Abstract
Retinitis pigmentosa (RP) is a group of inherited retinal disorders characterized by progressive photoreceptor degeneration. An accurate molecular diagnosis is essential for disease characterization and clinical prognoses. A retinal capture panel that enriches 186 known retinal disease genes, including 55 known RP genes, was developed. Targeted next-generation sequencing was performed for a cohort of 82 unrelated RP cases from Northern Ireland, including 46 simplex cases and 36 familial cases. Disease-causing mutations were identified in 49 probands, including 28 simplex cases and 21 familial cases, achieving a solving rate of 60 %. In total, 65 pathogenic mutations were found, and 29 of these were novel. Interestingly, the molecular information of 12 probands was neither consistent with their initial inheritance pattern nor clinical diagnosis. Further clinical reassessment resulted in a refinement of the clinical diagnosis in 11 patients. This is the first study to apply next-generation sequencing-based, comprehensive molecular diagnoses to a large number of RP probands from Northern Ireland. Our study shows that molecular information can aid clinical diagnosis, potentially changing treatment options, current family counseling and management.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Retinitis pigmentosa (RP; MIM#268000) refers to a group of inherited retinal diseases characterized by progressive photoreceptor apoptosis and retinal degeneration. RP is the most common form of hereditary retinal degeneration with a prevalence of approximately 1:3,500 to 1:4,000 (Wang et al. 2005; Haim 2002) affecting more than one million individuals worldwide (Chang et al. 2011). The typical clinical manifestations of RP include night blindness and tunnel vision. Some patients may eventually develop complete blindness. The phenotype of RP usually occurs alone, as nonsyndromic RP affecting only the eye. In some rare cases, RP can also be accompanied with other clinical symptoms affecting additional organs. For example, patients with Usher syndrome suffer both RP and hearing loss. RP is a highly genetically heterogeneous disease. First, more than 50 genes are known to be associated with nonsyndromic RP (RetNet; http://www.sph.uth.tmc.edu/Retnet/) and nearly 3,100 pathogenic mutations have been reported (Chang et al. 2011). Second, the inheritance pattern of RP involves all modes: autosomal-dominant (adRP), autosomal-recessive (arRP), X-linked (xlRP), and digenic forms (Anasagasti et al. 2013; Neveling et al. 2012; Kajiwara et al. 1994). Third, the molecular basis of RP overlaps with other retinal diseases. Different mutations in the same genes, or sometimes even the exact same mutations, can cause different retinal diseases (Wang et al. 2014).
Because of the heterogeneity of RP, accurate molecular diagnosis is essential for meaningful patient counseling as it can provide specific disease characterization and prognostic information. Hitherto, standard methods of genetic testing for RP include Sanger sequencing, arrayed primer extension (APEX) and next-generation sequencing (NGS). Sanger sequencing is the gold standard of sequencing, however, it is costly for large-scale sequencing. APEX only analyzes previously reported mutation loci and thus misses novel mutations, leading to a low diagnosis rate (Avila-Fernandez et al. 2010; Zeitz et al. 2009). NGS is currently considered the most efficient method for mutation screening. One approach of NGS is target sequencing, which limits testing to known disease-causing genes. For instance, our laboratory has developed a retinal capture panel to systematically test over 150 known retinal disease genes for pathogenic mutations in RP and Leber congenital amaurosis patients (Wang et al. 2013, 2014). The NGS-based targeted sequencing is superior in both time and cost compared to other methods, which makes it an optimal approach for the molecular diagnosis of RP.
It is known that the prevalence of causative genes and the mutation spectrum can vary significantly among different ethnicity groups. This is especially notable in relatively isolated populations or those with a higher consanguineous rate. For example, in Israeli and Palestinian patient populations, FAM161A mutations account for about 12 % of arRP cases (Bandah-Rozenfeld et al. 2010), whereas in North America FAM161A is responsible for only 1 % of arRP cases (Venturini et al. 2014). Furthermore, within a certain ethnic background, the frequency of a specific mutant allele may vary geographically. As an example, the well-known c.2299delG, p.(Glu767Serfs) mutation in USH2A is frequently found in European patients. This mutation accounts for 47.5 % of USH2A alleles in Denmark (Dreyer et al. 2008), while the allelic frequency is 31 % in the Netherlands (Pennings et al. 2004) and 10 % in France (Aller et al. 2010). The mutation frequency may become common as a result of the founder effect and may change due to genetic drift. Therefore, characterizing the mutation spectrum of a certain RP cohort can provide more comprehensive knowledge of the disease.
In this study, we performed NGS-based targeted sequencing in 82 unrelated RP cases from Northern Ireland; 46 were simplex cases and 36 were familial cases. The capture panel covered 55 RP genes and 131 other retinal disease genes. To our knowledge, this is the first study that performed NGS-based comprehensive molecular diagnosis on a large number of RP probands from Northern Ireland. Our study demonstrated that an NGS-based molecular diagnosis can facilitate a clinical diagnosis that better defines the disease and helps with family planning and patient management.
Materials and methods
Clinical diagnosis and sample collection
A cohort of 82 RP patients and other family members were ascertained at the Department of Ophthalmology (BHSCT) and Centre for Experimental Medicine (Belfast, UK). All patients had a detailed clinical history and underwent full ophthalmic evaluation including visual acuity testing, visual fields testing, fundal examination, and electroretinography. Retinitis pigmentosa was diagnosed on the basis of the typical fundal features (bone spicule retinal pigmentation, arteriolar attenuation, and optic disc pallor), visual field constriction, and an attenuated or abolished electroretinogram. Pedigrees are constructed based on interview. Available additional family members both affected and unaffected were also recruited. Genomic DNA of patients was extracted from peripheral blood. The research was conducted in accordance with the Tenets of the declaration of Helsinki. Ethical permission was granted through ORECNI and all patients gave written consent to participate in the study.
Retinal capture panel design
A capture panel of retinal disease genes was designed by our group which has been successfully applied for the molecular diagnosis of RP and Leber congenital amaurosis patients (Wang et al. 2013, 2014; Fu et al. 2013a). In this study, we updated the capture panel to include 23 newly reported retinal disease genes. The panel consisted of 994,088 bp covering 3,720 exons in 186 known retinal disease genes (RetNet; http://www.sph.uth.tmc.edu/Retnet/), including 55 known RP genes that had been reported at the time of panel design (Table S1).
Library preparation and capture sequencing
Pre-capture Illumina paired-end libraries were generated according to the manufacturer’s protocol. Briefly, ~1 μg of patient’s genomic DNA was sheared into 300–500 bp fragments. The DNA fragments were end-repaired and a single adenine base was added to the 3′ ends using Klenow exonuclease. Illumina Y-shape index adapters were ligated to the repaired ends, and DNA fragments were PCR amplified for eight cycles after ligation. The DNA libraries were quantified by the PicoGreen fluorescence assay kit (Invitrogen, Carlsbad, CA, USA). In each capture reaction, 50 pre-capture DNA libraries were pooled together. The targeted DNA was captured, washed and recovered using Agilent Hybridization and Wash Kits (Agilent Technologies, Santa Clara, CA, USA). Captured libraries were sequenced on Illumina HiSeq 2000 (Illumina, San Diego, CA, USA) as 100 bp paired-end reads, following the manufacturer’s protocol.
Bioinformatics analysis
Paired-end sequencing reads were obtained for each sample. Reads were mapped to human reference genome hg19 using Burrows–Wheeler Aligner (BWA version 0.6.1) (Li and Durbin 2009). Base quality recalibration and local realignment were performed using the Genome Analysis Tool Kit (GATK version 1.0.5974) (McKenna et al. 2010). AtlasSNP and AtlasIndel2 (Challis et al. 2012) were used to call single-nucleotide polymorphisms (SNPs) and small insertions and deletions (INDELs).
Since RP is a rare Mendelian disease, polymorphisms that appear at a higher than 0.5 % frequency (for recessive variants) or 0.1 % frequency (for dominant variants) in at least one of the following databases were considered too frequent to be pathogenic and therefore excluded from further analysis: the 1000 Genome (build 20110521 and 20101123) (Genomes Project C et al. 2010, 2012), dbSNP135 (Sherry et al. 2001), NHLBI exome sequencing database (Fu et al. 2013b), NIEHS exome sequencing database (Genomes Project C et al. 2010), and our internal control databases. After frequency-based filtering, ANNOVAR (Wang et al. 2010) was used to predict protein-coding changes and filter out synonymous variants. Furthermore, mutations known to cause retinal diseases were identified by searching against HGMD professional database (Stenson et al. 2013). Finally, dbNSFP (version 2.3) (Liu et al. 2013), a program that compiles prediction scores from six prediction algorithms [SIFT (Ng and Henikoff 2003), Polyphen2 (Adzhubei et al. 2010), LRT (Chun and Fay 2009), MutationTaster (Schwarz et al. 2010), MutationAssessor (Reva et al. 2011) and FATHMM (Shihab et al. 2013)] and three conservation scores [Phylop (Siepel et al. RECOMB 2006), GERP++ (Davydov et al. 2010) and Siphy (Garber et al. 2009; Lindblad-Toh et al. 2011)], was used to predict the pathogenicity of novel missense variants. The details of the method are described in supplementary material. The prediction of novel missense variants is listed in Table S2.
Causative mutation prioritization
For each patient, we looked for causative variants using the following prioritization strategy:
-
1.
Reported pathogenic variants in RP genes.
-
2.
Novel severe loss-of-function (LOF) variants (stop-gain, splicing, frameshift, fail-to-start) in RP genes.
-
3.
Novel missense variants in RP genes. The missense variants must be predicted to be deleterious by dbNSFP as described in the Sect. “Materials and methods”.
-
4.
Pathogenic variants in other retinal disease genes.
All the variants should be consistent with the known pattern of inheritance of the respective gene (i.e., homozygous/compound heterozygous for recessive genes and heterozygous for dominant genes). For the familial cases, we specifically looked for variants in genes that matched the inheritance patterns predicted from the pedigrees.
Sanger sequencing validation and family segregation test
All putative mutations identified by NGS were validated using Sanger sequencing and tested for co-segregation if additional affected family members are available. Primers were designed using Primer3 (Rozen and Skaletsky 2000). To ensure the quality of Sanger sequencing, the amplicons were designed to have a boundary around 100 bp away from the mutation. Then the amplicons (~400 bp) were Sanger sequenced on Applied BioSystems (ABI) 3,730 × l capillary sequencer (Applied Biosystems Inc., Foster City, CA, USA). The Sanger sequencing results were analyzed using Sequencher (version 5.0).
Results
82 Unrelated Northern Ireland families with RP patients were recruited
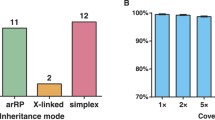
A total of 82 well-characterized RP families from Northern Ireland were recruited for this study. Among these families, 36 had two or more affected members, while the remaining 46 with only one affected member are considered as simplex cases. Based on the pedigree information, 26.8 % (22/82) of the families were arRP, 13.4 % (11/82) of the families were adRP, 3.7 % (3/82) were xlRP, and 56.1 % (46/82) were simplex.
High-quality NGS results were obtained
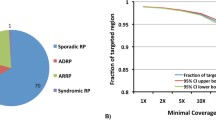
Capture NGS was performed on all 82 RP families. DNA from one affected member of each family was selected, captured and sequenced. Within the design region, an average of 141× coverage was achieved for all samples. 95.1 % of bases had coverage of >10×, 91.8 % of bases had coverage of >20× and 85.5 % of bases had coverage of >40×, indicating that sufficient coverage was achieved to enable high variant detection sensitivity (Fig. 1a). To test if the coverage of target region was evenly distributed, an evenness score was calculated for each sample as described previously (Fig. 1b) (Mokry et al. 2010). On average, the evenness score for all the 82 probands was 0.8, suggesting a nearly uniform distribution was achieved.
Pathogenic mutations were identified in 49 probands
An average of 732 variants, including 672 SNPs and 60 small INDELs, were initially identified for each sample in the targeted region. After all filtering and annotation steps (see Sect. “Materials and methods”), an average of 8.2 SNPs and 1.8 INDELs per patient remained and were therefore considered as candidate pathogenic variants. Through the mutation prioritization procedure (see Sect. “Materials and methods”), we identified pathogenic mutations in 49 probands, including 28 simplex cases and 21 familial cases, and achieved a solving rate of 60 % (49/82) (Tables 1 and 2).
Simplex cases
Out of the 46 simplex RP cases, 28 (61 %) were identified as carrying pathogenic or putative pathogenic mutations in known retinal disease genes. Overall, 41 mutations were identified in the simplex RP cases and 20 of them were novel. Among these novel mutations, six were LOF mutations, including four frameshift and two nonsense mutations. The remaining fourteen were novel missense variants that passed multiple frequency-based filters and were predicted to be pathogenic by dbNSFP (Table S2). Genotypes of the patients are detailed in Table 1.
According to the identified mutations, the inheritance pattern of two of the simplex probands was autosomal dominant (proband Rp25, proband Rp29), two probands were X-linked cases (proband Rp349B, proband Rp232A) rather than simplex, and the remaining 24 probands carry mutations in autosomal-recessive genes. In most cases, the diagnosis of simplex RP is strongly biased towards a recessive model; however, it is possible that the simplex cases are due to mutations in dominant RP genes. For proband Rp25 further assessment of family members was carried out. Both Rp25′s parents were deceased but reported as unaffected. However, a history of blindness was reported in the paternal grandfather and two great-uncles, making the inheritance pattern likely to be autosomal dominant.
A total of 16 causative genes were observed in our simplex cohort. The most prevalent mutated gene was USH2A, which explained disease in eight probands. Among the 16 causative genes, eight of them are known RP genes, which accounted for 18 (64 %) simplex cases. Interestingly, pathogenic mutations in eight other retinal disease genes (CDH23, VPS13B, MYO7A, CLRN1, RS1, CACNA1F, PHYH, and NPHP4) were found in 10 (36 %) probands, including five previously reported alleles, two novel LOF alleles, and eight novel missense alleles. The minor allele frequency (MAF) and pathogenicity predictions for the novel missense alleles are listed in Table S2. For these 10 simplex probands, the molecular information is inconsistent with the original clinical diagnosis. This could be due to the difficulty of assigning a more precise clinical diagnosis at the time of the initial visit, or a novel genotype–phenotype correlation as proposed in Wang et al. (2014).
To further investigate the two possibilities for these probands, we either reviewed the available clinical data and imaging or performed further clinical assessment. One proband (Rp131) was confirmed to be affected by RP, while the rest of nine probands (Rp78, Rp112, Rp399A, Rp83, Rp41, Rp76, Rp349B, Rp232A, and Rp58) were re-diagnosed to other retinal diseases (Table S4).
Proband Rp131 who carries compound heterozygous mutations in NPHP4 remained a diagnosis of RP after clinical reassessment. Mutations in NPHP4 are associated with nephronophthisis type 4, a renal disease, and with Senior–Loken syndrome type 4, a combination of nephronophthisis and retinitis pigmentosa (Hoefele et al. 2005; Otto et al. 2002). However, Rp131 did not show any clinical signs of nephrolithiasis; therefore, the mutation in NPHP4 must not be expressing clinically in the kidneys in this patient, and proband Rp131 was confirmed as RP.
Proband Rp58 is an interesting case of clinical re-diagnosis. The patient carries a putative pathogenic homozygous mutation c.403G > A, p.(Gly135Arg) in PHYH. PHYH was previously reported to cause Refsum disease (Jansen et al. 2004) which is characterized by early-onset RP with variable symptoms including, but not limited to, ataxia, neuropathy, hearing loss, and anosmia. Patients with Refsum disease usually have night blindness and retinal degeneration in their late childhood or early adulthood, and as the disease progresses, other symptoms may appear. Some patients will not develop other symptoms until their 40 or 50 s (Wanders et al. 1993). Therefore, it is very difficult to distinguish Refsum disease and RP if the disease is at the early stage. We revisited proband Rp58 and other available family members. Rp58 had developed mild cerebellar ataxia and hearing loss in later years. Interestingly, two sons of Rp58 showed learning disability and dyslexia (Fig. 2). Considering both the clinical reassessment and the molecular information, Rp58 was re-diagnosed to Refsum disease and dietary treatment was started.
Proband Rp83 carries compound heterozygous LOF mutations in VPS13B, which was reported to cause Cohen syndrome (Kolehmainen et al. 2004). The features of Cohen syndrome vary widely among affected individuals, and one of the features is retinal degeneration (Chandler et al. 2002), which is phenotypically similar to RP. We revisited patient Rp83 and other syndromic features were revealed, including learning difficulties, clumsiness, characteristic facial features, progressive retinochoroidal dystrophy, and myopia. Therefore, proband Rp83 was re-diagnosed to Cohen syndrome.
The remaining five re-diagnosed patients carry pathogenic mutations in genes that are known to cause Usher syndrome (Rp78 with CDH23 mutations, Rp112 with CDH23 mutations, Rp399A with CDH23 mutations, Rp41 with MYO7A mutations, and Rp76 with CLRN1 mutations). After clinical reassessment, all five probands were found to have a mild hearing loss in addition to RP, and were reclassified as Usher syndrome patients. Interestingly, patient Rp399A had posed a diagnostic difficulty. Although the patient had typical features of a pigmentary retinopathy, there was no history of nyctalopia. The patient’s mother had contracted rubella while pregnant with patient Rp399A and the family was keen to establish definitively whether the patient had nonprogressive retinopathy due to rubella or whether this was an inherited progressive disorder for the purposes of genetic counseling.
Familial cases
Out of 36 familial cases, 21 probands (58 %) were identified as carrying putative pathogenic mutations in known retinal disease genes, as shown in Table 2. These 21 solved familial cases are from 15 arRP, three adRP and three xlRP families. For the 15 solved arRP cases, there were in total 22 variants identified, including 15 previously reported variants and seven novel variants. The seven novel variants included four LOF mutations, one nonframeshift deletion and two missense mutations. The novel missense variants were filtered with 0.5 % frequency in multiple control databases, and predicted to be pathogenic by dbNSFP (Table S2). Among the three solved adRP cases, two probands carry previously reported mutations and one proband (RD120008) carries a fail-to-start mutation in dominant RP gene PRPH2. For the three xlRP cases, two probands carry reported mutations known to cause RP and one proband carries a LOF mutation in CHM.
For some familial cases, the inheritance modes obtained from the pedigree did not match with the mutations identified in the patients. For example, proband Rp73 was initially classified as xlRP according to the pedigree (Fig. 3a), as all the 5 patients were male and none of the female family members were affected. Since this family was at risk of xlRP, the male offspring of a carrier mother has a 50 % chance of having the disease. To prevent the transmission of RP, the daughters of affected members were undergoing embryonic testing. However, with the molecular diagnosis, proband Rp73 was found to carry a reported homozygous stop-gain mutation in CERKL on chromosome 2, which suggested that proband Rp73 in fact had arRP. To confirm this finding, we performed segregation on this family. The segregation test was consistent with the molecular diagnosis, saving the family from performing taxing offspring selection.
Pedigrees and mutations of proband Rp73, Rp278B, and Rp150. a Proband Rp73 carried a homozygous mutation c.847C > T, p.(Arg283*) in CERKL, and was refined to arRP from xlRP. b Proband Rp278 carried a heterozygous mutation c.1878G > C, p.(Gln626His) in PITPNM3, and was refined to cone dystrophy. c Proband Rp150 carried a heterozygous mutation c.498_499del, p.(Leu167Argfs) in CHM, and was refined to choroideremia
In our familial cases, we identified pathogenic mutations in two genes that have not been previously linked to RP (PIPTNM3, CHM) but are known to cause other retinal dystrophies. To resolve these ambiguous cases, we reviewed the patients and performed a clinical reassessment. After revisiting the patients, they were re-diagnosed to other retinal diseases (Table S4).
In the case of proband Rp278B (Fig. 3b), the pedigree appeared to show adRP. We identified a known heterozygous mutation c.1878G > C, p.(Gln626His) in PITPNM3. The mutation was reported to cause autosomal-dominant cone dystrophy (Kohn et al. 2007) and patient Rp278B was re-diagnosed to dominant cone dystrophy. However, when we performed a segregation test on other affected family members, this mutation was not shared by the patient’s affected mother and aunts. One possible explanation is that the affected members of this family have different types of retinal diseases that are caused by different genetic mutations.
Another case is proband Rp150 (female) (Fig. 3c), which was identified as carrying a heterozygous frameshift mutation in CHM. Mutations in this gene are known to cause choroideremia, an X-linked eye disorder characterized by progressive degeneration of the choroid, retinal pigment epithelium, and retina. A hemizygous mutated male is fully affected while the female heterozygous carriers usually show mild fundus abnormalities (irregular pigmentation of the retinal periphery) which are typically sub-clinical. Yet, some female carriers may also develop the full choroideremia phenotype (van den Hurk et al. 1997; Francois 1971). Choroideremia can be confused with RP since both have symptoms of night blindness and tunnel vision. The difference is that the loss of vision in choroideremia often starts as an irregular ring that gradually expands both centrally and out toward the extreme periphery (Coussa and Traboulsi 2012). In our case, proband Rp150 might be a female choroideremia carrier.
Collectively, as shown in Tables 1 and 2, 65 pathogenic mutations were identified in 49 probands, including 28 simplex cases and 21 familial cases. Twenty-nine (44.6 %) of 65 pathogenic mutations identified were novel (Table 3). Most of these mutations were nonsynonymous (61.5 %) while a significant proportion is frameshift (16.9 %) and stop-gain (12.3 %). As shown in Table S4, among all simplex and familial RP cases, there are in total 12 probands showing inconsistency between the molecular information and the original clinical diagnosis. After clinical reassessment, 11 of 12 subjects were reclassified in terms of their retinal disease on the basis of the mutation analysis.
Discussion
In this study, we performed an NGS-based molecular diagnosis on 82 well-characterized RP probands from Northern Ireland, including 46 simplex cases and 36 familiar cases. Our method successfully solved 49 out of 82 probands, achieving a solving rate of 60 %.
Our results demonstrate that NGS-based molecular information can contribute to precise clinical diagnoses enabling better disease management and accurate family counseling. Clinical manifestations of a number of retinal diseases are similar, especially for syndromic RP where some syndromes are late-onset and it can be difficult to distinguish these retinal diseases from nonsyndromic RP by clinical examination alone, even with a high index of clinical suspicion. Our approach can provide accurate molecular information to better define the disease manifestation. Patients with a precise diagnosis can then take advantage of any treatment available in a timely fashion. For example, Rp58 was re-diagnosed as Refsum disease. Unlike nonsyndromic RP, Refsum disease can be modified by diet, and preventative treatment can slow the neurological degeneration (Baldwin et al. 2010; Wanders et al. 1993); however, the clinical manifestations of Refsum disease are very subtle at an early stage. Therefore, a molecular diagnosis increases our understanding of how the patient’s disease will progress and allows the possibility of an earlier diagnosis and treatment in other family members. Further, as shown by proband Rp73, the characterization of genetic defects can help with family birth planning to minimize the risk of transmitting the disease to offspring. Moreover, the molecular testing of patient Rp399A helps resolve the diagnostic dilemma which was due to a history of maternal rubella, and confirmed a diagnosis of Usher Syndrome with mutations in the CDH23 gene. Finally, an accurate molecular diagnosis is the first step concerning eligibility for gene therapy (den Hollander et al. 2010).
It is also worth noting that simplex cases are often thought to be recessive since the parents of patients are assumed to be unaffected, however, 2/28 of our simplex cases were identified to carry heterozygous mutations in autosomal-dominant genes. One explanation could be a de novo mutation in the patient which results in only one affected member in the pedigree. It is also possible that patients carry dominant mutations inherited from their parents, but the mutation displays incomplete penetrance in the parents causing them not to manifest the disease phenotype. Here for example, proband Rp25 was identified to carry a heterozygous frameshift mutation in PRPF31 which is known to cause dominant RP. Both parents of the patient were deceased but reported as unaffected, however, a history of blindness was reported in the paternal grandfather and great-uncles. This suggests patient Rp25 is very likely to be adRP, and the unaffected parents could be due to incomplete penetrance.
Our patient cohort has a different mutation spectrum from patient cohorts of other ethnicities. For instance, mutations in EYS were frequently found in Chinese RP cases (Wang et al. Unpublished data), while we observed no pathogenic mutations in EYS. Furthermore, recurrent mutations were identified in our cohort. The most frequent mutations were c.4714C > T, p.(Leu1572Phe) and c.2299delG, p.(Glu767Serfs) in USH2A, shared by 4 probands (RD1200002, Rp311B, Rp159, Rp86). The genotypes of these 4 probands around this region are listed in Table S5. The shared SNPs may suggest specific haplotypes and indicate the founder effect. Recent studies on Irish population history suggested that a large proportion of Irish population was originated from northern Spain. Interestingly, the USH2A haplotype identified in our cohort is also found to be widespread in Spanish RP and Usher patients (Najera et al. 2002; Aller et al. 2010), which supports the close link between Irish and Spanish population.
In our cohort, we were able to solve a significantly lower fraction of adRP than xlRP or arRP patients. One reason is for this is that it is difficult to confidently verify that lone novel missense mutations cause disease. In the cases where DNA of other affected members was not available, a segregation test could not be performed. As a result, we could not confidently report the candidate mutations. Among our adRP cases, we did identify novel putative pathogenic missense mutations in three adRP families (Table S3). We also identified some novel missense mutations with lower confidence levels in unsolved simplex cases that failed to pass our rigorous criteria.
About 35 % of our cases do not have even low confidence candidates. For these unsolved patients, we have made every effort to ensure accurate clinical diagnoses and although it is possible that some cases are phenocopies, this is unlikely given that all cases have been followed clinically for many years and all show progression of their disease with the expected electrophysiological findings. Another explanation for this is that the disease-causing genes were not included in our designed panel. Therefore, we are performing whole-exome sequencing on all negative cases, the results of which will be presented in a future manuscript. A further possibility is that the patients’ phenotype is caused by novel disease genes. An additional explanation could be pathogenic intronic mutations that were not captured in our panel and copy number variations that were difficult to detect cause disease in these patients.
In summary, our approach identified the genetic cause of 60 % of disease in our patient cohort from Northern Ireland. A total of 31 novel mutations were found. Our study indicated that molecular information can aid clinical diagnosis and help with patient treatment and management, particularly highlighted by three patients and their families (Rp58, Rp73 and Rp399A). Further improvements in NGS technology together with the discovery of novel RP genes will undoubtedly boost the success rate of NGS-based diagnostic approaches in RP in the future.
References
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, Kondrashov AS, Sunyaev SR (2010) A method and server for predicting damaging missense mutations. Nat Methods 7(4):248–249. doi:10.1038/nmeth0410-248
Aller E, Jaijo T, Beneyto M, Najera C, Oltra S, Ayuso C, Baiget M, Carballo M, Antinolo G, Valverde D, Moreno F, Vilela C, Collado D, Perez-Garrigues H, Navea A, Millan JM (2006) Identification of 14 novel mutations in the long isoform of USH2A in Spanish patients with Usher syndrome type II. J Med Genet 43(11):e55. doi:10.1136/jmg.2006.041764
Aller E, Larrieu L, Jaijo T, Baux D, Espinos C, Gonzalez-Candelas F, Najera C, Palau F, Claustres M, Roux AF, Millan JM (2010) The USH2A c.2299delG mutation: dating its common origin in a Southern European population. Eur J Hum Genet 18(7):788–793. doi:10.1038/ejhg.2010.14
Allikmets R, Singh N, Sun H, Shroyer NF, Hutchinson A, Chidambaram A, Gerrard B, Baird L, Stauffer D, Peiffer A, Rattner A, Smallwood P, Li Y, Anderson KL, Lewis RA, Nathans J, Leppert M, Dean M, Lupski JR (1997) A photoreceptor cell-specific ATP-binding transporter gene (ABCR) is mutated in recessive Stargardt macular dystrophy. Nat Genet 15(3):236–246. doi:10.1038/ng0397-236
Anasagasti A, Barandika O, Irigoyen C, Benitez BA, Cooper B, Cruchaga C, Lopez de Munain A, Ruiz-Ederra J (2013) Genetic high throughput screening in Retinitis Pigmentosa based on high resolution melting (HRM) analysis. Exp Eye Res 116:386–394
Avila-Fernandez A, Cantalapiedra D, Aller E, Vallespin E, Aguirre-Lamban J, Blanco-Kelly F, Corton M, Riveiro-Alvarez R, Allikmets R, Trujillo-Tiebas MJ, Millan JM, Cremers FP, Ayuso C (2010) Mutation analysis of 272 Spanish families affected by autosomal recessive retinitis pigmentosa using a genotyping microarray. Mol Vis 16:2550–2558
Bader I, Brandau O, Achatz H, Apfelstedt-Sylla E, Hergersberg M, Lorenz B, Wissinger B, Wittwer B, Rudolph G, Meindl A, Meitinger T (2003) X-linked retinitis pigmentosa: RPGR mutations in most families with definite X linkage and clustering of mutations in a short sequence stretch of exon ORF15. Invest Ophthalmol Vis Sci 44(4):1458–1463
Baldwin EJ, Gibberd FB, Harley C, Sidey MC, Feher MD, Wierzbicki AS (2010) The effectiveness of long-term dietary therapy in the treatment of adult Refsum disease. J Neurol Neurosurg Psychiatry 81(9):954–957. doi:10.1136/jnnp.2008.161059
Bandah-Rozenfeld D, Mizrahi-Meissonnier L, Farhy C, Obolensky A, Chowers I, Pe’er J, Merin S, Ben-Yosef T, Ashery-Padan R, Banin E, Sharon D (2010) Homozygosity mapping reveals null mutations in FAM161A as a cause of autosomal-recessive retinitis pigmentosa. Am J Hum Genet 87(3):382–391. doi:10.1016/j.ajhg.2010.07.022
Baux D, Larrieu L, Blanchet C, Hamel C, Ben Salah S, Vielle A, Gilbert-Dussardier B, Holder M, Calvas P, Philip N, Edery P, Bonneau D, Claustres M, Malcolm S, Roux AF (2007) Molecular and in silico analyses of the full-length isoform of usherin identify new pathogenic alleles in Usher type II patients. Hum Mutat 28(8):781–789. doi:10.1002/humu.20513
Benaglio P, McGee TL, Capelli LP, Harper S, Berson EL, Rivolta C (2011) Next generation sequencing of pooled samples reveals new SNRNP200 mutations associated with retinitis pigmentosa. Hum Mutat 32(6):E2246–E2258. doi:10.1002/humu.21485
Bharadwaj AK, Kasztejna JP, Huq S, Berson EL, Dryja TP (2000) Evaluation of the myosin VIIA gene and visual function in patients with Usher syndrome type I. Exp Eye Res 71(2):173–181. doi:10.1006/exer.2000.0863
Bolz H, von Brederlow B, Ramirez A, Bryda EC, Kutsche K, Nothwang HG, Seeliger M, del CSCM, Vila MC, Molina OP, Gal A, Kubisch C (2001) Mutation of CDH23, encoding a new member of the cadherin gene family, causes Usher syndrome type 1D. Nat Genet 27(1):108–112. doi:10.1038/83667
Briggs CE, Rucinski D, Rosenfeld PJ, Hirose T, Berson EL, Dryja TP (2001) Mutations in ABCR (ABCA4) in patients with Stargardt macular degeneration or cone-rod degeneration. Invest Ophthalmol Vis Sci 42(10):2229–2236
Challis D, Yu J, Evani US, Jackson AR, Paithankar S, Coarfa C, Milosavljevic A, Gibbs RA, Yu F (2012) An integrative variant analysis suite for whole exome next-generation sequencing data. BMC Bioinform 13:8. doi:10.1186/1471-2105-13-8
Chandler KE, Biswas S, Lloyd IC, Parry N, Clayton-Smith J, Black GC (2002) The ophthalmic findings in Cohen syndrome. Br J Ophthalmol 86(12):1395–1398
Chang S, Vaccarella L, Olatunji S, Cebulla C, Christoforidis J (2011) Diagnostic challenges in retinitis pigmentosa: genotypic multiplicity and phenotypic variability. Curr Genomics 12(4):267–275. doi:10.2174/138920211795860116
Chun S, Fay JC (2009) Identification of deleterious mutations within three human genomes. Genome Res 19(9):1553–1561. doi:10.1101/gr.092619.109
Clark GR, Crowe P, Muszynska D, O’Prey D, O’Neill J, Alexander S, Willoughby CE, McKay GJ, Silvestri G, Simpson DA (2010) Development of a diagnostic genetic test for simplex and autosomal recessive retinitis pigmentosa. Ophthalmology 117(11):2169–2177 e2163. doi:10.1016/j.ophtha.2010.02.029
Coussa RG, Traboulsi EI (2012) Choroideremia: a review of general findings and pathogenesis. Ophthalmic Genet 33(2):57–65. doi:10.3109/13816810.2011.620056
Cremers FP, Kimberling WJ, Kulm M, de Brouwer AP, van Wijk E, te Brinke H, Cremers CW, Hoefsloot LH, Banfi S, Simonelli F, Fleischhauer JC, Berger W, Kelley PM, Haralambous E, Bitner-Glindzicz M, Webster AR, Saihan Z, De Baere E, Leroy BP, Silvestri G, McKay GJ, Koenekoop RK, Millan JM, Rosenberg T, Joensuu T, Sankila EM, Weil D, Weston MD, Wissinger B, Kremer H (2007) Development of a genotyping microarray for Usher syndrome. J Med Genet 44(2):153–160. doi:10.1136/jmg.2006.044784
Davydov EV, Goode DL, Sirota M, Cooper GM, Sidow A, Batzoglou S (2010) Identifying a high fraction of the human genome to be under selective constraint using GERP++. PLoS Comput Biol 6(12):e1001025. doi:10.1371/journal.pcbi.1001025
den Hollander AI, ten Brink JB, de Kok YJ, van Soest S, van den Born LI, van Driel MA, van de Pol DJ, Payne AM, Bhattacharya SS, Kellner U, Hoyng CB, Westerveld A, Brunner HG, Bleeker-Wagemakers EM, Deutman AF, Heckenlively JR, Cremers FP, Bergen AA (1999) Mutations in a human homologue of Drosophila crumbs cause retinitis pigmentosa (RP12). Nat Genet 23(2):217–221. doi:10.1038/13848
den Hollander AI, Black A, Bennett J, Cremers FP (2010) Lighting a candle in the dark: advances in genetics and gene therapy of recessive retinal dystrophies. J Clin Invest 120(9):3042–3053. doi:10.1172/JCI42258
Dreyer B, Brox V, Tranebjaerg L, Rosenberg T, Sadeghi AM, Moller C, Nilssen O (2008) Spectrum of USH2A mutations in Scandinavian patients with Usher syndrome type II. Hum Mutat 29(3):451. doi:10.1002/humu.9524
Estrada-Cuzcano A, Koenekoop RK, Senechal A, De Baere EB, de Ravel T, Banfi S, Kohl S, Ayuso C, Sharon D, Hoyng CB, Hamel CP, Leroy BP, Ziviello C, Lopez I, Bazinet A, Wissinger B, Sliesoraityte I, Avila-Fernandez A, Littink KW, Vingolo EM, Signorini S, Banin E, Mizrahi-Meissonnier L, Zrenner E, Kellner U, Collin RW, den Hollander AI, Cremers FP, Klevering BJ (2012) BBS1 mutations in a wide spectrum of phenotypes ranging from nonsyndromic retinitis pigmentosa to Bardet-Biedl syndrome. Arch Ophthalmol 130(11):1425–1432. doi:10.1001/archophthalmol.2012.2434
Eudy JD, Weston MD, Yao S, Hoover DM, Rehm HL, Ma-Edmonds M, Yan D, Ahmad I, Cheng JJ, Ayuso C, Cremers C, Davenport S, Moller C, Talmadge CB, Beisel KW, Tamayo M, Morton CC, Swaroop A, Kimberling WJ, Sumegi J (1998) Mutation of a gene encoding a protein with extracellular matrix motifs in Usher syndrome type IIa. Science 280(5370):1753–1757
Francois J (1971) Sex-linked chorioretinal heredodegenerations. Birth Defects Orig Artic Ser 7(3):99–116
Fu Q, Wang F, Wang H, Xu F, Zaneveld JE, Ren H, Keser V, Lopez I, Tuan HF, Salvo JS, Wang X, Zhao L, Wang K, Li Y, Koenekoop RK, Chen R, Sui R (2013a) Next-generation sequencing-based molecular diagnosis of a Chinese patient cohort with autosomal recessive retinitis pigmentosa. Invest Ophthalmol Vis Sci 54(6):4158–4166. doi:10.1167/iovs.13-11672
Fu W, O’Connor TD, Jun G, Kang HM, Abecasis G, Leal SM, Gabriel S, Rieder MJ, Altshuler D, Shendure J, Nickerson DA, Bamshad MJ, Project NES, Akey JM (2013b) Analysis of 6,515 exomes reveals the recent origin of most human protein-coding variants. Nature 493(7431):216–220. doi:10.1038/nature11690
Fujinami K, Lois N, Davidson AE, Mackay DS, Hogg CR, Stone EM, Tsunoda K, Tsubota K, Bunce C, Robson AG, Moore AT, Webster AR, Holder GE, Michaelides M (2013a) A longitudinal study of stargardt disease: clinical and electrophysiologic assessment, progression, and genotype correlations. Am J Ophthalmol 155(6):1075–1088 e1013. doi:10.1016/j.ajo.2013.01.018
Fujinami K, Zernant J, Chana RK, Wright GA, Tsunoda K, Ozawa Y, Tsubota K, Webster AR, Moore AT, Allikmets R, Michaelides M (2013b) ABCA4 gene screening by next-generation sequencing in a British cohort. Invest Ophthalmol Vis Sci 54(10):6662–6674. doi:10.1167/iovs.13-12570
Garber M, Guttman M, Clamp M, Zody MC, Friedman N, Xie X (2009) Identifying novel constrained elements by exploiting biased substitution patterns. Bioinformatics 25(12):i54–i62. doi:10.1093/bioinformatics/btp190
Genomes Project C, Abecasis GR, Altshuler D, Auton A, Brooks LD, Durbin RM, Gibbs RA, Hurles ME, McVean GA (2010) A map of human genome variation from population-scale sequencing. Nature 467(7319):1061–1073. doi:10.1038/nature09534
Genomes Project C, Abecasis GR, Auton A, Brooks LD, DePristo MA, Durbin RM, Handsaker RE, Kang HM, Marth GT, McVean GA (2012) An integrated map of genetic variation from 1,092 human genomes. Nature 491(7422):56–65. doi:10.1038/nature11632
Green PM, Saad S, Lewis CM, Giannelli F (1999) Mutation rates in humans. I. Overall and sex-specific rates obtained from a population study of hemophilia B. Am J Hum Genet 65(6):1572–1579. doi:10.1086/302651
Haim M (2002) Epidemiology of retinitis pigmentosa in Denmark. Acta Ophthalmol Scand Suppl 233:1–34
Siepel A, Pollard KS, Haussler D, RECOMB (2006) New methods for detecting lineage-specific selection. In: Proceedings of the 10th international conference on research in computational molecular biology, vol 3909. Springer, Berlin, pp 190–205
Hoefele J, Sudbrak R, Reinhardt R, Lehrack S, Hennig S, Imm A, Muerb U, Utsch B, Attanasio M, O’Toole JF, Otto E, Hildebrandt F (2005) Mutational analysis of the NPHP4 gene in 250 patients with nephronophthisis. Hum Mutat 25(4):411. doi:10.1002/humu.9326
Jansen GA, Waterham HR, Wanders RJ (2004) Molecular basis of Refsum disease: sequence variations in phytanoyl-CoA hydroxylase (PHYH) and the PTS2 receptor (PEX7). Hum Mutat 23(3):209–218. doi:10.1002/humu.10315
Kajiwara K, Berson EL, Dryja TP (1994) Digenic retinitis pigmentosa due to mutations at the unlinked peripherin/RDS and ROM1 loci. Science 264(5165):1604–1608
Kohn L, Kadzhaev K, Burstedt MS, Haraldsson S, Hallberg B, Sandgren O, Golovleva I (2007) Mutation in the PYK2-binding domain of PITPNM3 causes autosomal dominant cone dystrophy (CORD5) in two Swedish families. Eur J Hum Genet 15(6):664–671. doi:10.1038/sj.ejhg.5201817
Kolehmainen J, Wilkinson R, Lehesjoki AE, Chandler K, Kivitie-Kallio S, Clayton-Smith J, Traskelin AL, Waris L, Saarinen A, Khan J, Gross-Tsur V, Traboulsi EI, Warburg M, Fryns JP, Norio R, Black GC, Manson FD (2004) Delineation of Cohen syndrome following a large-scale genotype-phenotype screen. Am J Hum Genet 75(1):122–127. doi:10.1086/422197
Le Quesne Stabej P, Saihan Z, Rangesh N, Steele-Stallard HB, Ambrose J, Coffey A, Emmerson J, Haralambous E, Hughes Y, Steel KP, Luxon LM, Webster AR, Bitner-Glindzicz M (2012) Comprehensive sequence analysis of nine Usher syndrome genes in the UK National Collaborative Usher Study. J Med Genet 49(1):27–36. doi:10.1136/jmedgenet-2011-100468
Lewis RA, Shroyer NF, Singh N, Allikmets R, Hutchinson A, Li Y, Lupski JR, Leppert M, Dean M (1999) Genotype/Phenotype analysis of a photoreceptor-specific ATP-binding cassette transporter gene, ABCR, Stargardt disease. Am J Hum Genet 64(2):422–434. doi:10.1086/302251
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25(14):1754–1760. doi:10.1093/bioinformatics/btp324
Lindblad-Toh K, Garber M, Zuk O, Lin MF, Parker BJ, Washietl S, Kheradpour P, Ernst J, Jordan G, Mauceli E, Ward LD, Lowe CB, Holloway AK, Clamp M, Gnerre S, Alfoldi J, Beal K, Chang J, Clawson H, Cuff J, Di Palma F, Fitzgerald S, Flicek P, Guttman M, Hubisz MJ, Jaffe DB, Jungreis I, Kent WJ, Kostka D, Lara M, Martins AL, Massingham T, Moltke I, Raney BJ, Rasmussen MD, Robinson J, Stark A, Vilella AJ, Wen J, Xie X, Zody MC, Broad Institute Sequencing P, Whole Genome Assembly T, Baldwin J, Bloom T, Chin CW, Heiman D, Nicol R, Nusbaum C, Young S, Wilkinson J, Worley KC, Kovar CL, Muzny DM, Gibbs RA, Baylor College of Medicine Human Genome Sequencing Center Sequencing T, Cree A, Dihn HH, Fowler G, Jhangiani S, Joshi V, Lee S, Lewis LR, Nazareth LV, Okwuonu G, Santibanez J, Warren WC, Mardis ER, Weinstock GM, Wilson RK, Genome Institute at Washington U, Delehaunty K, Dooling D, Fronik C, Fulton L, Fulton B, Graves T, Minx P, Sodergren E, Birney E, Margulies EH, Herrero J, Green ED, Haussler D, Siepel A, Goldman N, Pollard KS, Pedersen JS, Lander ES, Kellis M (2011) A high-resolution map of human evolutionary constraint using 29 mammals. Nature 478 (7370):476–482. doi:10.1038/nature10530
Liu X, Jian X, Boerwinkle E (2013) dbNSFP v2.0: a database of human non-synonymous SNVs and their functional predictions and annotations. Hum Mutat 34(9):E2393–E2402. doi:10.1002/humu.22376
Maugeri A, van Driel MA, van de Pol DJ, Klevering BJ, van Haren FJ, Tijmes N, Bergen AA, Rohrschneider K, Blankenagel A, Pinckers AJ, Dahl N, Brunner HG, Deutman AF, Hoyng CB, Cremers FP (1999) The 2588G–>C mutation in the ABCR gene is a mild frequent founder mutation in the Western European population and allows the classification of ABCR mutations in patients with Stargardt disease. Am J Hum Genet 64(4):1024–1035
McGee TL, Seyedahmadi BJ, Sweeney MO, Dryja TP, Berson EL (2010) Novel mutations in the long isoform of the USH2A gene in patients with Usher syndrome type II or non-syndromic retinitis pigmentosa. J Med Genet 47(7):499–506. doi:10.1136/jmg.2009.075143
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M, DePristo MA (2010) The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20(9):1297–1303. doi:10.1101/gr.107524.110
McLaughlin ME, Ehrhart TL, Berson EL, Dryja TP (1995) Mutation spectrum of the gene encoding the beta subunit of rod phosphodiesterase among patients with autosomal recessive retinitis pigmentosa. Proc Natl Acad Sci 92(8):3249–3253
Mokry M, Feitsma H, Nijman IJ, de Bruijn E, van der Zaag PJ, Guryev V, Cuppen E (2010) Accurate SNP and mutation detection by targeted custom microarray-based genomic enrichment of short-fragment sequencing libraries. Nucleic Acids Res 38(10):e116. doi:10.1093/nar/gkq072
Najera C, Beneyto M, Blanca J, Aller E, Fontcuberta A, Millan JM, Ayuso C (2002) Mutations in myosin VIIA (MYO7A) and usherin (USH2A) in Spanish patients with Usher syndrome types I and II, respectively. Hum Mutat 20(1):76–77. doi:10.1002/humu.9042
Neveling K, Collin RW, Gilissen C, van Huet RA, Visser L, Kwint MP, Gijsen SJ, Zonneveld MN, Wieskamp N, de Ligt J, Siemiatkowska AM, Hoefsloot LH, Buckley MF, Kellner U, Branham KE, den Hollander AI, Hoischen A, Hoyng C, Klevering BJ, van den Born LI, Veltman JA, Cremers FP, Scheffer H (2012) Next-generation genetic testing for retinitis pigmentosa. Hum Mutat 33(6):963–972. doi:10.1002/humu.22045
Ng PC, Henikoff S (2003) SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res 31(13):3812–3814
Otto E, Hoefele J, Ruf R, Mueller AM, Hiller KS, Wolf MT, Schuermann MJ, Becker A, Birkenhager R, Sudbrak R, Hennies HC, Nurnberg P, Hildebrandt F (2002) A gene mutated in nephronophthisis and retinitis pigmentosa encodes a novel protein, nephroretinin, conserved in evolution. Am J Hum Genet 71(5):1161–1167. doi:10.1086/344395
Pennings RJ, Te Brinke H, Weston MD, Claassen A, Orten DJ, Weekamp H, Van Aarem A, Huygen PL, Deutman AF, Hoefsloot LH, Cremers FP, Cremers CW, Kimberling WJ, Kremer H (2004) USH2A mutation analysis in 70 Dutch families with Usher syndrome type II. Hum Mutat 24(2):185. doi:10.1002/humu.9259
Reva B, Antipin Y, Sander C (2011) Predicting the functional impact of protein mutations: application to cancer genomics. Nucleic Acids Res 39(17):e118. doi:10.1093/nar/gkr407
Rivera A, White K, Stohr H, Steiner K, Hemmrich N, Grimm T, Jurklies B, Lorenz B, Scholl HP, Apfelstedt-Sylla E, Weber BH (2000) A comprehensive survey of sequence variation in the ABCA4 (ABCR) gene in Stargardt disease and age-related macular degeneration. Am J Hum Genet 67(4):800–813. doi:10.1086/303090
Rivolta C, Sweklo EA, Berson EL, Dryja TP (2000) Missense mutation in the USH2A gene: association with recessive retinitis pigmentosa without hearing loss. Am J Hum Genet 66(6):1975–1978. doi:10.1086/302926
Rozen S, Skaletsky H (2000) Primer3 on the WWW for general users and for biologist programmers. Methods Mol Biol 132:365–386
Schwarz JM, Rodelsperger C, Schuelke M, Seelow D (2010) MutationTaster evaluates disease-causing potential of sequence alterations. Nat Methods 7(8):575–576. doi:10.1038/nmeth0810-575
Seyedahmadi BJ, Rivolta C, Keene JA, Berson EL, Dryja TP (2004) Comprehensive screening of the USH2A gene in Usher syndrome type II and non-syndromic recessive retinitis pigmentosa. Exp Eye Res 79(2):167–173. doi:10.1016/j.exer.2004.03.005
Sherry ST, Ward MH, Kholodov M, Baker J, Phan L, Smigielski EM, Sirotkin K (2001) dbSNP: the NCBI database of genetic variation. Nucleic Acids Res 29(1):308–311
Shihab HA, Gough J, Cooper DN, Stenson PD, Barker GL, Edwards KJ, Day IN, Gaunt TR (2013) Predicting the functional, molecular, and phenotypic consequences of amino acid substitutions using hidden Markov models. Hum Mutat 34(1):57–65. doi:10.1002/humu.22225
Shu X, Black GC, Rice JM, Hart-Holden N, Jones A, O’Grady A, Ramsden S, Wright AF (2007) RPGR mutation analysis and disease: an update. Hum Mutat 28(4):322–328. doi:10.1002/humu.20461
Simpson DA, Clark GR, Alexander S, Silvestri G, Willoughby CE (2011) Molecular diagnosis for heterogeneous genetic diseases with targeted high-throughput DNA sequencing applied to retinitis pigmentosa. J Med Genet 48(3):145–151. doi:10.1136/jmg.2010.083568
Song J, Smaoui N, Ayyagari R, Stiles D, Benhamed S, MacDonald IM, Daiger SP, Tumminia SJ, Hejtmancik F, Wang X (2011) High-throughput retina-array for screening 93 genes involved in inherited retinal dystrophy. Invest Ophthalmol Vis Sci 52(12):9053–9060. doi:10.1167/iovs.11-7978
Stenson PD, Mort M, Ball EV, Shaw K, Phillips AD, Cooper DN (2013) The Human Gene Mutation Database: building a comprehensive mutation repository for clinical and molecular genetics, diagnostic testing and personalized genomic medicine. Hum Genet. doi:10.1007/s00439-013-1358-4
Sun H, Smallwood PM, Nathans J (2000) Biochemical defects in ABCR protein variants associated with human retinopathies. Nat Genet 26(2):242–246. doi:10.1038/79994
Tuson M, Marfany G, Gonzalez-Duarte R (2004) Mutation of CERKL, a novel human ceramide kinase gene, causes autosomal recessive retinitis pigmentosa (RP26). Am J Hum Genet 74(1):128–138. doi:10.1086/381055
van den Hurk JA, Schwartz M, van Bokhoven H, van de Pol TJ, Bogerd L, Pinckers AJ, Bleeker-Wagemakers EM, Pawlowitzki IH, Ruther K, Ropers HH, Cremers FP (1997) Molecular basis of choroideremia (CHM): mutations involving the Rab escort protein-1 (REP-1) gene. Hum Mutat 9 (2):110–117. doi:10.1002/(SICI)1098-1004(1997)9:2<110::AID-HUMU2>3.0.CO;2-D
Venturini G, Di Gioia SA, Harper S, Weigel-Difranco C, Rivolta C, Berson EL (2014) Molecular genetics of FAM161A in North American patients with early-onset retinitis pigmentosa. PLoS One 9(3):e92479. doi:10.1371/journal.pone.0092479
Wada Y, Tada A, Itabashi T, Kawamura M, Sato H, Tamai M (2005) Screening for mutations in the IMPDH1 gene in Japanese patients with autosomal dominant retinitis pigmentosa. Am J Ophthalmol 140(1):163–165. doi:10.1016/j.ajo.2005.01.017
Wanders RJA, Waterham HR, Leroy BP (1993) Refsum Disease. In: Pagon RA, Adam MP, Bird TD, Dolan CR, Fong CT, Stephens K (eds) GeneReviews. Seattle
Wang DY, Chan WM, Tam PO, Baum L, Lam DS, Chong KK, Fan BJ, Pang CP (2005) Gene mutations in retinitis pigmentosa and their clinical implications. Clin Chim Acta 351(1–2):5–16. doi:10.1016/j.cccn.2004.08.004
Wang K, Li M, Hakonarson H (2010) ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res 38(16):e164. doi:10.1093/nar/gkq603
Wang X, Wang H, Sun V, Tuan HF, Keser V, Wang K, Ren H, Lopez I, Zaneveld JE, Siddiqui S, Bowles S, Khan A, Salvo J, Jacobson SG, Iannaccone A, Wang F, Birch D, Heckenlively JR, Fishman GA, Traboulsi EI, Li Y, Wheaton D, Koenekoop RK, Chen R (2013) Comprehensive molecular diagnosis of 179 Leber congenital amaurosis and juvenile retinitis pigmentosa patients by targeted next generation sequencing. J Med Genet 50(10):674–688. doi:10.1136/jmedgenet-2013-101558
Wang F, Wang H, Tuan HF, Nguyen DH, Sun V, Keser V, Bowne SJ, Sullivan LS, Luo H, Zhao L, Wang X, Zaneveld JE, Salvo JS, Siddiqui S, Mao L, Wheaton DK, Birch DG, Branham KE, Heckenlively JR, Wen C, Flagg K, Ferreyra H, Pei J, Khan A, Ren H, Wang K, Lopez I, Qamar R, Zenteno JC, Ayala-Ramirez R, Buentello-Volante B, Fu Q, Simpson DA, Li Y, Sui R, Silvestri G, Daiger SP, Koenekoop RK, Zhang K, Chen R (2014) Next generation sequencing-based molecular diagnosis of retinitis pigmentosa: identification of a novel genotype-phenotype correlation and clinical refinements. Hum Genet 133(3):331–345. doi:10.1007/s00439-013-1381-5
Webster AR, Heon E, Lotery AJ, Vandenburgh K, Casavant TL, Oh KT, Beck G, Fishman GA, Lam BL, Levin A, Heckenlively JR, Jacobson SG, Weleber RG, Sheffield VC, Stone EM (2001) An analysis of allelic variation in the ABCA4 gene. Invest Ophthalmol Vis Sci 42(6):1179–1189
Wiszniewski W, Zaremba CM, Yatsenko AN, Jamrich M, Wensel TG, Lewis RA, Lupski JR (2005) ABCA4 mutations causing mislocalization are found frequently in patients with severe retinal dystrophies. Hum Mol Genet 14(19):2769–2778. doi:10.1093/hmg/ddi310
Zeitz C, Labs S, Lorenz B, Forster U, Uksti J, Kroes HY, De Baere E, Leroy BP, Cremers FP, Wittmer M, van Genderen MM, Sahel JA, Audo I, Poloschek CM, Mohand-Said S, Fleischhauer JC, Huffmeier U, Moskova-Doumanova V, Levin AV, Hamel CP, Leifert D, Munier FL, Schorderet DF, Zrenner E, Friedburg C, Wissinger B, Kohl S, Berger W (2009) Genotyping microarray for CSNB-associated genes. Invest Ophthalmol Vis Sci 50(12):5919–5926. doi:10.1167/iovs.09-3548
Acknowledgments
We gratefully acknowledge all participating patients and their family members. R.C. is supported by grants from Retinal Research Foundation, Foundation Fighting Blindness (BR-GE-0613-0618-BCM) and the National Eye Institute (R01EY022356, R01EY018571). F.W. is supported by predoctoral fellowship: The Burroughs Wellcome Fund, The Houston Laboratory and Population Sciences Training Program in Gene Environment Interaction.
Conflict of interest
The authors declare no conflict of interest.
Ethical standards
This research was conducted in accordance with the Tenets of the declaration of Helsinki. Ethical permission was granted through ORECNI and all patients gave written consent to participate in the study.
Author information
Authors and Affiliations
Corresponding authors
Additional information
L. Zhao and F. Wang contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhao, L., Wang, F., Wang, H. et al. Next-generation sequencing-based molecular diagnosis of 82 retinitis pigmentosa probands from Northern Ireland. Hum Genet 134, 217–230 (2015). https://doi.org/10.1007/s00439-014-1512-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-014-1512-7