Abstract
Background
Gastric cancer is a common kind of malignancies, with yearly occurrences exceeding one million worldwide in 2017. Typically, ulcerous and cancerous tissues develop abnormal morphologies through courses of progression. Endoscopy is a routinely adopted means for examination of gastrointestinal tract for malignancy. Early and timely detection of malignancy closely correlate with good prognosis. Repeated presentation of similar frames from gastrointestinal tract endoscopy often weakens attention for practitioners to result in true patients missed out to incur higher medical cost and unnecessary morbidity. Highly needed is an automatic means for spotting visual abnormality and prompts for attention for medical staff for more thorough examination.
Methods
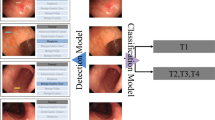
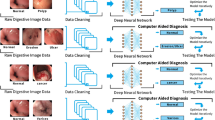
We conduct classification of benign ulcer and cancer for gastrointestinal endoscopic color images using deep neural network and transfer-learning approach. Using clinical data gathered from Gil Hospital, we built a dataset comprised of 200 normal, 367 cancer, and 220 ulcer cases, and applied the inception, ResNet, and VGGNet models pretrained on ImageNet. Three classes were defined—normal, benign ulcer, and cancer, and three separate binary classifiers were built—those for normal vs cancer, normal vs ulcer, and cancer vs ulcer for the corresponding classification tasks. For each task, considering inherent randomness entailed in the deep learning process, we performed data partitioning and model building experiments 100 times and averaged the performance values.
Results
Areas under curves of respective receiver operating characteristics were 0.95, 0.97, and 0.85 for the three classifiers. The ResNet showed the highest level of performance. The cases involving normal, i.e., normal vs ulcer and normal vs cancer resulted in accuracies above 90%. The case of ulcer vs cancer classification resulted in a lower accuracy of 77.1%, possibly due to smaller difference in appearance than those cases involving normal.
Conclusions
The overall level of performance of the proposed method was very promising to encourage applications in clinical environments. Automatic classification using deep learning technique as proposed can be used to complement manual inspection efforts for practitioners to minimize dangers of missed out positives resulting from repetitive sequence of endoscopic frames and weakening attentions.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Gastric cancer is a common kind of malignancies, with yearly occurrences exceeding one million worldwide in 2017 [1]. Almost one million new cases of stomach cancer were estimated to have occurred in 2012 (952,000 cases, 6.8% of the total), making it the fifth most common malignancy in the world, after cancers of the lung, breast, colorectum, and prostate [2]. Stomach cancer is the third leading cause of cancer death in both sexes worldwide (723,000 deaths, 8.8% of the total). For examination of gastrointestinal organs for cancer and other diseases, endoscopy is often the very first means employed. Manual examination for endoscopic data requires training for medical staff and tends to be time consuming with diagnosis results being subjective in nature, dependent on the level of expertise of person performing the procedure. Endoscopy for patients has to be performed over a sequence of seemingly similar frames of endoscopic data and typical clinical environments demand the same kinds of procedures be performed for multiple patients in series. Often reported are loss or weakening of attention for performing person resulting from the examination of long footage of endoscopy. They lower the quality of medical examination, falsely producing missed out negative cases for true patients and unnecessarily requiring re-examinations for healthy subjects [3].
In fact, ten studies involving 3,787 patients who were subjected to upper gastrointestinal endoscopy revealed that 11.3% of upper gastrointestinal cancers are missed up to 3 years before diagnosis [4]. Misses were dependent on the types and sites of gastric cancers and were more notable for examiners with fewer than 10 years of experience. Physical and mental conditions of endoscopists performing procedures also strongly affected miss rates [5]. Machine learning is a practice of developing automatic cognitive models using computerized means. Various kinds of tasks may be assigned including classification, regression, and clustering. Many cognitive and inferential tasks can be formulated as classification as in the case of normal vs cancer and cancer vs ulcer classification. Repeated presentation of data coupled with correct labels to predict until proper training objective is met is the usual procedure for training models. Diverse kinds of machine learning model were proposed. They can be characterized and distinguished apart in terms of the nature of data, kinds of the units comprising models, how units are combined into model, ways of presenting training data, and formulation of training objectives. Deep learning allows computational models that are composed of multiple processing layers to learn representations of data with multiple levels of abstraction [6]. A computer system employing laser-based endoscopy was developed for quantitative diagnosis of early gastric cancer. It was tested on 100 consecutive early gastric cancer in 95 patients to produce area under curve of 0.80 from sensitivity vs false positive rate plot [7].
Zhang et al. proposed a CNN-based gastric precancerous disease network for classification of polyp, erosion, and ulcer. It utilized iterative reinforcement learning method and fire modules from SqueezeNet for reductions of network size and computation time. Reported accuracy value was 88.90% [8]. Zhu used support vector machine (SVM) together with CNN for detection of gastrointestinal diseases. In addition to the raw image data, local binary pattern representations were used. Overall accuracy obtained was 80.0% [9]. The use of color wavelet features together with CNN features was proposed by Billah et al. [10]. The extracted features were then fed to SVM-based classifier for final classification results. Results on data from a slightly different imaging modality of bandwidth-limited imaging was experimented by Byrne et al. [11]. Red channel was mostly retained and the rest channels were suppressed therein. Inception module-based network was crafted to produce accuracy values ranging from 85 to 94%. An elaborate scheme of feature engineering involving color, texture, shape, and temporal information was experimented. Then separate classifiers for each of the features were trained and then combined [12]. Computer-based Automatic classification models can help the diagnosis efforts for gastrointestinal malignancy required of endoscopic examinations [13,14,15,16,17,18]. Alexnet revealed and demonstrated the potential of deep learning methods for the practical problem of automated image classification. Since then numerous architectures were introduced that further improved performance level—they include VGGNet, residual network, and inception net. However, there are few reports that compared the accuracies of CNN models for gastric malignancy. Here, we tried to develop a model for differentiating gastric ulcer and cancer using deep learning model.
Materials and methods
Datasets
All cases appearing either ulcerous or cancerous from visual inspection were histologically examined. Study was approved by IRB (IRB Number 2018-052: Diagnosis and cure for digestive tract diseases using machine learning). Patients with biopsy-proven, cancer cell-free ulcer were treated via medical management such as proton pump inhibitor. Follow-up examinations after treatments revealed all patients turned normal. These were defined as the benign ulcerative cases. Cases for negative histological examination results were labeled ulcer positive and cases with detected malignant cells were labeled cancer positive. We collected endoscopic data from Gil hospital which had 200 normal, 220 benign ulcer, and 367 cancer images. For training and testing purposes, we partitioned the data into 180, 200, and 337 image train sets and 20, 30, and 20 image test sets (Table 1). Frames constituting endoscopic sequences were all resized to 224 × 224 width and height for conformance to the common input dimension for ResNet, VGGNet, and inception network.
Image preprocessing
As a means to reduce image variations including brightness and contrast that are irrelevant to classification task, histogram equalization was used, i.e., adaptive histogram equalization (AHE) [19]. It differs from ordinary histogram equalization in that it computes several histograms, each corresponding to a distinct section of the image, and uses them to redistribute the lightness values of the image. It is therefore suitable for improving the local contrast and enhancing the definitions of edges in each region of an image. For the method of contrast-limited adaptive histogram equalization (CLAHE) [20], contrast amplification in the vicinity of a given pixel value is specified by the slope of a transformation function. It is set to be proportional to the slope of the neighborhood cumulative distribution function (CDF) and therefore to the value of the histogram at that pixel value. CLAHE sets a limit on the amplification by clipping the histogram at a predefined value before computing the CDF. The value at which the histogram is clipped, the so-called clip limit, depends on the normalization of the histogram and on the size of the neighborhood region. Common values for limiting the resulting amplification range between 3 and 4, and for our study, we used 3.5 for limit value (Fig. 1).
Differentiation utilizing deep neural networks
We used a number of architectures—inception network, ResNet, and VGGNet and fine tuned them using endoscopy images [21]. Alexnet was one of the first to explore the application of convolutional deep learning network models towards image classification task [22]. Substantial performance improvement was shown to be achievable over existing state of art in ImageNet ILSVRC 2012 challenge. Since its introduction, numerous network architectures were introduced. Contrary to the prior Lenet network featuring a single convolution layer, Alexnet employed a longer cascade of convolutional layers interlaced by pooling layers (Supplementary Material 1).
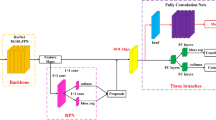
Three kinds of models were built for classification tasks—normal vs cancer, normal vs benign ulcer, and cancer vs benign ulcer. The inception network [23] model comprises 27 layers total including pooling layers; 22 of which are parameterized (Fig. 2). The inception module is an integral component for the model, which concatenates filters of different sizes and dimensions. Each “Inception” layer consists of six convolution layers and one pooling layer. The depth of representations is of importance for many visual recognition tasks. ResNet utilizes a building block of two convolutional layers featuring an identity connection skipping over them [24]. Particular ResNet network is instantiated by stacking such building blocks to a desired depth. Authors reported results for networks of depths of 18, 34, 50, 101, and 152. Canziani et al. [25] performed a comprehensive study for the relative performances of a range of deep learning networks for object classification task. The difference in accuracy between the ResNet-18 and ResNet-50 was substantial. For ResNet networks of depths greater than 50, increase in performance was rather marginal. Hence we experimented with the ResNet-50 architecture in this study. For VGGNet, the 16 layer and 19 layer variants were shown to exhibit rather insignificant performance difference while the 16 layer one entailed smaller amount of computational load, which consequently was our choice. The best performing network overall was found to be inception v4 with slight performance margin over those of ResNet network varieties.
Transfer learning
First, we utilized an inception v4 model pretrained on ImageNet and fine tuned it towards training endoscopic dataset. Transfer learning [26] refers to training a network on a base dataset and task and then fine tuning it towards a given task by allowing final layers to adjust. It has been applied in a variety of medical CAD cases including one performed by Andre Istiva team that used Imagenet-pretrained CNN architecture to classify skin cancers [27] (Supplementary Material 2).
Results
The accuracies of inception trained networks are as given in Tables 2 and 3. ResNet consistently showed the highest performance across the three classification tasks. The p value of significance was 5.8e−3. Also the standard deviation values in performance for the ResNet was substantially lower than those of inception v4, except for the case of cancer vs ulcer. This implies the learning results for ResNet are more stable than the rest of networks, necessitating less number of experimental drafting trial runs, which is another merit from practical training perspectives. Normal vs cancer and normal vs ulcer classifications produced accuracies above 0.90. In the case of cancer vs ulcer classification, performances were lower than the normal vs cancer and normal vs ulcer cases. The foremost step in the gastric endoscopy diagnosis is to determine whether a subject being seen is normal or has a disease. Hence the classification tasks involving normal-cancer and normal-ulcer have higher importance than the task of malignancy-only cancer vs ulcer cases. The nature of medical diagnosis typically demands high level of accuracies of computer-aided diagnosis systems. While the accuracies of 0.9649 and 0.9262 fall somewhat short of expected accuracies of CAD, overall the results show promise of deep learning-based models for automatic diagnosis based on gastric endoscopy. In Fig. 3, we give plots of AUROC curves for the three classifiers.
Discussion
Previously, feature-based approaches were mainly developed and applied to endoscopic examinations. Deep learning technique is expanding the performance envelopes for automatic diagnosis models and is being applied in increasing number of disciplines including medical sciences. We showed in this article an application of a number of widely used network architectures for the task of endoscopic image classification for malignancy. We trained the model with 10 epochs of training using batches of size 50, 0.001 for learning rate and 0.5 for dropout rate. Standard back-propagation algorithm was used. Data partitioning and model training were performed 100 times and classification performance values were averaged in order to obtain objective assessments. Normal vs ulcer classifier produced the highest AUC of 0.97, and the cancer vs ulcer classifier, the lowest 0.85. This high level of performance involving normal vs abnormal cases implies the promise of the proposed approach for clinical application (Figs. 4, 5, 6).
The computational speed of the classification routine was sufficiently fast (data not shown). It is conceivable that the three classifiers are combined into a composite diagnostic model to answer medical queries of whether a patient has a gastric disease or not and specifically whether it is ulcer or cancer. In studies to follow, we will focus on more rigorous testing and validation of performance results using larger datasets obtained from multiple clinics to improve reproducibility and objectivity in addition to performance improvement and network size reduction.
Differently from a previous study by Canziani et al. comparing performances of deep learning architectures, the ResNet architecture produced highest accuracy classification results. It suggests rather small training data size combined with the low number of classes to be predicted for gastric endoscopy which favors the use of ResNet over other alternatives. It will be interesting to investigate the performances of different networks resulting from utilization of larger datasets such as [28] in the future. We conclude that, despite the disparity between the ImageNet dataset and gastrointestinal endoscopy images, deep learning via transfer learning using well-known architectures is an efficient means for classifying such images. While rather small training dataset size used was a shortcoming of the present study, the test results obtained using the test dataset independent of the training set strongly implies overall soundness of our approach. Further cross and external validations are needed to strengthen the findings from our study.
References
World Cancer Report 2014 (2014) World Health Organization
http://globocan.iarc.fr. Accessed 23 Jan 2019
Watanabe K, Nagata N, Shimbo T, Nakashima R, Furuhata E, Sakurai T, Akazawa N, Yokoi C, Kobayakawa M, Akiyama J, Mizokami M, Uemura N (2013) Accuracy of endoscopic diagnosis of Helicobacter pylori infection according to level of endoscopic experience and the effect of training. BMC Gastroenterol 13:128
Menon S, Trudgill N (2014) How commonly is upper gastrointestinal cancer missed at endoscopy? A meta-analysis. Endosc Int Open 2(2):E46–E50
Almadi MA, Sewitch M, Barkun AN (2015) Adenoma detection rates decline with increasing procedural hours in an endoscopist’s workload. Can J Gastroenterol Hepatol 29(6):304–308
LeCun Y, Bengio Y, Hinton G (2015) Deep learning. Nature 521:436–444
Miyaki R, Yoshida S, Tanaka S, Kominami Y, Sanomura Y, Matsuo T, Oka S, Raytchev B, Tamaki T, Koide T, Kaneda K, Yoshihara M, Chayama K (2015) A computer system to be used with laser-based endoscopy for quantitative diagnosis of early gastric cancer. J Clin Gastroenterol 49(2):108–115
Zhang X, Hu W, Chen F, Liu J, Yang Y, Wang L, Duan H, Si J (2017) Gastric precancerous diseases classification using CNN with a concise model. PLoS ONE 12(9):e0185508
Zhu R, Zhang R, Xue D (2015) Gastric precancerous diseases classification using CNN with a concise model. In: 2015 8th International Congress on Image and Signal Processing (CISP) 14–16 Oct
Billah M, Waheed S, Rahman MM (2017) An automatic gastrointestinal polyp detection system in video endoscopy using fusion of color wavelet and convolutional neural network features. Int J Biomed Imaging. https://doi.org/10.1155/2017/9545920
Byrne MF, Chapados N, Soudan F, Oertel C, Linares PM, Kelly R, Iqbal N, Chandelier F, Rex DK (2017) Real-time differentiation of adenomatous and hyperplastic diminutive colorectal polyps during analysis of unaltered videos of standard colonoscopy using a deep learning model. Gut 68:94–100
Tajbakhsh N (2015) Automatic polyp detection in colonoscopy videos using an ensemble of convolutional neural networks. In: IEEE 12th International Symposium on Biomedical Imaging (ISBI)
Li B, Meng MQ (2009) Computer-aided detection of bleeding regions for capsule endoscopy images. IEEE Trans Biomed Eng 56(4):1032–1039
Tamai N, Saito Y, Sakamoto T, Nakajima T, Matsuda T, Sumiyama K, Tajiri H, Koyama R, Kido S (2017) Effectiveness of computer-aided diagnosis of colorectal lesions using novel software for magnifying narrow-band imaging: a pilot study. Endosc Int Open 5(8):E690–E694
Clements LM, Kockelman KM (2017) Economic effects of automated vehicles. J Transp Res Board 2606:2606–2614
Silver D, Huang A, Maddison CJ, Guez A, Sifre L, Driessche G, Schrittwieser J (2016) Mastering the game of Go with deep neural networks and tree search. Nature 529:484–489
Gerhardus D (2003) Robot-assisted surgery: the future is here. J Healthc Manag 48(4):242–251
Maroulis DE, Savelonas MA, Karkanis SA, Iakovidis DK, Dimitropoulos N (2005) Computer-aided thyroid nodule detection in ultrasound images. Proceedings of the 18th IEEE symposium on computer-based medical systems (CBMS’05)
Pizer SM, Amburn EP, Austin JD, Cromartie R, Geselowitz A, Greer T, Romeny B, Zimmerman JB, Zuiderveld K (1987) Adaptive histogram equalization and its variations. Comput Vis Graph Image Process 39(3):355–368
Zuiderveld K (1994) Contrast limited adaptive histogram equalization. Graphics gems IV. Academic Press Professional, Inc., San Diego, pp 474–485
Deng J, Dong W, Socher R, Li LJ, Li K, Fei-Fei L (2009) ImageNet: a large-scale hierarchical image database. In: Proceedings of IEEE conference on computer vision and pattern recognition, pp. 248–255
Krizhevsky A, Sutskever I, Hinton GE (2012) ImageNet classification with deep convolutional neural networks. NIPS 1:1097–1105
Szegedy C, Liu W, Jia Y, Sermanet P, Reed S, Anguelov D, Vanhoucke V, Rabinovich A (2015) Going deeper with convolutions. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp. 1–9
He K, Zhang C, Ren S, Sun J (2016) Deep residual learning for image recognition. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp. 770–778
Canziani A, Paszke A, Culurciello E (2016) An analysis of deep neural network models for practical applications. ArXiv:1605.07678
Dauphin GMY, Glorot X, Rifai S, Bengio Y, Goodfellow I, Lavoie E, Muller X, Desjardins G, Warde-Farley D, Vincent P, Courville A, Bergstra J (2012) Unsupervised and transfer learning challenge: a deep learning approach. JMLR 27:97–110
Esteva A, Kuprel B, Novoa RA, Ko J, Swetter SM, Blau HM, Thrun S (2017) Dermatologist-level classification of skin cancer with deep neural networks. Nature 542(7639):115–118
Pogorelov K, Randel KR, Griwodz C, Eskeland SL, de Lange T, Johansen D, Spampinato C, Nguyen DTD, Lux M, Schmidt PT, Riegler M, Halvorsen P (2017) Kvasir: a multi-class image dataset for computer aided gastrointestinal disease detection. In: Proceedings of the 8th ACM on Multimedia Systems Conference, Pages 164–169
Acknowledgements
This work was supported by Institute for Information & communications Technology Promotion (IITP) grant funded by the Korea government (MSIT) (2018-2-00861, Intelligent SW Technology Development for Medical Data Analysis) and the Gachon University Gil Medical Center (Grant No: 2018-5283). Authors Kim KG and Chung JW equally contributed to this work.
Funding
The authors state that this work has not received any funding.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Disclosure
Authors Jang Hyung Lee, Young Jae Kim, Yoon Woo Kim, Sungjin Park, Yoon Yi Choi, Yoon Jae Kim, Dong Kyun Park, Kwang Gi Kim, and Jun-Won Chung have no financial arrangement or affiliation with any product or services used or discussed in this paper, nor any potential bias against another product or service. The authors declare that there are no conflicts of interest regarding the publication of this paper.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lee, J.H., Kim, Y.J., Kim, Y.W. et al. Spotting malignancies from gastric endoscopic images using deep learning. Surg Endosc 33, 3790–3797 (2019). https://doi.org/10.1007/s00464-019-06677-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00464-019-06677-2