Abstract
Key message
Transcriptomes generated by laser capture microdissected abnormal staminodes revealed adoption of carpel programming during organ initiation with decreased expression of numerousHSPs,EgDEF1, EgGLO1but increasedLEAFYexpression.
Abstract
The abnormal mantled phenotype in oil palm involves a feminization of the male staminodes into pseudocarpels in pistillate inflorescences. Previous studies on oil palm flowering utilized entire inflorescences or spikelets, which comprised not only the male and female floral organs, but the surrounding tissues as well. Laser capture microdissection coupled with RNA sequencing was conducted to investigate the specific transcriptomes of male and female floral organs from normal and mantled female inflorescences. A higher number of differentially expressed genes (DEGs) were identified in abnormal versus normal male organs compared with abnormal versus normal female organs. In addition, the abnormal male organ transcriptome closely mimics the transcriptome of abnormal female organ. While the transcriptome of abnormal female organ was relatively similar to the normal female organ, a substantial amount of female DEGs encode HEAT SHOCK PROTEIN genes (HSPs). A similar high amount (20%) of male DEGs encode HSPs as well. As these genes exhibited decreased expression in abnormal floral organs, mantled floral organ development may be associated with lower stress indicators. Stamen identity genes EgDEF1 and EgGLO1 were the main floral regulatory genes with decreased expression in abnormal male organs or pseudocarpel initials. Expression of several floral transcription factors was elevated in pseudocarpel initials, notably LEAFY, FIL and DL orthologs, substantiating the carpel specification programming of abnormal staminodes. Specific transcriptomes thus obtained through this approach revealed a host of differentially regulated genes in pseudocarpel initials compared to normal male staminodes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tissue culture propagation is the only approach used to vegetatively propagate selected oil palm trees with desired traits. However, the mantled somaclonal variant is one of the prevailing problems arising from tissue culture (Corley et al. 1986). This abnormal phenotype is due to a feminization of the stamens and staminodes in the male and female inflorescences, respectively (Rival et al. 1996; Adam et al. 2005). Mantled fruits usually abort during development, leading to poor oil yields from these trees. Although mantling rates have decreased with stricter culling procedures and good practices applied during clonal propagation, the circumstances contributing to mantling remain to be elucidated.
As a monoecious plant, the oil palm produces separate male and female inflorescences in a single plant. Mantling mainly affects the male staminodes in female inflorescences whereby pseudocarpels are formed in place of arrested developing staminodes (Adam et al. 2005). However, there is minimal phenotypic difference between female carpels from normal and mantled female inflorescences. Occasionally, androgynous male inflorescences are observed on mantled palms, whereby staminate flowers develop as carpelloid structures. Mantling therefore morphologically affects the male floral organs in pistillate or staminate flowers through a feminization effect.
As the mantling phenotype is similar to that of B-type floral mutants in Arabidopsis thaliana and Antirrhinum majus (Coen and Meyerowitz 1991), the homeotic MADS-box genes, particularly the B-type MADS-box genes, were earlier hypothesized to be involved (Adam et al. 2007a). Recently, aberrant demethylation of a Karma transposon in the intron of a B-type MADS-box gene, EgDEF1, was attributed to the mantling phenotype (Ong-Abdullah et al. 2015). Karma demethylation associates with an increased expression of the kDEF1 isoform of EgDEF1 in female inflorescences at stage 3 floral development onward. At this developmental stage in normal flowers, staminode development is still active while carpels start to develop (Adam et al. 2005). Soon after this, the staminodes will arrest in development in normal female inflorescence but develop into pseudocarpels in mantled female inflorescence. Under anatomical observations, developing pseudocarpels and normal developing staminodes are generally indistinguishable at stage 3 floral development.
Previous studies on oil palm floral development generally employed the use of entire inflorescences. Hence, transcriptomes from this tissue would include those from the male and female floral organs as well as other parts, such as the bracts and rachis (Adam et al. 2005). The heterogeneity of the tissue increases the complexity of the transcriptomes obtained. To better understand the development of abnormal floral organs and gene expression signatures associated with mantling, studying the specifically affected floral organs itself would be advantageous to provide more precise information on the specific transcriptomes. Laser capture microdissection is one such technique to isolate highly pure or homogenous cell populations via direct visualization of the cells (Emmert-Buck et al. 1996).
In this study, a laser capture microdissection (LCM) coupled with RNA sequencing (RNA-seq) approach was used to profile transcriptomes in male and female floral organs from normal and mantled oil palm female inflorescences. As there were limited published reports on the use of LCM in plants and as far as we know, none from oil palm tissues, optimization of the processing steps for LCM was conducted as well. Subsequently, differential expression was analyzed from transcriptome sequences of the corresponding normal and abnormal floral organs, focusing especially on the differences between the normal and abnormal male floral organs. The results provided insights into the regulatory genes, especially transcription factors that are possibly involved in the early manifestation of mantling.
Materials and methods
Plant materials
Unopened oil palm inflorescences were sampled from two normal and three mantled clonal palms (nine years of age) at the Malaysian Palm Oil Board Station in Keratong, Pahang, Malaysia. Inflorescence length at stage 3 floral development (Adam et al. 2007a) was predicted using the ordinal logistic regression model for oil palm floral staging (Sarpan et al. 2015). Inflorescences with lengths of 7–15 cm were sampled and the floral development stage was subsequently verified through histological analysis (Fisher 1968; Sarpan et al. 2015).
Tissue processing
Spikelets were removed from the inflorescence, cut into ~ 0.5 cm in length and subsequently immersed in a 20% (w/v) sucrose solution (Chai et al. 2016). Infiltration was performed under vacuum for 30 min. The tissues were arranged in Tissue-Tek® Cryomold (Sakura, Torrance CA) and infiltrated with Tissue-Tek® O.C.T. mounting medium (Sakura, Torrance CA) under vacuum for 20 min. The tissues in the mounting medium were quickly solidified on liquid nitrogen and stored at − 20 °C until further use.
Cryosectioning
Temperatures for the Leica CM1950 cryostat (Leica Biosystems) were set at − 16 °C and − 18 °C for the chamber and holder, respectively. The embedded tissues were allowed to equilibrate in the cryostat prior to cryosectioning. Serial sections of 10 µm thickness were collected on polyethylene naphthalate (PEN) membrane slides (Applied Biosystems). The slides were then subjected to rehydration (95% (v/v) ethanol for 30 s and 75% (v/v) ethanol for 30 s), staining (0.5 g/L toluidine blue for 30 s), rinsing (0.1% DEPC-treated water several times) and dehydration process (75% (v/v) ethanol for 30 s, 95% (v/v) ethanol for 30 s, twice in 100% (v/v) ethanol for 1 min each, xylene (v/v) for 3 min and a final xylene (v/v) for 5 min). The slides were air-dried at room temperature for at least 20 min, prior to laser capture microdissection.
Laser capture microdissection (LCM)
The dried tissue sections on PEN membrane slides were immediately used for LCM using ArcturusXT™ Microdissection Instrument (Applied Biosystems). The microdissected tissues were captured on Arcturus® CapSure® HS LCM Caps (Applied Biosystems) using a dual laser function consisting of both infrared (IR) laser capture and ultraviolet (UV) laser cutting. LCM was conducted to isolate the male and female developing floral organs present in the oil palm female inflorescences.
RNA isolation, library construction and sequencing
Total RNA was extracted from laser capture microdissected tissues on LCM caps using Arcturus® PicoPure® RNA Isolation Kit (Applied Biosystems) in conjunction with RNase-free DNase (Qiagen) treatment, according to the manufacturers’ recommendations. RNA was eluted in 22 µl elution buffer and pooled from the respective tissues on LCM caps. The pooled samples were then vacuum concentrated until the volume was lower than 14 µl. The entire extract was then subjected to rRNA removal using Ribo-Zero rRNA Removal Kit (Plant Leaf) (Illumina Inc.) according to the manufacturer’s recommendations. Stranded RNA-seq libraries were then prepared using SMARTer® Stranded RNA-Seq Kit (Clontech Laboratories Inc.) according to the manufacturer’s instructions. Concentration and quality of the amplified libraries were assessed using Qubit® 2.0 Fluorometer (Life Technologies) and Bioanalyzer 2100 (Agilent Technologies). A total of 12 bar-coded LCM sample libraries were then sequenced to generate 150 bp paired-end reads (PE150) or 100 bp reads (PE100) on the Illumina HiSeq platform. A few libraries were resequenced using MiSeq platform (PE100) to obtain the required data quantity.
Differential gene expression analysis
Quality assessment of reads was conducted with FastQC (Andrews 2010). TrimGalore! v.0.4.0 (https://www.bioinformatics.babraham.ac.uk/projects/trim_galore/) at a Phred score of Q20 and cutadapt (Martin 2011) tools were used to filter and trim the first five to six bases from reads 1 and 2, respectively (the first three bases of read 1 correspond to part of the SMARTer Stranded Oligo, according to recommendations by Clontech Laboratories Inc.). Reads containing more than 35 consecutive A (or 35 T-bases from read 2) were also trimmed, and finally, reads shorter than 20 bases were removed using cutadapt. Clean reads from sequencing and resequencing corresponding to the same sample were concatenated. Read mapping of paired reads to the E. guineensis P5-build of an AVROS pisifera palm (Singh et al. 2013) was conducted with Tophat2 v.2.0.13 with intron length set at 30–50,000 nucleotides and fr-secondstrand option specified for library type (Kim et al. 2013). Read counts for CDS feature were obtained using HTSeq-Count ver 0.6.1p1 (Anders et al. 2015). All raw reads and read counts have been deposited in NCBI’s Gene Expression Omnibus (Edgar et al. 2002) and are accessible through GEO series accession number GSE115345. Differential expression was analyzed using DESeq2 (Love et al. 2014), with the normal group of samples generally specified as control or reference. The differentially expressed genes (DEGs) were annotated using BLASTX analysis to the RefSeq database with limit set to flowering plants and were designated by their palmXplore IDs (http://palmxplore.mpob.gov.my). GO annotations and GO-Slim ontologies were assigned using Blast2GOPro software. The pisifera P5 genome data can be accessed at the BioProject of the National Center for Biotechnology Information (NCBI) with accession ID PRJNA345530 or at genomsawit.mpob.gov.my. Neighbor-joining tree with JTT matrix-based and bootstrapping methods for MUSCLE sequence alignment was constructed using Mega X (Felsenstein 1985; Saitou and Nei 1987; Jones et al. 1992; Edgar 2004; Hall 2013; Kumar et al. 2018).
In situ RNA hybridization
FFPE sections from paraffin-embedded female inflorescence tissues were prepared as previously described (Ooi et al. 2012). Five genes were selected for design and synthesis of double DIG-labeled LNA™-enhanced probes (Exiqon). The probes were designed to the unique regions in exons associated with the RNA-seq reads. Sequences of the probes are as follows: 5′Dig-TGCTGGAGAACATGATAAGCGA-Dig3′ for EgDEF1 (first exon), 5′Dig-AGCAATCTCTCATTCTTCTTGA-Dig3′ for EgGLO1, 5′Dig-TCTAGCATTATTACTCTCTTCA-Dig3′ for BAG family molecular chaperone gene, 5′Dig-AGGTAGCATGAATAGAATCCGA-Dig3′ for HSP83 gene and 5′Dig-ACAGTAGTGGCAGCAGTAGC-Dig3′ for YABBY gene. The double DIG-labeled Scramble-ISH probe (5′Dig- GTGTAACACGTCTATACGCCCA-Dig3′) (Exiqon) was used as the negative control. In situ RNA hybridizations were carried out as previously described (Ooi et al. 2012, 2016) with incorporation of the TSA Plus DIG Kit (Perkin Elmer) according to the manufacturer’s instructions. Hybridizations were carried out at 45 °C overnight with 40-100 nM of probe. Hybridization with the Scramble-ISH probe was conducted at 55 °C. Washing and detection of hybridized tissue sections were carried out as previously described (Ooi et al. 2012).
Results
Establishment of LCM workflow for oil palm floral tissues
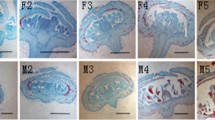
Female inflorescences with lengths of 7.2–12 cm from normal palms and from 8.0 to 15.2 cm from mantled (abnormal) palms were sampled, and their floral development stages were evaluated via histological analysis (Fig. 1). Inflorescences at approximately stage 3 of floral development were of 7.5 cm and 12 cm in length (both associated with frond 17) from normal palms while from the abnormal palms, inflorescences were 14 cm, 15 cm, 15.2 cm in length (fronds 19, 17 and 18, respectively) (Table 1). Sampling of inflorescences at stage 3 development in this study was made easier due to the use of a predictive model (Sarpan et al. 2015) as sampling could be halted after reaching the approximate predicted target length of inflorescence. Although this predictive model previously showed that there was no difference in inflorescence lengths between normal and mantled phenotypes at specific developmental stages, we observed in this study that mantled inflorescences were longer than normal inflorescences at stage 3 development.
Fruit phenotypes, inflorescence and florets of sampled palms. Fruit phenotypes from a mantled palms P273/123, b P273/144, c P291/193 and d normal palms P273/145, e P273/86. f A representative female inflorescence. Length of inflorescence is indicated by red arrow, and a spikelet is indicated by black arrow. Bar = 1 cm. Representatives of histologically stained sections of florets at stage 3 development for g normal floret, h mantled floret. c carpel, st staminodium, pc pseudocarpel initials. Bar = 100 µm
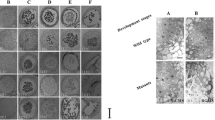
Prior to LCM, cryosections of spikelets were stained and screened microscopically for the presence of target floral organs. Once the cryosection was observed to have the target organs for LCM, three to four cryosections were prepared on PEN membrane slides. These staining steps followed by preparation of cryosections on PEN membrane slides were repeated for several cryoembedded tissues until the estimated target number of floral organs was achieved. The use of both UV and IR lasers was required to excise and capture the desired cell regions. Although IR laser is considered less damaging to cells, using it alone could not lift up the desired floral organs onto the HS LCM cap. It was important to obtain flat cryosections on the PEN membrane slide as bubbles or creases in sections can impede capture on LCM cap. Moreover, sections must be completely dried for LCM. A preliminary LCM of floral organs demonstrated a yield of 19.52 ng total RNA/mm2 section (10 µm thick). Therefore, the estimated average yield of total RNA from one staminodium was 0.133 ± 0.050 ng, while the estimated average yield from a developing carpel was 0.276 ± 0.027 ng. However, RNA yields varied among different extractions, ranging from 4.6 to 19.3 ng total RNA/mm2 (based on six different extractions). Overall, a total of 646 male organs (staminodes) and 295 female organs (developing carpels) were microdissected for RNA extraction (Table 1; Fig. 2). Reference samples were obtained by laser capture microdissection of the remaining tissues after excision of target floral organs. RNA yields and quality from microdissected floral organs were estimated from the microdissected reference, which demonstrated RIN values of approximately 7.0 (Supplementary Figure 1).
Laser capture microdissection of staminodes and carpels from cryosections of spikelets from female inflorescence. a Individual developing pistillate flower before microdissection, b After microdissection of staminodes (indicated by green outlined zones), c after microdissection of carpel (indicated by blue zone), d microdissected staminodes on Arcturus HS Cap, e microdissected carpels on HS Cap. Bar = 100 µm
Differential transcriptome analysis between abnormal and normal floral organs
Principal components analysis showed that the Illumina PE150 sequencing and PE100 resequencing reads clustered together and the normal and mantled groups of samples were reasonably separated (Supplementary Figure 2). PE150 and PE100 reads were quality filtered, trimmed and pooled. The Q20 percentages of the samples ranged from 83 to 91%. Quality assessment of the sequenced reads indicated a high percentage of reads with stretches of polyA or polyT (reverse reads). Evaluation of a good-quality oil palm RNA-seq dataset indicated that 0.13% of reads contain sequences with at least 35 bases of polyA, while a dataset from the LCM tissues contained 9.1–54% of this type of reads. We were unable to determine which part of the workflow that contributed to the polyA contamination. Attempts at read mapping with the quality-trimmed raw data were unsuccessful as processing became unresponsive possibly due to the high number of repetitive A–T stretches. Following that, reads with a minimum of 35 bases of polyA or 35 bases of polyT (reverse reads) were trimmed from the LCM datasets using cutadapt tool. The processed reads were then mapped to the E. guineensis pisifera reference genome using tophat2 read mapper. Unique paired-end reads ranging from 4.76 to 18.17 million reads (Table 2) were then used for subsequent differential transcriptomics analysis. RNA-seq data from two highly related biological replicates, N12M1 and N12M2, independently pooled floral organs that were microdissected from the same inflorescence, were highly correlated with a Pearson correlation coefficient of 0.86 (Supplementary Table 1). This indicated that the mapped RNA-seq data were likely usable, despite high initial amounts of unusable reads. Similarly, correlation between the biological replicates N12F1 and N12F2, M14F and M15F was highest for the female datasets (Supplementary Table 2).
Differential analysis between normal and abnormal male or female organs was conducted by HTSeq read count analysis (Anders et al. 2015) to predicted gene models (Chan et al. 2017), followed by DESeq2 analysis (Love et al. 2014). Significant differentially expressed genes (DEGs) with Benjamini–Hochberg padj < 0.05 were analyzed in more detail. BlastX search was conducted to the RefSeq protein database (limit to flowering plants), with a cutoff E value of 10−5, followed by GO annotation. Generally, more than half of the DEGs were down-regulated in abnormal compared to their normal counterparts. In male floral organs, 61.5% (88 of 143) of significantly expressed DEGs were down-regulated in abnormal organs while 90.9% (30 of 33) of DEGs from female datasets were down-regulated in abnormal female organs. Seventeen DEGs were identified in common from both the abnormal versus normal male and female datasets (Supplementary Table 3). All of these genes exhibited relatively lower expression levels in abnormal floral organs, irrespective of whether male or female organs. DEGs from abnormal versus normal female organs were more enriched for GO terms associated with metabolic and protein modification processes while DEGs from abnormal versus normal male organs were enriched for cell differentiation and flower development GO terms (Supplementary Figure 3).
To evaluate whether transcriptomes from abnormal male or pseudocarpel initials were similar to female carpels, differential analyses were also conducted for abnormal male versus normal female, abnormal female versus normal male and abnormal male versus abnormal female organs (Fig. 3). While 113 and 173 DEGs were identified from former two analyses, respectively, no DEGs were identified from the third analysis. Many of the DEGs in the former two analyses were also represented in the set of DEGs from abnormal versus normal male organs. While 17 DEGs were identified in normal male versus normal female organs, these DEGs will be discussed elsewhere (manuscript in prep.). As there was minimal biological variation between the abnormal male and female organs which were sampled from the same inflorescences, the absence of DEGs suggested that the transcriptomes of abnormal male and abnormal female organs were highly similar. This would then suggest that DEGs from abnormal versus normal male and abnormal female versus normal male would be similar as well, but the common genes between the two DEG sets was only 32% of total DEGs in the latter.
Differential expression analysis of LCM RNA-seq data. a Number of DEGs obtained from differential analyses of all datasets. Normal floral organs are colored green while mantled organs are colored red. b Neighbor-joining tree based on amino acid sequences of the differentially expressed YABBY genes in abnormal versus normal male organs with YABBY genes of Arabidopsis thaliana and Oryza sativa. Bootstrap values are indicated on branches (1000 replicates). The evolutionary distances are in the units of the number of amino acid substitutions per site. Oil palm YABBY genes (palmXplore IDs) are boxed, and accession numbers for the other genes are as labeled on the tree. c Expression heat map of DEGs (padj < 0.05) comprising HSPs and their co-chaperones, B-type MADS-box, LEAFY, YABBY and reproduction-associated genes, between mantled (abnormal) and normal, male and female organs. Genes are labeled by their PalmXplore annotations (refer Supplementary Table 3)
As the emphasis in this study was on the transcriptome differences between normal and abnormal male floral organs, we found that 20 of the 88 down-regulated DEGs in abnormal versus normal male organs encoded heat shock protein (HSP) genes and putative BAG family molecular chaperones (Fig. 3; Supplementary Table 3). These HSPs constituted to approximately 20% of down-regulated genes from not only the abnormal male, but also from the abnormal versus normal female. No HSPs or BAG genes were detected in the up-regulated DEGs.
Several YABBY genes involved in carpel development were also identified from the abnormal versus normal male DEGs. These were related to the Arabidopsis FILAMENTOUS FLOWER (FIL) and CRABS CLAW (CRC) from Arabidopsis or DROOPING LEAF (DL) from rice (Fig. 3). Of major importance are the B-type MADS-box floral organ identity genes, EgDEF1 and EgGLO1, which were lowly expressed in the abnormal feminized male organs as expected. However, the other oil palm GLO gene (Adam et al. 2006), EgGLO2 (Acc. No. AF411848), was not differentially expressed in these tissues.
The meristem identity LEAFY (LFY) gene expression was up-regulated not only in abnormal male organs, but also in abnormal and normal female carpels. Expression of EgDEF1 and EgGLO1 was also down-regulated in both abnormal and normal female carpels. Overall, the expression patterns of EgDEF1, EgGLO1, LFY, the four YABBY genes and several genes related to reproduction or flowering were similar in abnormal male and female as well as normal female organs, suggesting that floral patterning at this developmental stage in these three tissues was similar, in contrast to their expression patterns in normal male organs. The expression patterns of HSPs and their co-chaperones were, however, more similar between abnormal male and abnormal female organs, suggesting that normal floral development associates with higher stress states corresponding with higher expression of HSPs and their co-chaperones.
Transcript localization of selected DEGs through in situ RNA hybridization
Five DEGs with fold-change levels in abnormal versus normal male floral organs from 4.1 to 17.9 (Fig. 4) were selected for mRNA localization experiments via RNA in situ hybridization on female inflorescence FFPE sections. These genes encoded for EgDEF1 (palmXplore ID: p5.00_sc00322_p0006), EgGLO1 (p5.00_sc00051_p0055; Acc. No. AF227195), a BAG family molecular chaperone regulator (p5.00_sc00001_p0352) and a heat shock protein gene (HSP83) associated with stress response (p5.00_sc00062_p0021), and a YABBY gene (p5.00_sc00348_p0023).
Transcript levels of significant DEGs in normal and abnormal, male and female organs based on RNA-seq analysis and corresponding hybridized transcripts on developing female inflorescence FFPE sections at stage 3 floral development. a p5.00_sc00322_p0006 (EgDEF1); b p5.00_sc00051_p0055 (EgGLO1); c p5.00_sc00062_p0021 (HSP83); d p5.00_sc00001_p0352 (BAG family molecular chaperone regulator-like gene); e p5.00_sc00348_p0023 (YABBY gene). Error bars on bar charts are standard error (n = 3); *padj < 0.01. For inflorescence in situ hybridization images, bar = 100 µm for normal (N) and mantled (M) florets; corresponding negative control hybridization with Scramble probe (-ve) is below each N or M image. c carpel, st staminodium, pc pseudocarpel initials
From RNA-seq analysis, EgDEF1 transcript levels were ~ tenfold higher in normal male floral organs compared to abnormal male organs while EgGLO1 levels were 16-fold higher in normal male organs. The relative expression level of the BAG family molecular chaperone regulator-like gene was ~ sixfold higher in normal male and ~ fourfold higher in normal female floral organs compared with their abnormal counterparts (Fig. 4). In situ RNA hybridization results qualitatively verified the expression patterns detected from RNA-seq analysis, demonstrating higher signal intensities in normal organs compared to the abnormal floral organs (Fig. 4). EgDEF1, EgGLO1 and BAG molecular chaperone gene transcripts accumulated in the developing female carpel and also localized to the adaxial region of normal male staminodes. Transcripts of the heat shock protein gene (HSP83), which was detected at ~ 18-fold higher expression levels in normal male floral organs, appeared diffused over the entire normal floret. In contrast to the four genes above, the YABBY gene displayed fourfold higher expression level in abnormal male floral organs and its transcripts accumulated in the developing carpel and possibly abaxial epidermal cells of pseudocarpel initials in abnormal florets. The neighbor-joining tree for YABBY genes from the abnormal versus normal male DEGs suggested that this YABBY gene was closely related to OsYABBY5 and clustered with the Arabidopsis FIL (Fig. 3a).
Discussion
Microdissection and RNA sequencing of developing floral organs from female inflorescences
Laser capture microdissection coupled with transcriptomic analyses has yielded detailed information on specific cell types from various organisms. Important regulatory genes have been identified through this approach from studies on tumor clusters, stem cells, endosperm (Thakare et al. 2014), specific cell types in algae (Saint-Marcoux et al. 2015), giant cells in rice (Ji et al. 2013) and stem tissues from spruce (Abbott et al. 2010). In oil palm, stage 3 floral development involves the arrest of staminode development in normal female inflorescence while in mantled female inflorescence, pseudocarpels form in place of the staminodes (Adam et al. 2005).
As the target floral organs constitute only a small part of the entire inflorescence, the LCM approach with NGS technology provided specific transcriptomes from only the microdissected cell types, allowing for a more precise comparative analysis. Transcript diversity from microdissected floral organs consisting of a few cell types would be lower compared to that obtained from an entire inflorescence with more diverse and multiple cell types. In this study, 4.8–18.2 million uniquely mapped reads per sequencing library were used for differential expression analysis. A previous report on LCM of maize generated sequencing data of four to seven million reads (Thakare et al. 2014). Above one to three million uniquely mapped reads are possibly sufficient for mRNA-seq analysis from single cells (Ramskold et al. 2012; Wu et al. 2014) or low-input RNA-seq using SMART-seq (Bhargava et al. 2014).
Male floral organs are converted into pseudocarpels in mantled female inflorescences but the mantled female carpels are conserved and phenotypically similar to the normal carpels in the female inflorescence (Adam et al. 2005). Nevertheless, 33 DEGs were identified from female carpels of abnormal versus normal inflorescences while 143 DEGs were identified from abnormal versus normal male staminodes. No differentially expressed genes were detected between abnormal male and abnormal female organs, suggesting that their transcriptomes are highly similar at that developmental stage. Thus, it is probable that the main carpel in a severely mantled fruit bunch is molecularly similar to the pseudocarpels surrounding it. In addition, a normal carpel only differs from the abnormal carpel by the differential expression of 33 genes, of which a major subset encodes HSP genes. In mildly mantled oil palms, the main carpel is usually fertile but in severely mantled palms, the main carpel is parthenocarpic and sterile. Whether the differential expression of these 33 genes at this stage in floral development triggers molecular events leading to sterility will require further investigation.
Normal floral organs exhibited higher expression levels of HSPs and their co-chaperones
Most of the highly down-regulated DEGs in male and female abnormal floral organs relative to their normal counterparts were associated with stress response and redox regulation such as heat shock protein (HSP) genes, STI-like, BAG molecular chaperones, ascorbate oxidase, disulfide isomerase-like and glutathione-S-transferase. Moreover, HSP genes and their co-chaperones represented a large part of the down-regulated DEGs in abnormal floral organs, suggesting that suppression of HSPs expression may play important accompanying roles in the manifestation of the mantling abnormality during early floral organ development of the third and fourth whorls.
Flowers destined to be male or female begin as hermaphroditic flowers but later undergo a programmed degeneration of their gynoecium or androecium early in reproductive development (Smith and Zhao 2016). In pistillate flowers of the monoecious oil palm, staminodes at stage 3 rapidly arrest in development soon after and remain a rudimentary structure (Adam et al. 2005). The BAG family of proteins generally regulates diverse processes ranging from proliferation to growth arrest and cell death as well as cell differentiation (reviewed in Takayama and Reed 2001). The Arabidopsis BAG genes may inhibit programmed cell death (PCD) pathways or promote cell survival in response to stress (Doukhanina et al. 2006). BAG proteins are also known to regulate HSP70 proteins and can form complexes with various transcription factors (Zeiner and Gehring 1995; Wang et al. 1996; Takayama et al. 1997). Two BAG genes were down-regulated in pseudocarpel initials together with several HSP70 genes. Therefore, BAG and HSP70 genes may be co-expressed in pseudocarpel initials as BAG proteins have high binding affinity to HSP70 and modulate HSP70 chaperone activity (Takayama et al. 1997; Kabbage and Dickman 2008). Competition between BAG1 and HSP70 for binding to RAF1 protein may represent a molecular switch to encourage proliferation or to become quiescent (Takayama and Reed 2001).
Other than HSP70s, various types of HSPs were also represented in the down-regulated DEGs in abnormal male floral organs. Several putative HSP83 genes belonging to the HSP90 gene family (Conner et al. 1990; Felsheim and Das 1992) were identified as down-regulated in abnormal male organs. Recently, HSP90 was associated with vegetative-to-reproductive transition in Arabidopsis (Margaritopoulou et al. 2016). Depletion of HSP90 mRNAs in shoot apex disrupted flower formation and patterning. Strong interactions between HSP90 system and major flowering genes such as LFY, SOC1 and AGL24 were also observed. Aberrant levels of HSP90 affected flower organ formation into phenotypes resembling lfy, ap1 and ap3 mutants.
Differentially expressed flowering-related transcription factors in abnormal floral organs
The stamen identity B-type MADS-box genes, EgDEF1 and EgGLO1 (Adam et al. 2007b), were significantly up-regulated in normal staminodes compared to abnormal pseudocarpel initials (padj < 10−6). Their transcript levels in pseudocarpel initials or abnormal male floral organs were close to those in normal developing carpels. This is in line with the functional conservation of these two B-type MADS-box genes in oil palm (Beulé et al. 2011; Jaligot et al. 2014). However, EgGLO2 was not differentially expressed between these tissues, suggesting that only EgGLO1 expression is highly associated with EgDEF1 at this developmental stage. The expression patterns of EgDEF1 and EgGLO1 at stage 3 developing male and female floral organs further suggested that pseudocarpel identity or the feminized state of male floral parts is already present at this stage rather than a conversion in identity of the staminodes into pseudocarpels later. This is again supported by the absence of DEGs from the differential analysis between pseudocarpel initials and developing abnormal carpel.
LEAFY, which regulates AP3 and PI among other floral development genes (Parcy et al. 1998; Moyroud et al. 2010), was expressed at higher levels in pseudocarpel initials. Interaction of the LFY-UFO complex with AP3 promoter activates its expression for petal and stamen specification (Chae et al. 2008). AP3 and PI then autoregulate their expression through positive feedback loops in Arabidopsis (Liu and Mara 2010). As the oil palm kDEF1 transcript isoform detected in stage 3 mantled inflorescences probably leads to a truncated protein (Ong-Abdullah et al. 2015), this may break the positive feedback loop, therefore resulting in low levels of EgDEF1/EgGLO1 expression. Elevated LFY expression may then be required to regulate other genes such as AG for carpel development (Lohmann and Weigel 2002).
Besides EgDEF1 and EgGLO1, several other floral transcription factors were differentially expressed in abnormal and normal male floral organs. Increased expression of putative orthologs of several YABBY genes, SPATULA, SHORT INTERNODES 2 and BEL1-like genes was detected in abnormal male organs. Organ identity genes interact with multiple transcription factors to regulate networks that control adaxial–abaxial patterning, organ boundary formation and organ margin development (Sablowski 2015). The YABBY gene family are plant-specific transcription factors important for organ polarity determination. The Arabidopsis FIL, a YABBY gene, is required for normal flower development, and fil mutants have severe abaxial–adaxial polarity defects in their floral organs (Chen et al. 1999; Kumaran et al. 1999; Sawa et al. 1999; Siegfried et al. 1999). OsYAB1, a rice YABBY-gene member, is preferentially expressed in flowers, especially in stamen and carpel primordia (Jang et al. 2004). Transgenic rice overexpressing OsYAB1 was normal during vegetative growth period but exhibited abnormalities in their spikelets which contained supernumerary stamens and carpels compared with wild types. The mantling abnormality of oil palm exhibits a similar phenotype through development of pseudocarpel structures, and the oil palm putative FIL ortholog was highly expressed in pseudocarpel initials compared with normal staminodes.
SPATULA and CRC are necessary for carpel development, and disruption in function of these genes removed carpelloid properties (Alvarez and Smyth 1999). CRC is another member of the YABBY family (Bowman and Smyth 1999) and ortholog of the rice DL. This may explain the relatively higher transcript levels of SPATULA and DL orthologs in pseudocarpel initials and abnormal carpel regions. Moreover, function of the class B MADS-box genes negatively regulates CRC and DL expression in the third whorl (Bowman and Smyth 1999; Yamaguchi et al. 2004), suggesting that down-regulation of class B genes in oil palm pseudocarpels may lead to increased expression of DL orthologs. This further corroborates that the abnormal male organs have already assumed the carpel specification program at stage 3 floral development.
In addition to establishing a laser capture microdissection protocol for oil palm tissues, transcriptomics analysis of microdissected floral organs in this study demonstrated that normal floral organ development at the third and fourth floral whorls in oil palm is accompanied by high levels of stress indicators such as HSPs and their co-chaperones. The associated increased expression of BAG molecular chaperones supports the cytoprotective role of these genes (Doukhanina et al. 2006) during the postulated high stress state of floral development. Decreased expression of EgDEF1 and EgGLO1 and elevated expression of genes associated with carpel specification in pseudocarpel initials together with the transcriptome similarity between this floral organ and developing mantled carpels suggest that these two abnormal organs experience similar developmental programs leading to formation of pseudocarpels and abnormal carpels. All of these observations strongly indicate that pseudocarpel initials may have already adopted a feminized identity at stage 3 floral development. Further investigations would help to unravel the pathways involving these genes and their interactions with each other during reproductive organ development and formation of the mantled flower.
Author contribution statement
S-EO and MO-A conceived and designed the research. NS, S-EO and NAA optimized and conducted laser capture microdissection of the tissues of interest. NS generated the RNA-seq libraries. AN conducted the in situ RNA hybridization. S-EO carried out RNA-seq analysis and wrote the manuscript. All authors read and approved the manuscript.
References
Abbott E, Hall D, Hamberger B, Bohlmann J (2010) Laser microdissection of conifer stem tissues: isolation and analysis of high quality RNA, terpene synthase enzyme activity and terpenoid metabolites from resin ducts and cambial zone tissue of white spruce (Picea glauca). BMC Plant Biol 10:106. https://doi.org/10.1186/1471-2229-10-106
Adam H, Jouannic S, Escoute J, Duval Y, Verdeil JL, Tregear JW (2005) Reproductive developmental complexity in the African oil palm (Elaeis guineensis, Arecaceae). Am J Bot 92:1836–1852
Adam H, Jouannic S, Morcillo F, Richaud F, Duval Y, Tregear JW (2006) MADS box genes in oil palm (Elaeis guineensis): patterns in the evolution of the SQUAMOSA, DEFICIENS, GLOBOSA, AGAMOUS and SEPALLATA subfamilies. J Mol Evol 62:15–31
Adam H, Jouannic S, Morcillo F, Verdeil JL, Duval Y, Tregear JW (2007a) Determination of flower structure in Elaeis guineensis: do palms use the same homeotic genes as other species? Ann Bot 100:1–12
Adam H, Jouannic S, Orieux Y, Morcillo F, Richaud F, Duval Y, Tregear JW (2007b) Functional characterization of MADS box genes involved in the determination of oil palm flower structure. J Exp Bot 58:1245–1259
Alvarez J, Smyth DR (1999) CRABS CLAW and SPATULA, two Arabidopsis genes that control carpel development in parallel with AGAMOUS. Development 126:2377–2386
Anders S, Pyl PT, Huber W (2015) HTSeq—a python framework to work with high-throughput sequencing data. Bioinformatics 31:166–169
Andrews S (2010) FastQC: a quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc. Accessed 26 Apr 2017
Beulé T, Camps C, Debiesse S, Tranchant C, Dussert S, Sabau X, Jaligot E, Syed Alwee SSR, Tregear JW (2011) Transcriptome analysis reveals differentially expressed genes associated with the mantled homeotic flowering abnormality in oil palm (Elaeis guineensis). Tree Genet Genomes 7:169–182
Bhargava V, Head SR, Ordoukhanian P, Mercola M, Subramaniam S (2014) Technical variations in low-input RNA-seq methodologies. Sci Rep 4:3678
Bowman JL, Smyth DR (1999) CRABS CLAW, a gene that regulates carpel and nectary development in Arabidopsis, encodes a novel protein with zinc finger and helix-loop-helix domains. Development 126:2387–2396
Chae E, Tan QK, Hill TA, Irish VF (2008) An Arabidopsis F-box protein acts as a transcriptional co-factor to regulate floral development. Development 135:1235–1245
Chai S-K, Namasivayam P, Ho C-L, Norazlin AA, Ong-Abdullah M, Ooi S-E (2016) RNA from fresh frozen cryosections of oil palm inflorescences is superior to FFPE sections. J Oil Palm Res 28:154–160
Chan K-L, Rosli R, Tatarinova T, Hogan M, Firdaus-Raih M, Low E-TL (2017) Seqping: gene prediction pipeline for plant genomes using self-training gene models and transcriptomic data. BMC Bioinform 18:1426
Chen Q, Atkinson A, Otsuga D, Christensen T, Reynolds L, Drews GN (1999) The Arabidopsis FILAMENTOUS FLOWER gene is required for flower formation. Development 126:2715–2726
Coen ES, Meyerowitz EM (1991) The war of the whorls: genetic interactions controlling flower development. Nature 353:31–37
Conner TW, LaFayette PR, Nagao RT, Key JL (1990) Sequence and expression of a HSP83 from Arabidopsis thaliana. Plant Physiol 94:1689–1695
Corley RHV, Lee CH, Law IH, Wong CY (1986) Abnormal flower development in oil palm clones. Planter 62:233–240
Doukhanina EV, Chen S, van der Zalm E, Godzik A, Reed J, Dickman MB (2006) Identification and functional characterization of the BAG protein family in Arabidopsis thaliana. J Biol Chem 281:18793–18801
Edgar RC (2004) MUSCLE: multiple-sequence alignment with high accuracy and high throughput. Nucl Acids Res 32:1792–1797
Edgar R, Domrachev M, Lash AE (2002) Gene expression omnibus: NCBI gene expression and hybridization array data repository. Nucl Acids Res 30:207–210
Emmert-Buck MR, Bonner RF, Smith PD, Chuaqui RF, Zhuang Z, Goldstein SR, Weiss RA, Liotta LA (1996) Laser capture microdissection. Science 274:998–1001
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Felsheim RF, Das A (1992) Structure and expression of a heat-shock protein 83 gene of Pharbitis nil. Plant Physiol 100:1764–1771
Fisher DB (1968) Protein staining of ribboned epon sections for light microscopy. Histochemie 16:92–96
Hall BG (2013) Building phylogenetic trees from molecular data with MEGA. Mol Biol Evol 30:1229–1235
Jaligot E, Hooi WY, Debladis E, Richaud F, Beulé T, Collin M, Agbessi MDT, Sabot F, Garsmeur O, D’Hont A, Alwee SSRS, Rival A (2014) DNA methylation and expression of the EgDEF1 gene and neighboring retrotransposons in mantled somaclonal variants of oil palm. PLoS ONE 9:e91896. https://doi.org/10.1371/journal.pone.0091896
Jang S, Hur J, Kim S-J, Han M-J, Kim S-R, An G (2004) Ectopic expression of OsYAB1 causes extra stamens and carpels in rice. Plant Mol Biol 56:133–143
Ji H, Gheysen G, Denil S, Lindsey K, Topping JF, Nahar K, Haegeman A, De Vos WH, Trooskens G, Van Criekinge W, De Meyer T, Kyndt T (2013) Transcriptional analysis through RNA sequencing of giant cells induced by Meloidogyne graminicola in rice roots. J Exp Bot 64:3885–3898. https://doi.org/10.1093/jxb/ert219
Jones DT, Taylor WR, Thornton JM (1992) The rapid generation of mutation data matrices from protein sequences. Comput Appl Biosci 8:275–282
Kabbage M, Dickman MB (2008) The BAG proteins: a ubiquitous family of chaperone regulators. Cell Mol Life Sci 65:1390–1402. https://doi.org/10.1007/s00018-008-7535-2
Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL (2013) TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14:R36. https://doi.org/10.1186/gb-2013-14-4-r36
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549
Kumaran MK, Ye D, Yang WC, Griffith ME, Chaudhury AM, Sundaresan V (1999) Molecular cloning of ABNORMAL FLORAL ORGANS: a gene required for flower development in Arabidopsis. Sex Plant Reprod 12:118–122
Liu Z, Mara C (2010) Regulatory mechanisms for floral homeotic gene expression. Semin Cell Dev Biol 21:80–86
Lohmann JU, Weigel D (2002) Building beauty: the genetic control of floral patterning. Dev Cell 2:135–142
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-Seq data with DESeq2. Genome Biol 15:550. https://doi.org/10.1186/s13059-014-0550-8
Margaritopoulou T, Kryovrysanaki N, Megkoula P, Prassinos C, Samakovli D, Milioni D, Hatzopoulos P (2016) HSP90 canonical content organizes a molecular scaffold mechanism to progress flowering. Plant J 1:174–187. https://doi.org/10.1111/tpj.13191
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J 17:10–12. https://doi.org/10.14806/ej.17.1.200
Moyroud E, Kusters E, Monniaux M, Koes R, Parcy F (2010) LEAFY blossoms. Trends Plant Sci 15:346–352
Ong-Abdullah M, Ordway JM, Jiang N, Ooi S-E, Kok S-Y, Sarpan N, Azimi N, Hashim AT, Ishak Z, Rosli SK, Malike FA, Bakar NAA, Marjuni M, Abdullah N, Yaakub Z, Amiruddin MD, Nookiah R, Singh R, Low E-TL, Chan K-L, Azizi N, Smith SW, Bacher B, Budiman MA, Van Brunt A, Wischmeyer C, Beil M, Hogan M, Lakey N, Lim C-C, Arulandoo X, Wong C-K, Choo C-N, Wong W-C, Kwan Y-Y, Alwee SSRS, Sambanthamurthi R, Martienssen RA (2015) Loss of Karma transposon methylation underlies the mantled somaclonal variant of oil palm. Nature 525:533–537. https://doi.org/10.1038/nature15365
Ooi S-E, Lee F-C, Ong-Abdullah M (2012) A rapid and sensitive in situ RNA hybridisation method for oil palm tissues. J Oil Palm Res 24:1235–1239
Ooi S-E, Ramli Z, Alwee SSRS, Kulaveerasingam H, Ong-Abdullah M (2016) EgHOX1, a HD-Zip II gene, is highly expressed during early oil palm (Elaeis guineensis Jacq.) somatic embryogenesis. Plant Gene 8:16–25. https://doi.org/10.1016/j.plgene.2016.09.006
Parcy F, Nilsson O, Busch MA, Lee I, Weigel D (1998) A genetic framework for floral patterning. Nature 395:561–566
Ramskold D, Luo S, Wang Y-C, Li R, Deng Q, Faridani OR, Daniels GA, Khrebtukova I, Loring JF, Laurent LC, Schroth GP, Sandberg R (2012) Full-length mRNA-Seq from single cell levels of RNA and individual circulating tumor cells. Nat Biotechnol 30:777–782
Rival A, Aberlenc-Bertossi F, Morcillo F, Tregear J, Verdeil J-L, Duval Y (1996) Scaling-up in vitro clonal propagation through somatic embryogenesis: the case of the oil palm (Elaeis guineensis Jacq.). Plant Tissue Cult Biotechnol 3:74–83
Sablowski R (2015) Control of patterning, growth, and differentiation by floral organ identity genes. J Exp Bot 66:1065–1073
Saint-Marcoux D, Billoud B, Langdale JA, Charrier B (2015) Laser capture microdissection in Ectocarpus siliculosus: the pathway to cell-specific transcriptomics in brown algae. Front Plant Sci 6:54. https://doi.org/10.3389/fpls.2015.00054
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sarpan N, Kok S-Y, Chai S-K, Fitrianto A, Nuraziyan A, Zamzuri I, Ong-Abdullah M, Ooi S-E (2015) A model for predicting flower development in Elaeis guineensis Jacq. J Oil Palm Res 27:315–325
Sawa S, Watanabe K, Goto K, Kanaya E, Morita EM, Okada K (1999) FILAMENTOUS FLOWER, a meristem and organ identity gene of Arabidopsis, encodes a protein with a zinc finger and HMG-related domains. Genes Dev 13:1079–1088
Siegfried KR, Eshed Y, Baum SF, Otsuga D, Drews GN, Bowman JL (1999) Members of the YABBY gene family specify abaxial cell fate in Arabidopsis. Development 126:4117–4128
Singh R, Ong-Abdullah M, Low E-TL, Manaf MAA, Rosli R, Nookiah R, Ooi LC-L, Ooi S-E, Chan K-L, Halim MA, Azizi N, Nagappan J, Bacher B, Lakey N, Smith SW, He D, Hogan M, Budiman MA, Lee EK, DeSalle R, Kudrna D, Goicoechea JL, Wing RA, Wilson RK, Fulton RS, Ordway JM, Martienssen RA, Sambanthamurthi R (2013) Oil palm genome sequence reveals divergence of interfertile species in old and new worlds. Nature 500:335–339. https://doi.org/10.1038/nature12309
Smith AR, Zhao D (2016) Sterility caused by floral organ degeneration and abiotic stresses in Arabidopsis and cereal grains. Front Plant Sci 7:1503. https://doi.org/10.3389/fpls.2016.01503
Takayama S, Reed JC (2001) Molecular chaperone targeting and regulation by BAG family proteins. Nat Cell Biol. 3:E237–E241
Takayama S, Bimston DN, Matsuzawa S, Freeman BC, Aime-Sempe C, Xie Z, Morimoto RI, Reed JC (1997) BAG-1 modulates the chaperone activity of Hsp70/Hsc70. EMBO J 16:4887–4896
Thakare D, Yang R, Steffen JG, Zhan J, Wang D, Clark RM, Wang X, Yadegari R (2014) RNA-seq analysis of laser-capture microdissected cells of the developing central starchy endosperm of maize. Genomics Data 2:242–245
Wang M, Oppedijk BJ, Lu X, Van Duijn B, Schilperoort RA (1996) Apoptosis in barley aleurone during germination and its inhibition by abscisic acid. Plant Mol Biol 32:1125–1134
Wu AR, Neff NF, Kalisky T, Dalerba P, Treutlein B, Rothenberg ME, Mburu FM, Mantalas GL, Sim S, Clarke MF, Quake SR (2014) Quantitative assessment of single-cell RNA-sequencing methods. Nat Methods 11:41–46
Yamaguchi T, Nagasawa N, Kawasaki S, Matsuoka M, Nagato Y, Hirano H-Y (2004) The YABBY gene DROOPING LEAF regulates carpel specification and midrib development in Oryza sativa. Plant Cell 16:500–509
Zeiner M, Gehring U (1995) A protein that interacts with members of the nuclear hormone receptor family: identification and cDNA cloning. Proc Natl Acad Sci USA 92:11465–11469
Acknowledgements
We thank the Director-General of the Malaysian Palm Oil Board (MPOB) for permission to publish this study. Our deepest appreciation goes to Masniyana J., Zamzuri I., Feshah I., Rosna A., Roslan N. and Azizah M. of the Breeding and Tissue Culture Unit, MPOB, Nor Azwani A. B. of MPOB Keratong and the Bioinformatics Unit, MPOB, for their invaluable technical support and advice throughout this study. This study was funded by the Malaysian Palm Oil Board.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of competing interest.
Additional information
Communicated by Dolf Weijers.
Siew-Eng Ooi and Meilina Ong-Abdullah are joint corresponding authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ooi, SE., Sarpan, N., Abdul Aziz, N. et al. Differential expression of heat shock and floral regulatory genes in pseudocarpel initials of mantled female inflorescences from Elaeis guineensis Jacq.. Plant Reprod 32, 167–179 (2019). https://doi.org/10.1007/s00497-018-0350-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00497-018-0350-5