Abstract
To produce maltobionic acid (MBA) from maltose in Escherichia coli, we recombinantly expressed a glucose dehydrogenase gene (gdh1) from Enterobacter cloacae and a pyrroloquinoline quinone (PQQ) synthesis gene cluster (pqqFABCDEMIH) from Pseudomonas taetrolens. Although the recombinant E. coli strain (E. coli [pKK-ECGDH1 + pACYC-PQQ]) successfully produced MBA from maltose, the yield of MBA was rather low, indicating that E. coli has other maltose utilization pathways. Amylomaltase (MalQ) is the first enzyme in the maltose utilization pathway in E. coli. To investigate the potential role of MalQ on MBA production, E. coli malQ was inactivated. The culturing of the recombinant E. coli strain (E. coli ∆malQ [pKK-ECGDH1 + pACYC-PQQ]) in a flask resulted in higher MBA production titer, yield, and productivity (209.3 g/L, 100%, and 1.1 g/L/h, respectively) than those of E. coli [pKK-ECGDH1 + pACYC-PQQ] (162.1 g/L, 77.4%, and 0.5 g/L/h, respectively), indicating that the MalQ inactivation was highly effective in improving the MBA production ability of E. coli. After fermentation using 5-L bioreactor, MBA production titer, yield, and productivity of the recombinant E. coli strain were 209.3 g/L, 100%, and 1.5 g/L/h, respectively, which were 1.3-, 1.3-, 2.3-fold higher than those of E. coli [pKK-ECGDH1 + pACYC-PQQ] (167.3 g/L, 79.9%, and 0.65 g/L/h), respectively. Thus, our results provide an important foundation for efficient MBA production using recombinant E. coli strain.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Maltobionic acid (MBA) is an aldonic acid obtained from the oxidation of maltose. MBA has been a component of the human diet since it was first discovered in honey [1]. MBA is a stereoisomer of lactobionic acid (LBA), a well-known aldonic acid that has been used for various industrial applications [2, 3]. Similar to LBA, MBA is also utilized in a variety of industries such as foods, cosmetics, and pharmaceuticals [1, 4, 5] because of excellent antioxidant, metal-chelating, and moisturizing properties [6,7,8].
MBA is almost exclusively produced by chemical oxidation. However, this chemical method requires harmful and costly metal catalysts and generates undesired by-products, such as maltulose and 2-keto MBA [9]. In contrast, biological MBA production methods have recently received considerable attention, because they do not require toxic catalysts and does not generate by-products. In a previous study, we succeeded in producing MBA at a high concentration (200 g/L) and productivity (9.52 g/L/h) using genetically modified Pseudomonas taetrolens [10]. Moreover, using this P. taetrolens strain as a whole-cell biocatalyst (WCB), we successfully produced MBA with a high production titer (200 g/L) and improved productivity (18.18 g/L/h) [11]. However, P. taetrolens cannot grow to a high cell density, which could be a weakness in the preparation of WCB for MBA production. Therefore, a new MBA-producing strain that can grow to a high cell density is required for efficient MBA production via whole-cell biocatalysis.
Among various bacteria, E. coli is a candidate MBA-producing species. It is the most extensively used bacterium for producing chemicals and proteins. This species can grow at high cell densities using inexpensive substrates such as glycerol. Moreover, the genetic tools for E. coli are plentiful, and their metabolic pathways are well understood [12, 13]. Therefore, E. coli can be used as a WCB for MBA production. Because E. coli cannot inherently produce MBA from maltose, we attempted to produce MBA by constructing a recombinant E. coli strain in this study. In our previous studies, we found that GDH from P. taetrolens could not only convert lactose into LBA but also maltose into MBA [10, 14]. The GDH uses PQQ as a cofactor. The PQQ synthesis gene cluster of P. taetrolens consists of nine genes (pqqFABCDEMIH). Using the GDH gene and PQQ synthesis gene cluster from P. taetrolens, we constructed a recombinant E. coli strain (E. coli [pKK-PTGDH + pACYC-PQQ]) with LBA-producing ability [15]. We found that GDH1 from Enterobacter cloacae also converts lactose to LBA [16]. Thus, we constructed another recombinant E. coli strain (E. coli [pKK-ECGDH1 + pACYC-PQQ]) showing LBA-producing ability using GDH1 from E. cloacae and the PQQ synthesis gene cluster from P. taetrolens [15]. In this study, we investigated whether this recombinant strain could also produce MBA. In addition, we tried to enhance the MBA-producing ability of E. coli by engineering the metabolic pathways of E. coli.
Material and methods
Materials
Maltose (EP grade, 92% purity) was bought from Daejung Chemical and Metals Co. Ltd. (Korea). MBA and other reagents were obtained commercially from Sigma-Aldrich (USA) and BD Difco (USA). All DNA manipulation tools including restriction and DNA-modifying enzymes, were bought from New England Biolabs (USA), Solgent (Korea), or Sigma-Aldrich.
Bacterial strains and plasmids
Escherichia coli JM109 was used as the host strain for MBA production. The strain was grown in Luria–Bertani (LB) medium at 37 °C for 24 h and was kept frozen in 30% glycerol at –80 °C. Plasmid pKK-ECGDH1 was used for expressing the GDH gene from E. cloacae in E. coli. The GDH was cloned into pKK223-3, the expression vector for E. coli which contains the tac promoter. Plasmid pACYC-PQQ was used for expressing a pyrroloquinoline quinone (PQQ) synthesis gene cluster (pqqFABCDEMIH) from P. taetrolens in E. coli. The PQQ synthesis gene cluster was cloned into pACYC184, the expression vector for E. coli which contains the tet promoter [15].
Knock-out of amylomaltase gene in E. coli
The deletion of malQ (GenBank accession number CAD6000741.1) in E. coli JM109 was carried out using the Quick and Easy Conditional Knockout Kit (Germany). To delete malQ, a ∆malQ loxP-neo-loxP cassette was made by PCR using the following pair of designed primers: MalQ_Del_F 5′-ATGGAAAGCAAACGTCTGGATAATGCCGCGCTGGCGGCGGGGATTAGCCCCAATTACATCTACCGTTCGTATAGCATACA-3′, MalQ_Del_R 5′CTACTTCTTCTTCGCTGCAGCTCTGCGCCGTCTGTCCAAATCCTTCAGCAACTTGTTCACTACCGTTCGTATAATGTATG-3′ to use FRT-PGK-gb2-neo-FRT as the template DNA.
The malQ gene of E. coli JM109 was replaced with the ∆malQ loxP-neo-loxP (chloramphenicol, Cm) cassette [17]. After being cultivated in LB medium, E. coli cells reached a cell density of 0.6 at an OD600nm, and the pRED/ET recombinase gene was induced by adding 10% l-arabinose. After incubation for 1 h at 37 °C, the cells were harvested and rinsed thrice with ice-cold 10% glycerol. Approximately 400 ng of the PCR products were transformed via electroporation at 2.5 kV, 25 μF, and 200 Ω using a 2-mm cuvette. The deletion mutant was chosen in an LB medium containing 30 μg/mL Cm. The deletion of malQ was confirmed by PCR. The Cm cassette was removed from E. coli ∆malQ using a Cre expression plasmid pJW.
Culture conditions
To construct two recombinant E. coli strains, E. coli [pKK-ECGDH1 + pACYC-PQQ] and E. coli ∆malQ [pKK-ECGDH1 + pACYC-PQQ], pKK-ECGDH1 and pACYC-PQQ were transformed into E. coli JM109 and E. coli JM109 ∆malQ using a standard heat-shock transformation protocol.
Single colonies of E. coli strains were cultivated in 10 mL of nutrient broth (NB, 1 g/L beef extract, 2 g/L yeast extract, 5 g/L peptone, and 5 g/L NaCl). The cells were cultivated at 25 °C and 200 rpm for 24 h and then used as seed cultures for shaker flask and bioreactor experiments.
Shaker-flask culture experiments were conducted in 300-mL baffled flasks containing 50 mL of NB medium with 200 g/L maltose, 30 g/L CaCO3, and proper antibiotics. Pre-cultured bacteria were inoculated into 50 mL of NB medium and grown in a shaking incubator at 25 °C and 200 rpm.
Bioreactor conditions
Batch fermentation was carried out in a 5 L fermenter (BioCNS, Korea) containing 2 L of NB medium with 200 g/L maltose and 30 g/L CaCO3 at 25 °C. The agitation speed was adjusted from 200 to 1000 rpm to maintain the dissolved oxygen (DO) level at 30%. The added CaCO3 helped to keep the pH of the growth media between 6.0 and 7.2.
Analytical methods
Cell growth was assayed by a spectrophotometer (UV-2600, Shimadzu, Japan) at a wavelength of 600 nm (OD600nm). The concentrations of maltose and MBA in the culture medium were determined using high-performance liquid chromatography (HPLC) equipment (Agilent 1260, USA). The HPLC system was equipped with an ICSep ICE-ION-300 column (Transgenomic Inc., New Haven, CT, USA) and a refractory index detector (1260 Infinity II RID, Agilent). HPLC analysis conditions were as follows: a mobile phase of 0.5 mM H2SO4 was used at a flow rate of 0.3 mL/min, and the column temperature was set to 75 °C.
Results
MBA production by recombinant E. coli strain
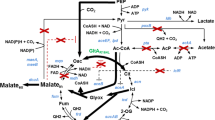
In this study, we first investigated whether E. coli [pKK-ECGDH1 + pACYC-PQQ] could produce MBA. As shown in Figure S1, E. coli [pKK-ECGDH1 + pACYC-PQQ] successfully produced MBA. However, in contrast to our previous results [10], the production yield (77.4%) of MBA from maltose obtained from this E. coli strain was lower than that obtained from P. taetrolens strain used in our previous study (100%; data not shown). This result implies that some maltose was not utilized for MBA synthesis in E. coli. Therefore, we attempted to identify the maltose utilization gene in E. coli. From a literature survey, we found that malQ is the first gene of the maltose utilization pathway in E. coli (Fig. 1) [18,19,20]. Therefore, in subsequent experiments, we attempted to inactivate malQ and investigated the effect of amylomaltase (MalQ) inactivation on MBA production.
Effect of MalQ inactivation on maltose utilization in E. coli
To investigate the effect of MalQ inactivation on maltose utilization in E. coli, the malQ gene in E. coli was knocked out, and this knockout strain (E. coli ∆malQ) was incubated in NB medium with maltose. In contrast to wild-type E. coli, E. coli ∆malQ cannot consume maltose (Fig. 2), confirming that MalQ plays a critical role in maltose utilization in E. coli. Using E. coli ∆malQ [pKK-ECGDH1 + pACYC-PQQ], we attempted to produce MBA from maltose in subsequent experiments.
Comparison of maltose consumption between the wild-type E. coli and E. coli ∆malQ in NB medium. Each strain was incubated in a 300 mL baffled flask containing 50 mL NB medium with 200 g/L maltose and 30 g/L CaCO3 with 200 rpm rotary agitation at 25 °C. Time course of cell growth (A) and maltose consumption (B). Error bars represent the standard deviation derived from three independent experiments
MBA production by recombinant E. coli ∆malQ strain
Escherichia coli ∆malQ [pKK-ECGDH1 + pACYC-PQQ] and E. coli [pKK-ECGDH1 + pACYC-PQQ] were cultivated in flasks containing NB medium containing maltose. The MBA production titer, yield, and productivity of E. coli ∆malQ [pKK-ECGDH1 + pACYC-PQQ] were 209.3 g/L, 100%, and 1.1 g/L/h, which were 1.3-, 1.3-, 2.2-fold higher than those of E. coli [pKK-ECGDH1 + pACYC-PQQ] (162.1 g/L, 77.4%, and 0.5 g/L/h), respectively (Fig. 3). This result indicates that all the available maltose was converted into MBA by E. coli ∆malQ [pKK-ECGDH1 + pACYC-PQQ], in contrast to E. coli [pKK-ECGDH1 + pACYC-PQQ]. Therefore, we found that E. coli ∆malQ [pKK-ECGDH1 + pACYC-PQQ] was more suitable for producing MBA from maltose.
Comparison of MBA production between E. coli [pKK-ECGDH1 + pACYC-PQQ] and E. coli ∆malQ [pKK-ECGDH1 + pACYC-PQQ] in NB medium. Each strain was incubated in a 300 mL baffled flask containing 50 mL NB medium with 200 g/L maltose and 30 g/L CaCO3 with 200 rpm rotary agitation at 25 °C. Time course of cell growth (A), maltose consumption (B), and MBA production (C). MBA maltobionic acid
To scale up MBA production, batch fermentation of E. coli ∆malQ [pKK-ECGDH1 + pACYC-PQQ] was conducted in a 5-L fermenter. After fermentation, the MBA production titer, yield, and productivity were 209.3 g/L, 100%, and 1.5 g/L/h, respectively (Fig. 4). This experiment established a successful MBA production scaled up in a 5-L fermenter.
Discussion
In the present study, we successfully produced MBA using a recombinant E. coli strain for the first time. Moreover, we established that the inactivation of MalQ, the first enzyme of the maltose utilization pathway in E. coli, could improve the MBA-producing capacity of the recombinant E. coli strain (Fig. 5). However, the MBA-producing ability of this E. coli strain was relatively low compared to that of P. taetrolens obtained in our previous study [10]. Therefore, further improvement in the MBA-producing ability of the E. coli strain is needed. In a previous study, we enhanced the LBA-producing ability of an E. coli strain by changing the type of GDH [15]. To secure a new E. coli strain with improved MBA-producing ability, our future work will involve expressing various new GDHs recombinantly in E. coli and assessing their MBA-producing abilities. Further work will focus on improving the MBA-producing ability by the protein engineering of GDH1 from E. cloacae.
In addition, we are trying to improve the MBA-producing ability of the recombinant E. coli strain by enhancing the PQQ synthesis. In this study, the PQQ synthesis gene cluster was cloned into pACYC184, which was a low copy number plasmid having a weak constitutive tet promoter, and thus, the expression level of PQQ synthesis gene cluster might be rather low. Hence, the synthesis level of PQQ in the E. coli strain might also be low, which might be insufficient to activate all the GDH expressed in the E. coli strain for efficient MBA production. In our previous works, we tried to produce LBA using E. coli [pKK-ECGDH1], in which the PQQ synthesis gene cluster (pACYC-PQQ) was not co-expressed [15]. When 10 μM PQQ was added, E. coli [pKK-ECGDH1] successfully produced 209.3 g/L LBA with a productivity of 2.75 g/L/h. In contrast, E. coli [pKK-ECGDH1 + pACYC-PQQ] produced 209.3 g/L LBA with a productivity of 0.85 g/L/h, of which MBA productivity was only 30.9% of that obtained from E. coli [pKK-ECGDH1] with external PQQ addition. This result implied that PQQ was the limiting factor for LBA production. Although the accurate PQQ concentration synthesized in E. coli [pKK-ECGDH1 + pACYC-PQQ] was unknown, the PQQ concentration might be lower than 10 μM, implying the PQQ synthesis level of E. coli [pKK-ECGDH1 + pACYC-PQQ] was insufficient. From these results, enhancing the PQQ concentration in the E. coli strain by increasing the expression level of the PQQ synthesis gene cluster during MBA production might improve the MBA-producing ability of the E. coli strain. Thus, we are constructing a recombinant E. coli strain to increase the synthesis level of PQQ.
Recently, consumers have preferred plant-derived cosmetic components over animal-derived components because of concerns regarding virus and prion contamination of animal-derived products. Consequently, materials derived from plant are generally regarded as safer for humans than animal-derived materials. LBA has been widely used in cosmetics as a moisturizing and peeling agent. However, LBA is generated from animal milk, because LBA is made from lactose. MBA is a plant-derived substance produced from maltose, a starch-derived disaccharide. Because the physicochemical properties of MBA are highly similar to those of LBA, MBA can also be used as a moisturizing and peeling agent in cosmetics. MBA is expected to be an alternative for LBA in the future because it can appeal to cosmetic consumers.
To further expand the use of MBA, lowering the price is highly desirable. The price of raw materials, particularly maltose, accounts for a substantial portion of the MBA production price. In our previous studies, we tried to produce MBA using cheap substrates. Using high-maltose corn syrup (HMCS), with a price approximately 1.1% of that of pure maltose, we successfully produced MBA [10]. Moreover, MBA can be produced using waste-cooked rice, which is significantly cheaper than HMCS and pure maltose [21]. Our study can help lower the production cost of MBA and extend the applications in industrial areas.
Conclusions
In the present study, a recombinant E. coli strain capable of producing MBA from maltose without the addition of PQQ was created by co-expressing the heterologous GDH gene and the PQQ synthesis gene cluster. The MBA-producing ability of this E. coli strain was enhanced by the inactivation of the maltose utilization pathway in E. coli. After batch fermentation of the E. coli strain using a 5-L fermenter, the MBA production titer, yield, and productivity of the strain were 209.3 g/L, 100%, and 1.5 g/L/h, respectively. Our future work will involve the cultivation of this recombinant strain at a high cell density and the production of MBA via whole-cell biocatalysis.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Abbreviations
- GDH1:
-
Glucose dehydrogenase
- HMCS:
-
High-maltose corn syrup
- HPLC:
-
High-performance liquid chromatography
- LBA:
-
Lactobionic acid
- MalQ:
-
Amylomaltase
- MBA:
-
Maltobionic acid
- PQQ:
-
Pyrroloquinoline quinone
- WCB:
-
Whole-cell biocatalyst
References
Suehiro D et al (2019) Maltobionic acid enhances intestinal absorption of calcium and magnesium in rats. Biosci Biotechnol Biochem 83(9):1766–1773
Alonso S, Rendueles M, Díaz M (2013) Bio-production of lactobionic acid: current status, applications and future prospects. Biotechnol Adv 31(8):1275–1291
Gutiérrez L-F, Hamoudi S, Belkacemi K (2012) Lactobionic acid: a high value-added lactose derivative for food and pharmaceutical applications. Int Dairy J 26(2):103–111
Suehiro D et al (2020) Maltobionic acid accelerates recovery from iron deficiency-induced anemia in rats. Biosci Biotechnol Biochem 84(2):393–401
Green BA, Yu RJ, Van Scott EJ (2009) Clinical and cosmeceutical uses of hydroxyacids. Clin Dermatol 27(5):495–501
Sarenkova I, Ciprovica I (2018) The current status and future perspectives of lactobionic acid production : a review. Res Rural Dev 1:233–239
Cardoso T et al (2019) Lactobionic acid as a potential food ingredient: recent studies and applications. J Food Sci 84(7):1672–1681
Fukami K et al (2016) Effect of Water Content on the glass transition temperature of calcium maltobionate and its application to the characterization of non-arrhenius viscosity behavior. Food Biophys 11(4):410–416
Mirescu A, Prüße U (2007) A new environmental friendly method for the preparation of sugar acids via catalytic oxidation on gold catalysts. Appl Catal B 70(1):644–652
Oh Y-R et al (2020) High-level production of maltobionic acid from high-maltose corn syrup by genetically engineered Pseudomonas taetrolens. Biotechnol Rep 28:e00558
Oh Y-R et al (2022) Whole-cell biocatalysis using genetically modified Pseudomonas taetrolens for efficient production of maltobionic acid from pure maltose and high-maltose corn syrup. Bioprocess Biosyst Eng 45(5):901–909
Rosano GL, Ceccarelli EA (2014) Recombinant protein expression in Escherichia coli: advances and challenges. Front Microbiol 5:172–172
Sharma A, Chaudhuri TK (2017) Revisiting Escherichia coli as microbial factory for enhanced production of human serum albumin. Microb Cell Fact 16(1):173
Oh Y-R et al (2020) Enhancement of lactobionic acid productivity by homologous expression of quinoprotein glucose dehydrogenase in Pseudomonas taetrolens. J Agric Food Chem 68(44):12336–12344
Han HJ et al (2022) Efficient production of lactobionic acid using Escherichia coli capable of synthesizing pyrroloquinoline quinone. J Agric Food Chem 70(6):1962–1970
Han HJ, Oh Y-R, Eom GT (2022) Isolation and characterization of a new superior lactobionic acid-producing bacterium, Enterobacter cloacae KRICT-1, from environmental soil samples. ACS Food Sci Technol 2(1):66–74
Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci 97(12):6640–6645
Boos W, Shuman H (1998) Maltose/maltodextrin system of Escherichia coli: transport, metabolism, and regulation. Microbiol Mol Biol Rev 62(1):204–229
Park JT et al (2011) Role of maltose enzymes in glycogen synthesis by Escherichia coli. J Bacteriol 193(10):2517–2526
Weiss SC, Skerra A, Schiefner A (2015) Structural basis for the interconversion of maltodextrins by MalQ, the amylomaltase of Escherichia coli. J Biol Chem 290(35):21352–21364
Oh Y-R, Jang Y-A, Eom GT (2022) Valorization of waste cooked rice into value-added maltobionic acid using genetically engineered Pseudomonas taetrolens. ACS Sustain Chem Eng 10(2):810–815
Funding
This work was supported partly by the R&D programs of MOTIE/KEIT (20018375), KRICT (SS2242-10, BSF22-515), and Ulsan-KRICT (US22-03).
Author information
Authors and Affiliations
Contributions
CC: Investigation, validation, data curation, writing–original draft. GTE: Conceptualization, project administration, supervision, writing–original draft, writing–review and editing, funding acquisition, resources.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no financial or commercial conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cho, C., Eom, G.T. Enhanced production of maltobionic acid by a metabolically engineered Escherichia coli incapable of maltose utilization. Bioprocess Biosyst Eng 46, 507–513 (2023). https://doi.org/10.1007/s00449-022-02835-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00449-022-02835-4