Abstract
The homeodomain transcription factors (TFs) Pax6 (OMIM: 607108) and Prox1 (OMIM: 601546) critically regulate gene expression in lens development. While PAX6 mutations in humans can cause cataract, aniridia, microphthalmia, and anophthalmia, among other defects, Prox1 deletion in mice causes severe lens abnormalities, in addition to other organ defects. Furthermore, the optimal dosage/spatiotemporal expression of these key TFs is essential for development. In lens development, Pax6 expression is elevated in cells of the anterior epithelium compared to fiber cells, while Prox1 exhibits the opposite pattern. Whether post-transcriptional regulatory mechanisms control these precise TF expression patterns is unknown. Here, we report the unprecedented finding that the cataract-linked RNA-binding protein (RBP), Celf1 (OMIM: 601074), post-transcriptionally regulates Pax6 and Prox1 protein expression in lens development. Immunostaining shows that Celf1 lens-specific conditional knockout (Celf1cKO) mice exhibit abnormal elevation of Pax6 protein in fiber cells and abnormal Prox1 protein levels in epithelial cells—directly opposite to their normal expression patterns in development. Furthermore, RT-qPCR shows no change in Pax6 and Prox1 transcript levels in Celf1cKO lenses, suggesting that Celf1 regulates these TFs on the translational level. Indeed, RNA-immunoprecipitation assays using Celf1 antibody indicate that Celf1 protein binds to Pax6 and Prox1 transcripts. Furthermore, reporter assays in Celf1 knockdown and Celf1-overexpression cells demonstrate that Celf1 negatively controls Pax6 and Prox1 translation via their 3′ UTRs. These data define a new mechanism of RBP-based post-transcriptional regulation that enables precise control over spatiotemporal expression of Pax6 and Prox1 in lens development, thereby uncovering a new etiological mechanism for Celf1 deficiency-based cataract.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Defects in gene regulation during lens development result in ocular abnormalities such as congenital cataract (Anand and Lachke 2017; Cvekl and Zhang 2017; Shiels and Hejtmancik 2019). Indeed, several key transcription factors (TFs) have been identified to control the gene regulatory network in lens development (Lachke and Maas 2010; Cvekl and Zhang 2017). The lens comprises of anteriorly localized epithelial cells that exit the cell cycle in what is termed the “transition zone” to differentiate into posteriorly localized fiber cells. Specific transcription factors that control expression of key downstream targets in the anterior epithelium and fiber cells have been identified, and their mutations have been linked to human ocular defects. For example, mutations in the anterior epithelium-expressed TF genes PAX6 and FOXE3 are linked to a variety of ocular defects in humans including congenital cataract and Peters anomaly, among others (Cvekl and Callaerts 2017; Anand et al. 2018a). On the other hand, the fiber cell-expressed TF genes PROX1 and MAF are linked to cataract in human or animal models (Wigle et al. 1999; Audette et al. 2016; Anand et al. 2018a). Furthermore, precise dosage and spatiotemporal expression of key TFs is necessary for normal lens development, disruption of which causes ocular defects. For example, optimal dosage of Pax6 is critical for lens and eye development. Indeed, PAX6 deficiency or mutation results in a range of ocular phenotypes, and its overexpression causes defects in lens fiber cells, resulting in cataract (Glaser et al. 1994; Schedl et al. 1996; Duncan et al. 2004).
However, the mechanisms that regulate the precise dosage and cell-specific expression of Pax6 or Prox1 in the lens are still not comprehensively understood. While some aspects of signaling and transcription-based control over Pax6 have been described, it is currently unknown whether RNA-binding protein (RBP) mediated post-transcriptional gene expression control—defined as control of alternative splicing, localization, decay, or translation of mRNA—is involved in regulation of Pax6 expression. Even less is known about Prox1 regulation in the lens. This represents a critical knowledge-gap as post-transcriptional regulation offers an additional layer of control for achieving optimal spatiotemporal expression and dosage of proteins (Dash et al. 2016).
Recently, we showed that deficiency of an iSyTE-predicted conserved lens-enriched RBP Celf1—involved in post-transcriptional gene expression control—causes lens defects and/or cataract (Kakrana et al. 2018; Siddam et al. 2018; Anand et al. 2018b; Aryal et al. 2020). Celf1 protein harbors three RNA recognition motifs (RRMs) that allows binding to GU-rich element (GRE) in target mRNAs. Celf1 protein binding to its targets can result in distinct post-transcriptional regulatory outcomes such as alternative splicing, mRNA decay, and translational control (Dasgupta and Ladd 2012; Vlasova-St Louis et al. 2013). Previously, we uncovered the molecular mechanism that explains the fiber cell denucleation defects in Celf1 knockout mice (Siddam et al. 2018). Specifically, we showed that Celf1 directly regulates the nuclease Dnase2b and the tumor suppressor and cyclin-dependent kinase inhibitor p27Kip1 to facilitate nuclei degradation in fiber cell maturation. However, we had not examined the role of Celf1 in mediating post-transcriptional gene expression control over key lens transcription factors.
Here, we report the surprising finding that Celf1 functions to control the dosage and cell-specific expression of the conserved TFs Pax6 and Prox1 in lens development. Lens-specific conditional deletion of Celf1 in mice causes misexpression of Pax6 in mature fiber cells and misexpression of Prox1 in anterior epithelial cells—directly opposite to their normal expression pattern in lens development. Furthermore, the dosage of both proteins—but not their transcripts—is strikingly elevated in these cells, suggesting that Celf1 likely regulates these TFs by a post-transcriptional control mechanism that operates at the level of translation inhibition. In support of this hypothesis, RNA-immunoprecipitation assays demonstrate that Celf1 protein binds to Pax6 and Prox1 mRNAs in the lens. Furthermore, reporter assay using the 3′ UTR of these mRNAs coupled with Celf1 knockdown and overexpression shows that Celf1 represses the translation of these proteins via their 3′ UTRs. These data provide critical new insights on how a conserved post-transcriptional regulatory RNA-binding protein controls the spatiotemporal expression and dosage of key transcription factors in the lens, in turn serving to define a new regulatory pathway implicated in cataract pathology.
Materials and methods
Animal studies
The animal protocols were reviewed and approved by the University of Delaware Institutional Animal Care and Use Committee (IACUC). Animal experiments were performed according to the Association for Research in Vision and Ophthalmology (ARVO) statement for the use of animals in ophthalmic and vision research. Celf1 lens-specific conditional knockout mice were generated as previously described (Siddam et al. 2018). In brief, mice carrying Celf1 conditional knockout allele in which Celf1 exon five is flanked by loxP sites (Celf1flox) were crossed with mice carrying Celf1 germline targeted knockout allele (Celf1lacZKI) and Pax6GFPCre transgenic mouse line that express Cre recombinase in the embryonic day (E) 9.5 lens placode to generate Celf1 compound conditional knockout mice (Pax6GFPCre±:Celf1flox/lacZKI; referred to as Celf1cKO). Mouse embryonic stages were determined by designating the day that the vaginal plug was observed as embryonic day (E) 0.5. Postnatal mouse stages were determined by designating the day of birth as postnatal day 0 (P0). As controls, the following mice were used, Celf1flox/flox, Pax6GFPCre±:Celf1+/+ genotype or C57BL/6 wild type, none of which exhibited lens defects. All experiments were performed in three biological replicates unless stated otherwise.
Immunofluorescence
Mouse head tissue at stages E14.5, E16.5, and mouse eye tissue at P10 were fixed in 4% PFA for 30 min on ice followed by incubation in 30% sucrose at 4 °C overnight and mounted in OCT (Tissue-Tech, Doral, FL). Frozen tissue from comparable depths in the eye was sectioned at 14 μm thickness. Tissue sections were blocked in either 5% chicken serum (Abcam, Cambridge, UK; for Prox1 staining) or 2% BSA (Sigma-Aldrich, St.Louis, MO; for Pax6 staining) along with 0.1% Tween at room temperature for 1 h. The primary antibodies were diluted in the blocking buffer as follows: Celf1 (Abcam #129115, 1:500 dilution), Prox1 (Covance #238C, 1:1000 dilution), and Pax6 (Millipore #Ab2237, 1:200 dilution). After 1 h blocking, the sections were incubated with the primary antibody overnight at 4 °C. The following day, the sections were washed and incubated for 1 h at room temperature with the appropriate secondary antibody conjugated to Alexa Fluor 594 (1:200) (Life Technologies, Carlsbad, CA) mixed with 1:1000 the nuclear stain DAPI (40,6-diamidine-2-phenylidole-dihydrochloride; Life Technologies, Carlsbad, CA; #D21490). Slides were washed, mounted using mounting media (Reed et al. 2001), and imaged using the Zeiss LSM 880 Confocal microscope configured with Diode/Argon laser (405 nm, 488 nm and 594 excitation lines) (Carl Zeiss Inc, Oberkochen, Germany). Optimal adjustment of brightness/contrast was performed uniformly across control and knockout samples in Adobe Photoshop (Adobe, San Jose, CA, USA). The Fiji ImageJ software (NIH, Bethesda, MD) was used to quantify the differences in the fluorescence signal intensity of Pax6 and Prox1 between E16.5 control and Celf1cKO lens (n = 3 biological replicates) as follows. Images were split into single channel and the fluorescence intensity of the region of interest, in this case, representing 20 individual nuclei, was measured in the red channel (representing Pax6 or Prox1) and the blue channel (representing DNA). The Student’s two sample t test was used for measuring the significance of the normalized difference between control and Celf1cKO samples.
Western blot analysis
For cell lines, the lysates were prepared as previously described (Siddam et al. 2018). Briefly, 1 ml lysis buffer [50 mM Tris–HCl at pH 8, 150 mM NaCl, 1% nonidet P40, 0.1% sodium dodecyl sulfate (SDS), 0.5% sodium deoxycholate, and protease inhibitors (1 X dilution) (Thermo Fisher Scientific, Waltham, MA)] was added to the culture petri-dish and incubated for 30 min at 4 °C. This was followed by centrifugation at 14,000 RPM for 30 min at 4 °C. The Pierce BCA protein kit (Thermo Fisher Scientific, Waltham, MA) was used to estimate protein concentration and the lysate total protein (25–50 μg) was resolved on TGX stain-free polyacrylamide gels (Bio-Rad, Hercules, CA Hercules, CA). This was transferred onto PVDF membrane (Thermo Fisher Scientific, Waltham, MA) and blocked with 5% non-fat dry milk for 1 h at room temperature. This was followed by overnight incubation at 4 °C with the following primary antibodies: Celf1 (Abcam ab-9549, 1:500 dilution), Pax6 (Millipore #Ab2237, 1:200 dilution), and Prox1 (ProteinTech #11067–2-AP, 1:500 dilution). The membrane blots were incubated with secondary antibodies conjugated to horseradish peroxidase (Cell Signaling Technology, Danvers, MA) for 1 h at room temperature, and the resulting signals were detected using the SuperSignal™ West Femto Maximum Sensitivity Substrate (Thermo Fisher Scientific, Waltham, MA).
RNA isolation and RT-qPCR
The micro-dissected lens tissues (each biological replicate comprised two lenses; total three biological replicates) were collected from control and Celf1cKO lenses at stage P0 and total RNA was extracted using RNeasy mini kit (Qiagen, Hilden, Germany Qiagen #74104). For isolated lens epithelium, which was micro-dissected as previously described (West-Mays et al. 2010), RNA extraction was performed using the PicoPure RNA Isolation Kit (Thermo Scientific #KIT0204). The cDNA was synthesized using iScript cDNA synthesis kit (Bio-Rad #1708890EDU) and RT-qPCR was performed as described on QuantStudio™ 6 Flex Real-Time PCR System, 96-well Fast, laptop system (Applied Biosystems, Foster City, CA) using Power Syber Green PCR Master Mix (Invitrogen Life Technologies, Carlsbad, CA # 4367659). Signals for test genes were normalized to those of the housekeeping gene Gapdh. Differential expression was determined using ΔΔCT method (Agrawal et al. 2015). The following primers were used for RT-qPCR assays: Celf1-F-5′-ACAGATGAAGCCTGCTGACA-3′ and Celf1-R-5′-CTCTGCTCAAGCCATCAGGT-3′; Gapdh-F-5′-GGGTGTGAACCACGAGAAATA-3′ and Gapdh-R-5′-CTGTGGTCATGAGCCCTTC-3′; Pax6-F-5′-GCACATGCAAACACACATGA-3′ and Pax6-R-5′-ACTTGGACGGGAACTGACAC-3′. Prox1-F-5′-AGTAAGACATCACCGCGTGCGC-3′ and Prox1-R-5′-GCTGGGCACAGCTCAAGAATCCC-3′. Statistical significance for RT-qPCR data was determined using nested ANOVA as previously described (Bookout and Mangelsdorf 2003; Audette et al. 2016; Siddam et al. 2018).
RNA immunoprecipitation
RNA immunoprecipitation (RIP) using the Celf1 antibody and IgG control was performed according to the manufacturer’s instructions (EMD Millipore, Billerica, MA, 17-700). Briefly, C57BL/6 wild-type mouse lens lysates at stage P15 were used for RIP (n = 15 lenses per replicate; total 3 biological replicates). Pre-conjugation of the Celf1 antibody (EMD Millipore, Billerica, MA, 03–104) and the IgG antibody (control) with magnetic beads was performed for 45 min at room temperature and the unbound antibody was removed by washing with the provided buffer. This was followed by subjecting the lens protein lysate to the bead–antibody complex and overnight incubation at 4 °C. The bound-RNA was isolated by phenol–chloroform extraction was assayed by RT-PCR. Beta-2-Microglobulin (B2M) (representing negative control) and p27Kip1 (representing positive control) were used as additional controls in RIP-RT-PCR as previously described (Zheng and Miskimins 2011; Siddam et al. 2018).
Cell culture
The mouse lens epithelium-derived cell line 21EM15 was obtained from Dr. John Reddan (Oakland University, MI) and cultured as previously described (Terrell et al. 2015). Briefly, 21EM15 or NIH3T3 cells were cultured at 37 °C in a humid chamber with 5% CO2 on 100 mm cell culture treated plates (Eppendorf) in DMEM media [10 mL of DMEM with 4.5 g/L glucose, l-glutamine, and sodium pyruvate included (Corning Cellgro, Manassas, VA, 10–013-CV)], 10% Fetal Bovine Serum (Fisher Scientific, Pittsburg, PA, 03-600-511), and 1% penicillin–streptomycin (GE Healthcare Life Sciences, Logan, UT, SV30010). Stable Celf1 knockdown was achieved by Celf1-specific shRNA lentiviral transduction in the mouse lens cell line 21EM15 and validated by Western blot analysis (Siddam et al. 2018). Transient Celf1 KD was performed in NIH3T3 cells using siRNAs. Transient Celf1 overexpression was performed in 21EM15 and NIH3T3 cell lines as described (Siddam et al. 2018).
Luciferase reporter assays
To test translational repression of Pax6 or Prox1, 2055 bp sequence from Pax6 3′ UTR (mm10 coordinates chr2:105,696,356–105,698,410) or 1439 bp from Prox1 3′ UTR (mm10 coordinates chr1:190,121,781–190,123,219) was cloned downstream of firefly luciferase in the pmirGLO Dual-Luciferase reporter vector (Promega, catalog no. E1330). This vector was transfected into Celf1 knockdown and control cells (NIH3T3 cell line), and in a separate experiment, it was also co-transfected with the Celf1 overexpression vector using Lipofectamine 3000 (Thermo Fisher Scientific, catalog no. L3000008). After 48 h, lysate was collected and dual-luciferase quantification was performed using Promega Dual-Luciferase Reporter Assay System (Promega, catalog no. E1910) and the PromegaTM GloMaxTM 20/20 Luminometry System (Promega, Madison, WI). Firefly luciferase activity was normalized to Renilla luciferase activity. Statistical significance for luciferase assays was determined by two-tailed Student’s t test.
Results
Celf1cKO mouse lens exhibits abnormal expression of Pax6 and Prox1 proteins
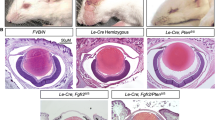
We recently showed that Celf1cKO mice exhibit early onset cataract accompanied by fiber cell defects, including absence of de-nucleation during fiber cell maturation (Siddam et al. 2018). Compared to control (Fig. 1a), the lens defects in Celf1cKO mice are detected early in embryonic development (Fig. 1b). In our previous study, we identified that misregulation of p27Kip1 and Dnase2b served to explain the nuclear degradation defects in Celf1cKO mice (Siddam et al. 2018). However, we did not examine the impact of Celf1 deficiency on the key lens TFs, Pax6, and Prox1, which exhibit a specific spatiotemporal expression pattern in lens development. Indeed, Pax6 protein is highly expressed in the lens anterior epithelium and its expression is found to be reduced in late differentiating fiber cells in normal lens development (Fig. 1c). Conversely, Prox1 protein is highly expressed in fiber cells and is absent in the lens anterior epithelium in normal lens development (Fig. 1d). Therefore, we sought to investigate the effect of Celf1 deficiency on these key lens TFs. Immunostaining shows that Celf1cKO mouse lens exhibits abnormally high levels of Pax6 protein (Fig. 2a). This defect is observed across multiple embryonic stages starting from E14.5 (Fig. 2a). Furthermore, the elevation of Pax6 protein in Celf1cKO lens was most striking in fiber cells compared to epithelial cells (Fig. 2b). Fluorescence quantification shows a significant increase in Pax6 protein in Celf1cKO mouse lenses, specifically in fiber cells, but not in the epithelium (Fig. 2c). Moreover, immunostaining shows that Celf1cKO mouse lens exhibits abnormally high levels of Prox1 protein (Fig. 3a). Please note that the Prox1 expression appears lower than usual in normal lenses, because the microscopy settings had to be adjusted to optimally detect the highly elevated Prox1 protein levels in the Celf1cKO lens. Prox1 protein was found to be generally high in the transition zone and fiber cells of Celf1cKO mouse lens (Fig. 3b). But more interestingly, compared to its absence in the anterior epithelium of control lenses, Prox1 protein was strikingly high in the anterior epithelium of Celf1cKO mouse lens (Fig. 3b). This was observed as early as embryonic day E14.5 as well as in early post-natal stages (Fig. 3a, b). Fluorescence quantification shows a significant increase in Prox1 protein in Celf1cKO epithelium and fiber cells (Fig. 3d). To further validate these data, we performed immunostaining on micro-dissected anterior epithelium from adult Celf1cKO and control lenses. This independently validated that Prox1 protein was elevated in isolated anterior epithelium of Celf1cKO mouse lens (Fig. 3c). Together, these data showed that Celf1 deficiency led to abnormal elevation of Pax6 and Prox1 proteins in Celf1cKO mouse lens.
Lens defects in the Celf1cKO mouse and expression of Pax6 and Prox1 in normal lens development. a, b Comparative histological analysis of control and Celf1cKO mouse shows severe lens defects (asterisk) at embryonic day (E) 16.5. Scale bar 100 μm. c Immunostaining shows that Pax6 protein is expressed in the anterior epithelium (asterisk) and early differentiating fiber cells in E16.5 lens development. d In contrast, immunostaining shows that Prox1 protein is expressed in the fiber cells (asterisk) and not in the anterior epithelium of the lens at E16.5. Abbr.: e anterior epithelium, f fiber cells. Scale bar 75 μm
Celf1cKO mouse lens exhibits abnormally high levels of Pax6 in fiber cells. a Immunostaining at embryonic (E14.5, E16.5) and postnatal (P10) stages shows that Celf1cKO mice exhibit abnormally high Pax6 protein levels (asterisk) starting from stage E16.5. Scale bar 100 μm. b High magnification of area indicated by dotted line box in a shows that compared to epithelial cells, Pax6 protein levels are abnormally high (asterisk) in fiber cells of Celf1cKO mouse lens. Abbr.: e anterior epithelium, f fiber cells. Scale bar 50 μm. c Fluorescence quantification of E16.5 lens shows significant increase of Pax6 protein specifically in Celf1cKO lens fiber cells (asterisk represents p value < 0.004), but not in the epithelium (NS not significant)
Celf1cKO mouse lens exhibits abnormally elevated levels of Prox1 in the anterior epithelium. a Immunostaining at embryonic (E14.5, E16.5) and postnatal (P10) stages shows that Celf1cKO mice exhibit abnormally high Prox1 protein levels (asterisk) starting from stage E14.5. Scale bar in E14.5 is 100 μm, E16.5 is 20 μm and P10 is 100 μm. Please note that in normal lenses, Prox1 expression appears lower than usual only, because microscopy settings had to be adjusted to optimally detect the comparatively highly elevated Prox1 protein levels in Celf1cKO lens, and the same settings were used for both control and Celf1cKO samples. b High magnification of area indicated by dotted line box in a shows that Prox1 protein levels are high (asterisk) in both epithelium and fiber cells in Celf1cKO mouse. However, compared to fiber cells, Prox1 protein levels are abnormally high in anterior epithelium (asterisk) of Celf1cKO mouse lens. Scale bar in E14.5 is 20 μm, E16.5 is 10 μm and P10 is 20 μm. c Immunostaining of micro-dissected isolated lens anterior epithelium shows that Prox1 protein is strikingly high in Celf1cKO mice. High magnification of area indicated by dotted line box in c shows high levels of Prox1 protein in nuclei of the anterior epithelium (asterisk) of Celf1cKO mouse lens. Abbr.: e anterior epithelium, f fiber cells, tz transition zone. Scale bar is 20 μm. d Fluorescence quantification of E16.5 lens epithelium (top) and fiber (bottom) shows a significant increase of Prox1 protein in Celf1cKO lens epithelium and fiber cells compared to control (asterisk represents p value < 0.001 for epithelium and p value < 0.02 for fiber cells)
Pax6 and Prox1 transcript levels are unchanged in Celf1cKO mouse lens
We next sought to examine whether the elevation of Pax6 and Prox1 proteins in the Celf1cKO mouse lens was due to abnormal upregulation of their transcripts. RT-qPCR analysis demonstrates no difference in the transcript levels of Pax6 or Prox1 between the Celf1cKO and control lenses (Fig. 4a, b). To further validate this, we performed RT-qPCR on isolated anterior lens epithelium and isolated fiber cells. These analyses showed that Prox1 transcript levels were unchanged in Celf1cKO mouse lens isolated epithelium or fiber cells (Fig. 4c). Together, these data indicate that the abnormal elevation of Pax6 and Prox1 is due to misregulation on the protein level, rather than the mRNA (or transcription level) level in Celf1cKO mouse lens.
Pax6 and Prox1 transcript levels are unchanged Celf1cKO mouse lens. a RT-qPCR showing no significant change in Pax6 (p value = 0.80) and b Prox1 (p value = 0.78) mRNAs between Celf1cKO and control mouse lens at P0. c Compared to control, RT-qPCR analysis of isolated anterior lens epithelium shows no change in Prox1 mRNA levels in Celf1cKO mice (p value = 0.46). Abbr.: f.c. fold-change, NS not significant
Celf1 protein binds to Pax6 and Prox1 transcripts in mouse lens
The above data suggested that misexpression of Pax6 and Prox1 in Celf1cKO mouse lens was due to their misregulation at the post-transcriptional level. To examine if Celf1 was directly involved in the regulation of these TFs, we first sought to determine whether Celf1 protein could directly bind to Pax6 and Prox1 transcripts. We performed RNA immunoprecipitation (RIP) using a Celf1-specific antibody on early postnatal wild-type mouse lenses (stage P15) followed by RT-PCR using Pax6-specific primers. This analysis demonstrated that both Pax6 and Prox1 transcripts are highly enriched in Celf1 RIP compared to IgG control (Fig. 5a, b). These data suggest that Celf1 protein directly interacts with Pax6 and Prox1 transcripts.
Lens cell lines recapitulate Celf1-based regulation of Pax6 and Prox1
We next sought to gain mechanistic insights into the nature of Celf1-based control over Pax6 and Prox1. We first used the mouse lens cell line, 21EM15 (Sax et al. 1995), which expresses several lens markers (Terrell et al. 2015), and which was previously used to establish stable Celf1 knockdown (Celf1 KD) (Siddam et al. 2018). Both Western blot and immunostaining confirmed knockdown of Celf1 and showed that Pax6 and Prox1 protein are upregulated in Celf1-KD cells (Fig. 6). Similar to Celf1cKO mouse lens, the elevation of these TFs at the protein level was not due to the up-regulation of their transcripts (Fig. 6). Furthermore, Celf1 overexpression in 21EM15 cells led to reduced expression of Pax6 and Prox1 protein expression (Fig. 6). Together, these findings established that Celf1 negatively regulates Pax6 and Prox1 protein expression.
Celf1 negatively regulates Pax6 and Prox1 protein expression in lens cell culture. a Western blot and b immunofluorescence show that Celf1 knockdown (KD) in the mouse lens epithelial cell line, 21EM15, causes elevation of Pax6 and Prox1 proteins (asterisks), similar to that observed in Celf1cKO mice. c Western blot and d immunofluorescence show that Celf1 overexpression (OE) in 21EM15 cells causes a reduction of Pax6 and Prox1 proteins (asterisks). Scale bar 50 μm. e RT-qPCR shows that similar to Celf1cKO mice, Pax6 and Prox1 mRNA levels are not significantly altered in Celf1-KD cells. For Pax6, p value = 0.37 and for Prox1 p value = 0.19. Abbr.: f.c. fold-change, NS not significant
Reporter assays uncover Celf1-based mechanism of control over Pax6 and Prox1 translation via their 3′ UTRs
Previous reports indicate that majority of the exonic CELF1-binding sites are located in the 3′ UTRs of target genes (Le Tonquèze et al. 2016). Therefore, we hypothesized that Celf1 protein binds to Pax6 and Prox1 mRNAs via their 3′ UTRs. Next, we sought to determine the outcome of this interaction. To do so, we performed a luciferase reporter assay using mouse Pax6 and Prox1 3′ UTR sequences that were cloned downstream of the firefly luciferase sequence in the pmirGLO dual-luciferase vector (Fig. 7a). Reporter expression in cells transfected with the Pax6 and Prox1 3′ UTR constructs was significantly elevated in Celf1 knockdown conditions (Fig. 7b). Conversely, upon Celf1 overexpression, the transfection of Pax6 and Prox1 3′ UTR constructs showed significantly reduced reporter expression (Fig. 7c). These reporter assays indicate that Celf1 negatively controls Pax6 and Prox1 translation via their 3′ UTRs, in turn providing new insight into the mechanistic basis of its post-transcriptional control in lens cells.
Reporter assays show that Celf1 negatively regulates Pax6 and Prox1 translation via their 3′ UTRs. a Reporter constructs with either the Pax6 or Prox1 3′ UTRs for use in luciferase assay. b Transfection of Pax6 or Prox1 3′UTRs constructs shows significantly elevated reporter activity in Celf1-KD cells, compared to control. c Conversely, transfection of Pax6 or Prox1 3′UTRs constructs with Celf1 overexpression construct results in significantly reduced reporter activity. d Asterisks indicate p value < 0.05
Discussion
RBPs and microRNAs are recognized as key factors involved in mediating post-transcriptional control of gene expression (Pasquinelli 2012; Castello et al. 2012). Intriguingly, while the human genome encodes over 1500 non-ribosomal RBPs, relatively few have been directly linked to developmental defects or disease, thus highlighting the urgent need to characterize their function in the context of specific tissue development and its associated defects (Lukong et al. 2008; Castello et al. 2012, 2013; Gerstberger et al. 2014; Manning and Cooper 2017; Hentze et al. 2018). Indeed, in the context of lens development and cataract, post-transcriptional control of gene expression remains largely unexplored except for a few recent studies involving Caprin2, Celf1, Tdrd7, Rbm24, and miR-204 (Lachke et al. 2011; Lachke and Maas 2011; Shaham et al. 2013; Choudhuri et al. 2013; Wolf et al. 2013; Xie et al. 2014; Dash et al. 2015, 2016, 2020; Siddam et al. 2018; Shao et al. 2020; Barnum et al. 2020). While the signaling and transcription control of the expression of the key homeodomain TF Pax6—mutations in which are linked to a variety of human eye defects—in early eye and lens development has been well characterized, its regulation by post-transcriptional mechanisms is not well understood (Dash et al. 2016). This is also the case regarding the homeodomain TF Prox1, which is involved in development of multiple organ and tissues, in addition to being critical for lens transparency (Wigle et al. 1999). It is known that the optimal dosage and/or precise spatiotemporal expression of these TFs is necessary for lens development (Cvekl and Zhang 2017). Specifically, Pax6 protein expression is higher in the anterior epithelium and is restricted to early differentiating fiber cells, while the opposite expression pattern needs to be achieved for Prox1 protein, whose expression is restricted to differentiating fiber cells and is absent in the anterior epithelium. How post-transcriptional control may contribute to the generation of these precise expression patterns in the lens is unknown.
Therefore, to address these knowledge deficiencies, we sought to answer a fundamental question, namely, do RBPs involved in post-transcriptional gene expression control regulate the expression of these key TFs in the lens? Our findings show that lens-specific conditional knockout of the RBP, Celf1, in mice results in misexpression of both Pax6 and Prox1 proteins in the lens. Although Pax6 is essential for initiation of fiber cell differentiation (Shaham et al. 2013), its expression needs to be sharply reduced in later stages of differentiating fiber cells to achieve proper levels of crystallin proteins (Duncan et al. 1998, 2004; Cui et al. 2004). Indeed, ectopic expression of Pax6 in fiber cells leads to profound lens defects and cataract, and interestingly, the severity of the cataract defects correlates with the extent of Pax6 elevated expression (Duncan et al. 2004). The mechanism that controls this precipitous reduction in Pax6 expression in normal differentiating fiber cells was unknown. Similarly, while Prox1 mRNA has shown to be expressed in the lens anterior epithelium as well as in fiber cells (Hoang et al. 2014; Zhao et al. 2018), its protein is robustly detected only in fiber cells and not in epithelial cells. Again, the mechanism of how Prox1 protein is absent in the epithelium despite of the presence of its mRNA was unknown. Findings in this report demonstrate that Celf1 protein binds to Pax6 and Prox1 mRNAs and directly inhibits their translation into protein. Moreover, we show that Celf1-based inhibition of these TFs is mediated via their mRNA 3′ UTRs. Thus, these findings represent a major breakthrough in our understanding of the complexity of molecular control over the expression of these TFs in the lens, while also advancing our understanding of the pathology of Celf1 deficiency associated congenital cataract.
However, these findings also open new and intriguing questions. For example, it is interesting to consider how Celf1 protein may suppress Prox1 abundance to a greater extent in the anterior epithelium as compared to fiber cells. Several scenarios that may lead to this outcome can be entertained. For example, it can be hypothesized that Celf1 protein inhibits translation of Prox1 mRNA in both epithelium and fiber cells, but the impact of this inhibitory function is sharply evident only in epithelial cells. This scenario considers that the overall ratio of Prox1 mRNA:Celf1 protein is higher in fiber cells compared to epithelial cells. Indeed, there is evidence to support this, which is as follows. While Celf1 protein is higher in fiber cells (compared to epithelial cells) in early lens development, its abundance is similar in both epithelial and fiber cells in later stages (by E16.5) of lens development (Siddam et al. 2018). Furthermore, there is evidence that Prox1 mRNA is several-fold highly abundant in lens fiber cells compared to epithelial cells (Hoang et al. 2014; Zhao et al. 2018). Thus, the stoichiometric difference between Celf1 protein and Prox1 mRNA is smaller in epithelial cells (i.e., the overall ratio of Prox1 mRNA:Celf1 protein is similar). Consequently, there is sufficient Celf1 protein to effectively inhibit majority of the Prox1 mRNA molecules in epithelial cells. This results in a more effective outcome (in terms of lower abundance of Prox1 protein) due to Celf1-based inhibition of Prox1 mRNA translation in epithelial cells. In contrast, because there is comparatively higher abundance of Prox1 mRNA in fiber cells (i.e., the overall ratio of Prox1 mRNA:Celf1 protein is higher), Celf1 protein cannot inhibit translation on majority of the Prox1 mRNAs molecules, resulting in comparatively higher accumulation of Prox1 protein in fiber cells. This also serves to explain how in addition to transcriptional control, post-transcriptional regulation functions in fine-tuning gene expression dosage in lens cells. Thus, it is not sufficient to produce a certain level of transcripts (based on transcriptional activation), it is also necessary to translate the resulting mRNAs to optimal levels.
An analogous but opposite scenario can be proposed for explaining the effective inhibitory effect of the Celf1 protein on Pax6 abundance in fiber cells but not in epithelial cells. This scenario considers that the overall ratio of Pax6 mRNA:Celf1 protein is higher in epithelial cells compared to fiber cells. Indeed, Pax6 mRNA is several-fold highly abundant in lens epithelial cells compared to fiber cells (Hoang et al. 2014; Zhao et al. 2018). Therefore, the smaller stoichiometric difference between Celf1 protein and Pax6 mRNA in fiber cells results in more effective Celf1-based inhibition of Pax6 mRNA translation—in terms of abundance of Pax6 protein—in this cell type. Because Pax6 mRNA in fiber cells progressively decreases with development (Zhao et al. 2018), this serves to explain why the impact of Celf1 on the abundance of Pax6 protein is higher after E14.5 (Fig. 2b).
Another explanation can be offered in terms of combinatorial control, analogous to that of transcription factors that can function as either activators or repressors depending on specific partner proteins. In this scenario, there are different RBPs (or other post-transcriptional regulatory factors such as miRNAs) expressed in epithelial and fiber cells that can partner with Celf1 to mediate coordinate control over distinct mRNAs. In epithelial cells, the presence of a specific co-repressor RBP allows Celf1 to inhibit Prox1 translation, while, in fiber cells, a different co-repressor RBP allows Celf1 to inhibit Pax6 translation. Indeed, a recent study has shown that there are specific RBPs that are expressed in lens epithelial and fiber cells (Zhao et al. 2019). Furthermore, Celf1 protein can be post-translationally modified. For example, Celf1 protein can be phosphorylated by Protein kinase C (PKC) leading to its increased stability in Myotonic Dystrophy 1 mouse models and patient cells (Kuyumcu-Martinez et al. 2007). Furthermore, phosphorylated Celf1 exhibits increased binding to p21 and C/EBPβ mRNAs in differentiating myoblasts (Salisbury et al. 2008). These findings suggest that post-translational modifications can impact Celf1 stability and function, and future studies can address the nature of Celf1 modifications (or their absence) in the lens epithelium and fiber cells.
From the new findings reported here, we propose a model for the post-transcriptional regulation of Pax6 and Prox1 in the lens (Fig. 8). This model, supported by our reporter assays, shows that Celf1 proteins likely binds to the 3′ UTR of Pax6 and Prox1 mRNAs and inhibits their translation into protein. While not addressed in the present study, it is possible that the 5′ UTRs of Pax6 and Prox1 may also have a role in their translational control by Celf1 protein. The final levels of these proteins may be influenced by several factors. As explained above, these include the differences in stoichiometric levels of Celf1 protein compared to the Pax6 and Prox1 mRNAs and/or presence of different co-repressor partner proteins (not shown) in epithelial and fiber cells. This model serves to explain how Celf1 protein controls the precise dosage and spatiotemporal expression of Pax6 and Prox1 proteins in lens development.
Model for Celf1-based regulation of Pax6 and Prox1 in lens development and cataract. Celf1 protein is expressed at comparable levels in both epithelial and fiber cells in the lens. Celf1 protein can directly bind Pax6 and Prox1 mRNA and inhibit their translation via their 3′ UTRs. The stoichiometric differences between Celf1 protein and Pax6 and Prox1 mRNAs in epithelial vs. fiber cells contribute to effective cell-specific inhibition of translation of Pax6 in fiber cells and of Prox1 in epithelial cells. This model serves to explain how misexpression of the key transcription factors Pax6 and Prox1 contributes to the cataract pathology resulting from Celf1 deficiency
While our previous study using genome-wide microarray analysis showed that Celf1 deficiency results in misexpression of several mRNAs (Siddam et al. 2018), the present study serves to reiterate that Celf1 deletion also results in profound changes in protein levels without an analogous change in their mRNA levels, and moreover, that these changes can be specific to epithelial vs. fiber cells. Thus, the total number of factors impacted by Celf1 deficiency is likely higher than that indicated by transcript profiling analysis. Therefore, it will be interesting to examine the isolated epithelium and isolated fiber cells from Celf1cKO lenses using transcriptomics and proteomics approaches. It will also be interesting to investigate which proteins/factors may partner with Celf1 to co-ordinately mediate post-transcriptional control. Furthermore, the presence of different RBPs may influence the outcome of Celf1-based regulation. Indeed, Celf1 and the RBP Elavl1 can both control translation of CDH1 and MYC mRNAs by interacting with their 3′ UTRs, wherein Celf1 functions as a repressor, while Elavl1 functions as positive regulator (Liu et al. 2015; Yu et al. 2016). Interestingly, Elavl1 protein is also reported to bind to an AU-rich element in the Celf1 mRNA, thereby negatively controlling its levels (Gu et al. 2017), which may add to the complexity of their downstream regulatory impact on the cellular proteome. Finally, besides the mechanisms that have been defined in the present study, it is possible that the changes in protein turnover in Celf1cKO lenses may also contribute to accumulation of Pax6 and Prox1 proteins in the lens, which can be examined in the future.
Together, these new data provide novel molecular insights—involving misregulation of Pax6 and Prox1—into the pathobiology of congenital cataract in Celf1cKO lenses. In addition to Caprin2, Tdrd7 and Rbm24, the present study further advances our understanding of the role of post-transcriptional regulatory RBPs in eye development and defects. Finally, because both Pax6 and Prox1 function in the development and homeostasis of several tissues besides the lens (e.g., Pax6 is critical for sensory organ development, pancreas, and the brain, while Prox1 is critical for the brain, heart, liver, pancreas, and the lymphatic system), and because Celf1 and/or other Celf-family proteins are also expressed in many of these tissues, it will be important to examine whether these new regulatory modules are co-opted in other cell/tissue types and their associated defects.
Conclusion
The RNA-binding protein Celf1 is linked to cataract. However, the mechanism of Celf1 function in normal lens development, and how its perturbation leads to cataract are not comprehensively understood. This report provides new mechanistic evidence, indicating that Celf1 functions at the post-transcriptional level to control the dosage and spatiotemporal expression of the key ocular transcription factors Pax6 and Prox1, both of which are linked to eye defects in human and/or animal models. Because Pax6 and Prox1 are critical to development of other tissues/organs, and because Celf1 and other Celf-family proteins are widely expressed, these new regulatory modules uncovered in the present study will have broader impact on understanding the pathobiology of other developmental defects, in addition to cataract.
References
Agrawal SA, Anand D, Siddam AD et al (2015) Compound mouse mutants of bZIP transcription factors Mafg and Mafk reveal a regulatory network of non-crystallin genes associated with cataract. Hum Genet 134:717–735. https://doi.org/10.1007/s00439-015-1554-5
Anand D, Agrawal SA, Slavotinek A, Lachke SA (2018a) Mutation update of transcription factor genes FOXE3, HSF4, MAF, and PITX3 causing cataracts and other developmental ocular defects. Hum Mutat 39:471–494. https://doi.org/10.1002/humu.23395
Anand D, Kakrana A, Siddam AD et al (2018b) RNA sequencing-based transcriptomic profiles of embryonic lens development for cataract gene discovery. Hum Genet 137:941–954. https://doi.org/10.1007/s00439-018-1958-0
Anand D, Lachke SA (2017) Systems biology of lens development: A paradigm for disease gene discovery in the eye. Exp Eye Res 156:22–33. https://doi.org/10.1016/j.exer.2016.03.010
Aryal S, Anand D, Hernandez FG et al (2020) MS/MS in silico subtraction-based proteomic profiling as an approach to facilitate disease gene discovery: application to lens development and cataract. Hum Genet 139:151–184. https://doi.org/10.1007/s00439-019-02095-5
Audette DS, Anand D, So T et al (2016) Prox1 and fibroblast growth factor receptors form a novel regulatory loop controlling lens fiber differentiation and gene expression. Development 143:318–328. https://doi.org/10.1242/dev.127860
Barnum CE, Al Saai S, Patel SD et al (2020) The Tudor-domain protein TDRD7, mutated in congenital cataract, controls the heat shock protein HSPB1 (HSP27) and lens fiber cell morphology. Hum Mol Genet. https://doi.org/10.1093/hmg/ddaa096
Bookout AL, Mangelsdorf DJ (2003) Quantitative real-time PCR protocol for analysis of nuclear receptor signaling pathways. Nucl Recept Signal 1:e012. https://doi.org/10.1621/nrs.01012
Castello A, Fischer B, Eichelbaum K et al (2012) Insights into RNA biology from an atlas of mammalian mRNA-binding proteins. Cell 149:1393–1406. https://doi.org/10.1016/j.cell.2012.04.031
Castello A, Fischer B, Hentze MW, Preiss T (2013) RNA-binding proteins in Mendelian disease. Trends Genet 29:318–327. https://doi.org/10.1016/j.tig.2013.01.004
Choudhuri A, Maitra U, Evans T (2013) Translation initiation factor eIF3h targets specific transcripts to polysomes during embryogenesis. Proc Natl Acad Sci USA 110:9818–9823. https://doi.org/10.1073/pnas.1302934110
Cui W, Tomarev SI, Piatigorsky J et al (2004) Mafs, Prox1, and Pax6 can regulate chicken betaB1-crystallin gene expression. J Biol Chem 279:11088–11095. https://doi.org/10.1074/jbc.M312414200
Cvekl A, Callaerts P (2017) PAX6: 25th anniversary and more to learn. Exp Eye Res 156:10–21. https://doi.org/10.1016/j.exer.2016.04.017
Cvekl A, Zhang X (2017) Signaling and Gene Regulatory Networks in Mammalian Lens Development. Trends Genet 33:677–702. https://doi.org/10.1016/j.tig.2017.08.001
Dasgupta T, Ladd AN (2012) The importance of CELF control: molecular and biological roles of the CUG-BP, Elav-like family of RNA-binding proteins. Wiley Interdiscip Rev RNA 3:104–121. https://doi.org/10.1002/wrna.107
Dash S, Brastrom LK, Patel SD et al (2020) The master transcription factor SOX2, mutated in anophthalmia/microphthalmia, is post-transcriptionally regulated by the conserved RNA-binding protein RBM24 in vertebrate eye development. Hum Mol Genet 29:591–604. https://doi.org/10.1093/hmg/ddz278
Dash S, Dang CA, Beebe DC, Lachke SA (2015) Deficiency of the RNA binding protein Caprin2 causes lens defects and features of Peters anomaly. Dev Dyn. https://doi.org/10.1002/dvdy.24303
Dash S, Siddam AD, Barnum CE et al (2016) RNA-binding proteins in eye development and disease: implication of conserved RNA granule components. Wiley Interdiscip Rev RNA 7:527–557. https://doi.org/10.1002/wrna.1355
Duncan MK, Haynes JI, Cvekl A, Piatigorsky J (1998) Dual roles for Pax-6: a transcriptional repressor of lens fiber cell-specific beta-crystallin genes. Mol Cell Biol 18:5579–5586. https://doi.org/10.1128/mcb.18.9.5579
Duncan MK, Xie L, David LL et al (2004) Ectopic Pax6 expression disturbs lens fiber cell differentiation. Invest Ophthalmol Vis Sci 45:3589–3598. https://doi.org/10.1167/iovs.04-0151
Gerstberger S, Hafner M, Tuschl T (2014) A census of human RNA-binding proteins. Nat Rev Genet 15:829–845. https://doi.org/10.1038/nrg3813
Glaser T, Jepeal L, Edwards JG et al (1994) PAX6 gene dosage effect in a family with congenital cataracts, aniridia, anophthalmia and central nervous system defects. Nat Genet 7:463–471. https://doi.org/10.1038/ng0894-463
Gu L, Wang H, Wang J et al (2017) Reconstitution of HuR-inhibited CUGBP1 expression protects cardiomyocytes from acute myocardial infarction-induced injury. Antioxid Redox Signal 27:1013–1026. https://doi.org/10.1089/ars.2016.6880
Hentze MW, Castello A, Schwarzl T, Preiss T (2018) A brave new world of RNA-binding proteins. Nat Rev Mol Cell Biol. https://doi.org/10.1038/nrm.2017.130
Hoang TV, Kumar PKR, Sutharzan S et al (2014) Comparative transcriptome analysis of epithelial and fiber cells in newborn mouse lenses with RNA sequencing. Mol Vis 20:1491–1517
Kakrana A, Yang A, Anand D et al (2018) iSyTE 2.0: a database for expression-based gene discovery in the eye. Nucleic Acids Res 46:D875–D885. https://doi.org/10.1093/nar/gkx837
Kuyumcu-Martinez NM, Wang G-S, Cooper TA (2007) Increased steady-state levels of CUGBP1 in myotonic dystrophy 1 are due to PKC-mediated hyperphosphorylation. Mol Cell 28:68–78. https://doi.org/10.1016/j.molcel.2007.07.027
Lachke SA, Alkuraya FS, Kneeland SC et al (2011) Mutations in the RNA granule component TDRD7 cause cataract and glaucoma. Science 331:1571–1576. https://doi.org/10.1126/science.1195970
Lachke SA, Maas RL (2010) Building the developmental oculome: systems biology in vertebrate eye development and disease. Wiley Interdiscip Rev Syst Biol Med 2:305–323. https://doi.org/10.1002/wsbm.59
Lachke SA, Maas RL (2011) RNA granules and cataract. Expert Rev Ophthalmol 6:497–500. https://doi.org/10.1586/eop.11.53
Le Tonquèze O, Gschloessl B, Legagneux V et al (2016) Identification of CELF1 RNA targets by CLIP-seq in human HeLa cells. Genom Data 8:97–103. https://doi.org/10.1016/j.gdata.2016.04.009
Liu L, Ouyang M, Rao JN et al (2015) Competition between RNA-binding proteins CELF1 and HuR modulates MYC translation and intestinal epithelium renewal. Mol Biol Cell 26:1797–1810. https://doi.org/10.1091/mbc.E14-11-1500
Lukong KE, Chang K, Khandjian EW, Richard S (2008) RNA-binding proteins in human genetic disease. Trends Genet 24:416–425. https://doi.org/10.1016/j.tig.2008.05.004
Manning KS, Cooper TA (2017) The roles of RNA processing in translating genotype to phenotype. Nat Rev Mol Cell Biol 18:102–114. https://doi.org/10.1038/nrm.2016.139
Pasquinelli AE (2012) MicroRNAs and their targets: recognition, regulation and an emerging reciprocal relationship. Nat Rev Genet 13:271–282. https://doi.org/10.1038/nrg3162
Reed NA, Oh DJ, Czymmek KJ, Duncan MK (2001) An immunohistochemical method for the detection of proteins in the vertebrate lens. J Immunol Methods 253:243–252
Salisbury E, Sakai K, Schoser B et al (2008) Ectopic expression of cyclin D3 corrects differentiation of DM1 myoblasts through activation of RNA CUG-binding protein, CUGBP1. Exp Cell Res 314:2266–2278. https://doi.org/10.1016/j.yexcr.2008.04.018
Sax CM, Dziedzic DC, Piatigorsky J, Reddan JR (1995) Analysis of alpha-crystallin expression in cultured mouse and rabbit lens cells. Exp Eye Res 61:125–127
Schedl A, Ross A, Lee M et al (1996) Influence of PAX6 gene dosage on development: overexpression causes severe eye abnormalities. Cell 86:71–82. https://doi.org/10.1016/s0092-8674(00)80078-1
Shaham O, Gueta K, Mor E et al (2013) Pax6 regulates gene expression in the vertebrate lens through miR-204. PLoS Genet 9:e1003357. https://doi.org/10.1371/journal.pgen.1003357
Shao M, Lu T, Zhang C et al (2020) Rbm24 controls poly(A) tail length and translation efficiency of crystallin mRNAs in the lens via cytoplasmic polyadenylation. Proc Natl Acad Sci USA 117:7245–7254. https://doi.org/10.1073/pnas.1917922117
Shiels A, Hejtmancik JF (2019) Biology of inherited cataracts and opportunities for treatment. Annu Rev Vis Sci 5:123–149. https://doi.org/10.1146/annurev-vision-091517-034346
Siddam AD, Gautier-Courteille C, Perez-Campos L et al (2018) The RNA-binding protein Celf1 post-transcriptionally regulates p27Kip1 and Dnase2b to control fiber cell nuclear degradation in lens development. PLoS Genet 14:e1007278. https://doi.org/10.1371/journal.pgen.1007278
Terrell AM, Anand D, Smith SF et al (2015) Molecular characterization of mouse lens epithelial cell lines and their suitability to study RNA granules and cataract associated genes. Exp Eye Res 131:42–55. https://doi.org/10.1016/j.exer.2014.12.011
Vlasova-St Louis I, Dickson AM, Bohjanen PR, Wilusz CJ (2013) CELFish ways to modulate mRNA decay. Biochim Biophys Acta 1829:695–707. https://doi.org/10.1016/j.bbagrm.2013.01.001
West-Mays JA, Pino G, Lovicu FJ (2010) Development and use of the lens epithelial explant system to study lens differentiation and cataractogenesis. Prog Retin Eye Res 29:135–143. https://doi.org/10.1016/j.preteyeres.2009.12.001
Wigle JT, Chowdhury K, Gruss P, Oliver G (1999) Prox1 function is crucial for mouse lens-fibre elongation. Nat Genet 21:318–322. https://doi.org/10.1038/6844
Wolf L, Gao CS, Gueta K, et al (2013) Identification and characterization of FGF2-dependent mRNA: microRNA networks during lens fiber cell differentiation. G3 (Bethesda) 3:2239–2255. https://doi.org/10.1534/g3.113.008698
Xie Q, Ung D, Khafizov K et al (2014) Gene regulation by PAX6: structural-functional correlations of missense mutants and transcriptional control of Trpm3/miR-204. Mol Vis 20:270–282
Yu T-X, Gu B-L, Yan J-K et al (2016) CUGBP1 and HuR regulate E-cadherin translation by altering recruitment of E-cadherin mRNA to processing bodies and modulate epithelial barrier function. Am J Physiol Cell Physiol 310:C54–65. https://doi.org/10.1152/ajpcell.00112.2015
Zhao Y, Wilmarth PA, Cheng C et al (2019) Proteome-transcriptome analysis and proteome remodeling in mouse lens epithelium and fibers. Exp Eye Res 179:32–46. https://doi.org/10.1016/j.exer.2018.10.011
Zhao Y, Zheng D, Cvekl A (2018) A comprehensive spatial-temporal transcriptomic analysis of differentiating nascent mouse lens epithelial and fiber cells. Exp Eye Res 175:56–72. https://doi.org/10.1016/j.exer.2018.06.004
Zheng Y, Miskimins WK (2011) CUG-binding protein represses translation of p27Kip1 mRNA through its internal ribosomal entry site. RNA Biol 8:365–371
Funding
This work was supported by National Institutes of Health/National Eye Institute [R01 EY021505 and R01 EY029770 to S.L.], and a grant from Retina France to LP. S.A. and A.D. were supported by a Fight for Sight Summer Student Fellowship and S.A. was also supported by a Sigma Xi award. B.A.T.W. was supported by the Delaware Governor’s Bioscience Fellowship and the Milton H. Stetson Memorial Award. Support from the University of Delaware Core Imaging Facility was made possible through the Institutional Development Award (IDeA) from the National Institutes of Health/National Institute of General Medical Sciences INBRE Program Grant [Grant number P20 GM103446]. Acquisition of the confocal microscope used in this study was funded by the National Institutes of Health/National Center for Research Resources Grant [1S10 RR027273].
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Aryal, S., Viet, J., Weatherbee, B.A.T. et al. The cataract-linked RNA-binding protein Celf1 post-transcriptionally controls the spatiotemporal expression of the key homeodomain transcription factors Pax6 and Prox1 in lens development. Hum Genet 139, 1541–1554 (2020). https://doi.org/10.1007/s00439-020-02195-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-020-02195-7