Abstract
In the current study, the effect of different available nitrogen sources on erythromycin fermentation by Saccharopolyspora erythraea No. 8 is evaluated. Three different combinations of corn steep liquor and yeast powder were developed to investigate their impacts on erythromycin production. The results indicate that the optimal combination of available nitrogen sources was 10.0 g/L corn steep liquor and 4.0 g/L yeast power, generating a maximum yield of erythromycin of 13672 U/mL. To explore the effects of nitrogen perturbations on cell metabolism, metabolic flux analyses were performed and compared under different conditions. A high flux pentose phosphate pathway provided more NADPH for erythromycin synthesis via nitrogen optimization. Moreover, high n-propanol specific consumption rate enhanced erythromycin synthesis and n-propanol flowed into the central carbon metabolism by methylmalonyl-CoA node. These results indicate that the selection of an appropriate organic nitrogen source is essential for cell metabolism and erythromycin synthesis, and this is the first report of the successful application of available nitrogen source combinations in industrial erythromycin production.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Erythromycin is a type of macrocyclic antibiotic with a 14-membered ring molecular structure. Due to structural similarity, erythromycin is divided into four main components, i.e., erythromycin A, B, C, and D. Erythromycin A has the strongest antibacterial activity and is the major product during the erythromycin fermentation process. In contrast, erythromycin B and C are by-products, which can be removed in a downstream separation process. Erythromycin A was first isolated from Saccharomyces in 1952 [1, 2]. It is clinically used as a treatment for Gram-positive bacterial infections. The mechanism of erythromycin antibacterial activity is dependent on to its binding to the 50S subunit of the ribosomes in bacteria, subsequently inhibiting the protein translation process and associated cellular functions [3, 4].
Improvement in the erythromycin production by microbial fermentation process is urgently needed due to the increased global demand for erythromycin in clinical applications. In recent years, genetic engineering technology has developed as a powerful and promising tool which could change microbial metabolic properties and intensify synthesis pathways of target metabolites efficiently, complementing traditional strain evolution by mutagenesis [5]. Chen et al. [6] knocked out the coding gene mutB which encodes the beta subunit of methylmalonyl-CoA mutase in the modification of Saccharopolyspora erythraea HL3168 E3, and the erythromycin A titer was substantially improved. Wang et al. [7] integrated S-adenosyl methionine (SAM) synthetase gene from Streptomyces spectabilis into the genome of S. erythraea to supply methyl precursor in erythromycin A synthesis. Consequently, the erythromycin A titer was increased 132% in the reconstructed strain, whereas the erythromycin B content was reduced by 30%. In addition, Minas et al. [8] integrated the Vitreoscilla hemoglobin gene (vhb) into S. erythraea to produce 70% more erythromycin in this way.
In contrast, medium and process controlling optimizations are the major methods to improve the final titer in industrial antibiotic fermentation process. El-Enshasy et al. [9] used inexpensive cane molasses as the sole carbon source and corn steep liquor (CSL) and ammonium sulfate as the nitrogen sources to obtain an erythromycin titer of 600 mg/L. Subsequently, an optimized n-propanol feeding strategy was adopted to further enhance the titer to 720 mg/L. Devi et al. [10] used bagasse as a carbon source to increase erythromycin yield by 28% compared to glucose by S. erythraea MTCC 1103. Zou et al. [11] applied a combined strategy of medium optimization by response surface methodology and process controlling optimization through on-line physiological parameter regulation to improve erythromycin production, generating a titer of 10622 U/mL.
Nitrogen is essential for cell growth and erythromycin production. It has been reported that erythromycin biosynthesis is initiated when amino nitrogen or phosphorus are in a restricted state [12, 13]. Some complex nitrogen sources such as soybean meal, CSL, and yeast extract can enhance antibiotic production by Streptomycetes [14]. These three nitrogen sources have also been applied in erythromycin fermentation. Zou et al. [15] used CSL and yeast extract in erythromycin fermentation to enhance the erythromycin A titer. They reported that a higher erythromycin A titer and ratio of erythromycin A to erythromycin C was obtained by maintaining a suitable feeding rate of ammonium sulphate, and this strategy has been successfully scaled up from laboratory scale to a 25 m3 fermenter. Viscosity is an important parameter for characterizing rheology properties in the filamentous fermentation process. High broth viscosity leads to mass transfer limitation (oxygen and nutrients) and environmental heterogeneities, which further affect cell growth and product synthesis [16]. Low dissolved oxygen (DO) tension can result in the accumulation of erythromycin B [17]. Consequently, viscosity is an important process parameter which should be carefully considered in erythromycin fermentation process. Chen et al. [18] explored the effects of nitrogen sources on broth viscosity and glucose consumption in erythromycin fermentation. The addition of ammonium sulphate decreases broth viscosity and glucose consumption in the fermentation process.
Metabolic flux analysis contributes to a more comprehensive understanding of microbial metabolic properties. Chen et al. [19] designed glucose and n-propanol feeding strategies during erythromycin fermentation process, and the relationship of glucose and n-propanol consumption was explored via quantitative metabolic flux analysis. The results revealed that n-propanol could flow into central carbon metabolism by the succinyl-CoA node in S. erythraea ZL1004. The deletion of the mutB gene verified that the relative ratio of n-propanol flowing into erythromycin synthesis was increased significantly [6].
Available nitrogen sources play a vital role in the regulation of antibiotic fermentation. There are very few studies investigating the optimization of different combinations of available organic nitrogen sources in erythromycin fermentation. In this study, the initial experiments investigated available organic nitrogen source optimization in erythromycin fermentation process. To study the effect of available nitrogen source combination on cell metabolism, we conducted quantitative metabolic flux analysis and investigated the flux distribution of glucose and n-propanol under different nitrogen conditions.
Materials and methods
Microorganism
S. erythraea No.8 was kindly provided by Yidu HEC Biochem. Co. Ltd. (Hubei province, China).
Media and culture
Preliminary seed medium contained (g/L): sugar 30.0, CSL 16.0, CaCO3 7.0, (NH4)2SO4 2.0, soybean oil 2.0, antifoam agent 0.6, pH 7.0.
Secondary seed medium contained (g/L): starch 20.0, dextrin 30.0, soybean meal 35.0, glucose 15.0, CSL 5.0, yeast powder 3.0, (NH4)2SO4 1.5, NaCl 2.0, CaCO3 5.0, antifoam agent 0.2, pH 7.0.
Fermentation medium contained (g/L): starch 50.0, soybean meal 35.0, (NH4)2SO4 2.0, NaCl 2.0, CaCO3 5.0, soybean oil 5.0, antifoam agent 0.2, pH 7.0.
Agar slants were incubated at 34 °C for 7 days, and the matured spores were then used for seed culture inoculation. The primary seed culture was conducted in 500 mL shake flasks containing 50 mL preliminary seed medium. The primary seed was transferred into a 15 L fermenter containing 8 L of the secondary seed medium, and then cultivated for 45 h at 34 °C. The fermentation was conducted in a 50 L fermenter (FUS-50, Shanghai Guoqiang Bioengineering Equipment Co. Ltd., Shanghai, China) containing 25 L fermentation medium at 34 °C for 150 h. Dissolved oxygen (DO) was monitored with polarographic DO electrode (Mettler Toledo, Switzerland) and was kept above 40% air saturation by adjusting agitation and aeration during fermentation. Process pH was measured with a pH electrode (Mettler Toledo, Switzerland).
Optimization of nitrogen sources
Based on the principle of equal total nitrogen, three modes of nitrogen source combination in the initial media were designed as listed in Table 1.
Analytical methods
Determination of packed mycelium volume (PMV), total sugar and amino nitrogen
Ten milliliters of fermentation broth were centrifuged at 4000 rpm for 15 min. The PMV value was the percentage of precipitate volume to broth volume (v/v). Total sugar concentration was determined by the DNS method [15]. The concentration of amino nitrogen was measured by the formaldehyde titration method [20]. Ten mL of broth was filtered through a pre-weighed filter paper and dried at 80 °C for 24 h [19].
Determination of erythromycin, n-propanol, and soybean oil concentrations
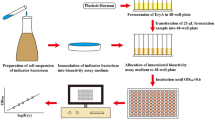
The chemical titer of erythromycin was determined by the modified phosphoric acid colorimetric method [11]. The components of erythromycin were analyzed by high performance liquid chromatography (HPLC) (Agilent 1200, USA) with Waters Xbridge C18 column (4.6 mm × 250 mm, 5 μm, Waters Corporation, Ireland). The mobile phase was a 5.7 g/L solution of dipotassium hydrogen phosphate solution and acetonitrile with a ratio of 60% and 40%. The flow rate and column temperature were 0.9 mL/min and 35 °C, respectively. A UV detector was applied at a wavelength of 215 nm, and the injection volume was 20 mL [21]. The concentration of n-propanol was measured with an electronic nose instrument [22]. The residual soybean oil was measured by the low field nuclear magnetic resonance method (PQ001-010, Niumag, Shanghai, China) [6].
Determination of oxygen and carbon dioxide concentrations in exhaust gas
The concentrations of oxygen and carbon dioxide in the exhaust gas were determined by a process mass spectrometer (MAX300-LG, Extrel, USA). Oxygen uptake rate (OUR), carbon dioxide evolution rate(CER) and respiratory quotient (RQ) were calculated online by the Biostar software.
Metabolic flux analysis
The stoichiometric model adopted in this study was based on the results described by Chen et al. [19] as shown in Table S1.
Results and discussion
Comparison of biological activities under different nitrogen conditions
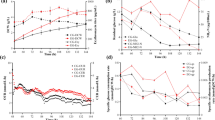
As shown in Fig. 1a, there was little difference in PMV between the three modes. The PMV was increased to above 40.0% before 50 h, and different available nitrogen combinations had little influence on cell growth. Figure 1b, c shows that total sugar and amino nitrogen decreased significantly before 40 h, as the cells mainly conducted growth metabolism in this phase. Figure 1e indicates that the viscosity of mode 1 and mode 3 were much higher than that in mode 2. During most of fermentation process, the viscosities of mode 1 and mode 3 were both above 200 s, and the viscosity of mode 1 approached 800 s at 80 h. In contrast, the viscosity of mode 2 stayed within a relatively reasonable range during the entire fermentation process. High broth viscosity has been reported to have negative effects on erythromycin fermentation [18]. Thus, it might have contributed to the high erythromycin titer of mode 2, which was 27.8% and 34.9% higher than those in mode 1 and mode 3, respectively (Fig. 1d). Figure 1f shows that the glucose consumption rate of mode 1 and propanol consumption rate of mode 2 were quite high among the three modes. The available nitrogen combination of mode 2 enhanced the utilization of n-propanol by cells. It could be speculated that more n-propanol flowed into the erythromycin synthesis pathway, leading to a high erythromycin production rate after 80 h in mode 2 (Fig. 1d). Based on these analyses, it can be concluded that an available nitrogen source combination of 10.0 g/L CSL and 4.0 g/L yeast powder is favorable for erythromycin synthesis. In addition, the highest erythromycin titer obtained in our previous work was less than 12500 U/mL [6, 19], while the current study substantially improved erythromycin production to approximately 14000 U/mL. Nitrogen optimization had no influence on the composition of erythromycin component, and thus, the erythromycin titer was used to assess the effect of available nitrogen source optimization.
Comparison of cell respiratory metabolism under different nitrogen conditions
During the cell growth phase (0–40 h), OURs of mode 1 and mode 3 were clearly higher than that of mode 2 (Fig. 2a). The high oxygen consumption rate indicates that the cell respiration rates of mode 1 and mode 3 were elevated in the early fermentation phase. Once the cells entered into the secondary metabolism, the main cellular physiological activity transitioned to metabolism maintenance and erythromycin synthesis. During the rapid erythromycin production period after 69 h, the CER of mode 2 was lower than mode 1 and mode 3 (Fig. 2b). In addition, RQ was adopted to characterize the carbon sources utilization by cells. A low RQ trend appears when cells consume reductive carbon sources. During this period, the RQ value of mode 2 was lower than mode 1 and mode 3 (Fig. 2c), suggesting that the cell might metabolize more reductive ingredients in mode 2 [19]. The reductive nutrient ingredients in the medium included n-propanol and soybean oil. There was a small change in specific consumption rate of soybean oil during fermentation, and the high n-propanol consumption rate was consistent with a low level of CER during rapid erythromycin synthesis phase in mode 2. As a result, it is concluded that CER fluctuation was caused by the consumption of n-propanol, and more n-propanol flowed into the erythromycin synthesis pathway in mode 2. Figure 1d indicates that the erythromycin production rate of mode 2 increased rapidly at 80 h compared with other modes. Thus, it is speculated that the cells utilized n-propanol rapidly to synthesize erythromycin in this period, resulting in the highest erythromycin titer in mode 2.
Metabolic flux analysis analyses under different nitrogen perturbation conditions
The above analyses of metabolic parameters in fermentation process indicate that nitrogen perturbations enhance n-propanol consumption to synthesize erythromycin. However, the detailed intracellular metabolic responses to nitrogen perturbations are still unclear, and metabolic flux analysis can be used to investigate the cellular metabolism.
Metabolic flux analysis was calculated based on the principles of metabolite balances. The period selected for the metabolic flux calculation was based on the balance of carbon atoms. This study analyzed the recoveries of the carbon atoms in three different modes at different time periods. During 69–81 h, the recoveries of the carbon atoms of mode 1, mode 2, and mode 3 were in the range of 95.4–101.3%, meeting the criteria for further metabolic flux calculations.
Metabolic flux calculation results are shown in Fig. 3. Overall, the specific glucose consumption rates of mode 1, mode 2, and mode 3 decreased gradually. The specific n-propanol consumption rate of mode 2 was the highest, and the specific consumption rates of soybean oil were similar among three modes. From the perspective of glucose metabolism, the glucose of mode 2 flowing into the pentose phosphate (PP) pathway, which is the main source of NADPH supply, was as high as 0.466 mmol g−1 day−1, which is 60.1% and 42.1% higher than those in mode 1 and mode 3, respectively. Among the n-propanol metabolism, the maximum flux flowing into erythromycin synthesis (r17) was 0.085 mmol g−1 day−1. The proportions of n-propanol participating in erythromycin synthesis were similar in three modes, which were all about 10.0%. The higher n-propanol specific consumption rate resulted in higher propanol flux into erythromycin, subsequently promoting erythromycin synthesis. Zou et al. [15] enhanced erythromycin production by feeding different available nitrogen sources, and Maeda et al. [23] improved cellulase production by available nitrogen sources optimization. These studies suggest that nitrogen perturbation is an effective approach to improve the production of target products.
Methylmalonyl-CoA is a key node affecting the synthesis of erythromycin. One molecule of erythromycin A is synthesized from one molecule of propionyl-CoA and six molecules of methylmalonyl-CoA [24]. Reeves et al. [25] increased the copy number of the methylmalonyl-CoA mutase operon in wild-type strains, increasing the erythromycin titer by 50%. Another study also suggested that methylmalonyl-CoA represents an important precursor for macrocyclic polyketide antibiotics [26]. The synthesis rate of methylmalonyl-CoA is the rate-limiting step of erythromycin biosynthesis. From the metabolic flux calculation results, the flux rates from the methylmalonyl-CoA node to succinyl-CoA (r16) of three modes were 0.263 mmol g−1 day−1, 0.373 mmol g−1 day−1 and 0.198 mmol g−1 day−1, respectively. More n-propanol flowed towards the tricarboxylic acid (TCA) cycle via methylmalonyl-CoA node in mode 2. Comparing with n-propanol flux rates of three modes into the succinyl-CoA node, the flux of mode 2 was the highest. There were two ways to supply methylmalonyl-CoA for erythromycin synthesis pathway in S. erythraea: the reversible isomerization of succinyl-CoA and the carboxylation of propionyl-CoA. It has been reported that the latter is the primary approach to fill the methylmalonyl-CoA pool in S. erythraea [27]. Therefore, n-propanol, as the direct donor of propionyl-CoA, can significantly affect cellular metabolism in erythromycin fermentation. This result indicates that a high consumption rate of n-propanol results in high n-propanol flux into the TCA cycle. Erythromycin titers suggest that the n-propanol metabolism of mode 2 was more efficient for erythromycin synthesis. It is concluded that the nitrogen allocation of mode 2 stimulates cells to consume more n-propanol. If the n-propanol flowing into the TCA cycle is reduced, erythromycin production will be further improved.
There were two methods to improve the n-propanol flux towards erythromycin synthesis, i.e., improvement of the specific consumption rate of n-propanol and reduction of the n-propanol flux to the TCA cycle. The former mainly depended on medium optimization in industrial fermentation. Previous reports have verified that reducing the n-propanol flux to the TCA cycle can be achieved by genetic engineering technology [6, 28]. Knockdown of the mutB gene encoding the methylmalonyl-CoA mutase completely block the flow of n-propanol to the succinyl-CoA from methylmalonyl-CoA. One equivalent of propionyl-CoA and six equivalents of methylmalonyl-CoA can be catalyzed by polyketide synthase to synthesize 6-Deoxyerythronolide B (6-DEB), an intermediate in the erythromycin synthesis pathway. Therefore, accelerating bioconversion of methylmalonyl-CoA and propionyl-CoA to 6-DEB could also reduce the n-propanol flux towards the TCA cycle. Overexpression of the DEBS gene clusters (eryAI–III) can improve the yield of 6DEB [29]. By increasing copy number of the PKS gene clusters, methylmalonyl-CoA is rapidly drained to synthesize 6-DEB, rather than succinyl-CoA. Thus, the most of methylmalonyl-CoA in the limited methylmalonyl-CoA pool would be distributed to the erythromycin synthesis pathway. However, n-propanol could be diverted to flow towards the erythromycin synthesis pathway.
Multi-omics analysis revealed that an alternative branched-chain amino acid biosynthesis pathway could supply methylmalonyl-CoA in high yield S. erythraea in which the several relevant genes of TCA cycle was downregulated [30]. For nitrogen source utilization, organic nitrogen sources provide a large number of branched-chain amino acids for cells, e.g., leucine, valine, and isoleucine. Thus, nitrogen perturbation represents a potential pathway to promote metabolic flow towards erythromycin synthesis.
The specific generation and consumption rates of ATP and NADPH are shown in Table 2. ATP is an energy substance required for the growth and maintenance of cells, and NADPH is required for anabolism and erythromycin synthesis. According to the results of each metabolic reaction flux calculation, there was little difference in the total ATP synthesis rates of the three modes. This indicates that the combination of different available nitrogen sources concentrations has little effect on energy supply. More than 53.1% of ATP was produced from glucose, indicating that the glucose was used as the main energy source and greater than 28.4% of ATP was provided by soybean oil, suggesting that soybean oil was the second alternative energy source in the erythromycin fermentation process. NADPH acting as reducing power for cell anabolism affected the synthesis of metabolites. The yield of target product can be increased by improving the synthesis of NADPH [31, 32]. Erythromycin synthesis requires 9 molecules of NADPH for the synthesis of each molecule of erythromycin A. The NADPH synthesis rate was the highest in mode 2, and about 82.3% of NADPH was used for erythromycin synthesis. It can be concluded from the metabolic flux analysis that the nitrogen combination of mode 2 strengthened the PP pathway, and a greater percentage of NADPH participated in erythromycin synthesis. The high NADPH and its ratio to erythromycin were important factors resulting in the high erythromycin titer of mode 2.
Metabolic flux distributions of glucose and n-propanol in cells are shown in Table 3. Glucose is the main energy source, and most of the glucose was consumed by the central metabolism. Likewise, NADPH is required for erythromycin synthesis, as 2 molecules of glucose are required for the synthesis of each molecule of erythromycin A. 6.3% of consumed glucose entered into erythromycin synthesis in mode 2, the highest ratio among three modes. The ratio of glucose to erythromycin and PP pathway reached 23.5%, suggesting that glucose metabolism and flux allocation promotes erythromycin synthesis. In the present three modes, a large proportion of n-propanol entered into the TCA cycle. N-propanol was the precursor of erythromycin [33], and more than 50.2% of n-propanol was used in the erythromycin synthesis.
Nitrogen perturbations are an effective strategy in erythromycin production. There was report about that feeding CSL and yeast powder enhanced erythromycin synthesis. Zou et al. [15] increased erythromycin titer by 11.3% and 11.6%, respectively, by feeding CSL and yeast powder. In addition, feeding CSL and yeast powder had no effect on the ratio of erythromycin A to impurity component. This was consistent with the above results. Moreover, the optimization in this paper provided one promising strategy in erythromycin fermentation by nitrogen combination in initial media. The erythromycin titer was improved by more than 20%. On the other hand, there was study about carbon source and precursor combination regulation in erythromycin fermentation in our lab. Chen et al. [19] explored the relationship of glucose and n-propanol consumption by the strategy of feeding the combination of glucose and n-propanol. The glucose and n-propanol uptake by cells can be regulated to promote the consumption of n-propanol and the flux of n-propanol to erythromycin was also increased. The results of this experiment revealed that glucose consumption by cells was increased during available organic nitrogen sources optimization process; consequently, the glycolysis pathway and pentose phosphate pathway were promoted. They were indispensable for intracellular erythromycin synthesis pathway. Guo et al. [34] optimized the feed rate of propanol during erythromycin fermentation process to increase the erythromycin titer to 10950 U/mL. Therefore, nitrogen source or carbon source regulation influences erythromycin synthesis in an effective way. In addition, Chen et al. [35] found that glucose respiratory quotient (RQ)-feedback control improved erythromycin titer and erythromycin A concentration more effective than glucose pH-feedback control strategy. From the above study about process technology optimization, nitrogen media optimization and carbon or precursor feed strategy optimization provided higher selectivity and feasibility for erythromycin fermentation improvement. Therefore, adding CSL and yeast powder in basal media combined with the optimized feed strategy of glucose and n-propanol in process was worth exploring in erythromycin fermentation. Related to this experiment, Chen et al. [6] improved the erythromycin output by the knockout the coding gene mutB. It was different from nitrogen source optimization that the erythromycin purity was improved significantly. Chen et al. [36] constructed two engineered strains by increasing the copy number of eryK and eryG, which are responsible for encoding methylase and hydroxylase, respectively. This led to the 25% improvement of erythromycin yield and the elimination of by-products. The process optimization of erythromycin recombinant strain was important topic in the fermentation field. Systematic erythromycin fermentation improvement can be achieved by genetic modification combined with fermentation process optimization when inheritance and expression stability of strain is intensified.
Based on the previous studies, GlnR, a global regulator of nitrogen metabolism in S. erythraea, has a central role in erythromycin synthesis [37]. Liao et al. [38] reported that GlnR also regulates the utilization of multiple types of carbohydrates in erythromycin producing strains by interacting with the carbohydrate ATP-binding cassette (ABC) transporter. ABC-transported carbohydrates include maltose, the hydrolysis product of starch. These results identify a correlation between nitrogen source and carbon source utilization in S. erythraea. Therefore, nitrogen optimization in erythromycin has a global impact on cellular metabolism. This supports the development of organic nitrogen combination strategies in industrial erythromycin fermentation.
Conclusion
The yield of erythromycin was improved by combining CSL and yeast powder in basal medium. The erythromycin titer was improved to 13672 U/mL, which is the highest value ever reported. Metabolic flux analysis revealed that higher n-propanol and NADPH fluxes flowing into erythromycin synthesis in optimal nitrogen source combinations contribute to improved erythromycin production. In addition, n-propanol flowing into the TCA cycle at the succinyl-CoA node is an inefficient pathway in erythromycin fermentation which could be improved by genetic engineering.
References
Wiley PF, Gerzon K, Flynn EH, Weaver O, Quarck UC, Chauvette RR, Monahan R (1957) Erythromycin. X1. Structure of erythromycin. J Am Chem Soc 79(22):6062–6070
Mironov VA, Sergienko OV, Nastasyak IN, Danilenko VN (2004) Biogenesis and Regulation of Biosynthesis of Erythromycins in Saccharopolyspora erythraea. Appl Biochem Microbiol 40(6):531–541
Taubman SB, Young FE, Corcoran JW (1963) Antibiotic glycosides, IV. studies on the mechanism of erythromycin resistance in Bacillus subtilis. Proc Natl Acad Sci USA 50(5):955–962
Taubman SB, Jones NR, Young FE, Corcoran JW (1966) Sensitivity and resistance to erythromycin in Bacillus subtilis 168: the ribosomal binding of erythromycin and chloramphenicol. Biochim Biophys Acta 123(2):438–440
Baltz RH (2006) Molecular engineering approaches to peptide, polyketide and other antibiotics. Nat Biotechnol 24(12):1533–1540
Chen CC, Hong M, Chu J, Huang MZ, Ouyang LM, Tian XW, Zhuang YP (2017) Blocking the flow of propionate into TCA cycle through a mutB knockout leads to a significant increase of erythromycin production by an industrial strain of Saccharopolyspora erythraea. Bioprocess Biosyst Eng 40(2):201–209
Wang Y, Wang YG, Chu J, Zhuang YP, Zhang LX, Zhang SL (2007) Improved production of erythromycin A by expression of a heterologous gene encoding S-adenosylmethionine synthetase. Appl Microbiol Biotechnol 75(4):837–842
Minas W, Brunker P, Kallio PT, Bailey JE (1998) Improved erythromycin production in a genetically engineered industrial strain of Saccharopolyspora erythraea. Biotechnol Prog 14(4):561–566
El-Enshasy HA, Mohamed NA, Farid MA, El-Diwany AI (2008) Improvement of erythromycin production by Saccharopolyspora erythraea in molasses based medium through cultivation medium optimization. Bioresour Technol 99(10):4263–4268
Devi CS, Saini A, Rastogi S, Naine SJ, Mohanasrinivasan V (2015) Strain improvement and optimization studies for enhanced production of erythromycin in bagasse based medium using Saccharopolyspora erythraea MTCC 1103. 3 Biotech 5(1):23–31
Zou X, Hang HF, Chen CF, Chu J, Zhuang YP, Zhang SL (2008) Application of oxygen uptake rate and response surface methodology for erythromycin production by Saccharopolyspora erythraea. J Ind Microbiol Biotechnol 35(12):1637–1642
Potvin J, Péringer P (1994) Ammonium regulation in Saccharopolyspora erythraea. Part I: Growth and antibiotic production. Biotechnol Lett 16(1):63–68
Martin JF, Demain AL (1980) Control of antibiotic biosynthesis. Microbiol Rev 44(2):230–251
Rafieenia R (2013) Effect of nutrients and culture conditions on antibiotic synthesis in Streptomycetes. Asian J Pharm Hea Sci 3(3):810–815
Zou X, Hang HF, Chu J, Zhuang YP, Zhang SL (2009) Enhancement of erythromycin A production with feeding available nitrogen sources in erythromycin biosynthesis phase. Bioresour Technol 100(13):3358–3365
Lara AR, Galindo E, Ramírez OT, Palomares LA (2006) Living with heterogeneities in bioreactors. Mol Biotechnol 34(3):355–381
Chen CF, Qi XC, Qian JC, Zhuang YP, Chu J, Zhang SL (2009) Effects of the dissolved oxygen on the erythromycin components of recombinant strain Saccharopolyspora erythraea ZLl004 fermentation. Chin J Antibio 34(11):659–663
Chen Y, Wang ZJ, Chu J, Zhuang YP, Zhang SL, Yu XG (2013) Significant decrease of broth viscosity and glucose consumption in erythromycin fermentation by dynamic regulation of ammonium sulfate and phosphate. Bioresour Technol 134(2):173–179
Chen Y, Huang MZ, Wang ZJ, Chu J, Zhuang YP, Zhang SL (2013) Controlling the feed rate of glucose and propanol for the enhancement of erythromycin production and exploration of propanol metabolism fate by quantitative metabolic flux analysis. Bioprocess Biosyst Eng 36(10):1445–1453
Zou X, Hang HF, Chu J, Zhuang YP, Zhang SL (2009) Oxygen uptake rate optimization with nitrogen regulation for erythromycin production and scale-up from 50 L to 372 m3 scale. Bioresour Technol 100(3):1406–1412
Zou X, Zeng W, Chen CF, Qi XC, Qian JC, Chu J, Zhuang YP, Zhang SL, Li WJ (2010) Fermentation optimization and industrialization of recombinant Saccharopolyspora erythraea strains for improved erythromycin a production. Biotechnol Bioprocess Eng 15(6):959–968
Zhao HT, Pang KY, Lin WL, Wang ZJ, Gao DQ, Guo MJ, Zhuang YP (2016) Optimization of the n-propanol concentration and feedback control strategy with electronic nose in erythromycin fermentation processes. Process Biochem 51(2):195–203
Maeda RN, Silva MMPD, Anna LMMS Jr (2010) Nitrogen source optimization for cellulase production by Penicillium funiculosum, using a sequential experimental design methodology and the desirability function. Appl Biochem Biotechnol 161(1–8):411–422
Corcoran JW (1981) Biochemical mechanisms in the biosynthesis of the erythromycins. Springer, Berlin
Reeves AR, Brikun IA, Cernota WH, Leach BI, Gonzalez MC, Weber JM (2007) Engineering of the methylmalonyl-CoA metabolite node for increased erythromycin production in oil-based fermentations of Saccharopolyspora erythraea. Metab Eng 9(3):293–303
Jung WS, Yoo YJ, Park JW, Park SR, Han AR, Ban YH, Kim EJ, Kim E, Yoon YJ (2011) A combined approach of classical mutagenesis and rational metabolic engineering improves rapamycin biosynthesis and provides insights into methylmalonyl-CoA precursor supply pathway in Streptomyces hygroscopicus ATCC 29253. Appl Microbiol Biotechnol 91(5):1389–1397
Li YY, Chang X, Yu WB, Li H, Ye ZQ, Yu H, Liu BH, Zhang Y, Zhang SL, Ye BC (2013) Systems perspectives on erythromycin biosynthesis by comparative genomic and transcriptomic analyses of S. erythraea E3 and NRRL23338 strains. BMC Genom 14(1):523
Reeves AR, Brikun IA, Cernota WH, Leach BI, Gonzalez MC, Weber JM (2006) Effects of methylmalonyl-CoA mutase gene knockouts on erythromycin production in carbohydrate-based and oil-based fermentations of Saccharopolyspora erythraea. J Ind Microbiol Biotechnol 33(7):600–609
Kumpfmüller J, Methling K, Fang L, Pfeifer BA, Lalk M, Schweder T (2016) Production of the polyketide 6-deoxyerythronolide B in the heterologous host Bacillus subtilis. Appl Microbiol Biotechnol 100(3):1209–1220
Karničar K, Drobnak I, Petek M, Magdevska V, Horvat J, Vidmar R, Baebler Š, Rotter A, Jamnik P, Fujs Š (2016) Integrated omics approaches provide strategies for rapid erythromycin yield increase in Saccharopolyspora erythraea. Microb Cell Fact 15(1):93
Lee WH, Chin YW, Han NS, Kim MD, Seo JH (2011) Enhanced production of GDP-L-fucose by overexpression of NADPH regenerator in recombinant Escherichia coli. Appl Microbiol Biotechnol 91(4):967–976
Ahmad I, Shim WY, Jeon WY, Yoon BH, Kim JH (2012) Enhancement of xylitol production in Candida tropicalis by co-expression of two genes involved in pentose phosphate pathway. Bioprocess Biosyst Eng 35(1–2):199–204
Potvin J, Péringer P (1993) Influence of N-propanol on growth and antibiotic production by an industrial strain of Streptomyces erythreus under different nutritional conditions. Biotechnol Lett 15(5):455–460
Guo Q, Chu J, Zhuang YP, Gao Y (2016) Controlling the feed rate of propanol to optimize erythromycin fermentation by on-line capacitance and oxygen uptake rate measurement. Bioprocess Biosyst Eng 39(2):255–265
Chen Y, Wang Z, Chu J, Xi B, Zhuang Y (2015) The glucose RQ-feedback control leading to improved erythromycin production by a recombinant strain Saccharopolyspora erythraea ZL1004 and its scale-up to 372-m3 fermenter. Bioprocess Biosyst Eng 38(1):105–112
Chen Y, Deng W, Wu JQ, Qian JC, Chu J, Zhuang YP, Zhang SL, Liu W (2008) Genetic modulation of the overexpression of tailoring genes eryK and eryG leading to the improvement of erythromycin a purity and production in Saccharopolyspora erythraea fermentation. Appl Environ Microbiol 74(6):1820–1828
Yao LL, Liao CH, Huang G, Zhou Y, Rigali S, Zhang BC, Ye BC (2014) GlnR-mediated regulation of nitrogen metabolism in the actinomycete Saccharopolyspora erythraea. Appl Microbiol Biotechnol 98(18):7935–7948
Liao CH, Yao L, Xu Y, Liu WB, Zhou Y, Ye BC (2015) Nitrogen regulator GlnR controls uptake and utilization of non-phosphotransferase-system carbon sources in actinomycetes. Proc Natl Acad Sci USA 112(51):15630–15635
Acknowledgements
This work was financially supported by the National Key Research and Development Program (2017YFB0309302), the National Basic Research Program of China (973 Program) (No. 2012CB721006), National Natural Science Foundation of China (No. 21276081) and the Fundamental Research Funds for the Central Universities (22221818014).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, Q., Hang, H., Tian, X. et al. Combined available nitrogen resources enhanced erythromycin production and preliminary exploration of metabolic flux analysis under nitrogen perturbations. Bioprocess Biosyst Eng 42, 1747–1756 (2019). https://doi.org/10.1007/s00449-019-02171-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00449-019-02171-0