Abstract
Preimplantation genetic testing for aneuploidy (PGT-A) is widely used in IVF and aims to improve outcomes by avoiding aneuploid embryo transfers. Chromosomal mosaicism is extremely common in early development and could affect the efficacy of PGT-A by causing incorrect embryo classification. Recent innovations have allowed accurate mosaicism detection in trophectoderm samples taken from blastocysts. However, there is little data concerning the impact of mosaicism on viability, and the optimal clinical pathway for such embryos is unclear. This study provides new information concerning the extent to which mosaic preimplantation embryos are capable of producing pregnancies and births. Archived trophectoderm biopsy specimens from transferred blastocysts were analyzed using next generation sequencing (NGS). Unlike other PGT-A methods, NGS accurately detects mosaicism in embryo biopsies. 44 mosaic blastocysts were identified. Their clinical outcomes were compared to 51 euploid blastocysts, derived from a well-matched, contemporary control group. Mosaic embryos were associated with outcomes that were significantly poorer than those of the control group: implantation 30.1 versus 55.8% (P = 0.038); miscarriage rate 55.6 versus 17.2% (P = 0.036); and ongoing pregnancy 15.4 versus 46.2% (P = 0.003). 61% of the mosaic errors affected whole chromosomes and 39% were segmental aneuploidies. Embryo viability is compromised by the presence of aneuploid cells. However, a minority of affected embryos can produce successful pregnancies. Hence, such embryos should not necessarily be excluded, but given a lower priority for transfer than those that are fully euploid. It is recommended that pregnancies established after mosaic embryo transfers be subjected to prenatal testing, with appropriate patient counselling.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In recent years, there has been an unprecedented explosion in the utilization of preimplantation genetic testing for aneuploidy (PGT-A; sometimes referred to as preimplantation genetic screening), a methodology which aims to reveal chromosome abnormalities in embryos produced during in vitro fertilization (IVF) treatment. In current practice, this usually involves the sampling of approximately five trophectoderm (TE) cells at the blastocyst stage (5–6 days after fertilization of the oocyte) and detection of aneuploidy affecting any chromosome using methods, such as array comparative genomic hybridization (aCGH) or next generation sequencing (NGS) (Maxwell et al. 2016). Aneuploidy is extremely common in human embryos and is believed to be the principal cause of unsuccessful IVF treatment, responsible for most implantation failures as well as the majority of miscarriages. In theory, PGT-A should have the capacity to improve certain IVF outcomes by ensuring that embryos transferred to the uterus are chromosomally normal and, therefore, more likely to be compatible with successful implantation and viable pregnancy. The recent publication of several randomised controlled trials, supporting this hypothesis (Yang et al. 2012; Forman et al. 2013; Scott et al. 2013), coupled with innovations that have cut the costs of embryo testing (Wells et al. 2014), has led the number of IVF treatments in the USA that utilize this approach to increase approximately fourfold in the space of just 4 years. It is estimated that over 20% of all IVF treatments in the USA now include PGT-A and adoption continues to increase at a rapid rate.
While most published studies utilizing modern embryo testing methods have suggested that PGT-A is beneficial, controversy over the efficacy of the strategy persists and has begun to crystallise around one issue—chromosomal mosaicism—defined as the presence of two or more cell lines of different karyotype within the same embryo (Delhanty et al. 1993, 1997). Multiple investigations, going back over two decades, have suggested that mosaicism is common in preimplantation embryos produced using IVF (Delhanty et al. 1993, 1997; Magli et al. 2000; Munne et al. 2002; Coonen et al. 2004; Daphnis et al. 2005; Fragouli et al. 2008, 2011). Some mosaic embryos do not contain any chromosomally normal cells (termed mosaic aneuploid), being composed of a mixture of different aneuploid lines. Such embryos should not be problematic for PGT-A as any cells sampled will lead to the same conclusion—the embryo is ‘abnormal’. However, more challenging are those embryos consisting of both normal and aneuploid cells, the existence of which prompts a variety of questions—What is the incidence of such embryos? What is their fate if transferred to the uterus? Do they negatively impact the accuracy and efficacy of PGT-A? With aneuploidy testing rapidly becoming a routine tool for IVF treatment, and already considered standard-of-care in a growing number of clinics, it is vital that we improve our understanding of the clinical consequences of mosaicism.
Until recently, many of the questions posed by mosaicism could not be addressed through the analysis of samples taken during the course of routine PGT-A, because testing strategies typically involved the removal of just one cell (blastomere) from cleavage stage embryos. Consequently, only one cell line could be detected and the embryo was classified as normal or abnormal based on that single result. The move to sampling several TE cells at the blastocyst stage has provided an opportunity to identify mosaicism within the biopsy specimen. However, the TE samples are not separated into individual cells, but instead treated as a single entity for the purposes of genetic analysis and receive a single designation (aneuploid or chromosomally normal). Methods commonly used for the purpose of PGT-A, such as aCGH or quantitative real-time PCR (Fragouli et al. 2011; Treff and Scott, 2013), are capable of examining the entire chromosome complement of TE biopsy specimens, but have suboptimal sensitivity for the detection of low-level mosaicism. Given these limitations, historically, the only way to determine whether an embryo is mosaic has been to take several distinct biopsies and test each separately to reveal whether all specimens are cytogenetically equivalent. Unfortunately, multiple biopsies are detrimental to the embryo, and thus, we find ourselves in a situation, where the procedures required for accurate characterization of the embryo alter the key factor that we want to measure—viability.
This paper presents clinical data obtained using a PGT-A method, based upon NGS (Wells et al. 2014; Fiorentino et al. 2014; Kung et al. 2015; Maxwell et al. 2016). We aimed to examine whether mosaic diploid–aneuploid blastocysts were capable of implanting and leading to ongoing pregnancies, and to assess whether pregnancy outcomes were similar to those of embryos with no detected mosaicism. The obtained results are novel and have important advantages over previously published data concerning mosaic embryo viability. In addition, the current study more than triples the amount of clinical outcome data following mosaic diploid–aneuploid embryo transfers. The accurate identification of cytogenetically heterogeneous biopsy samples allows us to address questions about embryo viability, and about the impact of mosaicism on PGT-A’s clinical utility.
Results
Next generation sequencing analyses produced approximately 700,000–1,000,000 DNA sequence reads per sample, with an average of ~800,000 reads successfully aligned to the human genome. To simulate the range of possible mosaic types seen in a TE biopsy, analysis of 85 samples, each consisting of five cells derived from karyotypically stable aneuploid cell lines, mixed in different ratios with euploid cells, was carried out. Seven distinct aneuploidies were evaluated in this manner, with analysis taking place blindly. Trisomies and monosomies (whole and segmental) were reliably detected with NGS even when only present in 20% of the cells (i.e., one aneuploid cell and four normal cells). Different proportions of aneuploid cells (0, 20, 40, 60, 80, and 100%) could be distinguished from one another in all cases, indicating that the NGS method is not only capable of detecting mosaicism, but also has the potential to quantify the proportion of aneuploid cells within a sample the size of a typical TE biopsy (Table 1; Fig. 1).
Examples of NGS and aCGH analysis of groups of aneuploid (47,XY, +18)–euploid (46,XY) mixes of cells in different ratios. a, b Chromosome abnormality present in 40% of cells (two aneuploid cells mixed with three euploid cells); c, d chromosome abnormality present in 80% of cells (four aneuploid cells with one euploid cell)
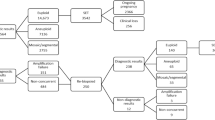
To determine the impact of mosaicism on embryo viability, an attempt was made to identify mosaic blastocysts that had been transferred to the uterus and for which a clinical outcome was known. A retrospective review of 150 clinical PGT-A results associated with transferred embryos led to the identification of 44 (29.3% in the examined group of results) with slight deviations of aCGH profiles for one or more chromosomes, too subtle to be characterized as aneuploid during the initial analysis, but possibly indicative of mosaicism. These 44 embryos were generated by a total of 39 patients (“Methods”, “Patient details”, and Table 2). NGS assessment of surplus amplified DNA from these embryos provided results consistent with the presence of mosaicism in all 44 TE biopsies. Additional analysis of the specimens at several hundred thousand SNP loci, scattered throughout the genome, confirmed that the results obtained were not technical artefacts related to contamination of the sample or triploidy.
In 18 (18/44, 41%) of the TE biopsy specimens, mosaicism only affected whole chromosome(s), 14 (14/44, 32%) carried only segmental chromosome abnormalities in mosaic form, while a combination of mosaic whole and segmental aneuploidy was scored for the remaining 12 (12/44, 27%). Sixteen (16/44, 36%) of the mosaic TE samples were characterized as ‘complex’ abnormal (3 or more chromosome errors present), a further 15 (15/44, 34%) carried two abnormalities, and 13 (13/44, 30%) had a single mosaic chromosome error.
Taking all 44 samples together, a total of 110 mosaic abnormalities were identified, affecting almost all chromosomes. Thirty-nine (39/110, 35.4%) were whole chromosome trisomies; 19 (19/110, 17.2%) were segmental trisomies; 28 (28/110, 25.4%) were whole chromosome monosomies; and 24 (24/110, 21.8%) were segmental monosomies. Segmental abnormalities involved the gain or loss of chromosomal fragments ranging in size from 4.5 to 96.8 Mb.
In an attempt to estimate the proportion of aneuploid cells in the mosaic TE samples, we compared the numerical expression of chromosome copy obtained from the BlueFuse analytical software (Illumina, UK) to the values ascertained during the aneuploid–euploid cell mixing experiments described above. This comparison suggested a range of 20–60% aneuploid cells for 43 of the 44 TE samples, with a single sample estimated to contain 80% abnormal cells. Across the entire set of mosaic samples, the proportion of abnormal cells was predicted to average 34%. The mosaic abnormalities scored during the NGS assessment of these TE samples are summarised in Table 2. Figure 2 illustrates the aCGH and NGS profiles of two different mosaic diploid–aneuploid TE samples.
aCGH and NGS profiles of two different mosaic TE samples. a aCGH image for sample 38-1 (Table 2). A slight deviation from the green line (chromosome copy number 2) was observed for the profile of chromosome 7; b NGS re-analysis of sample 38-1 demonstrated a mosaic loss of chromosome 7. Transfer of the corresponding embryo led to an ongoing pregnancy. c aCGH image for sample 21-5 (Table 2). A slight deviation from the green line (chromosome copy number 2) was observed for a segment of the profile of chromosome 6; d NGS re-analysis of sample 21-5 demonstrated a mosaic segmental loss of chromosome 6 (short arm affected). The corresponding embryo failed to implant after transfer (color figure online)
The embryos associated with the 44 mosaic TE samples were transferred to 39 patients in 40 procedures. There were 36 single embryo transfers (SETs) and four double embryo transfers (DETs, patients 12, 16, 24, and 34). One patient (number 22) underwent two separate SETs. Together, these transfers led to 17 implantations, 15 from SETs, and a pair of implantations from a DET (overall implantation rate of 38.6%, 17 of the 44 transferred embryos). There were five losses of singleton implantations, and consequently, the number of children born was 12 (27.3% of embryos produced a live birth, 12 of the 44 transferred embryos).
The transferred embryos could be subdivided into several categories. In 14 (32%) of the 44 mosaic blastocysts, the only abnormalities detected involved deletion or duplication affecting fragments of chromosome (segmental mosaics). In this group, eight of the embryos implanted, there were no losses, and the ultimate birth rate was, therefore, 57.1% (8/14) per embryo transferred. Another category consisted of embryos in which mosaicism affected one or more whole chromosomes. This was detected in 30 (68%) of the 44 mosaic embryos. In 12 of these embryos, mosaicism for a segmental aneuploidy was observed as well as for whole chromosomes. The implantation rate in this group of embryos was 30% (9/30), five losses occurred, thus resulting in a birth rate per embryo of 13.3% (4/30). A subset of this group, comprising 16 embryos, were categorised as ‘complex’ mosaics, containing populations of cells with mosaic aneuploidy affecting three or more different chromosomes. This group displayed the lowest birth rates (1/16, 6.25%). The differences in pregnancy loss and birth rates between embryos with a mosaic segmental abnormality and those with a mosaic aneuploidy affecting a whole chromosome were statistically significant (P = 0.004). The clinical outcomes after the transfer of embryos with mosaic TE samples are summarised in Table 2.
To determine the implantation potential of embryos with mosaic chromosomal abnormalities relative to embryos with uniformly euploid biopsy specimens, the outcomes of IVF cycles in which such embryos had been transferred were compared to those of a control group in which neither aCGH nor NGS had detected any evidence of mosaicism. The control group consisted of 51 transferred embryos generated by 47 couples clinically indistinguishable from, and being treated contemporaneously with, the patients of the study group. Similar to the mosaic group, the majority (43/47) of the transfers in the control group involved a single embryo, while two embryos were transferred in four cases (patients 5, 10, 12, and 16 in Table 3). Twenty-six pregnancies were established, with a total of 29 embryos (29/51, 56.9%) successfully implanting. Five of the embryos that implanted failed to progress, leading to a birth rate of 47% (24/51) per blastocyst transferred. Details of the control group patients and their clinical outcomes are summarised in Table 3. Comparison of clinical outcomes showed that embryos associated with TE biopsies with mosaicism affecting one or more whole chromosomes had implantation and pregnancy rates reduced to approximately one-third of those achieved using blastocysts with a non-mosaic euploid biopsy sample (P = 0.023 and P = 0.0036, respectively). The implantation rate for embryos with mosaic segmental abnormalities was no different to the control group.
Discussion
In the past 5 years, PGT-A has rapidly moved from a niche application, used for small numbers of narrowly defined patients undergoing IVF, to a mainstream intervention, considered to be a standard tool in a growing number of fertility clinics around the world. Given the unprecedented increase in utilization, it is imperative that lingering controversies associated with PGT-A are urgently examined. Modern methods for the testing of chromosomes in the small number of cells removed from each embryo have a high degree of technical accuracy, but have struggled to overcome confounding biological factors, chiefly chromosomal mosaicism. Until recently, mosaicism, which is common in human preimplantation embryos, and potentially results in the biopsied cells being unrepresentative of the remainder of the embryo, has been difficult to detect. This has led to concerns that PGT-A may sometimes provide an erroneous diagnosis, resulting in viable embryos being inappropriately discarded or the inadvertent transfer to the uterus of aneuploid embryos.
In the current study, the latest cytogenetics technologies were utilized to provide new data on the frequency of mosaicism and its impact on pregnancy outcome, allowing conclusions to be drawn about the extent to which mosaicism impacts PGT-A strategies that seek to facilitate the identification and preferential transfer of euploid embryos. Aneuploidy and mosaicism were detected using NGS—an umbrella term covering a variety of powerful technologies that have in common the ability to generate large quantities of DNA sequence information, rapidly and at a low-cost per base. The validation and clinical application of NGS for the purposes of screening IVF embryos for aneuploidy have been previously described by our group and others, and have helped to significantly improve patient access to PGT-A by reducing the cost of the procedure (Yin et al. 2013; Wells et al. 2014; Fiorentino et al. 2014; Kung et al. 2015).
Following a series of advances in embryology, PGT-A has shifted from analysis of a single cell, taken at the cleavage stage of development (3 day post-fertilization) to the sampling of approximately five cells at the blastocyst stage (day-5 or -6). In the United States, over 90% of all PGT-A now involves blastocyst biopsy. The current study conclusively demonstrates the ability of NGS to detect mosaic aneuploidy affecting entire chromosomes and also parts of chromosomes in blastocyst biopsies. We initially showed this in an experimental model of samples composed of five cells and subsequently in clinical samples. Mosaicism was reliably detected even when only one cell out of five was abnormal. As well as identification of mosaicism, NGS was confirmed to provide highly accurate diagnosis of non-mosaic aneuploidy in embryo samples previously tested using a well-validated aCGH method (>99.9% concordance).
Studies of blastocysts donated for research purposes, which have been biopsied and tested at multiple sites, clearly demonstrate that mosaicism can persist to the final stage of preimplantation development, but at a decreased frequency compared with the cleavage stage (Fragouli et al. 2008, 2011; Novik et al. 2014). It is also evident from such studies that most mosaic embryos are devoid of euploid cells, carrying different types of aneuploidies in all of their cells (referred to as mosaic aneuploid). In a previous study, using less sensitive methods than described here, we concluded that approximately 10% of blastocysts of high morphological grade are mosaic for a mixture of normal and aneuploid cells (Fragouli et al. 2011). Comprehensive cytogenetic assessment of TE and inner cell mass samples derived from the same embryos, carried out by our group and others, found no evidence of preferential allocation of mosaic abnormalities to the TE of the blastocyst (Evsikov and Verlinsky 1998; Fragouli et al. 2008). Similarly, Capalbo et al. (2013) did not observe diagnostic discrepancies between the TE and ICM or the preferential presence of mosaicism in either of the two tissues in 20 good quality blastocysts. The authors of this particular study postulated that the incidence of mosaic diploid/aneuploid embryos is even lower (4%) during the final stage of preimplantation development (Capalbo et al. 2013). The data obtained herein suggest that the true incidence of mosaic diploid–aneuploid blastocysts has been underestimated and may exceed 20%. The high frequency is only apparent now that highly sensitive NGS methods are available, capable of detecting a single-affected cell within a biopsy specimen, and revealing both whole chromosome aneuploidy and segmental abnormalities.
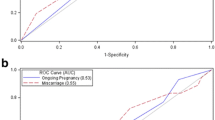
Importantly, the data from the current study clearly demonstrate that mosaic embryos are viable in some instances, although the likelihood of successful implantation was significantly less (38.6%) than for embryos associated with a non-mosaic, euploid biopsy specimen (56.9%). In addition, pregnancy loss rates were higher for embryos with mosaic PGT-A results (29.4 versus 20.7%), ultimately leading to a birth rate of 27.3% for mosaic embryos, versus 47% for embryos with an entirely euploid biopsy specimen.
Interestingly, not all forms of mosaicism had the same impact on embryo viability. Indeed, blastocysts with mosaic segmental abnormalities were associated with outcomes that were similar to embryos with completely normal trophectoderm biopsy specimens. Data obtained during comprehensive cytogenetic analysis of embryos at different developmental stages have shown that aneuploidies affecting fragments of chromosomes are very common at the cleavage stage (Wells and Delhanty 2000; Voullaire et al. 2000; Vanneste et al. 2009), but decline as development proceeds (Fragouli et al. 2013). This may be due to the fact that segmental abnormalities are associated with breakage of DNA strands, a form of genetic damage that typically triggers checkpoints responsible for inducing cell cycle arrest and, if repair does not occur, activating apoptotic pathways. Thus, affected cells may be eliminated, leaving behind only those cells with intact chromosomes. In addition, segmental abnormalities involving pieces of chromosome that lack a centromere (acentric fragments) are unable to attach to the spindle during mitosis and tend to be lost as cells divide. For this reason, it is likely that an embryo carrying an additional acentric fragment will ultimately end up with a normal chromosome complement. It is reasonable to suppose that most embryos from which abnormal cells have been eliminated will have an implantation potential similar to that seen for euploid embryos.
In contrast to embryos with mosaic segmental abnormalities, the implantation rate for those with mosaicism affecting whole chromosomes was only 13.3%, falling to just 6.3% if multiple chromosomes were involved. Five pregnancies achieved after transfer of a mosaic embryo ultimately miscarried, and in all cases, the embryo had mosaicism affecting a whole chromosome. These findings, therefore, suggest that unlike segmental aneuploidy, mosaic whole chromosome abnormalities tend to persist in developing blastocysts, affecting their ability to implant and lead to a viable pregnancy. Of note, we did not observe any relationship between the implantation ability of a mosaic blastocyst and the number of aneuploid cells in the TE biopsy. This is probably because the biopsy specimen represents a small, random sample of trophectoderm tissue, and does not necessarily reflect the proportion of chromosomally abnormal cells in the remainder of the embryo.
Considering the ongoing debate over the efficacy of embryo selection based upon PGT-A, these insights into the developmental potential of mosaic diploid–aneuploid blastocysts are of great value. It should be noted that all patients and their samples were de-identified prior to the initiation of this study, so we were unable to follow up on the karyotype of the live births. However, no chromosomally abnormal births have ever been reported following PGT-A at the clinic involved in this study.
Reduced viability of mosaic embryos was recently described in another study that involved the transfer of a small number of mosaic diploid–aneuploid blastocysts (Greco et al. 2015). The cytogenetic analysis of biopsied TE samples took place via aCGH, a less sensitive method than NGS in terms of the detection of mosaicism. In that investigation, approximately 5% of blastocysts were classified as mosaic. Transfers of affected embryos were undertaken for 18 women, leading to the establishment of six ongoing chromosomally normal pregnancies (33% pregnancy rate), significantly lower than embryos associated with an entirely euploid biopsy specimen.
While mosaicism is common during preimplantation development, it is rare at birth. The incidence of mosaic chromosome errors is estimated to be approximately 2% in prenatal samples and/or viable fetuses (Malvestiti et al. 2015). We have previously compared the aneuploidies seen during three different stages of development—the fertilised oocyte, cleavage stage embryo, and blastocyst stage embryo (Fragouli et al. 2013). The results obtained suggested that embryos carrying meiotic aneuploidies, originating (mostly) in the oocyte, frequently persist to the blastocyst stage and sometimes past implantation. These findings were in agreement with cytogenetic data from studies of products of conception (i.e., miscarriages), which have concluded that most aneuploidies detected once an embryo has implanted originate from errors occurring during female meiosis (reviewed in Hassold and Hunt 2001; Hassold et al. 2007). The absence of detectable aneuploid cell lines in most individuals is likely explained by the lethality of mosaicism prior to implantation or early in pregnancy. However, a minority of mosaic embryos are capable of producing chromosomally normal pregnancies, presumably due to loss of aneuploid cell lines, either via active mechanisms (e.g., apoptosis of abnormal cells) or passive processes (e.g., growth advantage of euploid cells).
The sensitivity of NGS and its increasing clinical use for the purposes of PGT-A have led to a significant increase in the detection of mosaic preimplantation embryos. If identified during IVF cycles, patients should receive counselling concerning the likely outcome of transfer. Embryos with a mosaic TE specimen should be given a lower priority for transfer than those associated with an entirely euploid biopsy. The data obtained during the current study suggest that, regardless of the proportion of abnormal cells within a TE sample, blastocysts with one or more mosaic segmental chromosome abnormalities have an implantation potential similar to that of euploid blastocysts. Consequently, in the absence of any completely euploid blastocysts, such embryos could be considered for transfer after appropriate patient counselling. Conversely, blastocysts with 1–2 mosaic errors affecting whole chromosomes, irrespective of the percentage of abnormal cells in the TE sample, were generally associated with poor clinical outcomes after transfer, suggesting that they should be given a lower transfer priority. Embryos characterized as having complex mosaic aneuploidy rarely implanted and could be excluded from transfer altogether. In addition, it is advisable to avoid transfer of embryos with mosaicism affecting chromosomes commonly associated with abnormal pregnancy, miscarriage or chromosomally aneuploid births (for example, but not limited to 13, 16, 18, 21, and 22).
It is recommended that patients with pregnancies established after transfer of a mosaic embryo should be offered prenatal testing, ideally amniocentesis, to confirm the absence of aneuploid cells in the foetus. More detailed recommendations were recently published by the Preimplantation Genetic Diagnosis International Society (PGDIS (2016), http://www.pgdis.org/docs/newsletter_071816.html). While the likelihood of a successful pregnancy is significantly reduced when mosaicism is detected, the fact that a minority of affected embryos produce apparently healthy births suggests that the blanket exclusion of such embryos may not be appropriate. Indeed, the failure to transfer potentially viable, mosaic embryos, could ultimately harm rather than enhance IVF success rates, a vitally important consideration for PGT-A.
Methods
Ethical approval
All procedures performed in this study were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. Specifically, assessment of all embryo samples via NGS took place only after written informed consent was obtained from all patients. Ethical approval was granted to the participating IVF clinic by their internal Institutional Review Board [9.baby Family and Fertility Center (previously Tecnobios Procreazione)].
Study design
To assess the capacity of NGS to detect mosaicism, cells were isolated from well-defined karyotypically abnormal fibroblast cell lines. Aneuploidies affected entire chromosomes or smaller chromosome segments. Different numbers of chromosomally normal male (46,XY) lymphocytes and aneuploid fibroblast cells were added to microcentrifuge tubes, to simulate different degrees of diploid–aneuploid mosaicism. In all cases, a total of five cells was evaluated, since this is the number typically found in a TE specimen: 5/5 cells aneuploid (100% abnormal); 4/5 (80%); 3/5 (60%); 2/5 (40%); 1/5 (20%); and 0/5 (entirely euploid). Details of the fibroblast cell line karyotypes are shown in Table 1. All samples were tested using aCGH, the most widely applied technique for PGT-A, as well as a well-validated NGS method (Fiorentino et al. 2014; Maxwell et al. 2016). Analysis was undertaken blindly, and a second independent individual decoded the results.
Following validation of mosaicism detection, a retrospective review was undertaken of 150 previous PGT-A results. In all cases, the data had been obtained using aCGH applied to TE biopsies and the embryos had been characterized as euploid and transferred. The clinical outcomes of these transfers were known. All 150 aCGH images were de-identified prior to review. The de-identification took place by an independent individual (i.e., not associated with the participating laboratories) in a way that only information about female patient age and clinical outcome after embryo transfer was available. In 44 cases, slight deviations in the aCGH profiles had been observed at the time of initial analysis, but these did not exceed the validated thresholds for calling chromosomes ‘aneuploid’, and consequently, the embryos had not been excluded from transfer.
Surplus DNA samples from these specimens were re-assessed via NGS in the hope that harnessing the superior sensitivity of that method would allow the presence of mosaicism to be confirmed or excluded. NGS took place using the VeriSeq protocol (Illumuina, UK), according to manufacturer’s instructions. As well as chromosomal analysis, samples were subjected to genotyping of ~300,000 single nucleotide polymorphisms (SNPs) using a microarray, as described by Konstantinidis et al. (2015). This approach allows detection of triploid samples and reveals whether a biopsy specimen has been contaminated with extraneous DNA/cells. Excluding these possibilities is important since, both can make a sample with a uniform karyotype appear as though it were mosaic after NGS.
To consider the effect of mosaicism on embryo implantation potential, the clinical outcomes of transfers involving mosaic blastocysts were compared to a carefully matched group of patients who had transfers of embryos that showed no evidence of mosaicism in their associated TE specimen. This patient group was also de-identified in the same way as the patients who received possibly mosaic embryos. The two different groups of patients were from the same clinic, were indistinguishable in terms of clinical parameters, had the same average female age, and were receiving treatment during the same time period.
Patient details
The 44 TE samples re-assessed by NGS were biopsied from embryos generated by 39 couples (average female age 37.2 years, range 26–43 years). The control group consisted of 51 embryos generated by 47 couples (average female age 37.9 years, range 29–43 years).
Mixing experiments
To define the limits of mosaicism detection via NGS, aneuploid and euploid cells were combined together. The aneuploid cells were isolated from fibroblast cell lines obtained from the NIGMS Human Genetic Cell Repository at the Coriell Institute for Medical Research (USA). These were combined with euploid male (46,XY) lymphocytes in different ratios, the proportion of abnormal cells ranging from 0 to 100% (see above). Entire chromosome gains were assessed with the use of two cell lines carrying trisomy 13 (47,XY,+13: GM00526), and trisomy 18 (47,XY,+18: GM01359), and an XXY, trisomy 21 cell line (48,XXY,+21: GM04965). Entire chromosome losses were assessed with the use of a 45,X0 cell line (45,X0: GM00857). Segmental chromosome losses and gains were assessed with the use of three cell lines carrying a partial loss of 10p [46,XY,del(10)(p14p12): GM03047], a partial loss of 13q [46,XX,del13(pter>q14): GM00509], and an unbalanced translocation between chromosomes 21 and X [46,XX,der(21)(21qter>21p11::Xqter): GM01730]. Mixing experiments were repeated three times for five of the cell lines, but the lines carrying the segmental errors affecting chromosomes 13 (GM00509) and X (GM01730) grew very poorly and could only be analyzed once each.
Ovarian stimulation and embryo culture
Controlled ovarian stimulation was induced using an antagonist (Orgalutran, Organon, Rome, Italy; or Cetrotide, Serono, Rome, Italy) and urinary or recombinant follicle-stimulating hormone (FSH) (Meropur, Ferring, Milan, Italy; or Gonal-F, Serono, Rome, Italy). A dose of 10,000 IU human chorionic gonadotrophin (hCG) (Gonasi, Amsa, Rome, Italy) or one ampoule of recombinant hCG (r-hCG; Ovitrelle, Serono, Rome, Italy) was administered when one or more follicles reached a maximum diameter of >23 mm. Oocyte collection was performed transvaginally, under ultrasound guidance, 36 h after hCG injection. Retrieved oocytes were rinsed and placed in Sydney IVF Fertilization Medium (Cook IVF, Brisbane, Australia) at 37 °C, 6% CO2, 5% O2, 89% N2 for at least 4 h. Fertilization was achieved by intracytoplasmic sperm injection (ICSI), following the standard techniques. Embryo culture was carried out in EmbryoScope (UnisenseFertiliTech, Denmark), an integrated embryo-culture time-lapse microscopy system with N2/CO2/O2 (89:6:5, v/v) at 37 °C without control of humidity. Immediately after ICSI, oocytes were placed inside pre-equilibrated slides (EmbryoSlide; Unisense Fertilitech) containing 12 droplets each of 25 μl fresh cleavage medium (Cook IVF) covered by 1.2 ml of mineral oil (SAGE, Biocare Europe, Rome, Italy). On day-3, cleavage medium was replaced with blastocyst medium (Cook IVF) for culture until day-5 or 6. Embryo transfers were carried out in fresh PGT-A cycles or after freeze-all cycles. Blastocysts were cryopreserved using a vitrification protocol with a closed-system device (HSV straw, Cryo Bio System, France) and Kitazato solutions (Kitazato BioPharma Co., Japan).
Embryo sampling and preparation
TE biopsy involved the removal of approximately five cells taking place either on day-5 or day-6 of preimplantation development, depending on embryo expansion. Specifically, at 113–142 h after insemination, all fully expanded blastocysts having a visible blastocoele, where an inner cell mass could be identified and with at least a few cells forming the trophectoderm epithelium, underwent trophectoderm biopsy. All biopsy procedures were conducted on a heated stage in a dish prepared with two 20 μl droplets of Sydney IVF Blastocyst Medium (Cook IVF, Brisbane, Australia) overlaid with pre-equilibrated mineral oil. Each blastocyst had its own dish, where it was left until transfer or cryopreservation, so that the identification procedures for each biopsied embryo were safer and the time used to carry out the biopsy was shorter (pre-equilibrated medium has been used during biopsy). Three-to-eight trophectoderm cells were aspirated into the biopsy aspiration pipette (Cook, Brisbane, Australia), followed by laser-assisted or mechanical removal of the trophectoderm cells from the epithelium. A diode laser (Saturn 3, Research Instruments, Cornwall TR11 4TA, UK) was used to assist the opening of a 10–20 μm hole in the zona pellucida on day-3 or 5 and for trophectoderm biopsy. Cell preparation for chromosome analysis took place as described previously (Fragouli et al. 2011, 2013). Clinical chromosome assessment had initially taken place via the application of a single, highly validated platform for microarray comparative genomic hybridisation (aCGH) (Fragouli et al. 2011; Wells et al. 2014).
Array CGH
Array CGH analysis was carried out using 24SureTM Cytochip V3 microarrays (Illumina, UK). The protocol used was as described in Fragouli et al. (2011, 2013), with modifications according to the manufacturer (Illumina, UK Resulting aCGH images were analyzed using the BlueFuse software (Illumina, UK).
Next generation sequencing
The NGS analysis of all TE samples took place with the use of the VeriSeq strategy (Illumina, UK), and as described previously (Fragouli et al. 2015). Cytogenetic assessment occurred with the use of the BlueFuse Multi v3 software (Illumina, UK).
Determination of mosaicism levels
Cytogenetic assessment of samples via NGS revealed the copy number of each of the 23 pairs of chromosomes. The BlueFuse Multi v3 software was capable of providing clear numerical values to determine chromosome copy number during sample analysis. Hence, a chromosome with two copies in an analyzed sample had a value of 2, a chromosome with three copies had a value of 3, a chromosome with one copy had a value of 1, whereas the presence of a nullisomy had a value of 0. As mosaic diploid–aneuploid samples consisted of a combination of chromosomally normal and abnormal cells, the values given by the software were either between 2 and 3 for mosaic chromosome gains, or between 2 and 1 for mosaic chromosome losses. Using the numerical values given by the Bluefuse Multi v3 software during analysis of the mixing experiments, we were able to determine the level of mosaicism in the TE biopsies. The numerical values obtained during the mixing experiments are shown in Table 1.
Statistical analysis
Unless stated otherwise, statistical evaluations were carried out using Fisher’s exact test.
References
Capalbo A, Wright G, Elliott T, Ubaldi FM, Rienzi L, Nagy ZP (2013) FISH reanalysis of inner cell mass and trophectoderm samples of previously array-CGH screened blastocysts shows high accuracy of diagnosis and no major diagnostic impact of mosaicism at the blastocyst stage. Hum Reprod 28:2298–2307. doi:10.1093/humrep/det245
Coonen E, Derhaag JG, Dumoulin JC, van Wissen LC, Bras M, Janssen M, Evers JL, Geraedts JP (2004) Anaphase lagging mainly explains chromosomal mosaicism in human preimplantation embryos. Hum Reprod 19:316–324
Daphnis DD, Delhanty JD, Jerkovic S, Geyer J, Craft I, Harper JC (2005) Detailed FISH analysis of day 5 human embryos reveals the mechanisms leading to mosaic aneuploidy. Hum Reprod 20:129–137
Delhanty JD, Griffin DK, Handyside AH, Harper J, Atkinson GH, Pieters MH, Winston RM (1993) Detection of aneuploidy and chromosomal mosaicism in human embryos during preimplantation sex determination by fluorescent in situ hybridisation (FISH). Hum Mol Genet 2:1183–1185
Delhanty JD, Harper JC, Ao A, Handyside AH, Winston RM (1997) Multicolour FISH detects frequent chromosomal mosaicism and chaotic division in normal preimplantation embryos from fertile patients. Hum Genet 99:755–760
Evsikov S, Verlinsky Y (1998) Mosaicism in the inner cell mass of human blastocysts. Hum Reprod 13:3151–3155
Fiorentino F, Bono S, Biricik A, Nuccitelli A, Cotroneo E, Cottone G, Kokocinski F, Michel CE, Minasi MG, Greco E (2014) Application of next-generation sequencing technology for comprehensive aneuploidy screening of blastocysts in clinical preimplantation genetic screening cycles. Hum Reprod 29:2802–2813. doi:10.1093/humrep/deu277
Forman EJ, Hong KH, Ferry KM, Tao X, Taylor D, Levy B, Treff NR, Scott RT Jr (2013) In vitro fertilization with single euploid blastocyst transfer: a randomized controlled trial. Fertil Steril 100(100–7):e1. doi:10.1016/j.fertnstert.2013.02.056
Fragouli E, Lenzi M, Ross R, Katz-Jaffe M, Schoolcraft WB, Wells D (2008) Comprehensive molecular cytogenetic analysis of the human blastocyst stage. Hum Reprod 23:2596–2608. doi:10.1093/humrep/den287
Fragouli E, Alfarawati S, Daphnis DD, Goodall NN, Mania A, Griffiths T, Gordon A, Wells D (2011) Cytogenetic analysis of human blastocysts with the use of FISH, CGH and aCGH: scientific data and technical evaluation. Hum Reprod 26:480–490. doi:10.1093/humrep/deq344
Fragouli E, Alfarawati S, Spath K, Jaroudi S, Sarasa J, Enciso M, Wells D (2013) The origin and impact of embryonic aneuploidy. Hum Genet 321:1001–1013. doi:10.1007/s00439-013-1309-0
Fragouli E, Spath K, Alfarawati S, Kaper F, Craig A, Michel CE, Kokocinski F, Cohen J, Munne S, Wells D (2015) Altered levels of mitochondrial DNA are associated with female age, aneuploidy, and provide an independent measure of embryonic implantation potential. PLoS Genet 11:e1005241. doi:10.1371/journal.pgen.1005241
Greco E, Minasi MG, Fiorentino F (2015) Healthy babies after intrauterine transfer of mosaic aneuploid blastocysts. N Engl J Med 373:2089–2090. doi:10.1056/NEJMc1500421
Hassold T, Hunt P (2001) To err (meiotically) is human: the genesis of human aneuploidy. Nat Rev Genet 2:280–291
Hassold T, Hall H, Hunt P (2007) The origin of human aneuploidy: where we have been, where we are going. Hum Mol Genet 16:R203–R208
Konstantinidis M, Prates R, Goodall NN, Fischer J, Tecson V, Lemma T, Chu B, Jordan A, Armenti E, Wells D, Munné S (2015) Live births following Karyomapping of human blastocysts: experience from clinical application of the method. Reprod Biomed Online 31:394–403. doi:10.1016/j.rbmo.2015.05.018
Kung A, Munné S, Bankowski B, Coates A, Wells D (2015) Validation of next-generation sequencing for comprehensive chromosome screening of embryos. Reprod Biomed Online 31:760–769. doi:10.1016/j.rbmo.2015.09.002
Magli MC, Jones GM, Gras L, Gianaroli L, Korman I, Trounson AO (2000) Chromosome mosaicism in day 3 aneuploid embryos that develop to morphologically normal blastocysts in vitro. Hum Reprod 15:1781–1786
Malvestiti F, Agrati C, Grimi B, Pompilii E, Izzi C, Martinoni L, Gaetani E, Liuti MR, Trotta A, Maggi F, Simoni G, Grati FR (2015) Interpreting mosaicism in chorionic villi: results of a monocentric series. Prenat Diagn 35:1117–1127. doi:10.1002/pd.4656
Maxwell SM, Colls P, Hodes-Wertz B, McCulloh DH, McCaffrey C, Wells D, Munné S, Grifo JA (2016) Why do euploid embryos miscarry? A case-control study comparing the rate of aneuploidy within presumed euploid embryos that resulted in miscarriage or live birth using next-generation sequencing. Fert Steril 106:1414–1419. doi:10.1016/j.fertnstert.2016.08.017
Munne S, Sandalinas M, Escudero T, Marquez C, Cohen J (2002) Chromosome mosaicism in cleavage-stage human embryos: evidence of a maternal age effect. Reprod Biomed Online 4:223–232
Novik V, Moulton EB, Sisson ME, Shrestha SL, Tran KD, Stern HJ, Mariani BD, Stanley WS (2014) The accuracy of chromosomal microarray testing for identification of embryonic mosaicism in human blastocysts. Mol Cytogenet 7:18. doi:10.1186/1755-8166-7-18
PGDIS (2016) Guidelines for mosaic embryos. http://www.pgdis.org/docs/newsletter_071816.html
Scott RT Jr, Upham KM, Forman EJ, Hong KH, Scott KL, Taylor D, Tao X, Treff NR (2013) Blastocyst biopsy with comprehensive chromosome screening and fresh embryo transfer significantly increases in vitro fertilization implantation and delivery rates: a randomized controlled trial. Fertil Steril 100:697–703. doi:10.1016/j.fertnstert.2013.04.035
Treff NR, Scott RT Jr (2013) Four-hour quantitative real-time polymerase chain reaction-based comprehensive chromosome screening and accumulating evidence of accuracy, safety, predictive value, and clinical efficacy. Fertil Steril 99:1049–1053. doi:10.1016/j.fertnstert.2012.11.007
Vanneste E, Voet T, Le Caignec C, Ampe M, Konings P, Melotte C, Debrock S, Amyere M, Vikkula M, Schuit F, Fryns JP, Verbeke G, D’Hooghe T, Moreau Y, Vermeesch JR (2009) Chromosome instability is common in human cleavage-stage embryos. Nat Med 15:577–583. doi:10.1038/nm.1924
Voullaire L, Slater H, Williamson R, Wilton L (2000) Chromosome analysis of blastomeres from human embryos by using compar- ative genomic hybridization. Hum Genet 106:210–217
Wells D, Delhanty JD (2000) Comprehensive chromosomal analysis of human preimplantation embryos using whole genome ampli- fication and single cell comparative genomic hybridization. Mol Hum Reprod 6:1055–1062
Wells D, Kaur K, Grifo J, Glassner M, Taylor JC, Fragouli E, Munne S (2014) Clinical utilisation of a rapid low-pass whole genome sequencing technique for the diagnosis of aneuploidy in human embryos prior to implantation. J Med Genet 51:553–562. doi:10.1136/jmedgenet-2014-102497
Yang Z, Liu J, Collins GS, Salem SA, Liu X, Lyle SS, Peck AC, Sills ES, Salem RD (2012) Selection of single blastocysts for fresh transfer via standard morphology assessment alone and with array CGH for good prognosis IVF patients: results from a randomized pilot study. Mol Cytogenet 5:24. doi:10.1186/1755-8166-5-24
Yin X, Tan K, Vajta G, Jiang H, Tan Y, Zhang C, Chen F, Chen S, Zhang C, Pan X, Gong C, Li X, Lin C, Gao Y, Liang Y, Yi X, Mu F, Zhao L, Peng H, Xiong B, Zhang S, Cheng D, Lu G, Zhang X, Lin G, Wang W (2013) Massively parallel sequencing for chromosomal abnormality testing in trophectoderm cells of human blastocysts. Biol Reprod 88:69. doi:10.1095/biolreprod.112.106211
Acknowledgements
This study was supported by institutional funding. Dagan Wells is supported by the Oxford NIHR Biomedical Research Centre.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors state no conflict of interest.
Rights and permissions
About this article
Cite this article
Fragouli, E., Alfarawati, S., Spath, K. et al. Analysis of implantation and ongoing pregnancy rates following the transfer of mosaic diploid–aneuploid blastocysts. Hum Genet 136, 805–819 (2017). https://doi.org/10.1007/s00439-017-1797-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-017-1797-4