes RNNs useful to deal with sequential data, where inputs depends on each other in a streaming manner (sentence consisting of several words). RNNs are the best chosen networks for speech recognition and automatic machine translation systems [9]. The architecture of a RNN is shown in Fig. 2.
Recurrent neural networks [8]
2.3 Restricted Boltzmann machine (RBM)
Boltzmann machine (BM) is an ANN where all neurons from visible and hidden layers are connected to each other resulting high complexity, slow learning speed, and enlarged learning time. Therefore, restricted Boltzmann machine (RBM) was introduced restricting connections between neurons within the same layer [10]. Figure 3 shows structure of the RBM. Restricted Boltzmann machines are probabilistic models, i.e., the model assigns probabilities. RBMs architectures consist of one input layer and one or more hidden layer(s). Activation functions and neurons corresponding biases vectors are the core of RBM function. The absence of an output layer is obvious. Here, the biases or weights represent the filters parameters.
My opinion is that RBM has an advantage in creating filters that have picked out the strongest features in the input data. RBM could be used in applications where the transformation part (features) is needed, such as dimensionality reduction, classification, regression, and features' learning, as will be explained in Sect. 6 .
3 Introducing deep neural networks for medical image analysis
3.1 Motivations and challenges
Computer-aided diagnosis (CAD) based on DNN has emerged over the past 3 years. Advance in computer hardware architecture, efforts in DL software toolkits, and improved image quality from different medical imaging sources have all facilitated such area of research. CAD aided with DNN could reduce errors and enable efficient measurements when compared to physicians or traditional CAD systems. It is evident that different medical image computing fields host an increasing number of annual publications based on DNNs. Figure 4 shows achievements of DNNs in image computing areas. However, the transition of CAD systems with DL from laboratory to bedside faces difficulties for many reasons. This is time-consuming and labor intensive. Moreover, researchers must aggregate medical case studies with proven pathology. Medical image databases are another challenge, there exists several well-known databases such as ImageNet, Visual Object Classes (VOC), and Microsoft Common Object Context (COCO) with millions or hundreds of thousands of images; however, they lack medical image annotations. Employing CNN requires a large amount of annotated training dataset. To solve the problem of training powerful and effective DNN with only hundreds or thousands of patient scans or images, new trends use data preprocessing, innovative network designs, and different evaluation strategies. DNNs have the ability to learn medical image features during training [11]. Through multiple convolutional and data reduction layers, learning process could be easier and use adequate datasets [12]. For example, recent DNN could use hundreds or less dataset size to reach very low errors and improve the sensitivity of CAD systems by 13 to 34% in a variety of medical imaging applications [13].
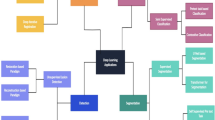
3.2 Applying deep neural networks in medicine
Recently, DNN is emerging in computer vision and medical imaging especially in areas such as mammography X-rays, cardiovascular CT/MRI scans, or microscopy images. In the incoming sections, four different clinical areas will be reviewed regarding intervention of DNN. Figure 5 shows those areas.
3.2.1 Some examples of DNN contributing in clinical images
-
Mammogram Image Analysis
Screening is the only way to reduce breast cancer risks and achieve early detection in women. Mammography is the most safe and adequate way for screening. Besides, the whole process comprises several classes. Miscellaneous tissues are detected, lesions are analyzed, and mass calcifications are monitored to classify the tumor grade and surgery decisions [14]. Manual process for such detection usually reaches sensitivity from 84 to 91% as proven by Zhang [15]. CAD systems that use CNN offer analysis of breast lesions from mammograms in three main steps starting from lesion detection, segmentation, to classification. This achieves an automated end-to-end CAD system.
In 2015, a breast mass segmentation method based on CNN was presented [16]. This used several potential functions; however, authors concluded that DL stand-alone models could not achieve high accuracy due to small training data set. They suggested integrating DL with a structured output model that gave assumptions about appearance and shape of the masses. More efforts were presented in breast cells detection in [17]. These achieved high accuracy without intervention of any integrated system and using adequate data set size. This concluded that the use of DL allowed accuracy improvements when compared to [16] in terms of classification of tumors.
From my opinion, mammogram analysis using DNN achieved a lot in the past few years due to data set preprocessing and noise elimination using morphological operations adopted by the authors. Furthermore, a paradigm shift in mammograms analysis reduces the classical triple-steps methodology to a one-step lesion detection and classification, trained with smaller data sets.
-
Cardiovascular Image Analysis
Cardiac observations comprise several imaging techniques. Ultrasound imaging is the most one used in case of monitoring cardiac functionality analysis, CT is widely used for coronary artery imaging, and fluoroscopy/ angiography is the dominant in case of interventions. Cardiac image analysis used traditional segmentation techniques such as region growing, thresholding, and watershed. More advanced methods used active contours or level sets. However, DL changed cardiac image analysis in the last few years [18]. In [19], a CNN was presented to detect left ventricular bounding box from MRI. They used level a set function proposed in that work combined with an energy term, a region-based term, and prior shape calculations. Good results were achieved. Zhen et al. [20] presented an effective technique to estimate the ventricular volume without segmentation. They proposed an RBM, where each layer was fully connected to its former layer. This type of full connectivity led to more network parameters when compared to a CNN and risked overfitting.
The main achievements are that the network was trained using unlabeled data set and the trained network was considered as an image feature extractor.
-
Vessel Segmentation
In this area of vessel segmentation, DNN are usually used to perform what is called pixel-wise classification. The network is well trained in order to obtain the segmentation mask. In [21], Wangetal used a CNN to segment retinal vessel. This proposed a multilayer CNN as a trainable feature extractor. The hierarchical method achieved good results and high accuracy even with only hundreds images training data sets.
-
Retinal Diseases
The retina and retinal structure of human eyes in diabetes is highly affected. This area needs more attention from computer scientists. In [22], a new supervised method for vessel segmentation from retinal images was presented for image diagnosis of ophthalmologic diseases. A wide and deep neural network to monitor this transformation and an efficient training strategy were presented. This outperformed state-of-the-art works in terms of sensitivity, computation, and accuracy. Authors used cross-training (a semi- supervised learning method) which required no preprocessing step and the training data set focused on diabetes retinal images. However, high accuracy was obtained in case of larger training databases.
The previous methods adopted pixel-wise classification which is time-consuming. In [23], a method that combined pixel classification and vessel tracking was presented. They started from a seed point and moved toward vessel particles. Those particles are given scores for being vessel belonging or not through a trained CNN. According to those scores, vessels’ particles were selected. Combining pixel classification and vessel tracking achieved a speed up of 2X, compared to previous methods.
3.2.2 DNN for pathological image analysis
Pathological and clinical diseases are highly supported by microscopic image analysis. This plays an important role in CAD systems. Large amount of microscopic daily image makes manual analysis inefficient. Deep learning finds the way in this area, due to many reasons. Firstly, DL requires huge amount of labeled images for training which is easily found in microscopic images. Secondly, pathological analysis is usually based on predefined models and structures which are easily detected through machine learning techniques. Finally, accuracy in such area is more important compared to computational time, which is achieved through selected type of DNN [24].
Therefore, from previous overview, DNNs have biased both clinical and pathological image analysis.
In the incoming sections, we will start focusing on specific MI areas: detection, localization, automatic diagnosis, classification, and segmentation; monitor achievements and comment on obtained results.
4 CNN applied for detection and localization
Manual detection suffers many problems that could lead to drastic consequences for both patients and physicians. Thus, automatic localization and detection prevents missing parts during MI analysis. Bowl [25] introduced a detection of cancerous lung from CT lung scans. He used two- stages CNN; the first for image enhancement and feature extraction, and the second for classification of cancer probability. To train the proposed network 2000 CT scans were used to obtain an accuracy of 98%. This accuracy is reached due to the proposed cascaded design; however, no studies related to time or algorithm complexities were presented. Another study was presented by Shin in [26] in the same area of lymph nodes cancer detection from CT images, where a CNN was used. They used ImageNet database (ImageNet: an image database with thousands of annotated images) and achieved an adequate accuracy of 95%. Yang et al. [27] did an effort in kidney cancer detection from histopathological images. They used a CNN with seven convolutional layers and used a set of 500 images for training to achieve an accuracy of 98%. Their training problem was easy enough since they only classified images as tumor or non-tumor. In [28], Shin et al. used an unsupervised learning method based on restricted Boltzmann machine (RBM) applied on a set of 78 MRI scans. The scans regions were containing liver or kidney tumors. They succeeded to detect tumors from both image categories and the RBM was able to learn features. They achieved accuracy of up to 79% based on the organ. It could be highlighted that unsupervised learning methods achieve lower accuracy compared to those obtained from CNN.
In the incoming subsections, highly selected researchers’ efforts and state-of-the-art works in medical image detection and localization will be surveyed deeply. Comments and discussions will enrich presented works.
4.1 Solving false positive detection in CAD systems using CNN
False positive (FP) in medical image detection means considering a few normal pixels as abnormalities. These usually reduce the sensitivity of automatic CAD systems and lead to wrong invasive interventions. Several works used cascaded classifiers for FP reduction [29]. This was achieved either using a post-processing filter that can eliminate FPs based on statistical analysis or using manual methods. However, these methods are not effective and contradict with automatic diagnosis systems. A better method is to acquire new image features at the candidate location and use such features to train new classifying methods. New features can lead to missing information within the first stage and thus could lead to a better classification; FPs could be then reduced to the minimum extent [30].
In [31] (more than 90 Scopus citations), authors presented a FP detection solution for CAD systems keeping high sensitivities. In this work, CNN is used to differentiate hard false positives from true positives. To let the CNN better sees 2D images, random rotation, translation, and multiscaling analysis were applied around a coordinate center. Sensitivity has increased from an average value of 57% to an average value of 75% regarding FP localization.
The following paragraphs describe method, algorithm, and CNN implementation. Finally, we will comment on results.
4.1.1 Data set preparation
Since CNN effectiveness is highly dependent on the size of training data, authors in [31] presented a simple and efficient way to produce an increased number of training dataset in number and diversity. The 3D volumetric raw images are first sliced into 2D images, three different transformations are then applied to each 2D slice: translation along a random vector v, rotation around a center coordinate with angle in the range [0° 360°], and scaling. The number of random translations, rotations, and scales are Nt, Nr, and Ns, respectively. It is mandatory to keep the same number of pixels/voxels during such transformations. Finally, this preprocessing stage generates N sample 2D images (N = Ns × Nt × Nr) for each region of interest (ROI). To obtain labeled images for the prepared data set, ground truth data are used. Observations on pixels under investigations as ‘FP’ or ‘TP’ depends on whether it belongs to a true lesion (object of interest) or not. Resultant labeled images are successfully used to train the CNN in a fully supervised manner.
4.1.2 CNN implementation
Three radiological data sets were chosen comprising different clinical applications: spine images for sclerotic metastases detection, cancer detection from lymph images, and cancer detection from colonic images. The proposed CNN was run on an NVIDIA GeForceGTX TITAN (6GBmemory) hardware environment. Training time while considering a number of 1200 optimization epochs ranged from nine to thirty hours. Supervised learning methods with labeled MI data sets may take large training time and thus it is essential to use GPU cores [32], as will be explained later.
4.1.3 Comments and observations
From my opinion, the proposed method succeeded in solving FP detection problem using CNN. Two important observations could be monitored. The first is the proposed image preprocessing that created a huge training dataset using scaling, rotation, and translation. The second is the study of the same 2D slice from different views and scales that increased the effectiveness of the CNN as a classifier and thus led to an increasing sensitivity.
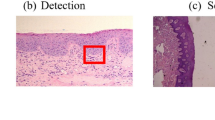
4.2 Mitosis detection from breast cancer pathological images
Pathology quantitative tissue analysis could help in better understanding cancer behavior and localization. State-of-the-art work in cell and nucleus detection usually considered thresholding and morphological operations [33], region growing [34], level sets [35], K-means [36], or active contours [37]. However, recent researches consider DL techniques to test and validate larger number of histopathological images [38]. Mitotic count is a good indicator for breast cancer aggressiveness. This is manually performed by pathologists, which is dangerous and time-consuming. A multistage DL method for mitotic cells detection from histopathology images was presented [39] (37 citations in Scopus in 8 months). The proposed method [39] has two main objectives: deep detection network for localization of mitotic cells using contextual based information, and a deep verification network for removal of false positive detection, as will be shown below. A state-of-the-art well-known breast cancer dataset was used during experimental results [40]. The performance of such systems is evaluated according to only correct counts, irrelevant from the shape of the mitosis [39]. Details are given in the incoming paragraphs.
4.2.1 Data set preparation
The system in [39] used 1696 High Power Field (HPF), i.e., area visible under the maximum magnification power of the electron microscope, images at 40X magnification. Each HPF had a size of 1539 × 1376 pixels. The training data consisted of 1200 with only 749 labeled images. The testing dataset considered the rest of images. Dataset augmentation used cropping, rotation, and mirroring. It started by cropping into 512 × 512 pixels from original images then images were rescaled to 1024 × 1024. Rotation and mirroring were both applied to the original HPF images to produce more training samples. Rotation was applied with a step size of 45°.
4.2.2 CNN implementation
The core component of this proposed system is the deep detection (DeepMitosis). This utilized a 50-layers CNN, trained over 12,000 iterations, and learning rate of 0.01. This CNN generated reference boxes during the last convolutional feature map layer. Those reference boxes were called anchors. Two fully connected layers (pooling) were designed to classify anchors and reduce bounding box sizes. The refining detection came as a second phase, took the detected boxes from the detection CNN as input and estimated a probability score for each anchor being ‘true positive’ or ‘false positive’. The system was implemented on Caffe DL framework using Python and C + + . Experiments were carried out on Lunix server with NVIDIA GeoForce GTX TITAN X GPU and results are shown in Fig. 6.
Results of Two DNNs: Localization and Verification [39]
4.2.3 Comments and observations
Since, mitotic count and not the mitotic shape is the most critical item when estimating breast cancer from pathological images, the proposed work with detection and refinement is highly appreciated.
From my point of view, the method used to form image patches is inadequate, since mitotic cells in the boundary of patches could be split into two or more patches and thus increase the counts. Since pathological images are usually analyzed in labs and not real time, authors did not give any study regarding time performance while using two cascaded CNNs one with 50 layers.
5 Classification and diagnosis using deep neural networks
Deep learning diagnosis convolves several areas. In [41], electrocardiogram (ECG) beat classification has been analyzed aided with deep learning. Since ECG beat data lies on high-dimension manifold, this work proposed a novel “local deep field” for classifying the devil in the details of such complex variations of ECG data. This method learnt different deep models to be able to detect the hidden class information within local distributions. The results showed good accuracy in classifying ECG that surpassed cardiologists work.
Another outstanding area is the early diagnosis of Renal Transplant Rejection (RTR), where the current diagnostic technique is renal biopsy that is not preferred due to its invasiveness, time recovery, and complications. A computer-aided diagnostic (CAD) system for early Automatic RTR (ARTR) detection from 3D magnetic resonance imaging (MRI) data was presented in [42]. The CAD process started from kidney tissue segmentation using level-set-based segmentation. A B-spline-based 3D data alignment was employed to overcome local deviations due to breathing and heart beating. Then, empirical cumulative distribution functions of apparent diffusion coefficients of the segmented tissue were collected as discriminatory transplant status features. Finally, a deep-learning-based classifier with autoencoders was employed to distinguish between rejected and non-rejected renal transplants. Experiment was applied on 100 subjects, 97.0% were correctly classified.
There is a high demand for developing CAD tools to help pathologists making accurate diagnosis. CAD systems from histopathology are possible since emergence of digital pathology [43 and 44]. Recently, interest has been given to the application of DL techniques to implementing CAD systems that are able to classify and take decisions aided with big data images. Another research area where DNNs assisted CAD systems remarkably is mammogram image classification and diagnosis. Since, it was found difficult to segment mammogram image accurately due to low contrast between normal and abnormal lesion tissues, in [45], a CNN was used to better learn features of an initial contour of mammograms and micro calcifications located through a Chan–Vese level set method. To increase the classification accuracy and reduce the false positives, a relaxation network classifier was used in the last stage of the proposed CNN. Three performance measures were applied. Accuracy, sensitivity, and specificity reached 99%, 0.9875, and 1.0, respectively. These results proved how DNNs could improve CAD systems with annotated data.
In the incoming sections, focus will be given to three research articles to show DNNs performance in different clinical areas.
5.1 A skin cancer classification approach from photographic images
Skin cancer is visually diagnosed. Beginning with an initial clinical screening and followed potentially by dermoscopic analysis, biopsy, and histopathological examination. The intervention of DL in MI analysis can open another view and facilitate detection of the most common human malignancy [46 and 47]. Previous work in dermatological CAD systems focused only on either dermoscopy or histological images. The former needs a specialized instrument, while the latter uses invasive biopsy and microscopy [48].
An outstanding research was presented in [49] (214 citations in Scopus). This work presented an end-to-end well trained CNN for skin cancer classification from direct skin images. Photographic images exhibit few problems: zoom, angle and lighting, or blurring. This makes classification a challenging problem [50 and 51].
The proposed method in [49] overcame such problem using a data driven approach, i.e., training a million of photographic images over CNN transforms image features into learnt data via the CNN and makes it robust for photographic variability.
5.1.1 Data set preparation
The data set used came from a combination of open access dermatology and Stanford Hospital. Data set contained images representing same lesions form multiple view points for the same person, i.e., image scaling, rotation, and flipping were used with random probabilities. Blurry images were removed from testing and validation pool but kept in training phase. The overall data set consisted of 129,450 images representing about 2000 visual skin appearances.
5.1.2 CNN implementation
The taxonomy presented in [49] described a tree structure with two main classes. The first class comprised: benign – malignant – non neoplastic lesions. The second class represented the major diseases nodes as shown in Fig. 7. The paper used 2014 ImageNet CNN already trained but replacing the final classification layer according to the skin cancer problem. All images were adjusted to 299 × 299 pixels to fit with the CNN. Google TensorFlow DL framework was used.
Proposed Taxonomy Classes [49]
5.1.3 Results
The proposed CNN achieved 73% sensitivity compared to 66% in traditional analysis methods (dermatologists: dermoscopy or histological images).
5.1.4 Comments and discussions
In my opinion, the proposed work has opened a new trend for future DL in medical image analysis for many reasons:
-
Classification from photographic images, where it could be possible, was impressing and achieved good results.
-
The author extended their work to a mobile application used by dermatologists outside clinics.
-
The CNN evaluation is highly outstanding, a group of 21 board-certified dermatologists approved obtained results. This is an important achievement regarding the real existence and approval of CAD systems by physicians.
The impressing number of citations and the quality of the paper (Nature publication) makes it a role model for CNN applications in medical imaging.
5.2 Lung diseases CAD system using CNN
Lung diseases comprise more than 100 chronic lung disorders characterized by inflammation of the lung tissue [52]. Till now, the diagnosis of lung diseases involves questioning the patient, performing physical examinations, and image scans via chest X-ray or CT. Those scans are examined through physicians using visual inspections leading to wrong diagnosis in many cases. Rare CAD systems for lung assessment comprise the following steps: lung segmentation – lung disease quantification – diagnosis or classification. A few classifiers were presented in the literature based on: k-nearest neighbors, ANN, support vector machine, or random forest [53 and 54]. Some attempts have recently used DL techniques, especially CNN in lung tissue analysis [55].
In [56] (85 citations in Scopus), a CNN was proposed for lung diseases patterns classification. The proposed CNN consisted of five convolutional layers followed by an average pooling layer following the number of diseases classes. In their work, seven classes were selected: healthy, ground glass opacity (GGO), micronodules, consolidation, reticulation, honeycombing, and a combination of GGO/reticulation.
5.2.1 Data preparation
The data used for training and validation were acquired from two main sources: Swiss University Hospital (94 scans) and Bern University (26 scans), leading to a total of 120 patients’ scans, each of size 512 × 512 pixels. Images comprised healthy and unhealthy tissues.
A new trend was applied to augment data size. Each scan was partitioned into a 32 × 32 pixels image patch, i.e., one CT scan gave 256 image patches. A total of 30,720 (120 × 256) image patches were then obtained for the whole data set. However, physicians excluded non-ROIs and bronchovascular patches resulting a total of 14,969 image patches for training and evaluation. This new trend of subdividing the image was adopted for two reasons. First, one CT scan for a patient could have more than one disease. Therefore, each part in the image scan is of great importance. Second, focusing on every part let the CNN learn features better. Figure 8 presents example of generating image patches.
Generating Image Patches from One CT Slide [56]
5.2.2 CNNs architectures
The input image to the CNN was 32 × 32, and five convolutional layers were then used. The size of the kernels in each layer was chosen the minimal (2 × 2), as smaller kernels lead to deeper CNNs. An average pooling layer followed the convolutional layers with size 7 (representing the classes). Three CNNs were implemented for results comparisons, all with similar architecture but with different kernels sizes, number of convolutional layers, and loss functions. The proposed algorithm was implemented using Theano framework [57], and experiments were performed under Lunix OS on a core i7 machine with GPU NVIDIA GeForce Titan.
5.2.3 Results
The number of kernels affected the convergence time and each training epoch became slower by more than 20X. By altering the number of convolutional layers, it could be concluded that five to six layers gave the best results. Comparison with state-of-the-art work showed that the proposed CNN proved superior performance. Accuracy achieved was 0.86. Furthermore, the accuracy achieved in this method surpassed VGG-Net [57] and AlexNet [58] by 8% and 12%, respectively.
5.2.4 Comments
The number of convolutional layers plays an important role, as increasing this number led to overfitting and smaller numbers reduced the accuracy.
The proposed data augmentation method represents a new trend, since splitting the features map into multiple pooled regions leads to more features view in different areas of the same image and thus facilitate the CNN to study such features.
To the best of my knowledge, if the authors used Wavelet Transform (WT) prior CNN, they may have achieved better performance. Since WT emphasizes image features and could help in image partitioning in both spatial and frequency domains, it could be an asset to the previous work.
5.3 Alzheimer’s diseases classification using RBM
According to Alzheimer’s Disease International, nearly 44 million people have Alzheimer worldwide. Only 1-in-4 people with Alzheimer’s disease has been diagnosed. Alzheimer’s is most common in Western Europe and North America. On the other hand, it is least prevalent in Sub-Saharan Africa and Asia. Alzheimer’s is considered the top cause for disabilities in later life [59].
5.3.1 Alzheimer’s classification challenges
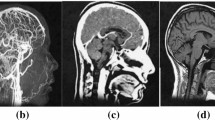
Early diagnosis plays an important role in preventing dramatic drawbacks of Alzheimer’s Disease (AD). This is based on classifying extracted features from brain images. The problem is very different when compared to tumors or calcifications detection, since such features have to monitor variations of anatomical brain structures, such as, ventricles size, shape, tissue thickness, or brain volume. In [60], a deep 3D-CNN was proposed to capture AD biomarkers, learn generic features, and predict AD. The 3D-CNN was pre-trained to capture anatomical shape variations in structural brain MRI scans. Experiments showed good results over the proposed MRI dataset with no skull-stripping preprocessing. To diagnose AD and its prodromal stage, namely, Mild Cognitive Impairment (MCI), Suk et al. [61] proposed a DL method for finding high-level latent and shared features from two imaging modalities: MRI images and Positron Emission Tomography (PET) images. In their study, a restricted Boltzmann machine (RBM) was used to find a latent hierarchical feature representation from a 3D patch (a joint feature representation from the paired patches of MRI and PET) with a multimodal RBM. In the multimodal deep Boltzmann machine, a Gaussian RBM was trained to transform the paired patches into binary vectors. After finding high-level latent and shared features by using the paired patches and trained multimodal deep Boltzmann machine, an image-level classifier was developed to perform the final classification.
5.3.2 RBM structure and training
Restricted Boltzmann machines are probabilistic models. RBMs have one of the easiest architectures; it consists out of one input layer, called the visible layer and one or more hidden layer(s). The absence of an output layer is obvious in this proposed model, since the predictions are made in a different manner; the biases or weights represent the filters parameters. These filters can be visualized as a grayscale image, as explained before in Sect. 3.3. Dataset was partitioned into ten subsets, each including 10% of the total data. Nine subsets were used for training and the rest for testing. They defined a preprocessor that effectively converted MR tissue densities or PET voxel intensities into 500-dimensional binary vectors. Those vectors were used to train the RBMs. The proposed RBM consisted of three-layers for MRI and PET (PET-DBM) respectively, and four-layers for MRI + PET. Both the MRI-RBM and the PET-RBM were structured with 500 (visible), 500 (hidden), and 500 (hidden) neurons. The MRI + PET had a final layer with 1,000 hidden units.
5.3.3 Results
To validate the effectiveness of the proposed method, authors performed experiments on ADNI dataset and compared with the state-of-the-art methods. In a binary classification problem of AD against healthy Normal Control (NC), a maximum accuracy of 95.35% was obtained, outperforming the state-of-the-art work in this area. By visual inspection of the trained model, it could be observed that the proposed method could hierarchically discover the complex latent patterns inherent in both MRI and PET.
5.3.4 Comments
From the previous state of the art works, it could be concluded that RBM surpasses CNN and other DNN techniques in classifying Alzheimer. The main reason is that the challenge within AD is that we are searching for textures rather than abnormalities within the image. The effect of RBM as filter banks helped so much in this area.
6 Deep neural networks for medical image segmentation
Automatic tissue and region of interest (ROI) segmentation in medical images is of great importance for different clinical routines. Segmentation is sometimes a preprocessing stage for several medical analysis. MI segmentation encounters many challenges. For example, automatic and reliable segmentation techniques for removing brain tumors are required since this can affect patients’ health and shorten their life. However, such tumors have large spatial variability and structural complexity [62]. Several state-of-the-art works used DL in brain tumor segmentation’s methods [63, 64]. For example, Pereira et al. [65] used a CNN with reduced convolutional kernels with the aim to segment gliomas (the most common and aggressive brain tumors). In their research, authors used small kernels and thus deep networks for more features’ observation.
Other DL methods focused on segmentation of various tissues to differentiate between three important ROIs in an MRI brain image: Gray Matter (GM), White Matter (WM), and Cerebrospinal Fluid (CSF) [66]. Therefore, CNN architectures were designed according to different input patch sizes. Different convolutional network architectures with variable number of convolutional layers were used for comparison purposes and to obtain resulted different feature map levels.
MI segmentation for measurement of cardiac ventricle parameters plays a crucial role in clinical assessment, i.e., ventricular volume, wall thickness, and ejection fraction, and functionality. Therefore, DL methods have been proposed to reach an accurate automatic segmentation [67]. For example, to segment the LV from MRI images, Avendi et al. [19] proposed a methodology, as explained earlier in this survey, which combined DL architecture and deformable models.
6.1 Brain tumors segmentation using two-pathways CNNs
Although surgery is sometimes essential for brain tumors treatment, there are tumors that cannot be physically removed. Radiation and chemotherapy are used to slow the growth of those tumors. MRI is one of the most common tests for brain tumors’ diagnosis and surgery decision. Automatic brain tumor segmentation has great impact on growth rate prediction and treatment planning. As described above, healthy brains are consisting of three types of tissues: WM, GM, and the CSF. The aim of brain tumor segmentation is to detect the active tumorous tissue, or the location and extension of the tumor regions [68]. In the last 5 years, the use of deep CNNs for brain tumor segmentation was discussed in several big medical conferences. Davyetal [69], Zikicetal [70], and Urban et al. [71] divided the 3D MR images into 2D or 3D patches [71] and trained a CNN to predict its center pixel class. Urban et al. [71] as well as Zikic et al. [70] implemented a common CNN consisting of a series of convolutional layers, a nonlinear activation function between each layer and a softmax output layer.
In [68] (345 citations in Scopus), a fully automatic brain tumor segmentation method based on CNNs was presented. The proposed networks were tailored to glioblastomas pictured in MR images: tumors that can appear anywhere in the brain with different kind of shapes, sizes, and contrasts.
6.1.1 Motivations
The motivation within this work was to explore different CNN architectures and thus present a novel architecture that could exploit both local features as well as global contextual features simultaneously. Furthermore, they used a fully connected layer as an output layer; which allowed a 40X speed up in the overall training and testing algorithms. Finally, they explored a cascaded architecture in which the output of the CNN was treated as an additional source of information for a subsequent CNN. The results reported on the 2013 BRATS test dataset when compared with [68] revealed that this architecture improved segmentation performance.
6.1.2 CNN architecture
-
Two-pathway Architecture
The architecture in [68] consisted of two main streams: a pathway with 7 × 7 receptive fields and another with 13 × 13 receptive fields.
Pathways were named: ‘local pathway’ and ‘global pathway’, respectively. The motivation for this architectural choice was the correct prediction of the label of a pixel influenced by two aspects: the visual details of the region around that pixel and its larger “context”, i.e., exploit both local features as well as global contextual features simultaneously.
-
Cascaded Architecture
The idea was based on feeding the output probabilities of the first CNN as additional inputs to the layers of a second CNN. The outcome was to increase the efficiency of CNNs to specify the dependencies between adjacent labels. This technique was named ‘joint segmentation’. Figure 9 depicts the architecture.
Fig. 9 Two Pathways and Cascaded Architectures [68]
6.1.3 Implementation and results
The implementation was based on an open source machine-learning library specialized in DL algorithms, Pylearn2 [72]. It also supported the use of GPUs, which are nowadays essential for DL algorithms. Since CNN’s are able to learn useful features from scratch, Havaei et al. applied only minimal preprocessing. The preprocessing followed three steps: the 1% highest and lowest intensities were removed, a bias correction was applied, and finally the data were normalized within each input channel by subtracting the channel’s mean and dividing by the channel’s standard deviation. The training dataset contained 30 patient subjects all with pixel accurate ground truth (20 high grade and 10 low grade tumors); the testing dataset contained 10 (all high grade tumors). The training brains come with ground truth with five segmentation labels: non-tumor, necrosis, edema, non-enhancing tumor, and enhancing tumor. In total, the model iterated over about 2.2 million examples of tumorous patches.
6.1.4 Comments
The first achievement in this work is the performance when using the novel two-pathway architecture that was able to model both the local details and global context or modeling local label dependencies. From my opinion, the cascaded CNN could better be replaced by one or two additional convolutional layers.
The authors did not observe the disadvantage of cascaded layers since they implemented their system over highly performing distributed GPU cores.
6.2 Interactive MI segmentation using DL
Interactive segmentation methods are new trends that integrate user knowledge and non-visual image features to reach accurate results [73]. These trends are highly appreciated by most physicians. In [74], a novel DL interactive segmentation framework used CNNs to study a bounding box from a supervisor. The proposed framework was applied to segment human organs from 2D MRI slices. The experimental and simulation results showed that the proposed model was robust to segment unseen organs with high accuracy and little intervention from supervisor. The system succeeded when trained in an unsupervised learning manner.
6.2.1 Interactive segmentation challenges
Three challenges could be observed related to organs segmentation:
-
One MI contained several neighboring organs.
-
CNN usually does not generalize to previously unseen organs.
-
Interactive segmentation requires DL of a ROI then generalization using context variations outside this ROI.
-
Fast inference and memory efficiency are highly required for interactive segmentation.
6.2.2 Method
The proposed interactive framework [74] consisted of bounding box that represented the input to a CNN. This specified an initial organ segmentation. The segmentation was based on the fact that the CNN is capable of learning some common features, saliency, contrast, and hyper intensity across different objects. This process was applied to other organs for more generalization during training. The pre-trained CNN accepted unseen images and was capable of segmenting the organ existing in the bounding box.
6.2.3 Training phase
The proposed CNN consisted of five convolutional layers, one concatenation layer, and one softmax layer. The kernel size was varied during the five convolutional layers to 1, 2, 4, 8, and 16. The main reason was that to adapt the CNN to capture features at different scales. Since, the MRI input image contained several organs even inside the bounding box. Features from these five layers were concatenated and fed into layer six; that served as a classifier. Finally, a softmax layer was used to obtain probability-like outputs. In the testing phase, they updated the model to ensure efficient fine-tuning and fast response to user interactions. Features in the concatenation layer for the test image were stored before the fine-tuning.
6.2.4 Results
MR images from 18 patients were used. They performed data splitting at patient level and used images from 10, 2, 6 patients for training, validation, and testing, respectively. The training set consisted of 333 and 213—2D instances of the placenta and fetal brain. The validation set contained 70, 25, 36, and 41—2D instances of the placenta, fetal brain, fetal lungs, and maternal kidneys. The testing set consisted of 165, 80, 114, and 124 2D instances of the placenta, fetal brain, fetal lungs, and maternal kidneys, respectively. The CNN performed well on previously unseen fetal lungs and maternal kidneys.
6.2.5 Comments
From my view, two main observations could be seen within this study. The first is that authors succeeded to build a CNN that segmented totally unseen images. The second, is the user interaction framework. User interaction leads to weak learning and slower time; however, results proved high accuracy and fast response.
6.3 3D Medical image segmentation using CNN
Deep learning techniques emerged as powerful supervised learning tools with great model capacity and ability to learn highly discriminative features for different MI tasks. Usually segmentation of 3D images was performed by processing groups of 2D slice independently, which lacks the importance of volumetric medical image data [75]. Fully 3D CNNs come with an increased number of parameters, significant memory usage, and high computational requirements. Several works studied limitations while using 3D CNN for medical imaging [76]. The main reasons for that could be summarized in the following two considerations:
-
Convolution with 3D kernels are computationally expensive
-
3D-CNN architectures have a huge number of training parameters
In [77], a dual pathway, 11-layers deep, 3D- CNN was presented. The challenging task was a brain lesion segmentation. To overcome the computational problem within 3D MI scans, authors achieved three contributions:
-
An efficient dense training scheme that used adjacent image patches instead of using the whole image during training
-
The development of a deeper and more discriminative 3D-CNNs
-
Introducing a dual pathway architecture at multiple scales
This work improved the state-of-the-art work with top ranking performance on the public benchmarks BRATS 2015 [75].
6.3.1 3D- CNN architecture
3D- CNNs used voxel segmentation by classifying each 3D image voxel independently taking the neighborhood (the local and contextual image information) into account. This was achieved by sequential convolutions of the input with multiple filters at the successive layers of the proposed network. The neurons of higher layers combined the patterns extracted in previous layers, which resulted in the detection of increasingly more complex patterns. The activations of the neurons in the last layer (L) was related to particular segmentation class labels, thus the last layer was also considered as the classification layer. Figure 10 demonstrates a 3D CNN with kernel equal 5 × 5 × 5. Similarly to 2D convolution, the size of the resultant image is (L−K + 1), where L is the input image size and K is the kernel size.
6.3.2 Dense training on image segments in 3D- CNN
When the receptive field is fully enclosed within the input and captures only original content, i.e., the input segment dimensions are divided by the kernel size, the computational costs and memory loads will be reduced. In [77], repeated computations of convolutions on the same voxels in overlapping patches were avoided for the reason stated above and thus optimal performance was achieved. However, GPU memory constraint means that there is no sufficient space to deal with the complete input images and thus image patches were used to be small enough and fit into memory. Image patches (Segments) were analyzed instead of original images, where the number of patches was assumed as B. Larger Bs were preferred as they could approximate the whole data more accurately and led to better segmentation of the tumor lesions. However, a compromise should be considered when selecting B even while using GPUs.
6.3.3 Deeper CNN
In order to build a deeper 3D architecture, small kernels were adopted. Smaller kernels are faster to convolve with and contains less weights. In the work presented by Kostantinos [77], it was concluded that small kernels reduced both the element-wise multiplications and the number of trainable parameters, as well.
6.3.4 Multiscale parallel convolutional pathways
In order to incorporate both local and larger contextual information into the proposed 3D-CNN, a second parallel pathway was added. This operated on down-sampled images, thus, the 3D-CNN simultaneously processed the input image at multiple scales. Higher level features such as the location within the brain were learnt in the second pathway, while the detailed local appearance were learnt in the first pathway. The size of the pathways could be adjusted based on the existing computational capacity.
6.3.5 Comments
Deeper network variants that are more efficient can be designed by simply replacing each layer of common architectures with more layers that use smaller kernels. However, deeper networks are difficult to train. From my point of view, the use of 3D CNNs was efficient. It opened an area to future research in medical image analysis of 3D volumetric data.
7 Discussion and observations
From the previous sections, it could be concluded summarized the main highlights in Table 1. The aim is to describe in a deeper way some of the highly recognized efforts in MI detection, diagnosis, and segmentation using DNNs. Table 2 summarizes the challenges and their relevant solutions. The table presents six major challenges we faced while surveying hundreds of papers in the area, relevant solutions are given citing one of the most comprehensive research that solves such problem.
8 Deep neural networks implementation
The most well-known software frameworks in the past few years includes: Caffe, MXNet, Tensorflow, MatConvNet, Torch, and Theano. Caffe stands for “Convolutional Architecture for Fast Feature Embedding” [80]. MXNet, “Mix and Maximize Networks” [81], is a high-performance deep learning library with many systems-level design decisions. Tensorflow where its name is derived from the operations that such neural networks perform on multidimensional data arrays referred to as “tensors” [82], and “MATLAB Convolution Networks” (MatConvNet) [83] are two important frameworks. Torch [84] and Theano [85] could be classified as the least used DL tools nowadays. Tables 3 and 4 summarize a comparative study for well-known DNN frameworks.
Choosing the correct hardware for DL depends on the learning problem, the throughput requirements, and the available cost. Special hardware design and architectures have significantly increased the efficiency of DNNs for medical applications: development of graphical processing units (GPUs) and progress in distributed systems. GPUs play an important role in DL because of their effective highly parallel processing structure for both learning and inference algorithms. The typical application requires a host computer with a GPU board installed. Each GPU core includes tens of arithmetic logic units (ALUs). In CNN, large amounts of neurons will be processed by the same instructions at each layer [86]. Since the performance of a single GPU is not sufficient to manage large-scale deep learning applications, it is quite common to parallelize processing tasks across multiple GPUs. Distributed computing is an efficient parallel solution to increase the DL performance by exploiting more distributed resources [87].
Although GPU processing has solved most computational challenges in the area of medical image processing, the GPU efficiency is still around 20% of the maximum performance [88 and 89]. Both memory bandwidth and capacity have a great effect on training, validation, and testing performances [90]. To explain this problem, all network parameters are distributed toward layers (a sizeable amount of data that makes the network layers exhibit an incremental amount of data. The main problem with increasing this efficiency is related to the high bandwidth stacked memory [91 and 92]. Different approaches based on FPGA, GPU, and CPU are listed accordingly in Table 5. This table presents a comparison between different devices to facilitate the tradeoffs while choosing an approach for configuring your system designed using FPGA, GPU, or CPU devices. To conclude, each has its corresponding strong and weak areas, which means that still there are no clear one-size-fits-all solutions. It is all according to the application [93].
9 Future of CNNs in medical image computing
9.1 Summary of existing well-known CNN structures
With the increased performance from GPU achievement and big data, CNN researches experienced breakthroughs. One of the most classical CNN structures is AlexNet. AlexNet [58] when introduced used dual-GPU training process then moved to a single GPU with eight deep layers as a result of the advance in GPU computation. AlexNet is considered the root for several CNN structures. VGGNet comes as an upgraded CNN developed by Simonyan and Zisserman [57]. It uses repeatedly stacking convolutional layers and maximum pooling layer. This widely used network to extract image features using a number of 16 to 19 CNN layers. The innovation of VGGNet to extract image features is due to the use of a 3 × 3 convolution and 2 × 2 pooling kernels, respectively.
Utilizing more deep layers leads to negative effects: overfitting, gradient disappearance, or gradient explosion. GoogleNet [94] proposes another way which for a more efficient computation time, i.e., extracting more features with the same computation amount. The structure consists of several cascaded modules. This idea indicates that when two convolutions are put in series, more nonlinear features could be combined. Using a 1 × 1 convolution reduces the dimensionality, which in turns decreases the computational complexity.
ResNet was proposed by He, Zhang [95] in 2015. It utilizes residual units. ResNet trains a 152 layer CNN, and achieves the best result with a minimal number of parameters compared to VGGNet. ResNet structure is able to speed up the training process of the DNN with higher accuracy. If the DNN depth is continuously increasing, a degradation problem might occur, i.e., the accuracy rise, reach saturation, then decline.
UNet is a classical CNN with a U-shaped structure that is able to capture semantic information during down-sampling. The main advantage of this structure is its ability to be trained with a small number of images based on sliding Windows [96]. One of the main limitations of U-Net is that it uses SoftMax cross entropy loss to deal with the problem with medical images with similar target boundaries. Solutions suggested adding weights to each pixel while calculating the objective function for the network to be more able to specify boundaries.
The region-based CNN or the R-CNN [97] starts by extracting regions of interest from input images and warp them to a fixed size images. It aims focusing on possible target locations. These normalized regions are entered into the CNN to extract features. SVM is applied as a classifier to identify features with linear regression. Using low and high quality regions, R-CNN performs better than the traditional sliding window from accuracy point of view. The R-CNN is time-consuming due to repeated computations. Moreover, it takes larger memory size. Other versions have been implemented such as fast and faster R-CNN [98].
YOLO [99] CNN algorithm could be considered as a one-stage target detection algorithm. The main contributions within the YOLO are its high speed, less background errors, and good generalization performance. However, YOLO has a reduced performance in target positioning process leading to low detection accuracy.
SDD [100] is as an extended version of YOLO, as YOLO uses full-image features while SSD predicts locations by means of features nearby that location. SSD considers different scales in different image feature’s layers. SSD outputs a series of discrete boxes representing feature maps of different layers and different aspect ratios, a method that resemble multi0scale analysis.
9.2 Future CNN trends in MI applications
The main problem affecting the accuracy of DNNs applied for MI analysis is the amount of labeled data used for training. Due to the lack of available labeled medical data sets, recently, some researchers proposed several directions to overcome such problem. One practical image preprocessing stage was explained in Sect. 4.1, data augmentation. Simple augmentation techniques such as cropping, rotating, and flipping succeeded to generate new abnormal images.
On the other hand, CNN could be combined with transfer learning [101]. Transfer learning is a research technique that stores knowledge gained while solving a MI problem in an organ (e.g., brain tumor detection) and applies such knowledge into a different organ (e.g., lung tumor detection). The idea concerns with using the CNNs obtained parameters in the first application to train the second one. Integrating transfer learning into CNNs could be considered as an important future research direction that could solve the limiting number of labeled medical data.
Moreover, another possible idea for data set increment is to introduce the crowdsource mechanism [102]. Crowdsourcing for health challenges means sharing solutions (trained structures) from one research team to a group of people (public). This is also an interesting future research direction that shifts individual tasks to public tasks generating public benefit. Some unsupervised or semi-supervised learning methods could be used to deal with inconsistent training data [78 and 103]. The main difference between both methods is that the former works independently without any labeled data, while the latter needs to incorporate labeled data (small amount) with unlabeled data (large amount). Semi-supervised learning showed adequate results for a few medical areas, but still needs more and more efforts in the future.
Future CNN comprises three main trends: Pre-trained, frozen, and multimodal CNNs. Those could be summarized as:
Pre-trained Models: The availability of pre-trained networks to learn a complex model using data from a source with large-scale annotated images will be the future of DNNs when only a small number of annotated images are available.
Frozen Deep Networks: Reducing the number of learning parameters in the DNN could be achieved using freezing few of network layers to constant parameter values, those parameter values are directly learnt from other networks trained on similar tasks. The rest of the network that now has less parameters can then be trained for the target task as normal [104].
Multimodal Images: Learning from multi sources can give a milestone regarding in-depth understanding and thus error-free decisions [105]. Multimodal deep machine learning will be a multi-disciplinary field with big potential in the next decade, as it could acquire different source of images and combine them to reach a decision.
10 Conclusions
A comprehensive study of recent DNN techniques applied in Medical Imaging was introduced. Such techniques were classified according to either clinical or pathological analysis and according to image processing areas (classification – detection – localization – segmentation- and diagnosis). Both supervised and unsupervised learning DNNs were examined. On the other hand, different imaging technologies: X-Ray, MRI, CT, Ultrasound, PET, Fluorescene Angiography, and even photographic images were used. From the presented work, it could be concluded that DNNs are highly flexible modeling approaches that learn a comprehensive representation of the input data through optimizing a loss function to find millions of network weights. The CNN represents the largest percentage of published researches in this area for many reasons. CNN emphasizes image features which are extremely important for medical image analysis. Furthermore, it has a well-known architecture and many pre-designed networks could be found within related software frameworks. Finally, pre-trained CNNs are found in different environments and for several applications. Regarding DL implementation, it would be efficient to use GPUs for DNNs training due to their significant speed. However, for tasks like inference, it is usually believed that CPUs are sufficient and are more attractive due to their cost savings except when inference speed is important (real-time applications).
Concerning training and DNNs architecture, a huge number of training samples is needed and thus data augmentation is presented. Data augmentation could be achieved through cropping, rotation, and translation. Using image patches and segments could be another way. Another way to boost training samples is to use an open image database, especially in case of unsupervised methods such as RBM. However, CNN and RNN are supervised methods and require annotated data or manual labeling. Regarding DNN architecture, deeper networks require tremendous training time and may lead to overfitting while smaller networks could sometimes never converge and give unacceptable accuracies. Thus, it is of great importance to pre-train the network several times, using multi architectures before inference phase. CNN structures encountered several schemes starting from multilayers networks, moving toward complex structures such as dual-pathways and cascaded networks. The latter could be considered efficient solutions to study local and global features and thus increase the overall testing accuracy. To conclude, DL is becoming widespread, and will continue to grow in the near future in all fields of medical science.
Availability of data and material
N/A.
Code availability
N/A.
References
The Age of Intelligent Machines, Kurzweil, Ray, Cambridge, MA: MIT Press, 1990
Russell S, Norvig Artificial intelligence: a modern approach, Prentice Hall.
Artificial Intelligence: How knowledge is created, transferred, and used," Elsevier, Scopus Report https://www.elsevier.com/__data/assets/pdf_file/0010/823654/ACAD-RL-AS-RE-ai-report-WEB.pdf
Nath V, Levinson S (2014) Autonomous robotics and deep learning, ISBN: 978–3–319–05603–6, Springer, 2014.
Introduction to Machine Learning, Ethem Alpaydin, 3rd edition, MIT Press, 2015.
Lee J-G et al (2017) Deep learning in medical imaging: general overview. Korean J Radiol 18(4):570–584
Deng L, Dong Y (2014) Deep learning: methods and applications. Found Trends® Signal Process 7(3–4):197–387
Patterson J, Gibson A, O'Reilly (2017) Deep learning, Media- USA, 1st edition, 2017.
Ball JE, Anderson DT, Chan CS (2017) Comprehensive survey of deep learning in remote sensing: theories, tools, and challenges for the community. J Appl Remote Sens 11(4):042609
Donghwoon K et al (2017) "A survey of deep learning-based network anomaly detection." Cluster Computing, Springer, pp 1–13, https://doi.org/10.1007/s10586-017-1117-8.
Russakovsky O, Deng J, Su H, Krause J, Satheesh S, Ma S, Huang Z, Karpathy A, Khosla A, Bernstein M (2015) Imagenet large scale visual recognition challenge. Int J Comput Vision 115(3):211–252
Gulcehre C (2016) Deep learning software Links. http://deeplearning.net/software_links/. Accessed Feb 2019.
Wang X, Lu L, Shin H, Kim L, Bagheri M, Nogues I, Yao J, Summers RM (2017) “Unsupervised joint mining of deep features and image labels for large-scale radiology image annotation and scene recognition,” IEEE Winter Conf Appl Comput Vis (WACV), pp 998–1007.
Zhang X, Liu W, Dundar M, Badve S, Zhang S (2015) Towards large-scale histopathological image analysis: hashing-based image retrieval. IEEE Trans Med Imag 34(2):496–506
Zhang X, Xing F, Su H, Yang L, Zhang S (2015) High-throughput histopathological image analysis via robust cell segmentation and hashing. J Med Image Anal 26(1):306–315
Dhungel N, Carneiro G, Bradley AP (2015) Deep learning and structured prediction for the segmentation of mass in mammograms. Medical image computing and computer-assisted intervention–MICCAI 2015. Springer, Cham, pp 605–612
Dubrovina A, Kisilev P, Ginsburg B, Hashoul S, Kimmel R (2016) Computational mammography using deep neural networks. In: Workshop on deep learning in medical image analysis (DLMIA).
Zheng Y (2015) Model based 3D cardiac image segmentation with marginal space learning. Medical image recognition, segmentation and parsing: methods, theories and applications. Elsevier, Amsterdam, pp 383–404
Avendi MR, Kheirkhah A, Jafarkhani H (2016) A combined deep-learning and deformable model approach to fully automatic segmentation of the left ventricle in cardiac MRI. Med Image Anal 30:108–119
Zhen X, Wang Z, Islam A, Bhaduri M, Chan I, Li S (2016) Multi-scale deep networks and regression forests for direct bi-ventricular volume estimation. Med Image Anal 30:120–129
Wang S, Yin Y, Cao G, Wei B, Zheng Y, Yang G (2015) Hierarchical retinal blood vessel segmentation based on feature and ensemble learning. Neruocomputing 149:708–717
Li Q, Feng B, Xie L, Liang P, Zhang H, Wang T (2016) A cross-modality learning approach for vessel segmentation in retinal images. IEEE Trans Med Imag 35(1):109–118
Wu A, Xu Z, Gao M, Buty M, Mollura DJ (2016) Deep vessel tracking: a generalized probabilistic approach via deep learning. In: Proceedings of IEEE international symposium on biomedical, imaging, pp 1363–1367.
Xing F, Yang L (2016) Robust nucleus/cell detection and segmentation in digital pathology and microscopy images: a comprehensive review. IEEE Rev Biomed Eng 9:234–263
Kaggle B (2017). Kaggle Data Science Bowl (2017) [Online] https: //www.kaggle.com/c/data-science-bowl-2017
Shin H-C et al (2016) Deep convolutional neural networks for computer aided detection: CNN architectures, dataset characteristics and transfer learning. IEEE Trans Med Imag 35(5):1285–1298
Yang X et al. (2016) A deep learning approach for tumor tissue image classification. In: Proc. Int. Conf. Biomed. Eng., Calgary, Canada [Online] /https://doi.org/10.2316/P.2016.832-025.
Shin H-C, Orton MR, Collins DJ, Doran SJ, Leach MO (2013) Stacked autoencoders for unsupervised feature learning and multiple organ detection in a pilot study using 4D patient data. IEEE Trans Pattern Anal Mach Intell 35(8):1930–1943
Yao J, Li J, Summers RM (2009) Employing topographical height map in colonic polyp measurement and false positive reduction. Pattern Recogn 42(6):1029–1040
Roth HR et al (2016) Improving computer aided detection using convolutional neural networks and random view aggregation’’. IEEE Transact Med Imag 35(5):1170–1181
Roth H, Lu L et al (2017) Efficient false positive reduction in computer aided detection using convolutional neural networks and random view aggregation. In: Le L, Zheng Y, Carneiro G, Yang L (eds) Deep learning and convolutional neural networks for medical image computing. Springer, Cham
Ker J et al (2018) Deep learning applications in medical image analysis. IEEE Access 6:9375–9389
Veta M, Pluim J, van Diest P, Viergever M (2014) Breast cancer histopathology image analysis: a review. IEEE Trans Biomed Eng 61(5):1400–1411
Kuse M, Wang Y-F, Kalasannavar V, Khan M, Rajpoot N (2011) Local isotropic phase symmetry measure for detection of beta cells and lymphocytes. J Pathol Inform 2(2):2
Al-Kofahi Y, Lassoued W, Lee W, Roysam B (2010) Improved automatic detection and segmentation of cell nuclei in histopathology images. IEEE Trans Biomed Eng 57(4):841–852
Vink JP, Van Leeuwen M, Van Deurzen C, De Haan G (2013) Efficient nucleus detector in histopathology images. J Microsc 249(2):124–135
Ali S, Madabhushi A (2012) An integrated region-, boundary-, shape-based active contour for multiple object overlap resolution in histological imagery. IEEE Trans Med Imag 31(7):1448–1460
Cruz-Roa AA, Ovalle JEA, Madabhushi A, Osorio FAG (2013) A Deep learning architecture for image representation, visual interpretability and automated basal-cell carcinoma cancer detection, in Medical image computing and computer-assisted intervention–MICCAI, Springer, pp. 403–410.
Chao L, Xinggang W, Wenyu L, Longin L (2018) Deep mitosi: mitosis detection via deep detection, verification, and segmentation. Med Image Anal 45:121–133
MITOS-ATYPIA-14 (2014) Mitos-Atypia-14-dataset. https://mitos-atypia-14.grand-challenge.orgldataset/Online; accessed 03.03.2018.
Li W, Li J (2018) Local deep field for electrocardiogram beat classification. IEEE Sens J 18(4):1656–1664
Shehata M et al (2019) Computer-aided diagnostic system for early detection of acute renal transplant rejection using diffusion-weighted MRI. IEEE Trans Biomed Eng 66(2):539–552
Hamilton PW, Bankhead P, Wang YH, Hutchinson R, Kieran D, McArt DG, James J, SaltoTellez M (2014) Digital pathology and image analysis in tissue biomarker research. Methods 70(1):59–73
Rimm DL (2011) C-path: awatson-like visit to the pathology lab. Sci Trans Med 3:108
Duraisamy S, Emperumal S (2017) Computer-aided mammogram diagnosis system using deep learning convolutional fully complex-valued relaxation neural network classifier. Sci Trans Med 11(8):656–662
American Cancer Society. Cancer facts & figures 2016. Atlanta, American Cancer Society 2016. http://www.cancer.org/acs/groups/content/@research/documents/document/acspc-047079.pdf.
LeCun Y, Bengio Y, Hinton G (2015) Deep learning. Nature 521:436–444
Gutman D et al (2016) Skin lesion analysis toward melanoma detection. In: International symposium on biomedical imaging (ISBI), (International Skin Imaging Collaboration (ISIC), 2016).
Esteva A et al (2017) Dermatologist-level classification of skin cancer with deep neural networks. Nature 542:115–126
Ramlakhan K, Shang Y (2011) A mobile automated skin lesion classification system. In: 23rd IEEE international conference on tools with artificial intelligence (ICTAI-2011), pp. 138–141.
Celebi M, Schaefer G (2013) Color Medical Image Analysis Springer, pp 63–86.
B. T. Society (1999) The diagnosis, assessment and treatment of diffuse parenchymal lung disease in adults. Thorax, 54 (1).
Anthimopoulos M, Christodoulidis S, Christe A, Mougiakakou S (2014) Classification of interstitial lung disease patterns using local DCT features and random forest. In Proc. 36th Annual Int. Conf. IEEE Eng. Med. Biol. Soc., pp 6040–6043.
Li Q, Cai W, Feng DD (2014) Lung image patch classification with automatic feature learning. In Proc. 36th Annual Int. Conf. IEEE Eng. Med. Biol. Soc., pp 6079–6082.
Li Q et al (2014) Medical image classification with convolutional neural network. In: Proc. 13th Int. Conf. Control Automat. Robot. Vis., pp 844–848.
Anthimopoulos M, Ebner L, Christe A, Mougiakakou S (2016) Lung pattern classification for interstitial lung diseases using a deep convolutional neural network. IEEE Trans Med Imag 35(5):1207–1216
Simonyan K, Zisserman A (2015) Very deep convolutional networks for large-scale image recognition. In: International conference learning representation, San Diego, USA.
Krizhevsky A, Sutskever I, and Hinton G (2012) ImageNet classification with deep convolutional neural networks, Advanced Neural Inference Processing Systems.
https://www.alzheimers.net/resources/alzheimers-statistics/ last accessed date: 1st March, 2019
Suk HI, Lee SW, Shen DG (2014) Hierarchical feature representation and multimodal fusion with deep learning for AD/MCI diagnosis. Neuroimage 101:569–582
Hosseini-Asl E, Keynton R, El-Baz A (2016) Alzheimer’s disease diagnostics by adaptation of 3D convolutional network. In: Proc. 2016 IEEE Int. Conf. Image Processing (ICIP), Phoenix, AZ, USA, pp 126–130.
Liu J, Pan Y, Li M, Chen Z, Tang L, Wang J (2018) Applications of deep learning to MRI images: a survey. Big Data Mining and Analytics 1(1):1–18
Zikic D, Y. Ioannou Y, Criminisi A, Brown M (2014) Segmentation of brain tumor tissues with convolutional neural networks. In: Proceedings MICCAI Workshop on Multimodal Brain Tumor Segmentation Challenge, Boston, USA, pp 36–39
Havaei M, Davy A, Warde-Farley D, Biard A, Courville A, Bengio Y, Pal C, Jodoin PM, Larochelle H (2017) Brain tumor segmentation with deep neural networks. Med Image Anal 35:18–31
Pereira S, Pinto A, Alves V, Silva CA (2016) Braintumor segmentation using convolutional neural networks in MRI images. IEEE Trans Med Imag 35(5):1240–1251
Kleesiek J, Urban G, Hubert A, Schwarz D, MaierHein K, Bendszus M, Biller A (2016) Deep MRI brain extraction: a 3D convolutional neural network for skull stripping. Neuroimage 129:460–469
Ngo TA, Lu Z, Carneiro G (2017) Combining deep learning and level set for the automated segmentation of the left ventricle of the heart from cardiac cine magnetic resonance. Med Image Anal 35:159–171
Havaei M et al (2017) Brain tumor segmentation with deep neural networks. Med Image Anal 35:18–31
Davy A, Havaei M, Warde-Farley D, Biard A, Tran L, Jodoin PM, Courville A, Larochelle H, Pal C, Bengio Y (2014) Brain tumor segmentation with deep neural networks. In: Proc. of BRATS-MICCAI.
Zikic D, Ioannou Y, Brown M, Criminisi A (2014) Segmentation of brain tumor tissues with convolutional neural networks. In: Proc. of BRATS-MICCAI.
Urban G, Bendszus M, Hamprecht F, Kleesiek J (2014) Multi-modal brain tumor segmentation using deep convolutional neural networks. In: Proc. of BRATS-MICCAI.
Goodfellow I.J. et al. (2013) Pylearn2: a machine learning research library. arXiv preprint arXiv:1308.4214
Rajchl M et al (2017) DeepCut: Object segmentation from bounding box annotations using convolutional neural networks. IEEE Trans Med Imag 36(2):674–683
Wang G et al (2018) Interactive medical image segmentation using deep learning with image specific fine tuning. IEEE Trans Med Imag 37(7):1562–1572
Menze BH et al (2015) The multimodal brain tumor image segmentation benchmark (BRATS). IEEE Trans Med Imag 34(10):1993–2024
Roth HR et al (2014) A new representation for lymph node detection using random sets of deep convolutional neural network observations. In: Golland P, Hata N, Barillot C, Hornegger J, Howe R (eds) Medical Image Computing and Computer Assisted Intervention – MICCAI. Springer, Cham, pp 520–527
Kostantinos K et al (2017) Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation. Medical Image Analysis, Elsevier, Amsterdam
Sun W, Tseng TB, Zhang J, Qian W (2017) Enhancing deep convolutional neural network scheme for breast cancer diagnosis with unlabeled data. Comput Med Imag Graph 57:4–9
Sajjad M, Khan S, Muhammad K, Wu W, Ullah A, Baik SW (2019) Multi-grade brain tumor classification using deep CNN with extensive data augmentation. J Comput Sci 30:174–182
Jia Y et al (2014) Caffe: convolutional architecture for fast feature embedding. In: Proceedings of the 22nd ACM international conference on Multimedia.
Tianqi C et al (2015) MXNet: a flexible and efficient machine learning library for heterogeneous distributed systems, In: Neural information processing systems, workshop on machine learning systems.
Abadi M et al. (2016) Tensorflow: Large-scale machine learning on heterogeneous distributed systems, arXiv preprint arXiv:1603.04467.
Vedaldi A, Lenc K (2015) MatConvNet: convolutional neural networks for MATLAB. In Proc. 23rd ACM Int. Conf. Multimedia, Brisbane, Australia, pp 689–692.
Collobert R, Kavukcuoglu K, Farabet C (2011) Torch7: a matlab-like environment for machine learning. In BigLearn, NIPS Workshop, No. EPFL-CONF-192376.
Bergstra J, Breuleux O, Bastien F, Lamblin P, Pascanu R, Desjardins G, Turian J, Warde-Farley D, Bengio Y (2010). Theano: a CPU and GPU math expression compiler. In: Proceedings of the python for scientific computing conference (SciPy).
Ota K et al (2017) Deep learning for mobile multimedia: a survey. ACM Trans Multimed Comput Commun Appl 34:1–32
Choi W et al (2016) Hybrid network-on-chip architectures for accelerating deep learning kernels on heterogeneous many core platforms. In: Proceeding of compliers, architectures, and sythesis of embedded systems (CASES), 2016 IEEE International Conference.
Zhang J, Yeung SH, Shu Y, He B, Wang W. (2019) Efficient memory management for GPU-based deep learning systems; arXiv preprint arXiv:1903.06631.
Zhao H, Han Z, Yang Z, Zhang Q, Yang F, Zhou L, Yang M, Lau FC, Wang Y, Xiong Y, et al. Hived (2020) sharing a {GPU} cluster for deep learning with guarantees, 14th USENIX symposium on operating systems design and implementation (OSDI 20), pp 515–532.
Lin Y, Jiang Z, Gu J, Li W, Dhar S, Ren H, Khailany B, Pan DZ (2020) Dream place eep learning toolkit-enabled GPU acceleration for modern VLSI placemen. IEEE Trans Comput Aid Des Integr Circuits Syst 40:748–61
Hossain S, Lee DJ (2019) Deep learning-based real-time multiple-object detection and tracking from aerial imagery via a flying robot with GPU-based embedded devices. Sensors 19(15):3371
Castro FM, Guil N, Marin-Jimenez MJ, Perez-Serrano J, Ujaldon M (2019) Energy-based tuning of convolutional neural networks on multi-GPUs. Concurr Comput Pract Exp 31(21):4786
Alzubaidi et al (2021) Review of deep learning: concepts, CNN architectures, challenges, applications, future directions. J Big Data 8:53. https://doi.org/10.1186/s40537-021-00444-8
Szegedy C et al (2017) Inception-v4, inception-resnet and the impact of residual connections on learning, Thirty-first AAAI conference on artificial intelligence
He K et al (2016) Identity mappings in deep residual networks. Springer, European conference on computer vision
Ronneberger O, Fischer P, Brox T (2015) U-net: Convolutional networks for biomedical image segmentation, International CONFERENCE ON MEDICAL image computing and computer-assisted intervention, Springer.
Gkioxari G, Girshick R, Malik J (2015) Contextual action recognition with R-CNN, Proceedings of the IEEE international conference on computer vision.
Ren S et al (2015) Faster R-CNN: towards real-time object detection with region proposal networks, NIPS'15: Proceedings of the 28th international conference on neural information processing systems, Vol 1, pp 91–99.
Du J (2018) Understanding of object detection based on CNN family and YOLO. J Phys: Conf Ser 1004:012029
Liu W et al (2016) SSD: single shot multibox detector. Springer, European conference on computer vision
Khan SU, Islam N, Jan Z, Din IU, Rodrigues JJPC (2019) A novel deep learning based framework for the detection and classification of breast cancer using transfer learning. Pattern Recogn Lett 125:1–6
Zhang Q, Yang LT, Chen Z, Li P, Bu F (2018) An adaptive dropout deep computation model for industrial IoT big data learning with crowdsourcing to cloud computing. IEEE Trans Ind Inf 15(4):2330–2337
Hang Yu, Yang LT, Zhang Q, David Armstrong M, Deen J (2021) Convolutional neural networks for medical image analysis: state-of-theart, comparisons, improvement and perspectives. Neurocomputing 444:92–110
Altaf F et al (2019) Going deep in medical image analysis: concepts, methods, challenges and future directions. IEEE Access 7:99540
Baltrusaitis T, Ahuja C, Morency L (2019) Multimodal machine learning: a survey and taxonomy. IEEE Trans Pattern Anal Mach Learn 41(2):423–443
Funding
N/A.
Author information
Authors and Affiliations
Contributions
N/A.
Corresponding author
Ethics declarations
Conflict of interest
The author declares that there is not conflicts of interest.
Ethical approval
N/A.
Consent to participate
The author has consented to the submission of this survey to the journal.
Consent for publication
The author has consented to the publication of this survey to the journal.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Abdou, M.A. Literature review: efficient deep neural networks techniques for medical image analysis. Neural Comput & Applic 34, 5791–5812 (2022). https://doi.org/10.1007/s00521-022-06960-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00521-022-06960-9