Abstract
Nuclear export signal (NES) is a nuclear targeting signal within cargo proteins, which is involved in signal transduction and cell cycle regulation. NES is believed to be “born to be weak”; hence, it is a challenge in computational biology to identify it from high-throughput data of amino acid sequences. This work endeavors to tackle the challenge by proposing a computational approach to identifying NES using spiking neural P (SN P) systems. Specifically, secondary structure elements of 30 experimentally verified NES are randomly selected for training an SN P system, and then 1224 amino acid sequences (containing 1015 regular amino acid sequences and 209 experimentally verified NES) abstracted from 221 NES-containing protein sequences randomly in NESdb are selected to test our method. Experimental results show that our method achieves a precision rate 75.41 %, better than NES-REBS 47.2 %, Wregex 25.4 %, ELM, and NetNES 37.4 %. The results of this study are promising in terms of the fact that it is the first feasible attempt to use SN P systems in computational biology after many theoretical advancements.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
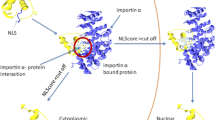
Dynamic trafficking of proteins from cell nucleus to cytoplasm mostly depends on transport factors in the Karyopherin-\(\beta\) family, which are named importins and exportins [1–3]. The direction of nuclear–cytoplasmic transport is mainly mediated by the targeting cell signal within cargo proteins, that is, nuclear localization signal and nuclear export signal [4, 5]. Nuclear export signals (NESs) are nuclear targeting signals within the cargo proteins, which is composed of four main hydrophobic residues that targets it for export from cell nucleus to cytoplasm through nuclear pore complex [6]. Since NESs were first identified in proteins HIV-1 Rev and cyclical AMP-dependent protein kinase inhibitor, many other NESs have been experimentally identified in more than 200 proteins, such as translation factors [7], cell cycle regulators [8], transcription factors [9] and viral proteins [10]. A well-known NES is the leucine-rich NES, which mediates binding to the receptor of the karyopherin exportin 1/chromosomal region maintenance 1 (CRM1), which has important application in replication of plenty of viruses that might cause human diseases [11–13].
Finding NES in cellular proteins is a challenging but important problem, and many efforts in cellular biology have been made to detect NESs. Experimental identification of NES-containing proteins has been an effective method to find NESs, but it is likely to take a lot of resources, time and effort to perform experiments on false NES from large-scale candidate protein sequences. In experiments for identifying NES-containing proteins, it is necessary to do cell culture and treatments, plasmid construction and some other tedious steps. It needs about 3 days in laboratory to determine whether an amino acid sequence is a NES or not. If a large number of NESs are planned to be detected experimentally, the cost of the experimental resources becomes huge, which is also very time-consuming [14, 15]. A possible way to solve this problem is developing computational approaches to predicting or identifying NES from protein sequences, which paves the way for a hot and promising research branch in Bioinformatics [16–18]. Among computational approaches, a general way is to construct prediction or identification models from particular biological features of NESs, including secondary structure [19], inner disorder [6], \(\alpha\)-helical structure [20], \(\alpha\)-helix-loop or all-loop structures binding in a hydrophobic groove on the convex surface [21, 22] and regular expressions [23]. In recent research, it is found that intelligent computing methods, such as machine learning-based strategy [24] and position-specific scoring matrix (PSSM [25]), performed well in predicting NESs from high-throughput data of amino acid sequences, with prediction rate around 60 %.
Neural networks are well-known computational models inspired by the central nervous system of animals. Spiking neural P systems are spiking neural-like computing models, which are inspired from the way the neurons’ spiking and communicating by means of spikes [26]. These systems are named SN P systems, for shortly. As a new candidate of spiking neural network models [27], SN P systems perform well in doing computation, that is, the systems and almost all of their variants can achieve Turing completeness. Notably, it is proved that SN P systems can generate and accept the set of Turing computable natural numbers [26], generate recursively enumerable languages [28] and compute the set of Turing computable functions [29]. Inspired by different biological phenomena and mathematical motivations, lots of variants of SN P systems have been proposed, such as SN P systems with anti-spikes [30, 31], SN P systems working in asynchronous mode [32], asynchronous SN P systems with local synchronized sets of neurons [33], SN P systems with astrocyte-like control [34], SN P systems with request rules [35, 36], homogeneous SN P systems [37, 38], sequential SN P systems [39, 40] and SN P systems with rules on synapses [41–43]. For applications, SN P systems are used to design logic gates, logic circuits [44] and operating systems [45], perform basic arithmetic operations [46], solve combinatorial optimization problems [47] and diagnose fault of electric power systems [48–50]. SN P systems with neuron division, budding or separation can generate exponential space with space–time trade-off strategy, thus providing a way to theoretically solve computationally hard problems in a feasible (polynomial or linear) time [51–54]. There are some significant simulators for SN P systems, see, for example, [55–57].
In artificial neural networks, a sigmoid function is used to imitate biological neuron’s spiking, while in SN P systems, spiking rules are used, denoted in the form of production in grammar of formal languages, to describe the neuron’s spiking behavior. The applications of spiking rules are controlled by the number of spikes contained in the neuron at certain moment, which determine the triggering conditions. A neuron may contain multiple rules, in the sense of having ability to select spiking conditions, and can send out different numbers of spikes by consuming different numbers of spikes.
In this work, we propose a computational approach to identify NES using SN P systems. Specifically, 30 experimentally verified NES which have the unique biological feature secondary structure elements are randomly selected for training the SN P system. Subsequently, 1224 amino acid sequences, composed of 1015 regular amino acid sequences and 209 experimentally verified NES, are selected from 221 NES-containing protein sequences randomly in NESdb [13] to test our method. The experimental results show that our method achieves a precision rate of 74.18 %, which performs better than NES-REBS with a precision rate of 47.2 % [58], Wregex with a precision rate of 25.4 % [59], ELM with a precision rate of 33.5 % [23] and NetNES with a precision rate of 37.4 % [60]. The results of this study are promising in terms of the fact that it is the first feasible attempt to use spiking neural P systems in computational biology after many theoretical advancements.
2 Spiking neural P system with Hebbian learning strategy
It is useful for readers to have some familiarity with basic concepts and notions in SN P systems [26, 61].
The formal definition of SN P system of degree \(m\ge 1\) is a construct of the form from [26]:
-
\(O=\{a\}\) is a singleton alphabet and a is called spike;
-
\(\sigma _1,\sigma _2,\ldots ,\sigma _m\) are neurons of the form \(\sigma _i=(n_i,R_i)\), with \(1\le i\le m\), where \(n_i\) is initial number of spikes in neuron \(\sigma _i\) and \(R_i\) is the set of rules in neuron \(\sigma _i\):
-
1.
spiking rule: \(E/a^c\rightarrow a^p\), where E is a regular expression over O, c the number of spikes to be consumed and \(c\ge p\ge 1\);
-
2.
forgetting rule: \(a^s\rightarrow \lambda\), with the restriction that \(a^s\notin L(E)\) for any spiking rule;
-
1.
-
\(syn\subseteq \{1,2,\dots ,m\}\times \{1,2,\dots ,m\}\) with \((i,i)\notin syn\) is the set of synapses between neurons;
-
\(i_{in}\) indicates the input neuron that reads spikes from the environment, and \(i_{out}\) indicates the output neuron that can also emit spikes into the environment.
A spiking rule is a rule of the form \(E/a^c\rightarrow a\). At certain moment, when neuron \(\sigma _i\) holds k spikes such that \(a^k\in L(E)\), \(k\ge c\), then spiking rule \(E/a^c\rightarrow a\) is applied. This means that c spikes are consumed (\(k-c\) spikes remaining in neuron \(\sigma _i\)) and neuron \(\sigma _i\) fires, producing one spike. This spike will be emitted from neuron \(\sigma _i\) to each of its neighboring neurons (a global clock is assumed, marking the time for the whole system; hence, the functioning of the neurons is synchronized). The applicability of a rule is controlled by the total number of spikes accumulated in the neuron. The work of each neuron is sequential: Only one rule can be applied in each time unit, but different neurons work in a parallel manner.
Forgetting rules are of the form \(a^s\rightarrow \lambda\) with \(s\ge 1\) and are applied only if the neuron contains exactly s spikes. By applying the forgetting rule, s spikes will be removed from the neuron and thus out of the system. We have the limitation that when a forgetting rule is used at a computation step, no spiking rule is enabled. It means that in any neuron, if a spiking rule is applicable, then no forgetting rule is enabled to use, and vice versa.
The study of incorporating Hebbian learning to SN P systems was initialed in [62], where a theoretical Hebbian learning SN P system model was developed. The systems were designed in a formal way to update their information, but no application was proposed. We construct here an SN P system with Hebbian learning strategy to identify nuclear export signals, where the systems are different from the ones in [62], but having the common biological facts.
The system will be given graphically by a directed graph, where rounded rectangles with the initial number of spikes and rules are used to represent neurons and edges represent the synapses. Input neurons have inputting synapses, by which they can read spikes from the environment; output neurons have outgoing synapses to emit spikes into the environment. The system consists of two modules, the input module and the predict module.
2.1 The input module
The input module is composed of an input neuron (reading spike trains from the environment), a trigger neuron (starting the module), 75 “transmitting” neurons labeled with \(A_1, A_2, \dots , A_{75}\) and 75 “gathering” neurons labeled with \(B_1, B_2,\) \(\dots , B_{75}\). The topological structure of the input module is shown in Fig. 1.
-
The input neuron has no initial spike inside and the unique spiking rule \(a\rightarrow a\). With the spiking rule at any moment, when the nput neuron has one spike inside, it fires with using spiking rule \(a\rightarrow a\), emitting one spike to its neighboring neurons (the ones having synapses pointing from the neuron Input). The function of the input neuron is to read spike trains (in the form of binary strings) from the environment one by one bit. The input neuron reads spike train as follows. Let \(w=w_1w_2\dots w_{75}\) is the spike train to be read with \(w_i\in \{0,1\}\), and at certain moment t the input neuron starts to read it. In each step, the input neuron reads one bit of spike train w. At any step \(t+p\), if \(w_p=1\), then the input neuron reads one spike from the environment; otherwise, the input neuron reads no spike.
-
The trigger neuron is used to start the computation of the module. In the input module, all the neurons initially have no spike inside, with the exception that the trigger neuron contains one spike. With the spike, the trigger neuron can fire at the first step of the computation, consuming the initially contained spike and emitting one spike to its neighboring neuron. Neuron \(\sigma _{A_1}\) is the unique neighboring neuron of the trigger neuron.
-
The 75 “transmitting” neurons labeled with \(A_1, A_2, \dots , A_{75}\) have spiking rule \(a\rightarrow a\). For any \(2\le i\le 74\), when neuron \(\sigma _{A_i}\) receives one spike from neuron \(\sigma _{A_{i-1}}\), it fires by using the spiking rule \(a\rightarrow a\), emitting one spike to neuron \(\sigma _{A_{i+1}}\). When neuron \(\sigma _{A_{75}}\) fires, it sends one spike to neuron \(\sigma _{A_1}\), and a new circle is started.
-
The 75 “gathering” neurons labeled by \(B_1, B_2, \dots , B_{75}\) have spiking rule \(a^2\rightarrow a\) and forgetting rule \(a\rightarrow \lambda\). This means that when neuron \(\sigma _{B_i}\) has two spikes, it fires by using the spiking rule \(a^2\rightarrow a\) emitting one spike to neuron \(\sigma _{C_i}\); when neuron \(\sigma _{B_i}\) has one spike, the spike is deleted by using forgetting rule \(a\rightarrow \lambda\). Each neuron \(\sigma _{B_i}\) has a synapses from the input neuron and “transmitting” neuron \(\sigma _{A_i}\), respectively. This means only when both of the input neuron and “transmitting” neuron \(\sigma _{A_i}\) fire, each of them sends one spike to neuron \(\sigma _{B_i}\). Neuron \(\sigma _{B_i}\) accumulates two spikes and fires by using spiking rule \(a^2\rightarrow a\) to send one spike to neuron \(\sigma _{C_i}\).
2.2 The predict module
The predict module consists of 75 “processing neurons”, which are labeled by \(C_1, C_2, \dots , C_{75}\), and four output neurons \(\sigma _{Output_1},\sigma _{Output_2},\sigma _{Output_3},\sigma _{Output_4}\). All the neurons in the predict module have a unique spiking rule \(aa^*/a\rightarrow a\), and the weights of all the synapses are initially set to be 1. The spiking rule \(aa^*/a\rightarrow a\) can be used when neuron \(\sigma _{C_i}\) has any number of spikes. For each transition step, one spike is consumed and one spike is emitted out. For example, if neuron \(\sigma _{C_i}\) accumulates k spikes inside, then it will fire for k times and emits in total k spikes out, one spike in each time spiking. The 75 “processing neurons” are separated into three layers: the inner layer, hidden layer and outermost layer. The topological structure of the predict module is like a “ripple” with three layers as shown in Fig. 2. The spikes transmit from the inner layer to the hidden layer and then to the outmost layer.
The inner layer consists of 11 neurons, framed up in a red dashed line, which have connections to each neuron in the hidden layer, framed up a blue dashed line. The hidden layer has four subgroups, named top, bottom, leftward and rightward subgroups. The top and bottom subgroups have 3 neurons each, while the leftward and rightward subgroups have 11 neurons. The outermost layer, framed in green dashed lines, is composed of 36 neurons, which is divided into four subgroups as well. The top and bottom subgroups of the outermost layer have 5 neurons each, and the leftward and rightward subgroups have 13 neurons. In Fig. 3, it shows the involved neurons in each layer.
Each neuron in the top (resp. bottom, leftward, rightward) subgroup of the hidden layer has a synapse connection to every neuron in the top (resp. bottom, leftward, rightward) subgroup of outermost layer. Four neurons are used to collect information from the neurons from the four (top, bottom, leftward and rightward) subgroups, respectively. The result of a computation is a 4-dimensional vector recording the number of spikes emitted into the environment from the 4 output neurons.
2.3 The Hebbian learning strategy
Each “processing” neuron \(\sigma _{C_i}\) has a synapse connection with “gathering” neuron \(\sigma _{B_i}\). Neuron \(\sigma _{C_i}\) can receive spikes from “gathering” neuron \(\sigma _{B_i}\), when the system reads spike trains through the input neuron one by one bit. With the spikes inside, neuron \(\sigma _{C_i}\) can fire and send spikes to its neighboring neurons. At any moment, when a neuron sends spikes along certain synapse, a Hebbian learning strategy is imposed on the synapse in the predict module, that is, the weight of the synapse will be increased by an augmenter \(\Delta w\) for each time passing some spikes. In general, at any moment, if neuron \(\sigma _{C_i}\) has fired t times and passed spikes along its synapse for t times, then the weight of the synapse is \(1+t\times \Delta w\). Note that, the weights on the synapses among neurons from the input module are fixed during the computation.
The weight on a synapse is a function which amplifies the spikes passing along it. Specifically, if at some moment the weight on certain synapse is w and k spikes pass along it, then in total \(w*k\) spikes are received by the target neuron. The received spikes will be accumulated in the neuron. Mathematically, we can use the following Eq. 1 to calculate the weight at certain moment \(t+1\) of the synapse connecting neuron \(\sigma _{C_i}\) and \(\sigma _{C_j}\).
The weights on synapses connecting each pair of neurons \(\sigma _{B_i}\) and \(\sigma _{C_i}\) are updated with the Hebbian learning strategy, while the weights on synapses from the input module are always 1.
In the computation, the topological structure the input module does not change during the computation, but the topological structure of the prediction module can be modified by the updating strategy of weights on synapses.
3 Identification of NES by the SN P system
In this section, NES identification using the SN P system with Hebbian learning is presented. It starts by explaining the way to encode secondary structure of NES into binary sequences, and then, the training strategy and prediction processes are elaborated.
3.1 Encoding secondary structure of NES into binary sequence
The information that the SN P system can read are encoded in form of spike trains, i.e., binary sequences. Before we use the SN P system to identify NESs, it is necessary to encode the secondary structure of NES into a binary sequence.
The secondary structure of a NES is usually a loop conformation or helix-loop conformations starting with an \(\alpha\)-helix. We use the secondary structure prediction tool PSIPRED to calculate the secondary structure of NESs, by which the secondary structure of a NES can be described by a string of letters. Each letter has a specific meaning of the structure, such as H (alpha helix), B (residues in isolated beta-bridge), G (3-helix), S (bend), I (5 helix), T (hydrogen bonded turn) and E (extend strand). There are in total 20 letters that are used to describe the secondary structure of NESs. Each letter is represented by a binary string of five bits in a disjoint manner. The binary strings for the letters describing the secondary structure of NESs are shown in Table 1.
With the encoding method, any NES can be represented by a binary string. Specifically, a NES is a sequence of amino acid, whose secondary structure can be obtained by PSIPRED and represented by a string of letters. With the encoding strategy in Table 1, the string of letters of secondary structure can be transformed into a binary string. An example of encoding NES “90-L R S E E V H W L H V D M G V-104” into binary string is given in Table 2.
3.2 The general process of identifying NESs
In general, the process of identifying NESs (represented by binary strings/spike trains) using the SN P system has four stages: reading stage, training stage, generating standard output and identifying unknown NESs.
3.2.1 Reading stage
A set of NESs are randomly selected and encoded into binary strings with their secondary structure. The binary strings will be read by the input neuron one by one bit. Since each NESs is of length 15 (having 15 amino acids), the string encoding a secondary structure has 15 letters. With the strategy shown in Table 1, each binary string is of length 75. That is why in the input module 75 “transmitting” neurons and 75 “gathering” neurons are designed.
3.2.2 Training stage
The input neuron reads binary strings one by one bit. Suppose the input neuron starts to read a binary string at a certain moment t. When it reads one spike from the environment at any step \(t+p\) (\(1\le p \le 75\)), it fires and sends one spike to the “gathering” neuron \(\sigma _{B_p}\). Meanwhile, “transmitting” neuron \(\sigma _{A_p}\) sends one spike to neuron \(\sigma _{B_p}\). With two spikes inside, neuron \(\sigma _{B_p}\) fires by using spiking rule \(a^2\rightarrow a\), sending one spike to neuron \(\sigma _{C_p}\). Having any number of spikes inside, neuron \(\sigma _{C_p}\) fires by using spiking rule \(a^*/a\rightarrow a\), emitting one spike to each of its neighboring neurons. Also, the weights on the synapses the spikes passing along with will be increased by \(\Delta w\).
If the input neuron reads no spike from the environment (indicating the bit of the binary string is 0), then the “gathering” neuron \(\sigma _{B_p}\) has only one spike from “transmitting” neuron \(\sigma _{A_p}\). In this case, neuron \(\sigma _{B_p}\) cannot fire and sends no spike out. Neuron \(\sigma _{C_p}\) cannot receive any spike, and remains inactive. The weights on synapses starting from neuron \(\sigma _{C_p}\) remain unchanged.
When the system finishes reading one binary string, the input module returns to its initial configuration and is ready to read the next binary string. The system can read multiple binary strings one by one. By reading binary strings from environment, the “precessing” neuron \(\sigma _{C_i}\) may fire and the weights on the synapses among “processing” neurons are updated with the Hebbian learning strategy. The four output neurons emit spikes into the environment, but will be ignored. When the system finishes reading the set of binary strings of NESs, it forms a specific topological structure by processing the input information.
3.2.3 Generating standard output
For each NES used in training the SN P system, the binary string representing its secondary structure is input into the “trained system.” In total, 30 four-dimensional vectors can be obtained, which record the numbers of spikes emitted by output neurons \(\sigma _{output_1},\sigma _{output_2},\sigma _{output_3},\sigma _{output_4}\) by reading the 30 NESs. The average vector of the 30 output vectors, denoted by \((stan_1,stan_2,stan_3,stan_4)\), is called the standard outputting vector of the letter.
3.2.4 Identifying unknown NESs
The task of identifying NESs is to judge whether an amino acid sequence is a NES. For any amino acid sequence, its binary string of the secondary structure is obtained by PSIPRED and then introduced into the trained SN P system. When the system halts, a 4-dimensional vector \((out_1,out_2,out_3,out_4)\) is generated recording the numbers of spikes emitted by the four output neurons. We calculate the variance between the outputting vector of the amino acid sequence and standard outputting vector. The variance is calculated by
where \((out_1,out_2,out_3,out_4)\) is the outputting vector of the unknown letter and \((stan_1,stan_2,stan_3,stan_4)\) is the standard outputting vector of a certain letter. If the value of the variance is lower than a threshold, then the amino acid sequence is determined as a potential NES.
4 Experimental results
In the experiments, secondary structure elements of 30 experimentally verified NES are randomly selected for training the SN P system. The 30 selected NESs are shown in Table 3.
In the process of training the SN P system, we set the unit increment \(\Delta w\) to be 0.1, that is, when a spike passes along a synapse from prediction module, its weight will be increased by \(\Delta w=0.1\). The threshold value is set to be 525, which is obtained by calculating the average variance of each pair of NESs used to train the system. For any amino acid sequence, if the variance between its output vector (calculated by the trained SN P system) and the standard outputting vector is less than 525, then the amino acid sequence is determined as NES; otherwise, it is determined as a regular or non-signal amino acid sequence. To test our method, we use the trained SN P system to identify 209 experimentally verified NESs from 1224 amino acid sequences, where 1015 regular amino acid sequences are randomly abstracted from 221 NES-containing protein sequences.
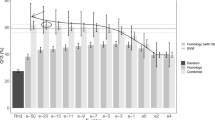
Experimental results show that the SN P system can identify correctly 114 of the 209 experimentally verified NESs and can also determine correctly 809 of the 1015 regular amino acid sequences. The distribution of the variance of the 209 experimentally verified NESs and 1015 regular amino acid sequences are shown in Figs. 4 and 5.
Hence, our method achieves the precision rate \(\frac{114+809}{209+1015}\approx 75.41\,\%\). While NES-REBS has a precision rate of 47.2 % [58], Wregex has a precision rate of 25.4 % [59], ELM has a precision rate of 33.5 % [23], and NetNES has a precision rate of 37.4 % [60].
For statistic analysis, the proposed method is used to identify 2530 randomly generated amino acid sequences. It identifies 1792 sequences as non-NESs, that is, it achieves a precision rate above 70 %, thus having statistical significance.
5 Conclusion
In this work, we address the challenge of identifying NESs from amino acid sequences using SN P system. An SN P system with Hebbian learning strategy is firstly constructed with input and predict modules. After that, secondary structure elements of 30 experimentally verified NES are randomly selected for training an SN P system. We use 1224 amino acid sequences to test our method, where 1015 regular amino acid sequences and 209 experimentally verified NESs are abstracted from 221 NES-containing protein sequences in NESdb randomly. Experimental results show that our method achieves a precision rate of 74.18 %, which performs better than NES-REBS, Wregex, ELM and NetNES.
In our method, the secondary structure elements of experimentally verified NESs is applied to train the SN P system. Actually, there are some other biochemical properties shown as follows from [24].
-
Secondary structure prediction of the regular expression match sequences.
-
Avg.predicted surface accessibility of the regular expression sequences.
-
Avg.predicted disorder score of the regular expression.
-
Hydrophobicity of the regular expression match sequences of negatively charged residues in the upstream flank.
-
Whether the first two residues are involved in a \(\beta\)-strand based on secondary structure.
-
Prediction of polar residues in the downstream flank.
-
Whether the first two residues are involved in a \(\beta\)-strand based on secondary structure prediction.
-
Distance to previous match of the regular expression divided by the protein length.
It is worth for further research of investigating the performances of other biological properties to train SN P system and other computing models. For example, biological networks [63–66] and machine learning methods [67–70] can be considered in this aspect.
In the SN P system, we use a simple Hebbian learning strategy to update the weights on synapses among neurons in predict module. There may be potential further research of designing complex learning strategy and involving the recently developed large-scale neural networks training algorithms, see, for example [71, 72], for the training task. It would be a quite interesting topic to find the inherent advantages of SN P systems comparing with some other models and methods.
Also, some variants of SN P systems, see, for example [31, 46, 73, 74], can be used to improve the performance of our method. The architecture of the SN P system in this work is designed from the biological observation of spiking a neural network from inside to outside. Specifically, the outer layer neurons are divided into four groups, and each neuron of the outer layer connects to some neurons in the outmost layer. As well, all of the outer layer neurons and the outmost layer neurons have been divided into four groups. At this moment, there is no theory to design SN P systems to do pattern recognition, but it would be an interesting topic for future research. Artificial intelligent models and algorithms have been used in solving problems in practice, see, for example [75–79]. It is of interests to use SN P systems to solve some other real-life problems.
References
Görlich D, Kutay U (1999) Transport between the cell nucleus and the cytoplasm. Annu Rev Cell Dev Biol 15:607–660
Conti E, Izaurralde E (2001) Nucleocytoplasmic transport enters the atomic age. Curr Opin Cell Biol 13:310–319
Ren X-X, Wang H-B, Li C, Jiang J-F, Xiong S-D, Jin X, Wu L, Wang J-H (2016) HIV-1 nef-associated factor 1 enhances viral production by interacting with CRM1 to promote nuclear export of unspliced HIV-1 gag mRNA. J Biol Chem 291:4580–4588
Weis K (2003) Regulating access to the genome: nucleocytoplasmic transport throughout the cell cycle. Cell 112:441–451
Strambio-De-Castillia C, Niepel M, Rout MP (2010) The nuclear pore complex: bridging nuclear transport and gene regulation. Nat Rev Mol Cell Biol 11:490–501
La Cour T, Kiemer L, Mølgaard A, Gupta R, Skriver K, Brunak S (2004) Analysis and prediction of leucine-rich nuclear export signals. Protein Eng Des Sel 17:527–536
Fischer U, Huber J, Boelens WC, Mattajt LW, Lührmann R (1995) The HIV-1 Rev activation domain is a nuclear export signal that accesses an export pathway used by specific cellular RNAs. Cell 82:475–483
Fischer U, Meyer S, Teufel M, Heckel C, Lührmann R, Rautmann G (1994) Evidence that HIV-1 rev directly promotes the nuclear export of unspliced RNA. EMBO J 13:4105
Ho JH-N, Kallstrom G, Johnson AW (2000) Nmd3p is a crm1p-dependent adapter protein for nuclear export of the large ribosomal subunit. J Cell Biol 151:1057–1066
Vissinga CS, Yeo TC, Warren S, Brawley JV, Phillips J, Cerosaletti K, Concannon P (2009) Nuclear export of NBN is required for normal cellular responses to radiation. Mol Cell Biol 29:1000–1006
Fornerod M, Ohno M, Yoshida M, Mattaj IW (1997) Crm1 is an export receptor for leucine-rich nuclear export signals. Cell 90:1051–1060
KIrlI K, Karaca S, Dehne HJ, Samwer M, Pan KT, Lenz C, Urlaub H, Görlich D (2016) A deep proteomics perspective on crm1-mediated nuclear export and nucleocytoplasmic partitioning. eLife e11466. doi:10.7554/eLife.11466
Xu D, Grishin NV, Chook YM (2012) NESdb: a database of nes-containing crm1 cargoes. Mol Biol Cell 23:3673
Diella F, Haslam N, Chica C, Budd A, Michael S, Brown NP, Travé G, Gibson TJ (2008) Understanding eukaryotic linear motifs and their role in cell signaling and regulation. Front Biosci 13:6580–6603
Iraia G-S, Sonia B, Jose AR (2012) A global survey of crm1-dependent nuclear export sequences in the human deubiquitinase family. Biochem J 441:209–217
Via A, Gould CM, Gemünd C, Gibson TJ, Helmer-Citterich M (2009) A structure filter for the eukaryotic linear motif resource. BMC Bioinform 10:351
Lee T-Y, Lin Z-Q, Hsieh S-J, Bretaña NA, Lu C-T (2011) Exploiting maximal dependence decomposition to identify conserved motifs from a group of aligned signal sequences. Bioinformatics 27:1780–1787
Van Berlo RJ, Wessels LF, De Ridder D, Reinders MJ (2007) Protein complex prediction using an integrative bioinformatics approach. J Bioinform Comput Biol 5:839–864
la Cour T, Gupta R, Rapacki K, Skriver K, Poulsen FM, Brunak S (2003) NESbase version 1.0: a database of nuclear export signals. Nucleic Acids Res 31:393–396
Xu D, Farmer A, Collett G, Grishin NV, Chook YM (2012) Sequence and structural analyses of nuclear export signals in the NESdb database. Mol Biol Cell 23:3677–3693
Dong X, Biswas A, Chook YM (2009) Structural basis for assembly and disassembly of the CRM1 nuclear export complex. Nat Struct Mol Biol 16:558–560
Güttler T, Madl T, Neumann P, Deichsel D, Corsini L, Monecke T, Ficner R, Sattler M, Görlich D (2010) NES consensus redefined by structures of PKI-type and Rev-type nuclear export signals bound to CRM1. Nat Struct Mol Biol 17:1367–1376
Gould CM, Diella F, Via A, Puntervoll P, Gemünd C, Chabanis-Davidson S, Michael S, Sayadi A, Bryne JC, Chica C et al (2009) ELM: the status of the 2010 eukaryotic linear motif resource. Nucleic Acids Res. doi:10.1093/nar/gkp1016
Fu S-C, Imai K, Horton P (2011) Prediction of leucine-rich nuclear export signal containing proteins with nessential. Nucleic Acids Res. doi:10.1093/nar/gkr493
Prieto G, Fullaondo A, Rodriguez JA (2014) Prediction of nuclear export signals using weighted regular expressions (wregex). Bioinformatics 30(9):1220–1227. doi:10.1093/bioinformatics/btu016
Ionescu M, Păun G, Yokomori T (2006) Spiking neural P systems. Fundam Inform 71:279–308
Maass W (1997) Networks of spiking neurons: the third generation of neural network models. Neural Networks 10:1659–1671
Chen H, Freund R, Ionescu M, Păun G, Pérez-Jiménez MJ (2007) On string languages generated by spiking neural P systems. Fundam Inform 75:141–162
Păun A, Păun G (2007) Small universal spiking neural P systems. BioSyst 90:48–60
Pan L, Paun G (2009) Spiking neural P systems with anti-spikes. Int J Comput Commun Control IV(3):273–282
Song T, Pan L, Jiang K, Song B, Chen W (2013) Normal forms for some classes of sequential spiking neural P systems. IEEE Trans NanoBiosci 12:255–264
Cavaliere M, Ibarra OH, Păun G, Egecioglu O, Ionescu M, Woodworth S (2009) Asynchronous spiking neural P systems. Theor Comput Sci 410:2352–2364
Song T, Pan L, Păun G (2012) Asynchronous spiking neural P systems with local synchronization. Inf Sci 219:197–207
Păun G (2007) Spiking neural P systems with astrocyte-like control. J Univ Comput Sci 13:1707–1721
Song T, Pan L (2016) Spiking neural P systems with request rules. Neurocomputing 193:193–200
Wang J, Peng H (2013) Adaptive fuzzy spiking neural P systems for fuzzy inference and learning. Int J Comput Math 90:857–868
Zeng X, Zhang X, Pan L (2009) Homogeneous spiking neural P systems. Fundam Inf 97:275–294
Song T, Wang X, Zhang Z, Chen Z (2014) Homogenous spiking neural P systems with anti-spikes. Neural Comput Appl 24(7–8):1833–1841. doi:10.1007/s00521-013-1397-8
Ibarra OH, Păun A, Rodríguez-Patón A (2009) Sequential SNP systems based on min/max spike number. Theor Comput Sci 410:2982–2991
Song T, Xu J, Pan L (2015) On the universality and non-universality of spiking neural P systems with rules on synapses. IEEE Trans NanoBiosci 14:960–966
Song T, Pan L, Păun G (2014) Spiking neural P systems with rules on synapses. Theor Comput Sci 529:82–95
Song T, Pan L (2015) Spiking neural P systems with rules on synapses working in maximum spikes consumption strategy. IEEE Trans NanoBiosci 1:38–44
Song T, Pan L (2015) Spiking neural P systems with rules on synapses working in maximum spiking strategy. IEEE Trans NanoBiosci 4:465–477
Ionescu M, Sburlan D (2007) Several applications of spiking neural P systems. In: Fifth brainstorming week on membrane computing, Sevilla
Adl A, Badr A, Farag I (2010) Towards a spiking neural P systems OS. arXiv preprint arXiv:1012.0326
Zeng X, Song T, Zhang X, Pan L (2012) Performing four basic arithmetic operations with spiking neural P systems. IEEE Trans NanoBiosci 11:366–374
Zhang G, Rong H, Neri F, Pérez-Jiménez MJ (2014) An optimization spiking neural P system for approximately solving combinatorial optimization problems. Int J Neural Syst 24(5):1440006
Wang T, Zhang G, Zhao J, He Z, Wang J, Pérez-Jiménez MJ (2014) Fault diagnosis of electric power systems based on fuzzy reasoning spiking neural P systems. IEEE Trans Power Syst 30:1182–1194
Peng H, Wang J, Pérez-Jiménez MJ, Wang H, Shao J, Wang T (2013) Fuzzy reasoning spiking neural P system for fault diagnosis. Inf Sci 235:106–116
Wang J, Shi P, Peng H, Pérez-Jiménez MJ, Wang T (2013) Weighted fuzzy spiking neural P systems. IEEE Trans Fuzzy Syst 21:209–220
Ishdorj T-O, Leporati A, Pan L, Zeng X, Zhang X (2010) Deterministic solutions to QSAT and Q3SAT by spiking neural P systems with pre-computed resources. Theor Comput Sci 411:2345–2358
Pan L, Păun G, Perez-Jimenez MJ (2011) Spiking neural P systems with neuron division and budding. Sci China Inform Sci 54:1596–1607
Wang X, Song T, Gong F, Zheng P (2016) On the computational power of spiking neural P systems with self-organization. Sci Rep. doi:10.1038/srep27624
Leporati A, Mauri G, Zandron C, Păun G, Pérez-Jiménez MJ (2009) Uniform solutions to SAT and subset sum by spiking neural P systems. Nat Comput 8:681–702
Macias-Ramos LF, Perez-Hurtado I, Garcia-Quismondo M, Valencia-Cabrera L, Perez-Jimenez MJ, Riscos-Nunez A (2012) A P-lingua based simulator for spiking neural P systems. Lect Notes Comput Sci 7184:257–281
Ramirez-Martinez D, Gutierrez-Naranjo MA (2007) A software tool for dealing with spiking neural P systems. In: Gutirrez-Naranjo MA (ed) Proceeding of the 5th brainstorming week on membrane computing, pp 299–313
Macias-Ramos LF, Perez-Jimenez MJ, Song T, Pan L (2015) Extending simulation of asynchronous spiking neural P systems in P-Lingua. Fundam Inform 136:253–267
Tingfang W, Xun W, Zheng Z, Faming G, Tao S, Zhihua C (2016) NES-REBS: a novel nuclear export signal prediction method using regular expressions and biochemical properties. J Bioinform Comput Biol (in press)
Prieto G, Fullaondo A, Rodriguez JA (2014) Prediction of nuclear export signals using weighted regular expressions (wregex). Bioinformatics 30:1220–1227
Carla HV, Chiodi G (2013) Structural characterization of netnes glycopeptide from Trypanosoma cruzi. Carbohydr Res 373:28–34
Păun G, Rozenberg G, Salomaa A (2010) The Oxford handbook of membrane computing. Oxford University Press, Oxford
Gutierrez-Naranjo MA, Perez-Jimenez MJ (2009) Hebbian learning from spiking neural P systems view. Lect Notes Comput Sci 5391:217–230
Zeng X, Zhang X, Zou Q (2016) Integrative approaches for predicting microRNA function and prioritizing disease-related microRNA using biological interaction networks. Brief Bioinform 17(2):193–203
Zou Q, Li J, Song L, Zeng X, Wang G (2016) Similarity computation strategies in the microRNA disease network: a survey. Brief Funct Genomics 15(1):55–64
Wang X, Song T, Wang Z, Su Y, Liu X (2013) MRPGA: motif detecting by modified random projection strategy and genetic algorithm. J Comput Theor Nanosci 10:1209–1214
Wang X, Miao Y, Cheng M (2014) Finding motifs in DNA sequences using low-dispersion sequences. J Comput Biol 21:320–329
Zou Q, Hu Q, Guo M, Wang G (2015) HAlign: fast multiple similar DNA/RNA sequence alignment based on the centre star strategy. Bioinformatics 31:2475–2481
Liu B, Chen J, Wang X (2015) Application of learning to rank to protein remote homology detection. Bioinformatics 31:3492–3498
Liu X, Li Z, Liu J, Liu L, Zeng X (2015) Implementation of arithmetic operations with time-free spiking neural P systems. IEEE Trans Nanobioscience 14(6):617–624
Zhang X, Pan L, Paun A (2015) On the universality of axon P systems. IEEE Trans Neural Netw Learn Syst 26:2816–2829
Zhang X, Tian Y, Jin Y (2015) A knee point driven evolutionary algorithm for many-objective optimization. IEEE Trans Evolut Comput 19(6):761–776
Zhang X, Tian Y, Cheng R, Jin Y (2015) An efficient approach to nondominated sorting for evolutionary multiobjective optimization. IEEE Trans Evol Comput 19:201–213
Song T, Pan L (2015) Spiking neural P systems with rules on synapses working in maximum spiking strategy. IEEE Trans NanoBiosci 14:465–477
Zeng X, Zhang X, Song T, Pan L (2014) Spiking neural P systems with thresholds. Neural Comput 26:1340–1361
Gu B, Sheng VS, Tay KY, Romano W, Li S (2015) Incremental support vector learning for ordinal regression. IEEE Trans Neural Netw Learn Syst 26:1403–1415
Gu B, Sun X, Sheng VS (2016) Structural minimax probability machine. IEEE Trans Neural Netw Learn Syst. doi:10.1109/TNNLS.2016.2544779
Wen X, Shao L, Xue Y, Fang W (2015) A rapid learning algorithm for vehicle classification. Inf Sci 295:395–406
Gu B, Sheng VS, Wang Z, Ho D, Osman S, Li S (2015) Incremental learning for support vector regression. Neural Netw 67:140–150
Xia Z, Wang X, Sun X, Wang Q (2016) A secure and dynamic multi-keyword ranked search scheme over encrypted cloud data. IEEE Trans Parallel Distrib Syst 27:340–352
Acknowledgments
This work was supported by National Natural Science Foundation of China (61272152, 61370105, 61402187, 61502535, 61572522 and 61572523), China Postdoctoral Science Foundation funded project (2016M592267), Program for New Century Excellent Talents in University (NCET-13-1031), 863 Program (2015AA020925), and Fundamental Research Funds for the Central Universities (R1607005A).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, Z., Zhang, P., Wang, X. et al. A computational approach for nuclear export signals identification using spiking neural P systems. Neural Comput & Applic 29, 695–705 (2018). https://doi.org/10.1007/s00521-016-2489-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00521-016-2489-z